Abstract
Key message
Blossom-End Rot is Quantitatively Inherited and Maps to Four Loci in Tomato.
Abstract
Blossom-end rot (BER) is a devastating physiological disorder that affects tomato and other vegetables, resulting in significant crop losses. To date, most studies on BER have focused on the environmental factors that affect calcium translocation to the fruit; however, the genetic basis of this disorder remains unknown. To investigate the genetic basis of BER, two F2 and F3:4 populations along with a BC1 population that segregated for BER occurrence were evaluated in the greenhouse. Using the QTL-seq approach, quantitative trait loci (QTL) associated with BER Incidence were identified at the bottom of chromosome (ch) 3 and ch11. Additionally, linkage-based QTL mapping detected another QTL, BER3.1, on ch3 and BER4.1 on ch4. To fine map the QTLs identified by QTL-seq, recombinant screening was performed. BER3.2, the major BER QTL on ch3, was narrowed down from 5.68 to 1.58 Mbp with a 1.5-LOD support interval (SI) corresponding to 209 candidate genes. BER3.2 colocalizes with the fruit weight gene FW3.2/SlKLUH, an ortholog of cytochrome P450 KLUH in Arabidopsis. Further, BER11.1, the major BER QTL on ch11, was narrowed down from 3.99 to 1.13 Mbp with a 1.5-LOD SI interval comprising of 141 candidate genes. Taken together, our results identified and fine mapped the first loci for BER resistance in tomato that will facilitate marker-assistant breeding not only in tomato but also in many other vegetables suffering for BER.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tomato (Solanum lycopersicum L.) is the second most produced and consumed vegetable in the world. The demand for this vegetable has increased over the years since the produce offers wide-ranging health benefits. This growing demand has led to a steady increase in tomato production in the world, exceeding 182 million tons in 2018 (FAOSTAT 2020). Yet, this crop faces major biotic and abiotic challenges that can lead to a substantial amount of the produce being lost. Among these, physiological disorders are abiotic syndromes that affect either the whole plant or specific parts of the plant such as fruits, roots, and leaves. These disorders render the vegetable or fruit unmarketable and thus result in significant yield losses and penalized market prices (Hagassou et al. 2019; Ikeda and Kanayama 2015). As one of the most common physiological disorders in tomato, blossom-end rot (BER) alone causes serious economic losses that may reach up to 50% in this vegetable (Taylor and Locascio 2004). Just as an example, in 2018, Hickory Hill Farm in Carlton GA, USA, experienced dramatic yield losses due to BER in organically grown hybrid tomatoes that reached up to 80% (Josh Johns and Gary Shaw, personal communication, August 9, 2018). Unfortunately, the unpredictability of BER occurrence and adverse weather conditions aggravate this problem since extreme weather events are becoming increasingly more prevalent (Barickman et al. 2014; Penella and Calatayud 2018). Despite its economic importance, the underlying causes of BER are not well understood. The occurrence of BER has been primarily attributed to calcium (Ca2+) deficiency (Adams and Ho 1993; Raleigh and Chucka 1944). Along this vein, aberrant regulation of cellular Ca2+ distribution and partitioning, especially in the distal placenta, appears to be linked to BER (de Freitas et al. 2011, 2012b, 2014). Ca2+ plays an important role as a structural component of cell walls and membranes, and previous studies have suggested that higher concentration of Ca2+ in the apoplastic space ([Ca2+]apo) affects cell wall strength and stability (de Freitas et al. 2012a; Thor 2019). Just as important is the role of calcium as an intracellular secondary messenger. Therefore, Ca2+ concentration in the cytosol ([Ca2+]cyt) is tightly regulated as well (Clapham 2007; Clarkson et al. 1993; Kudla et al. 2010; Thor 2019). Transient, sustained, or oscillatory elevations in the [Ca2+]cyt initiate a signaling cascade that orchestrates induction of downstream responses needed for given stimulus, such as defense and stress response gene expression, and Ca2+-controlled stomatal closure (Dodd et al. 2010; Ng et al. 2001). Even though a central role for Ca2+ in the development of BER has been postulated for many years, neither a consistent solution nor a direct link to fruit Ca2+ concentration has been conclusively demonstrated (Ho and White 2005). Therefore, other physiological links to the causes of BER such as reactive oxygen species (ROS) formation have recently gained prominence (Rached et al. 2018). Because of its destructive activity, excessive ROS release upon exposure of plants to stress conditions causes cell membrane lipid peroxidation, membrane leakage, and subsequently cell death, which can lead to the development of BER (Aktas et al. 2003; Rached et al. 2018; Van Breusegem and Dat 2006). Moreover, ROS production reaches a maximum, when the rate of cell expansion during fruit growth is at its maximum (Aktas et al. 2003). As a defense mechanism against ROS, plants produce antioxidants to neutralize or alleviate the negative impact of ROS. Hence, tomato varieties that feature high levels of antioxidants show resistance to BER (Rached et al. 2018). In addition to aberrant regulation of calcium and ROS, much emphasis has been placed on other physiological and genetic factors, such as accelerated fruit growth rate, phytohormones, salinity, drought, high light intensity, fruit weight and shape to explain BER (Hagassou et al. 2019; Ho and White 2005). Typically, fruit weight and elongated shape are positively correlated to BER occurrence (Ho and White 2005; Marcelis and Ho 1999). Yet not all large fruited or oval-shaped tomato varieties feature BER to the same degree. This implies that there may be genetic basis for the disorder that is hitherto unknown. Nonetheless, only a few genetic and mapping studies have been carried out for BER, despite the desire to identify resistance loci to utilize them for crop improvement (Uozumi et al. 2012). Hence, the objective of this study is to investigate the genetic basis of BER tomato. It is our expectation that these findings will ultimately provide novel knowledge about the causes of BER and to enable marker-assisted breeding not only in tomato but also in other crops that suffer from the disorder.
Material and methods
Plant materials and population construction
Two segregating F2 populations, 17S28 (n = 192) and 20S166 (n = 192), were generated by crossing BER-resistant accessions BGV007900 (Solanum lycopersicum var. cerasiforme) and BGV008224 (S. lycopersicum var. lycopersicum), respectively, with BER-susceptible accession BGV007936 (S. lycopersicum var. lycopersicum). Furthermore, a BC1 population (18S243, n = 144) was created using the susceptible accession (BGV007936) as the recurrent parent in the BGV007900 x BGV007936 F1. These phylogenetically closely related accessions (Razifard et al. 2020) were selected from the “Varitome” collection, and SNP data in this collection are publicly available at SGN (https://solgenomics.net/projects/varitome/). The 17S28 and 20S166 F2 mapping populations were, respectively, grown in Spring 2017 and Fall 2020 and included the F1 and parental controls. The 18S243 BC1 mapping population was grown in Spring 2018 without controls. Follow-up mapping populations were 19S499 (n = 171) and 20S74 (n = 192) F3:4 populations that were grown in Spring and Summer 2020, respectively. Only recombinant plants in the QTL interval on ch3 and ch11, respectively, were selected and grown with parental checks. All populations were grown in greenhouse where the irrigation, temperature, and supplemental light settings are Argus controlled at the University of Georgia (Athens, USA). Briefly, plants were grown in 3.79-L pots filled with a commercial potting mix (Sun Gro® Fafard® 3B Mix/Metro-Mix 830, Sun Gro Horticulture Inc, Agawam, MA) and fertilized with Nutricote controlled release fertilizer (CRF) (18N-6P-8K with 37.5 g/pot, Florikan, Sarasota, FL) and MEG-IRON V micronutrient mix (7.5 g/pot, Florikan, Sarasota, FL) following the manufacturers’ recommendations.
