Abstract
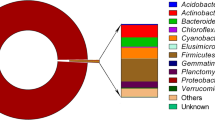
Gut microbiota of freshwater carp (Labeo rohita) was investigated by shotgun metagenomics to understand its taxonomic composition and functional capabilities. With the presence of 36 phyla, 326 families and 985 genera, the fish gut microbiota was found to be quite diverse in nature. However, at the phylum level, more than three-fourths of gut microbes belonged to Proteobacteria. Very low prevalence of commonly used probiotic bacteria (Bacillus, Lactobacillus, Streptococcus, and Lactococcus) in fish gut suggested the need to search for alternative probiotics for aquaculture use. Biosynthesis pathways were found to be the most dominant (51%) followed by degradation (39%), energy metabolism (4%) and fermentation (2%). In conformity with herbivorous feeding habit of L. rohita, gut microbiome also had pathways for the degradation of cellulose, hemicellulose, chitin, pectin, starch, and other complex carbohydrates. High prevalence of Actinobacteria and antibiotic biosynthesis pathways in the fish gut microbiome indicated its potential for bioprospecting of potentially novel natural antibiotics. Fifty-one different types of antibiotic resistance genes (ARGs) belonging to 15 antimicrobial resistance (AMR) gene families and conferring resistance against 24 antibiotic types were detected in fish gut. Some of the ARGs for multi-drug resistance were also found to be located on sequences of plasmid origin. The presence of pathogenic bacteria and ARGs on plasmid sequences suggested the potential risk due to horizontal gene transfer in the confined gut environment. The role of ARGs in fish gut microbiome needs further investigations.




Similar content being viewed by others
References
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments. Nat Rev Microbiol 8:251–259. https://doi.org/10.1038/nrmicro2312
Barka EA et al (2016) Taxonomy, physiology, and natural products of actinobacteria. Microbiol Mol Biol Rev 80:1–43. https://doi.org/10.1128/MMBR.00019-15
Bashir Y, Pradeep Singh S, Kumar Konwar B (2014) Metagenomics: an application based perspective. Chin J Biol 2014:7. https://doi.org/10.1155/2014/146030
Bengtsson-Palme J, Boulund F, Fick J, Kristiansson E, Larsson DG (2014) Shotgun metagenomics reveals a wide array of antibiotic resistance genes and mobile elements in a polluted lake in India. Front Microbiol 5:648. https://doi.org/10.3389/fmicb.2014.00648
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Buchfink B, Xie C, Huson DH (2015) Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. https://doi.org/10.1038/nmeth.3176
David LA et al (2014) Diet rapidly and reproducibly alters the human gut microbiome. Nature 505:559–563. https://doi.org/10.1038/nature12820
den Besten G, van Eunen K, Groen AK, Venema K, Reijngoud DJ, Bakker BM (2013) The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J Lipid Res 54:2325–2340. https://doi.org/10.1194/jlr.R036012
Ghanbari M, Kneifel W, Domig KJ (2015) A new view of the fish gut microbiome: advances from next-generation sequencing. Aquaculture 448:464–475. https://doi.org/10.1016/j.aquaculture.2015.06.033
Giatsis C et al (2016) Probiotic legacy effects on gut microbial assembly in tilapia larvae. Sci Rep 6:33965. https://doi.org/10.1038/srep33965
Govan JRW (2012) Pseudomonads and non-fermenters: Opportunist infection; cystic fibrosis; melioidosis. In: Greenwood D, Barer M, Slack R, Irving W (eds) Medical microbiology, 18th edn, Churchill Livingstone, Edinburgh, pp 298–304
Handelsman J, Rondon MR, Brady SF, Clardy J, Goodman RM (1998) Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products. Chem Biol 5:R245–R249
Hossein HS, Yun-Zhang S, Marlowe CC (2017) Short-chain fatty acids as feed supplements for sustainable aquaculture: an updated view. Aquac Res 48:1380–1391 doi. https://doi.org/10.1111/are.13239
Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinf 11:119. https://doi.org/10.1186/1471-2105-11-119
Jami M, Ghanbari M, Kneifel W, Domig KJ (2015) Phylogenetic diversity and biological activity of culturable Actinobacteria isolated from freshwater fish gut microbiota. Microbiol Res 175:6–15. https://doi.org/10.1016/j.micres.2015.01.009
Konig H, Li L, Frohlich J (2013) The cellulolytic system of the termite gut. Appl Microbiol Biotechnol 97:7943–7962. https://doi.org/10.1007/s00253-013-5119-z
Krawczyk PS, Lipinski L, Dziembowski A (2018) PlasFlow: predicting plasmid sequences in metagenomic data using genome signatures. Nucl Acids Res 46:e35–e35. https://doi.org/10.1093/nar/gkx1321
Li X, Yu Y, Feng W, Yan Q, Gong Y (2012) Host species as a strong determinant of the intestinal microbiota of fish larvae. J Microbiol 50:29–37. https://doi.org/10.1007/s12275-012-1340-1
Li T, Long M, Gatesoupe FJ, Zhang Q, Li A, Gong X (2015) Comparative analysis of the intestinal bacterial communities in different species of carp by pyrosequencing. Microbiol Ecol 69:25–36. https://doi.org/10.1007/s00248-014-0480-8
