Abstract
Key message
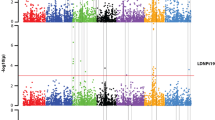
A total of 449 barley accessions were phenotyped for Pyrenophora teres f. teres resistance at three locations and in greenhouse trials. Genome-wide association studies identified 254 marker–trait associations corresponding to 15 QTLs.
Abstract
Net form of net blotch is one of the most important diseases of barley and is present in all barley growing regions. Under optimal conditions, it causes high yield losses of 10–40% and reduces grain quality. The most cost-effective and environmentally friendly way to prevent losses is growing resistant cultivars, and markers linked to effective resistance factors can accelerate the breeding process. Here, 449 barley accessions expressing different levels of resistance comprising landraces and commercial cultivars from the centres of diversity were selected. The set was phenotyped for seedling resistance to three isolates in controlled-environment tests and for adult plant resistance at three field locations (Belarus, Germany and Australia) and genotyped with the 50 k iSelect chip. Genome-wide association studies using 33,818 markers and a compressed mixed linear model to account for population structure and kinship revealed 254 significant marker–trait associations corresponding to 15 distinct QTL regions. Four of these regions were new QTL that were not described in previous studies, while a total of seven regions influenced resistance in both seedlings and adult plants.






Similar content being viewed by others
References
Abu Qamar M et al (2008) A region of barley chromosome 6H harbors multiple major genes associated with net type net blotch resistance. Theor Appl Genet 117:1261–1270. https://doi.org/10.1007/s00122-008-0860-x
Afanasenko O (1995) Characteristics of resistance of barley accessions to different Pyrenophora teres populations. Mycol Phytopathol 29:27–32
Afanasenko O, Makarova I, Zubkovich A (1999) The number of genes controlling resistance to Pyrenophora teres Drechs. strains in barley. Russ J Genet C/C Genet 35:274–283
Afanasenko O, Jalli M, Pinnschmidt H, Filatova O, Platz G (2009) Development of an international standard set of barley differential genotypes for Pyrenophora teres f. teres. Plant Pathol 58:665–676
Afanasenko O et al (2015) Mapping of the loci controlling the resistance to Pyrenophora teres f. teres and Cochliobolus sativus in two double haploid barley populations. Russ J Genet Appl Res 5:242–253
Afgan E et al (2016) The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Res 44:W3–W10. https://doi.org/10.1093/nar/gkw343
Afonin A, Greene S, Dzyubenko N, Frolov A (2008) Interactive Agricultural ecological Atlas of Russia and Neighboring Countries. Economic Plants and their Diseases, Pests and Weeds. http://www.agroatlas.ru/. Accessed 26 Mar 2019
Amezrou R et al (2018) Genome-wide association studies of net form of net blotch resistance at seedling and adult plant stages in spring barley collection. Mol Breed 38:1–14. https://doi.org/10.1007/s11032-018-0813-2
Anisimova A, Novikova L, Novakazi F, Kopahnke D, Zubkovich A, Afanasenko O (2017) Polymorphism on virulence and specificity of microevolution processes in populations of causal agent of barley net blotch Pyrenophora teres f teres. Mikol I Fitopatol 51:229–240
Bayer MM et al (2017) Development and evaluation of a barley 50 k iSelect SNP. Array Front Plant Sci 8:1792. https://doi.org/10.3389/fpls.2017.01792
Bellucci A et al (2017) Genome-wide association mapping in winter barley for grain yield and culm cell wall polymer content using the high-throughput CoMPP technique. PLoS ONE 12:e0173313. https://doi.org/10.1371/journal.pone.0173313
Bengtsson T, Manninen O, Jahoor A, Orabi J (2017) Genetic diversity, population structure and linkage disequilibrium in Nordic spring barley (Hordeum vulgare L. subsp. vulgare). Genet Resour Crop Evolut 64:2021–2033. https://doi.org/10.1007/s10722-017-0493-5
Berger GL et al (2013) Marker-trait associations in Virginia Tech winter barley identified using genome-wide mapping. Theor Appl Genet 126:693–710
Brandl F, Hoffmann G (1991) Differentiation of physiological races of Drechslera teres (Sacc.) Shoem., pathogen net blotch of barley. Zeitschrift fuer Pflanzenkrankheiten und Pflanzenschutz
Burlakoti RR et al (2017) Genome-Wide Association Study of spot form of net blotch resistance in the upper midwest barley breeding programs. Phytopathology 107:100–108. https://doi.org/10.1094/PHYTO-03-16-0136-R
Burleigh J, Tajani M, Seck M (1988) Effects of Pyrenophora teres and weeds on barley yield and yield components. Phytopathology 78:295–299
Cakir M et al (2003) Mapping and validation of the genes for resistance to Pyrenophora teres f. teres in barley (Hordeum vulgare L.). Aust J Agric Res 54:1369–1377
Cakir M et al (2011) Genetic mapping and QTL analysis of disease resistance traits in the barley population Baudin × AC Metcalfe. Crop Pasture Sci 62:152–161