BER phenotyping
To assess BER, three continuous assays and one nominal assay were developed. These assays were the following: (1) BER Severity 1, the ratio of the BER diameter to the whole fruit diameter (DBER/DFruit); (2) BER Severity 2, the ratio of the weight of the affected part of the fruit to the total fruit weight (WBER/WALL); (3) BER Incidence, the ratio of the number of fruit affected to the total number of fruit (AFN/TFN); and (4) BER Visual, where each fruit was scored by using a scale of 1–5 with 1 = healthy with no BER symptoms and 5 = extensive BER (Fig. 1). BER was evaluated using only the first 3–5 fruits on the first three inflorescence.
DNA isolation and library preparation for sequencing
The DNA extraction and library preparation were performed as described before (Illa-Berenguer et al. 2015). Briefly, the genomic DNA of the plants was extracted from young true leaves using the DNeasy Plant Mini Kit (Qiagen, Valencia, CA) following the manufacturer’s recommendation. For the recombinant screening, the genomic DNA of the plants was extracted from cotyledons following the Geno/Grinder method described by Zhang et al. (2012). In the 17S28 F2 population, 12 plants that represented the high BER Incidence and 19 plants that represented the low BER Incidence (resistant) were selected. Similarly, 10 plants that featured high BER Severity 2 were selected for a total of three pools (Supplementary Table 1). Prior to library preparation, the genomic DNA of the plants selected for each bulk was quantified using the Qubit 2.0 Fluorimeter (Invitrogen, Carlsbad, CA, USA). For the library preparation, the NEBNext Ultra™ II DNA Library Prep Kit (New England Biolabs, USA.) and three barcoded primers from the NEBNext® Multiplex Oligos for Illumina kits (New England Biolabs, USA.) were used. Libraries were subjected to whole genome sequencing using the Illumina NextSeq 550 (300 cycles) paired-end 150-bp (PE150) flow cells at the Georgia Genomics and Bioinformatics Core at University of Georgia (Athens, GA).
Genome sequence analysis for QTL-seq
The generated FASTQ files were merged and then assessed using the FastQC program (version 0.11.4) (Andrews 2010). Prior to further analysis, FASTQ files were filtered and trimmed using Trim Galore (version 0.4.5) for a quality value of at least 28 ( https://github.com/FelixKrueger/TrimGalore). The remaining 150-bp paired-end reads were aligned to the genomes of the inbred tomato cultivar “Heinz 1706; version SL4.0” (Sato et al. 2012) using Burrows-Wheeler Aligner (BWA) with the default parameters (Li and Durbin 2009). Average coverage for each bulk was calculated using SAM tools (Li et al. 2009). After alignment, the SAM files were converted into BAM files using SAM tools (Li et al. 2009). The BAM files were sorted and filtered using Picard software (version 2.17.4) ( https://broadinstitute.github.io/picard/). Next, the variant calling including SNP-calling was performed using Genome Analysis Toolkit (GATK) (version 3.4–0) (McKenna et al. 2010). The recommended default settings for GATK Haplotype caller were used, and SNPs with QUAL > 30 were kept (Van der Auwera et al. 2013). The final variant call format (VCF) file was converted into a tab-delimited table using the “VariantsToTable” function from GATK. The tab-delimited table format file was used for downstream analyses. R package “QTLseqR” was used to identify QTL (Mansfeld and Grumet 2018).
KASP marker development and genotyping
The parental lines were genotyped for known fruit weight and shape genes (Supplementary Table 2) including CNR (FW2.2), KLUH (FW3.2), CSR (FW11.3), OVATE (OVATE), OFP20 (SOV1), LOCULE NUMBER (LC), and FASCIATED (FAS) (Chakrabarti et al. 2013; Ramos 2018; Rodríguez et al. 2011; Wu et al. 2018). The 17S28 F2 population is segregating for FAS and FW3.2, whereas the 20S166 F2 population was fixed for all the known fruit weight and shape genes (Table 1). Additionally, fluorescence-based KASP markers were developed using SNPs data identified from the genome sequencing data using the Primer Express® Software version 3.0.1 (Applied Biosystems, Carlsbad, CA). The Tm of two allele-specific forward primers were selected in the range of 58–61 °C (optimum: 60 °C) with minimum total GC content of 30%. Tm difference between primer pairs was set to be maximum 1 °C, and the desired product size was determined to be between 60–200 bp. Moreover, each primer had less than five repeating nucleotides in a row and was at least 25 bp in length. Next, we BLASTed each allele-specific forward primers against the SL4.0 tomato reference genome assembly in SGN (http://solgenomics.net/tools/blast/), and primers that only corresponded to the target sequence were selected. IDT oligo analyzer tool (https://www.idtdna.com/calc/analyzer) was used to test possible secondary structures, such as hairpins and primer dimers. Primer3Plus software (http://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi) was used to design the reverse primer. Finally, allele-specific primers along with common primer were tested for possible cross-dimer formation in primer pairs using multiple primer analyzer function in Thermo Scientific Web Tools (https://www.thermofisher.com/… /thermo-scientific-web-tools.html). Either FAM™ (GAAGGTGACCAAGTTCATGCT) or HEX™ (GAAGGTCGGAGTCAACGGATT) unique tail sequences were attached to the 5’ end of the allele-specific primers. The KASP markers used in this study are summarized in Supplementary Table 2. KASP genotyping was conducted in 384-well plates with a total reaction volume of 5 μL, containing 2 μL of 20–100 ng/μL genomic DNA. A total of 3 μl of the KASP PCR reaction mix was dispensed into each well using Mantis® microfluidic liquid handler (FORMULATRIX®, Bedford, MA). KASP PCR mix and PCR conditions are summarized in Supplementary Table 3. Tecan Infinite M200 Pro microplate reader (Tecan, Group Ltd., Mannersdorf, Switzerland) was used for KASP fluorescent end-point readings after the amplification. Automated genotype calling was performed using KlusterCaller software (Version 3.4.1.39, LGC Genomics LLC) using the raw data imported from Tecan microplate reader.
Linkage map construction and QTL analysis
The R/QTL (version 1.46–2, (Broman et al. 2003)) was employed to estimate genetic distances and construct genetic linkage maps. The Kosambi map function (Kosambi 1943) was used to estimate mapping distance in centimorgan (cM) by converting recombination frequencies. The logarithm of odds (LOD) scores was estimated using nonparametric interval mapping (scanone function; model = “np”) in R/QTL since the BER Incidence data does not meet the normality assumption. To declare the presence of a significant QTL, a 99% significance threshold was determined using permutation test with 1000 permutations.
Statistical analysis
The assumption of normality was checked using the Shapiro–Wilk tests and quantile–quantile (Q-Q) plot. Significant differences were considered at p < 0.05. Histograms, scatter and box plots were created in R open-source software (version 1.2.5001, (R Core Team 2019)). Pearson correlation coefficient was calculated using JMP software (version 13.2.0 (SAS Institute Inc 2017)). The broad sense heritability of each trait (H2) was calculated as described by Kearsey and Pooni 1996. In brief, roughly six to nine F1 progenies and six to nine plants from each parent were grown with the populations, and the following formula was used to estimate H2;
where VF2 denotes the variation amongst F2 individuals, VP1 and VP2 represent the variation amongst parents and finally VF1 shows the variation amongst F1 plants. The phenotypic variance for F2 lines is due to the combination of both genetic and environmental factors. However, the phenotypic variance amongst parental lines and F1 progenies is due to only environmental factors.