Li D et al (2016) MEGAHIT v1.0: A fast and scalable metagenome assembler driven by advanced methodologies and community practices. Methods 102:3–11. https://doi.org/10.1016/j.ymeth.2016.02.020
Liu H et al (2016) The gut microbiome and degradation enzyme activity of wild freshwater fishes influenced by their trophic levels. Sci Rep 6:24340. https://doi.org/10.1038/srep24340
Manivasagan P, Venkatesan J, Sivakumar K, Kim SK (2014) Pharmaceutically active secondary metabolites of marine actinobacteria. Microbiol Res 169:262–278. https://doi.org/10.1016/j.micres.2013.07.014
Martinez Cruz P, Ibanez AL, Monroy Hermosillo OA, Ramirez Saad HC (2012) Use of probiotics in aquaculture. ISRN Microbiol 2012:916845. https://doi.org/10.5402/2012/916845
McArthur AG et al (2013) The comprehensive antibiotic resistance database. Antimicrob Agents Chemother 57:3348–3357. https://doi.org/10.1128/AAC.00419-13
Prakash T, Taylor TD (2012) Functional assignment of metagenomic data: challenges and applications. Brief Bioinf 13:711–727. https://doi.org/10.1093/bib/bbs033
Saha AK, Ray AK (1998) Cellulase activity in rohu fingerlings. Aquac Int 6:281–291. https://doi.org/10.1023/a:1009210929594
Satola B, Wübbeler JH, Steinbüchel A (2013) Metabolic characteristics of the species Variovorax paradoxus. Appl Microbiol Biotechnol 97:541–560. https://doi.org/10.1007/s00253-012-4585-z
Seemann T (2014) Prokka: rapid prokaryotic genome annotation. Bioinformatics 30:2068–2069. https://doi.org/10.1093/bioinformatics/btu153
Singh B, Tyagi A, Kumar BTN, Ansal MD (2018) Prevalence and antimicrobial resistance of vibrios of human health significance in inland saline aquaculture areas. Aquac Res 49:2166–2174 doi. https://doi.org/10.1111/are.13672
Tarnecki AM, Burgos FA, Ray CL, Arias CR (2017) Fish intestinal microbiome: diversity and symbiosis unravelled by metagenomics. J Appl Microbiol. https://doi.org/10.1111/jam.13415
Tyagi A, Singh B (2017) Microbial diversity in rohu fish gut and inland saline aquaculture sediment and variations associated with next-generation sequencing of 16S rRNA gene. J Fish Life Sci 2:8
Verschuere L, Rombaut G, Sorgeloos P, Verstraete W (2000) Probiotic bacteria as biological control agents in aquaculture. Microbiol Mol Biol Rev 64:655–671
Wang WL, Xu SY, Ren ZG, Tao L, Jiang JW, Zheng SS (2015) Application of metagenomics in the human gut microbiome. World J Gastroenterol 21:803–814. https://doi.org/10.3748/wjg.v21.i3.803
Wang AR, Chao R, Einar R, ZZ G (2017) Progress in fish gastrointestinal microbiota research. Rev Aquac 0:15 doi. https://doi.org/10.1111/raq.12191
Wang JH, Lu J, Zhang YX, Wu J, Luo Y, Liu H (2018) Metagenomic analysis of antibiotic resistance genes in coastal industrial mariculture systems. Bioresour Technol 253:235–243. https://doi.org/10.1016/j.biortech.2018.01.035
Wood DE, Salzberg SL (2014) Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol 15:R46. https://doi.org/10.1186/gb-2014-15-3-r46
Wu S, Wang G, Angert ER, Wang W, Li W, Zou H (2012) Composition, diversity, and origin of the bacterial community in grass carp intestine. PLoS One 7:e30440. https://doi.org/10.1371/journal.pone.0030440
Xing M, Hou Z, Yuan J, Liu Y, Qu Y, Liu B (2013) Taxonomic and functional metagenomic profiling of gastrointestinal tract microbiome of the farmed adult turbot (Scophthalmus maximus). FEMS Microbiol Ecol 86:432–443. https://doi.org/10.1111/1574-6941.12174
Ye Y, Doak TG (2009) A parsimony approach to biological pathway reconstruction/inference for genomes and metagenomes. PLoS Comput Biol 5:e1000465. https://doi.org/10.1371/journal.pcbi.1000465
Acknowledgements
The authors are grateful to the Dean (College of Fisheries, Guru Angad Dev Veterinary and Animal Sciences University, Ludhiana, India) for all necessary support during this study. Funding support from Science and Engineering Research Board, Department of Science and Technology (DST-SERB) Young Scientist Start-Up Research Grant (YSS/2014/000269) to Anuj Tyagi has been utilized to carry this work and it is gratefully acknowledged.
Author information
Authors and Affiliations
Contributions
AT conceived the study, performed the bioinformatics analysis and drafted the manuscript. BS collected and processed the samples. NKBT helped in sample collection, result interpretations, and manuscript drafting. NKS supported in data analysis and manuscript drafting. All authors contributed in final manuscript correction.
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest declared.
Ethical standards
No special permission was required for this study.
Additional information
Communicated by Erko Stackebrandt.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tyagi, A., Singh, B., Billekallu Thammegowda, N.K. et al. Shotgun metagenomics offers novel insights into taxonomic compositions, metabolic pathways and antibiotic resistance genes in fish gut microbiome. Arch Microbiol 201, 295–303 (2019). https://doi.org/10.1007/s00203-018-1615-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-018-1615-y