The gene action or degree of dominance (D/A) was calculated as the ratio between dominance and additive effects. Additive effect (A) was estimated as ½ (A1A1—A2A2), where A1A1 is the mean phenotypes of homozygous BGV007900 allele and A2A2 is the mean phenotypes of homozygous BGV007936. Dominance effect (D) was estimated as A1A2 – ½ (A1A1 + A2A2). The software QTL IciMapping (Version 4.1 (Meng et al. 2015)) was used to calculate/infer D/A values. Based on the estimates of dominance effect with those of additive effect, QTL were divided into additive effect (− 0.25 ≤ d/a ≤ 0.25), incomplete or partial dominance (± 0.25 ≤ d/a ≤ ± 0.75), complete dominance (± 0.75 ≤ d/a ≤ ± 1.25), overdominance (d/a > ± 1.25) (Tanksley 1993).
Results
Phenotypic evaluation of BER traits in the 17S28 F2 population
BER was quantified using four methods: BER Severity 1 (DBER/DFruit), BER Severity 2 (WBER/WALL), BER Incidence (AFN/TFN), and BER Visual (a scale of 1–5). The parents of the 17S28 F2 population showed striking differences in terms of BER Incidence and Severity. While the resistant parent BGV007900 showed consistently low Incidence and low Severity, the susceptible parent BGV007936 displayed an opposite trend (Fig. 2a, Supplementary Fig. 1). The distribution of F2 plants exhibited continuous variation for each trait, indicating BER Severity 2 that the traits were quantitatively inherited (Fig. 2b-e). Yet, all distributions (BER Severity 1, BER Severity 2 and BER Visual) were skewed toward BER-resistant parent BGV007900 except for BER Incidence where the distribution was skewed toward BER-susceptible parent. The broad sense heritability for all the four BER-related traits in the 17S28 F2 population ranged from H2 = 0.48 to H2 = 0.80, suggesting a strong genetic basis to the BER traits (Table 2). The BER traits were also found to be significantly correlated to one another ranging from r = 0.78 to r = 0.98 (Table 3).Correlation analysis between BER Incidence and BER Severity 2 indicated that some F2 plants displayed high BER Incidence while showing low BER Severity 2 (Fig. 2f). This suggested that BER Incidence and BER Severity 2 may be controlled by different loci. As shown in the BER Severity 2 frequency histogram (Fig. 2d), 101 F2 plants were slightly affected by BER (less than 0.1 BER Severity 2). However, 6 F2 individuals were completely consumed by BER. A Chi-square goodness-of-fit test shows that data are consistent with a 15:1 and two segregating loci scenario (χ2(0.01,1) = 0.075, Prob > ChiSq = 0.783 for BER Severity 2). With respect to BER Incidence, 48 F2 plants produced healthy fruits and featured less than 0.1 BER Incidence, whereas 17 plants produced fruit that were all affected by BER (Fig. 2b). The ratio of 48:17 is consistent with 3:1 and one segregating locus scenario (χ2(0.01,1) = 0.046, Prob > ChiSq = 0.829) for BER Incidence. Since we expected few loci for BER, a QTL-seq approach was used. For this purpose, genomic libraries were constructed using DNA from plants showing most extreme phenotypes from the 17S28 F2 population. The high BER Incidence and high BER Severity 2 plants in each pool did not overlap (Supplementary Table 1).
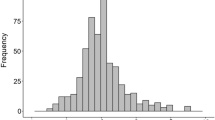
Phenotypic evaluations of BER parents and BER frequency distributions in 17S28 F2 population. a Phenotypic difference between BER-resistant parent BGV007900 (P1, with no BER Incidence), BER-susceptible parent BGV007936 (P2, with high BER Incidence), and their F1 generation 17S29. b BER Incidence frequency histogram in F2 progenies (17S28, n = 192). Gray and black triangles indicate the plants selected to generate Resistant Bulk (RB) and Susceptible Bulk (SB). White, gray, and black arrowheads on each histogram show the average of BGV007900, F1 and BGV007936 plants, respectively, for the trait of interest. Frequency distribution of c BER Severity 1, d BER Severity 2, and e BER Visual traits. f Correlation between BER Severity 2 and BER Incidence traits
Identification and mapping of QTL using QTL-seq
The next-generation sequencing (NGS)-based Illumina protocol generated between 224,961,232 and 394,052,926 million 150-bp paired-end reads that after filtering were mapped to tomato reference genome (Supplementary Table 4). A total of 434,022 and 424,900 high-quality SNPs were called by comparing the BER Incidence and BER Severity 2 bulks, respectively (Supplementary Tables 5, 6). The absolute Δ(SNP index) for “SNP-index_resistant Bulk—SNP-index_ Incidence _Bulk” and “SNP-index_resistant Bulk—SNP-index_ Severity _Bulk” with a statistical confidence of p < 0.05 was calculated. As a result, two significant genomic positions were identified for both BER Incidence and Severity 2 traits on ch1 and ch3 (Supplementary Fig. 2,3). For BER Incidence, another genomic region was identified on ch11, whereas a different genomic position was identified for BER Severity 2 on ch8 (Supplementary Figs. 2,3). To further examine additional putative small effect QTL (the average absolute Δ(SNP-index)) was close to the 95% confidence interval), additional KASP markers were developed for ch2, ch6, ch8, and ch10 for both BER Incidence and Severity 2 traits (Supplementary Figs. 2, 3; Supplementary Table 2).
Validation of identified QTL by SNP markers
To validate QTL(s) identified by QTL-seq, polymorphic SNPs were converted into molecular markers to flank and encompass the identified loci. Next, marker–trait association analyses for BER Incidence and Severity 2 traits were conducted using entire 17S28 F2 population. Once marker–trait associations were found to be significant, additional molecular markers were developed and QTL analysis was performed. Although the QTL-seq approach found three candidate regions, only QTL at ch3 and ch11 were validated using the entire 17S28 F2 population (Fig. 3). KASP markers developed for BER QTL on ch1 did not show significant association with both BER Incidence and Severity 2 traits (p > 0.05). Furthermore, most of the minor QTLs were not validated using marker trait association analysis except on ch4 (Fig. 3).
Mapping of BER Incidence in the 17S28 F2 population. a−d Mapping of BER3.2. e−h Mapping of BER4.1. i−l Mapping of BER11.1. a, e, i Absolute Δ(SNP-index) plots of a ch3, e ch4, and i ch11. X-axis shows the genomic position in Mb and Y-axis shows Absolute Δ(SNP-index). Tricube-weighted abs Δ(SNP-index) was used to facilitate visualization and interpretation of the graphs (solid blue line). Green and pink lines indicate the 99% and 95% confidence interval (CI), respectively, under the null hypothesis of no QTLs is present (p < 0.01 and 0.05. Triangles on the X-axis show the approximate genomic positions of the first and last genotyped marker in Mb. b, c, f, g, j, k Linkage-based QTL mapping. Partial genetic maps and map distances of b, c ch3, f, g ch4, and j, k ch11. The triangles on the x-axis show the position of the genotyped markers and their corresponding genetic distances in cM, and the y-axis represents the LOD scores. A logarithm of odds (LOD) threshold of α = 0.01 was found at 3.05 after 1000 permutation test (red dotted line). Bars show 1.0-LOD SI, and the whiskers represent 1.5-LOD SIs for each QTL. d, h, l Box plots of allelic effects of the highest associated SNP markers in each QTL interval for the trait of BER Incidence. The allelic effect of d BER11.1, h BER4.1, and l BER11.1. Comparisons for all pairs were made using Tukey–Kramer HSD, and levels not connected by the same letter are significantly different at α = 0.05 significance level. “RR” are plants homozygous for the resistant BGV007900 allele, “RS” are plants heterozygous at this locus, and “SS” are plants homozygous for the susceptible BGV007936 allele
Of the two analyzed traits, BER Incidence showed the highest association with molecular markers compared to BER Severity 2. Hence, only BER Incidence was further investigated in follow-up studies. The largest effect BER QTL on ch3, BER3.2, accounted for 16.35% phenotypic variation and exhibited a LOD score of 7.44 (Fig. 3b). To determine the confidence intervals of the identified QTLs, BER3.2 showed a 1.5-LOD SI extending from 15.7 to 42.1 cM (closest genetic markers 19EP596 and 18EP730, respectively), which corresponded to the physical positions of SL4.0 54,214,617 – SL4.0 59,891,210-bp (equaling 5.68 Mbp) region on the tomato reference genome of version SL4.0. Furthermore, the additive effect and D/A values for this QTL were -0.19 and 0.01, respectively. This indicated that BER3.2 acted in an additive manner (Table 4). QTL BER4.1 was flanked by markers 19EP885 (SL4.0ch4 5,481,420) and 18EP625 (SL4.0ch4 55,400,792) (Fig. 3f). The highest associated markers with BER4.1 were located near the centromeric region of ch4, and 1.5-LOD SI covered nearly the entire chromosome. BER4.1 explained a small portion of the phenotypic variance (8.57%) with a maximum LOD score of 3.74. The D/A was -0.16, suggesting that the alleles largely acted in an additive manner (Table 4). On ch11, BER11.1 explained 17.24% of the phenotypic variation with a LOD of 8.22. BER11.1 had a 1.5-LOD SI extending from 9.6 to 32.1 cM (closest genetic markers 18EP789 and 18EP1043, respectively), which corresponded to the physical positions of SL4.0 48,131,615—SL4.0 52,123,165 (equaling 3.99 Mbp) (Fig. 3j). The additive effect and D/A for this QTL were -0.18 and 0.38, respectively. In contrast to BER3.2 and BER4.1, BER11.1 exhibited an incomplete or partial dominance gene action for the BGV007900 allele (low BER occurrence). In addition, box plots of the highest associated SNP markers in each QTL interval are shown in Fig. 3d,h,l. Furthermore, digenic interactions of BER QTLs in the 17S28 F2 population were tested, but no significant epistatic or additive interactions were found among the loci (Prob > F = 0.1486 [ch3 and ch4, Fig. 4a], Prob > F = 0.5574 [ch3 and 11, Fig. 4b], Prob > F = 0.8959 [ch4 and ch11, Fig. 4c]).
BER mapping in the other two populations
Since most distributions in 17S28 F2 population were skewed toward BER-resistant parent BGV007900 and BER appeared to be additive, a backcross population 18S243 BC1 population was generated using the BER-susceptible BGV007936 as a recurrent parent. BER Incidence showed continuous variation in the BC1 population with a skewed distribution toward BER-susceptible parent BGV007936 (Fig. 5a). BER Visual also displayed continuous variation with a normal distribution (Shapiro–Wilk W test, W = 0.99, Prob < W = 0.66). Pearson correlation coefficient between BER Incidence and BER Visual was r = 0.81, indicating that both traits were highly correlated as expected. In the backcross population, BER11.1. has a 1.5-LOD SI extended from 16.7 to 40.4 cM (closest genetic markers 18EP951 and 18EP1117, respectively), corresponding to position SL4.0 50,569,217—SL4.0 54,182,901 bp (3.61 Mbp) (Fig. 5d,g). BER11.1. explained 13.75% of the phenotypic variance with a LOD score of 4.62 (Table 4). Box plots of allelic effects of the highest associated SNP marker 18EP879 are shown in Fig. 5j. In the 18S243 BC1 population, BER3.2 and BER4.1 were not segregating.
BER mapping in two additional populations. a−c Frequency distributions of BER traits. a BER Incidence distribution in 18S243 BC1, b BER Incidence, and c BER Visual distributions in the 20S166 F2 population. White, gray, and black arrowheads on each histogram show the average of BGV008224, F1 and BGV007936 plants, respectively, for the trait of interest. d−i Linkage-based QTL mapping. Partial linkage map and map distances of d, g ch11, e, h ch3, and f, i ch4. A logarithm of odds (LOD) threshold value for α = 0.01 was found to be 2.98 for 18S243 BC1 and 3.40 for 20S166 F2 after 1000 permutation test. j−l Box plots of allelic effects of the highest associated SNP markers. The allelic effect of j BER11.1, k BER3.1 and l BER4.1. “RR” shows plants homozygous for the resistant BGV008224 allele, “RS” plants heterozygous at this locus, and “SS” plants homozygous for the susceptible BGV007936 allele
Since two loci (fw3.2 and fas) associated with fruit weight variation were segregating in both 17S28 F2 and 18S243 BC1 populations, a new F2 mapping population was developed that did not segregate for any of the known fruit weight or shape genes. The F2 population 20S166 was derived from a cross between BER-resistant accession BGV008224 and the same BER-susceptible accession BGV007936 (Table 1). BER Incidence distribution of F2 plants showed skewed distribution toward BER-susceptible parent BGV007936. A total of 187 F2 plants were severely affected by BER (BER Incidence ≥ 0.50) ,whereas only five plants were slightly affected by BER (BER Incidence ≤ 0.50 [Fig. 5b]). For BER Visual, F2 plants showed continuous variation without normal distribution (Shapiro–Wilk W test, W = 0.97, Prob < W = 0.0011[Fig. 5c]). Because of the extremely high BER Incidence, the BER Visual trait was used to phenotype the 20S166 F2 population. Using the previously identified regions linked to BER, incomplete linkage maps of ch3, ch4, and ch11 were generated, showing only two loci that segregated in the 20S166 F2 population (Fig. 5e,f). Interestingly, the most significant markers on ch3 did not overlap with the map position of BER3.2 found in the other F2 population. Therefore, we named this locus BER3.1 because it mapped higher on the chromosome. BER3.1 explained 15.47% of the BER Visual variance with a LOD of 7.00 in 20S166 F2 population. The D/A was 0.02, suggesting that the alleles largely acted in an additive manner. The 1.5-LOD SI extended from 0 to 14.1 cM (closest genetic markers 20EP1015 and 18EP703, respectively), which corresponded to the physical interval of SL4.0ch3 47,418,933…53,495,792 bp (equaling 6.08 Mbp) (Fig. 5h). BER4.1 explained 19.12% of the phenotypic variation with a LOD of 8.85 and thus was the most significant BER QTL in this population (Table 4). BER4.1 had a 1.5-LOD SI extending from 49.6 to 66.1 cM (closest genetic markers 20EP139 and 18EP625, respectively), which corresponded to the physical positions of SL4.0 49,843,412—SL4.0 55,400,792 (equaling 5.56 Mbp) (Fig. 5f,i). Importantly, 1.5-LOD SI of BER4.1 in 20S166 F2 population partially overlapped with the 1.5-LOD SI of BER4.1 in 17S28 F2 population, suggesting that they were the same. Moreover, D/A for BER4.1 was 0.41, indicating that BER4.1 exhibited an incomplete or partial dominance gene action for the BGV007900 allele. Box plots of allelic effects of the highest associated SNP markers 20EP1012 and 20EP194 are shown in Fig. 5k,l. Finally, epistatic, or additive interaction between BER3.1 and BER4.1 was evaluated, and no significant interaction was found between the loci (Prob > F = 0.4212 [ch3 and ch4]) for the trait of BER Visual (Fig. 6).
Fine mapping BER3.2 and BER11.1
To further delineate the BER3.2 and BER11.1 intervals, recombinant screening was performed. A total of 768 F3:4 seedlings were genotyped with markers 18EP703 (SL4.0ch3 53,495,792) and 18EP1037 (SL4.0ch3 60,772,821) for BER3.2 and only recombinant plants were transplanted in the greenhouse (20S74 population; n = 192). After the selection, the recombinant plants were genotyped with additional KASP markers. The frequency histogram of 20S74 population for the BER Incidence trait showed continuous variation with a skewed distribution toward BGV007900 resistant parent (Fig. 7a). QTL mapping showed that BER3.2 was located between 20EP1033 (SL4.0ch3 58,308,917) and 18EP730 (SL4.0ch3 59,891,210), narrowing the locus down from 5.68 to 1.58 Mbp (Fig. 7b). This interval consists of 209 candidate genes including the fruit weight locus FW3.2/SlKLUH. The box plot of allelic effects of the SNP marker 19EP261 is shown in Fig. 7c.
Fine mapping of the BER3.2 and BER11.1. a−c Recombinant screening and fine mapping of BER3.2 d−f Recombinant screening and fine mapping of BER11.1 Frequency distribution of recombinant F3:4 plants for a BER3.2 in 20S74 and d BER11.1 in 19S499 F4 populations. White, gray, and black arrowheads on each histogram show the average of BGV007900, F1 and BGV007936 plants, respectively, for the trait of interest. b, e Linkage-based QTL mapping. b BER3.2 was further delineated to approximately 1.58-Mb region flanked by 20EP1033 (SL4.0ch3 58,308,917) and 18EP730 (SL4.0ch3 59,8912,10) markers. e BER11.1 was further fine mapped to 1.13-Mb region flanked by 18EP1043 (SL4.0ch11 52,123,165) and 18EP1114 (SL4.0ch11 53,258,120) markers. The numbers between genetic markers represent the number of recombinant plants. Bars show 1.0-LOD SI, and the whiskers represent 1.5-LOD SIs for each QTL c,f) Box plots of allelic effects of the most significant markers identified in the 1.5-LOD SI. The allelic effects of the most significant markers c 19EP2161 in BER3.1 region and f 20EP385 in BER11.1 region. “RR” indicates plants homozygous for the resistant BGV007900 allele, “RS” shows plants heterozygous at this locus, and “SS” plants homozygous for the susceptible BGV007936 allele
The same approach was applied to further map BER11.1. A total of 1152 F3:4 seedlings were genotyped with markers 18EP1049 (SL4.0ch11 51,268,187) and 18EP1114 (SL4.0ch11 53,258,120), and the 19S499 population comprising of 171 recombinants were transplanted to the greenhouse. BER Incidence showed continuous variation with a skewed distribution toward BGV007936-susceptible parent (Fig. 7d). In this population, BER11.1 mapped between 18EP1043 (SL4.0ch11 52,123,165) and 18EP1114 (SL4.0ch11 53,258,120), corresponding to a 1,134,955-bp interval (Fig. 7e). Consequently, the BER11.1 QTL was narrowed down from 3.69 to 1.13 Mbp interval, covering 141 candidate genes. Box plot of allelic effects of the SNP marker 20EP385 is shown in Fig. 7f.
Discussion
Despite extensive efforts to manage BER and related disorders in fruit and vegetable production, the underlying causes of this syndrome are poorly understood. While the genetic bases of physiological disorders have remained elusive, most emphasis has been placed on the physiological aspects of the syndromes. Nonetheless, these extensive efforts have delivered limited practical solutions to the problem and led to diminished production of tomato and other vegetables around the world (Taylor and Locascio 2004). Therefore, we sought to gain a better understanding about the genetic basis of BER with the expectation that these insights may lead to additional and potentially more cost-effective solutions to BER and related syndromes. In this study, we phenotyped a BER segregating population by first evaluating the methods to score the trait. Since it was expected that the disorder was quantitatively controlled and under environmental control, accurate phenotyping was deemed imperative to successfully map BER as in other complex traits (Bernardo 2020). Despite high correlations among the four traits, some F2 plants exhibited distinct patterns for BER Incidence and Severity 2, suggesting that these traits may be controlled by different loci. This conclusion was based on the observation that some plants carried fruits slightly affected by BER Severity 2, while exhibiting high BER Incidence values. Hence, these two traits were used to generate genomic pools for whole genome sequencing.
The QTL-seq approach has been employed to rapidly map QTL(s), and the results are often directly implemented in marker-assistant breeding (Clevenger et al. 2018; Das et al. 2015; Illa-Berenguer et al. 2015; Imerovski et al. 2019; Lu et al. 2014; Paudel et al. 2019a; Ramos et al. 2020; Wang et al. 2016). The power of QTL-seq relies on the population size, the number of SNPs between the parents and quantitative nature of the trait (Illa-Berenguer et al. 2015). In addition, the number of selected individuals in each bulk and sequencing depth also needs to be taken into consideration since these parameters significantly affect the power of QTL detection (Takagi et al. 2013). With a size of 192 plants and 778,685 SNPs that differentiate the parents, the QTL-seq approach led to the identification of BER3.2 and BER11.1 for the BER Incidence trait but missed the minor BER4.1. Therefore, traits controlled by many loci with a small effect, a higher population size (n ≥ 192) should be considered to capture the minor QTLs. In our study, additional markers that spanned the QTL-seq identified loci were mapped in the entire 17S28 F2 population, resulting in the confirmation of BER3.1 and BER11.1. The other QTL on ch1 and ch8 were false positives, which is not uncommon in QTL-seq experiments (Paudel et al. 2019b). Generally for QTL-Seq, the bulk size needs to be composed of at least 15% of the total F2 population to detect minor QTL that explain less than 10% of the percentage of total phenotypic variation explained (Takagi et al. 2013). In this study, in an effort to include only the most extreme phenotypes, the bulks were composed of 6–10% of the total F2 population. This may have resulted in less power to detect the minor QTL, especially for BER4.1 using the QTL-seq approach. BER11.1 was validated in the BC1 population, whereas BER3.2 and BER4.1 were not. The limited size of this population may have led to an underestimation of QTL numbers in this population (Beavis 1998; Melchinger et al. 1998; Vales et al. 2005). To further confirm BER3.2, BER4.1, and BER11.1, we created another F2 population (20S166) that was not segregating for any of the known fruit weight and shape genes. In this population, we confirmed BER4.1 and found an additional QTL on ch3 (BER3.1). Interestingly, BER3.1 may have been detected in the first F2 population as a minor QTL close to the major QTL BER3.2 (Fig. 3b).
We sought to refine and delineate BER3.2 and BER11.1, the QTLs that were first detected and found to explain more than one third of the total phenotypic variance. BER3.2 was narrowed down from 5.68 to 1.58 Mbp, flanked by 20EP1033 (SL4.0ch3 58,308,917) and 18EP730 (SL4.0ch3 59,891,210) markers. This region was comprised of 209 candidate genes (Supplementary Table 7) including the FW3.2/SlKLUH an ortholog of KLUH that regulates cell proliferation in developing organs in Arabidopsis (Anastasiou et al. 2007). It is known that increased fruit size and BER onset are strongly correlated despite the notion that certain large tomato varieties are resistant, whereas certain mid-sized elongated tomato varieties are susceptible to the disorder (Heuvelink and Körner 2001; Marcelis and Ho 1999). Moreover, to date, no conclusive association has been found between fruit weight genes and BER occurrence except Cell Size Regulator (FW11.3/CSR) that increases the fruit weight by increasing the cell size and results in high BER Incidence (Mu 2015). However, since FW3.2/SlKLUH was segregating in the population, we inferred that it was likely to be the gene underlying BER3.2. However, in addition to FW3.2/SlKLUH, other putative genes in this interval included Solyc03g113920, Solyc03g113940, Solyc03g113950, Solyc03g113960, Solyc03g113970, and Solyc03g113980 (Supplementary Table 7) encoding calmodulin binding proteins. These proteins may be involved in Ca2+ signaling and ROS scavenging, which play a role in maintaining cell membrane integrity by inhibiting the cell membrane lipid peroxidation (Dhindsa et al. 1981; Van Breusegem and Dat 2006). ROS scavenging pathways are downregulated under Ca2+-deficient conditions, but their activities are up-regulated via Ca2+/calmodulin signaling (Schmitz-Eiberger et al. 2002; Yang and Poovaiah 2002; Zeng et al. 2015). In addition, other candidates included Solyc03g114420, a gene encoding calmodulin and Solyc03g114110 and Solyc03g114450, genes related to Ca2+ sensing and transport (Supplementary Table 7). Hence, Ca2+-related genes may be considered good candidate genes for BER3.2. Furthermore, in the BER3.2 interval we also found genes encoding COBRA-like proteins (Solyc03g114880, Solyc03g114890, Solyc03g114900 and Solyc03g114910 (Supplementary Table 7) that are thought to be involved in secondary cell wall biosynthesis and fruit development. Mutations in the COBRA-Like genes (COBLs), such as cobl4 in Arabidopsis and brittle stalk 2 (bk2) in corn, affect cell wall thickness and cellulose content, especially in the sclerenchyma cells and vascular bundles (Brown et al. 2005; Ching et al. 2006). Strikingly, suppression of SlCOBRA-like gene in tomato leads to impaired cell wall integrity in developing fruit, coinciding with the first symptoms of BER (Cao et al. 2012). Finally, this interval included six expansion genes (Solyc03g115270, Solyc03g115300, Solyc03g115310, Solyc03g115320, Solyc03g115340, and Solyc03g115345) that may play an important role in cell wall modification and stress resistance in tomato (Lu et al. 2016; Minoia et al. 2016).
The fine mapping of BER11.1 led to the reduction of the interval from 3.99 to 1.13 Mbp flanked by 18EP1043 (SL4.0ch11 52,123,165) and 18EP697 (SL4.0ch11 53,250,673) markers. This region was comprised of 141 candidate genes including FASCIATED (fas), which can affect fruit weight by controlling locule number during flower development and is due to a regulatory mutation in SlCLV3 (Chu et al. 2019) (Supplementary Table 8). Even though fas segregated in the population, it is less likely a candidate for BER11.1 since the fruit shape allele is derived from the BER-resistant parent BGV007900. We also found two pectin methylesterase genes (PMEs) (Solyc11g070175 and Solyc11g070187). The role of PMEs in BER development has been studied previously, and suppression of PMEs decreases fruit susceptibility to BER (de Freitas et al. 2012a). Similar to BER3.2, the BER11.1 interval contained two genes (Solyc11g071740 and Solyc11g071750) encoding Ca2+-binding proteins. In addition to these candidates, other genes related to calcium (Solyc11g069580), cell division and expansion (Solyc11g069500, Solyc11g069570, and Solyc11g069720), and nutrient uptake (Solyc11g069735, Solyc11g069750 and Solyc11g069760) may be plausible candidates for BER11.1 (Supplementary Table 8).
Finally, we showed that further mapping of QTLs by evaluating only the recombinant plants resulted in excellent power to detect and fine map the loci. Other methods such as progeny testing, where 10 to 20 plants derived from a single recombinant parent are evaluated, work well also but relatively few recombinants are investigated at a time. The progeny testing often results in multiple years before a locus is fine mapped to a few candidates (Chakrabarti et al. 2013; Huang and van der Knaap 2011). Therefore, space and time limitations may be met by only analyzing the recombinants near and around the locus, especially in earlier generations. Furthermore, QTL mapping can tolerate some phenotypic outliers, whereas one outlier in a progeny test can obscure the proper interpretation of the data.
To conclude, we employed the QTL-seq and linkage-based QTL-mapping approaches to genetically map four loci in three populations that were associated with BER in tomato. BER3.2 and BER11.1 QTL were further mapped to intervals of less than 1.6 Mb, whereas BER3.1 and BER4.1 await further fine mapping. The eventual cloning of the underlying genes will facilitate marker assistant breeding not only in tomato but also offer knowledge to other breeders working on vegetables and fruits that suffer from BER.
References
Adams P, Ho LC (1993) Effects of environment on the uptake and distribution of calcium in tomato and on the incidence of blossom-end rot. Plant Soil 154(1):127–132. https://doi.org/10.1007/BF00011081
Aktas H, Karni L, Aloni B, Bar-Tal A (2003) Physiological and biochemical mechanisms leading to blossom-end rot in greenhouse-grown peppers, irrigated with saline solution. Acta Hortic 609:81–88. https://doi.org/10.17660/ActaHortic.2003.609.9
Anastasiou E, Kenz S, Gerstung M, MacLean D, Timmer J, Fleck C, Lenhard M (2007) Control of plant organ size by KLUH/CYP78A5-dependent intercellular signaling. Dev Cell 13(6):843–856. https://doi.org/10.1016/j.devcel.2007.10.001
Andrews S (2010) FastQC A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/. Accessed 14 Feb 2021
Barickman TC, Kopsell DA, Sams CE (2014) Foliar applications of abscisic acid decrease the incidence of blossom-end rot in tomato fruit. Sci Hortic 179:356–362. https://doi.org/10.1016/j.scienta.2014.10.004
Beavis WD (1998) QTL analyses: power, precision, and accuracy. In: Paterson AH (ed) Molecular dissection of complex traits, 1st edn. CRC Press, Boca Raton, pp 145–162
Bernardo R (2020) Reinventing quantitative genetics for plant breeding: something old, something new, something borrowed, something BLUE. Heredity 125(6):375–385. https://doi.org/10.1038/s41437-020-0312-1
Broman KW, Wu H, Sen S, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19(7):889–890. https://doi.org/10.1093/bioinformatics/btg112
Brown DM, Zeef LAH, Ellis J, Goodacre R, Turner SR (2005) Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics. Plant Cell 17(8):2281–2295. https://doi.org/10.1105/tpc.105.031542
Cao Y, Tang X, Giovannoni J, Xiao F, Liu Y (2012) Functional characterization of a tomato COBRA-like gene functioning in fruit development and ripening. BMC Plant Biol 12(1):211. https://doi.org/10.1186/1471-2229-12-211
Chakrabarti M, Zhang N, Sauvage C, Munos S et al (2013) A cytochrome P450 regulates a domestication trait in cultivated tomato. Proc Natl Acad Sci USA 110(42):17125–17130. https://doi.org/10.1073/pnas.1307313110
Ching A, Dhugga KS, Appenzeller L, Meeley R, Bourett TM, Howard RJ, Rafalski A (2006) Brittle stalk 2 encodes a putative glycosylphosphatidylinositol-anchored protein that affects mechanical strength of maize tissues by altering the composition and structure of secondary cell walls. Planta 224(5):1174–1184. https://doi.org/10.1007/s00425-006-0299-8
Chu Y-H, Jang J-C, Huang Z, van der Knaap E (2019) Tomato locule number and fruit size controlled by natural alleles of lc and fas. Plant Direct 3(7):e00142. https://doi.org/10.1002/pld3.142
Clapham DE (2007) Calcium signaling. Cell 131(6):1047–1058. https://doi.org/10.1016/j.cell.2007.11.028
Clarkson DT, Mansfield TA, Davies WJ, Leigh RA (1993) Roots and the delivery of solutes to the xylem. Philos Trans R Soc Lond, B, Biol Sci 341(1295):5–17. https://doi.org/10.1098/rstb.1993.0086
Clevenger J, Chu Y, Chavarro C, Botton S et al (2018) Mapping late leaf spot resistance in peanut (Arachis hypogaea) using QTL-seq reveals markers for marker-assisted selection. Front Plant Sci 9:83. https://doi.org/10.3389/fpls.2018.00083
Das S, Upadhyaya HD, Bajaj D, Kujur A et al (2015) Deploying QTL-seq for rapid delineation of a potential candidate gene underlying major trait-associated QTL in chickpea. DNA Res 22(3):193–203. https://doi.org/10.1093/dnares/dsv004
de Freitas ST, Handa AK, Wu Q, Park S, Mitcham EJ (2012a) Role of pectin methylesterases in cellular calcium distribution and blossom-end rot development in tomato fruit. Plant J 71(5):824–835. https://doi.org/10.1111/j.1365-313X.2012.05034.x
de Freitas ST, Jiang C-Z, Mitcham EJ (2012b) Mechanisms involved in calcium deficiency development in tomato fruit in response to gibberellins. J Plant Growth Regul 31(2):221–234. https://doi.org/10.1007/s00344-011-9233-9
de Freitas ST, McElrone AJ, Shackel KA, Mitcham EJ (2014) Calcium partitioning and allocation and blossom-end rot development in tomato plants in response to whole-plant and fruit-specific abscisic acid treatments. J Exp Bot 65(1):235–247. https://doi.org/10.1093/jxb/ert364
de Freitas ST, Padda M, Wu Q, Park S, Mitcham EJ (2011) Dynamic alternations in cellular and molecular components during blossom-end rot development in tomatoes expressing sCAX1, a constitutively active Ca2+/H+ antiporter from Arabidopsis. Plant Physiol 156(2):844–855. https://doi.org/10.1104/pp.111.175208
Dhindsa RS, Plumb-Dhindsa P, Thorpe TA (1981) Leaf senescence: correlated with increased levels of membrane permeability and lipid peroxidation, and decreased levels of superoxide dismutase and catalase. J Exp Bot 32(1):93–101. https://doi.org/10.1093/jxb/32.1.93
Dodd AN, Kudla J, Sanders D (2010) The language of calcium signaling. Annu Rev Plant Biol 61:593–620. https://doi.org/10.1146/annurev-arplant-070109-104628
FAOSTAT (2020) Global tomato production. http://www.fao.org/faostat/en/#data/QC. Accessed 01 Sept 2020
Hagassou D, Francia E, Ronga D, Buti M (2019) Blossom end-rot in tomato (Solanum lycopersicum L.): a multi-disciplinary overview of inducing factors and control strategies. Sci Hortic 249:49–58. https://doi.org/10.1016/j.scienta.2019.01.042
Heuvelink E, Körner O (2001) Parthenocarpic fruit growth reduces yield fluctuation and blossom-end rot in sweet pepper. Ann Bot 88(1):69–74. https://doi.org/10.1006/anbo.2001.1427
Ho LC, White PJ (2005) A cellular hypothesis for the induction of blossom-end rot in tomato fruit. Ann Bot 95(4):571–581. https://doi.org/10.1093/aob/mci065
Huang Z, van der Knaap E (2011) Tomato fruit weight 11.3 maps close to fasciated on the bottom of chromosome 11. Theor Appl Genet 123(3):465–474. https://doi.org/10.1007/s00122-011-1599-3
Ikeda H, Kanayama Y (2015) Blossom-end rot in fruit vegetables. In: Kanayama Y, Kochetov A (eds) Abiotic stress biology in horticultural plants. Springer Japan, Tokyo, pp 117–126
Illa-Berenguer E, Van Houten J, Huang Z, van der Knaap E (2015) Rapid and reliable identification of tomato fruit weight and locule number loci by QTL-seq. Theor Appl Genet 128(7):1329–1342. https://doi.org/10.1007/s00122-015-2509-x
Imerovski I, Dedić B, Cvejić S, Miladinović D et al (2019) BSA-seq mapping reveals major QTL for broomrape resistance in four sunflower lines. Mol Breed 39(3):1–15. https://doi.org/10.1007/s11032-019-0948-9
Kearsey MJ, Pooni HS (1996) The genetical analysis of quantitative traits. Chapman and Hall, London
Kosambi DD (1943) The estimation of map distance from recombination values. Ann Eugen 12(1):172–175. https://doi.org/10.1111/j.1469-1809.1943.tb02321.x
Kudla J, Batistič O, Hashimoto K (2010) Calcium signals: the lead currency of plant information processing. Plant Cell 22(3):541–563. https://doi.org/10.1105/tpc.109.072686
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A, Fennell T et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Lu H, Lin T, Klein J, Wang S et al (2014) QTL-seq identifies an early flowering QTL located near Flowering Locus T in cucumber. Theor Appl Genet 127(7):1491–1499. https://doi.org/10.1007/s00122-014-2313-z
Lu Y, Liu L, Wang X, Han Z, Ouyang B, Zhang J, Li H (2016) Genome-wide identification and expression analysis of the expansin gene family in tomato. Mol Genet Genomics 291(2):597–608. https://doi.org/10.1007/s00438-015-1133-4
Mansfeld BN, Grumet R (2018) QTLseqr: an R package for bulk segregant analysis with next-generation sequencing. The Plant Genome 11(2):180006. https://doi.org/10.3835/plantgenome2018.01.0006
Marcelis LFM, Ho LC (1999) Blossom-end rot in relation to growth rate and calcium content in fruits of sweet pepper (Capsicum annuum L.). J Exp Bot 50(332):357–363. https://doi.org/10.1093/jxb/50.332.357
McKenna A, Hanna M, Banks E, Sivachenko A et al (2010) The Genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. https://doi.org/10.1101/gr.107524.110
Melchinger AE, Utz HF, Schön CC (1998) Quantitative trait locus (QTL) mapping using different testers and independent population samples in maize reveals low power of QTL detection and large bias in estimates of QTL effects. Genetics 149(1):383–403
Meng L, Li H, Zhang L, Wang J (2015) QTL IciMapping: Integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J 3(3):269–283. https://doi.org/10.1016/j.cj.2015.01.001
Minoia S, Boualem A, Marcel F, Troadec C et al (2016) Induced mutations in tomato SlExp1 alter cell wall metabolism and delay fruit softening. Plant Sci 242:195–202. https://doi.org/10.1016/j.plantsci.2015.07.001
Mu Q (2015) The cloning and cellular basis of a novel tomato fruit weight gene: Cell Size Regulator (FW11.3/CSR). In: Dissertation, The Ohio State University
Ng CK-Y, Mcainsh MR, Gray JE, Hunt L, Leckie CP, Mills L, Hetherington AM (2001) Calcium-based signalling systems in guard cells. New Phytol 151(1):109–120. https://doi.org/10.1046/j.1469-8137.2001.00152.x
Paudel L, Clevenger J, McGregor C (2019a) Chromosomal locations and interactions of four loci associated with seed coat color in watermelon. Front Plant Sci 10:788. https://doi.org/10.3389/fpls.2019.00788
Paudel L, Clevenger J, McGregor C (2019b) Refining of the egusi locus in watermelon using KASP assays. Sci Hortic 257:108665. https://doi.org/10.1016/j.scienta.2019.108665
Penella C, Calatayud A (2018) Pepper crop under climate change: grafting as an environmental friendly strategy. In: Shanker A (ed) Climate resilient agriculture: strategies and perspectives, 1st edn. IntechOpen, London, pp 129–155
R Core Team (2019) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria
Rached M, Pierre B, Yves G, Matsukura C, Ariizumi T, Ezura H, Fukuda N (2018) Differences in blossom-end rot resistance in tomato cultivars is associated with total ascorbate rather than calcium concentration in the distal end part of fruits per se. Hort J 87(3):372–381. https://doi.org/10.2503/hortj.OKD-150
Raleigh SM, Chucka JA (1944) Effect of nutrient ratio and concentration on growth and composition of tomato plants and on the occurrence of blossom-end rot of the fruit. Plant Physiol 19(4):671–678. https://doi.org/10.1104/pp.19.4.671
Ramos A (2018) Mining QTL for fruit weight quality traits in uncharacterized tomato germplasm. In: Dissertation, University of Georgia
Ramos A, Fu Y, Michael V, Meru G (2020) QTL-seq for identification of loci associated with resistance to Phytophthora crown rot in squash. Sci Rep 10:5326. https://doi.org/10.1038/s41598-020-62228-z
Razifard H, Ramos A, Della Valle AL, Bodary C et al (2020) Genomic evidence for complex domestication history of the cultivated tomato in Latin America. Mol Biol Evol 37(4):1118–1132. https://doi.org/10.1093/molbev/msz297
Rodríguez GR, Muños S, Anderson C, Sim S-C et al (2011) Distribution of SUN, OVATE, LC, and FAS in the tomato germplasm and the relationship to fruit shape diversity. Plant Physiol 156(1):275–285. https://doi.org/10.1104/pp.110.167577
SAS Institute Inc. (2017) Using JMP®Version 13.2.0. SAS Institute Inc., Cary, NC, USA, 1989–2019
Sato S, Tabata S, Hirakawa H, Asamizu E et al (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485(7400):635–641. https://doi.org/10.1038/nature11119
Schmitz-Eiberger M, Haefs R, Noga G (2002) Calcium deficiency—influence on the antioxidative defense system in tomato plants. J Plant Physiol 159(7):733–742. https://doi.org/10.1078/0176-1617-0621
Takagi H, Abe A, Yoshida K, Kosugi S et al (2013) QTL-seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J 74(1):174–183. https://doi.org/10.1111/tpj.12105
Tanksley SD (1993) Mapping polygenes. Annu Rev Genet 27:205–233. https://doi.org/10.1146/annurev.ge.27.120193.001225
Taylor MD, Locascio SJ (2004) Blossom-end rot: A calcium deficiency. J Plant Nutr 27(1):123–139. https://doi.org/10.1081/PLN-120027551
Thor K (2019) Calcium-nutrient and messenger. Front Plant Sci 10:440. https://doi.org/10.3389/fpls.2019.00440
Uozumi A, Ikeda H, Hiraga M, Kanno H et al (2012) Tolerance to salt stress and blossom-end rot in an introgression line, IL8-3, of tomato. Sci Hort 138:1–6. https://doi.org/10.1016/j.scienta.2012.01.036
Vales MI, Schön CC, Capettini F, Chen XM et al (2005) Effect of population size on the estimation of QTL: a test using resistance to barley stripe rust. Theor Appl Genet 111:1260–1270. https://doi.org/10.1007/s00122-005-0043-y
Van Breusegem F, Dat JF (2006) Reactive oxygen species in plant cell death. Plant Physiol 141(2):384–390. https://doi.org/10.1104/pp.106.078295
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R et al. (2013) From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics 43(1110):11.10.11–11.10.33. https://doi.org/10.1002/0471250953.bi1110s43
Wang H, Cheng H, Wang W, Liu J et al (2016) Identification of BnaYUCCA6 as a candidate gene for branch angle in Brassica napus by QTL-seq. Sci Rep 6:38493. https://doi.org/10.1038/srep38493
Wu S, Zhang B, Keyhaninejad N, Rodríguez GR et al (2018) A common genetic mechanism underlies morphological diversity in fruits and other plant organs. Nat Commun 9(1):4734. https://doi.org/10.1038/s41467-018-07216-8
Yang T, Poovaiah BW (2002) Hydrogen peroxide homeostasis: activation of plant catalase by calcium/calmodulin. Proc Natl Acad Sci 99(6):4097–4102. https://doi.org/10.1073/pnas.052564899
Zeng H, Xu L, Singh A, Wang H, Du L, Poovaiah BW (2015) Involvement of calmodulin and calmodulin-like proteins in plant responses to abiotic stresses. Front Plant Sci 6:600–600. https://doi.org/10.3389/fpls.2015.00600
Zhang N, Brewer MT, van der Knaap E (2012) Fine mapping of fw3.2 controlling fruit weight in tomato. Theor Appl Genet 125(2):273–284. https://doi.org/10.1007/s00122-012-1832-8
Acknowledgements
The authors greatly appreciate the members of Esther van der Knaap laboratory for their help during the study and analysis and to Katherine Hardigree and Pam Lewis for plant care in the greenhouse. YT is the recipient of the graduate scholarship sponsored by the Ministry of National Education of Turkey and the John Ingle Innovation in Plant Breeding Award .
Funding
This research was supported by United States Department of Agriculture-National Institute of Food and Agriculture (USDA-NIFA) under grant Agreement No. [2020–67013-30912] to SN and EvdK, and the John Ingle Award to YT.
Author information
Authors and Affiliations
Contributions
YT created populations, carried out the experiments, performed SNP calling and QTL analysis, developed KASP markers, and wrote the manuscript; MS conducted the bioinformatic analysis for the QTL-Seq; EIB created the first mapping population (17S28 F2 population), extracted DNA for the QTL-seq analysis, genotyped the parents used in the study; EvdK and SN conceived the idea, initiated the project, supervised the research analysis, prepared the sequencing libraries for the QTL-seq, and edited the manuscript; MS and EIB read and reviewed the manuscript. All authors contributed and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The experiments carried out fulfill the laws of the USA.
Additional information
Communicated by Hong-Qing Ling.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Topcu, Y., Sapkota, M., Illa-Berenguer, E. et al. Identification of blossom-end rot loci using joint QTL-seq and linkage-based QTL mapping in tomato. Theor Appl Genet 134, 2931–2945 (2021). https://doi.org/10.1007/s00122-021-03869-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-021-03869-0