Campoy JA et al (2016) Genetic diversity, linkage disequilibrium, population structure and construction of a core collection of Prunus avium L. landraces and bred cultivars. BMC Plant Biol 16:49. https://doi.org/10.1186/s12870-016-0712-9
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ (2015) Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4:7. https://doi.org/10.1186/s13742-015-0047-8
Douglas G, Gordon I (1985) Quantitative genetics of net blotch resistance in barley. N Z J Agric Res 28:157–164
Earl DA, vonHoldt BM (2012) Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Flint-Garcia SA, Thornsberry JM, Buckler ESt (2003) Structure of linkage disequilibrium in plants. Annu Rev Plant Biol 54:357–374. https://doi.org/10.1146/annurev.arplant.54.031902.134907
Friesen TL, Faris JD, Lai Z, Steffenson BJ (2006) Identification and chromosomal location of major genes for resistance to Pyrenophora teres in a doubled-haploid barley population. Genome 49:855–859. https://doi.org/10.1139/g06-024
Graner A, Foroughi-Wehr B, Tekauz A (1996) RFLP mapping of a gene in barley conferring resistance to net blotch (Pyrenophora teres). Euphytica 91:229–234
Grewal TS, Rossnagel BG, Pozniak CJ, Scoles GJ (2008) Mapping quantitative trait loci associated with barley net blotch resistance. Theor Appl Genet 116:529–539. https://doi.org/10.1007/s00122-007-0688-9
Grewal TS, Rossnagel BG, Scoles GJ (2012) Mapping quantitative trait loci associated with spot blotch and net blotch resistance in a doubled-haploid barley population. Mol Breed 30:267–279
Gupta S et al (2004) Gene distribution and SSR markers linked with net type net blotch resistance in barley. In: 9th International Barley Genetics Symposium, 20–26 June. Czech Republic, Brno, pp 668–673
Gupta S et al (2010) Quantitative trait loci and epistatic interactions in barley conferring resistance to net type net blotch (Pyrenophora teres f teres) isolates. Plant Breed 129:362–368
Gupta S, Li C, Loughman R, Cakir M, Westcott S, Lance R (2011) Identifying genetic complexity of 6H locus in barley conferring resistance to Pyrenophora teres f teres. Plant Breed 130:423–429. https://doi.org/10.1111/j.1439-0523.2011.01854.x
Gurung S et al (2011) Identification of novel genomic regions associated with resistance to Pyrenophora tritici-repentis races 1 and 5 in spring wheat landraces using association analysis. Theor Appl Genet 123:1029–1041. https://doi.org/10.1007/s00122-011-1645-1
Gyawali S, Otte ML, Chao S, Jilal A, Jacob DL, Amezrou R, Verma RPS (2017) Genome wide association studies (GWAS) of element contents in grain with a special focus on zinc and iron in a world collection of barley (Hordeum vulgare L.). J Cereal Sci 77:266–274. https://doi.org/10.1016/j.jcs.2017.08.019
Islamovic E, Bregitzer P, Friesen TL (2017) Barley 4H QTL confers NFNB resistance to a global set of P. teres f. teres isolates. Mol Breed 37:29. https://doi.org/10.1007/s11032-017-0621-0
Kangas A, Jalli M, Kedonperä A, Laine A, Niskanen M, Salo Y, Vuorinen M, Jauhiainen L, Ramstadius E 2005 Viljalajikkeiden herkkyys tautitartunnoille virallisissa lajikekokeissa 1998-2005. Disease susceptibility of cereals in Finnish official variety trials. [English summary and titles] Agrifood Res Rep, MTT:n selvityksiä 96: 33
Khan T (1982) Occurence and pathogenicity of Drechslera teres isolates causing spot-type symptoms on barley in Western Australia. Plant Dis 66:423–425
Koladia V, Faris J, Richards J, Brueggeman R, Chao S, Friesen T (2017) Genetic analysis of net form net blotch resistance in barley lines CIho 5791 and Tifang against a global collection of P. teres f teres isolates. Theor Appl Genet 130:163–173
König J, Perovic D, Kopahnke D, Ordon F (2013) Development of an efficient method for assessing resistance to the net type of net blotch (Pyrenophora teres f teres) in winter barley and mapping of quantitative trait loci for resistance. Mol Breed 32:641–650. https://doi.org/10.1007/s11032-013-9897-x
König J, Perovic D, Kopahnke D, Ordon F, Léon J (2014) Mapping seedling resistance to net form of net blotch (Pyrenophora teres f teres) in barley using detached leaf assay. Plant Breed 133:356–365. https://doi.org/10.1111/pbr.12147
Lex J, Ahlemeyer J, Friedt W, Ordon FJ (2014) Genome-wide association studies of agronomic and quality traits in a set of German winter barley (Hordeum vulgare L.) cultivars using Diversity Arrays Technology (DArT). J Appl Genet 55:295–305
Lightfoot DJ, Able AJ (2010) Growth of Pyrenophora teres in planta during barley net blotch disease. Australas Plant Pathol 39:499–507
Lipka AE et al (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399
Liu Z, Ellwood SR, Oliver RP, Friesen TL (2011) Pyrenophora teres: profile of an increasingly damaging barley pathogen. Mol Plant Pathol 12:1–19. https://doi.org/10.1111/j.1364-3703.2010.00649.x
Liu Z, Holmes DJ, Faris JD, Chao S, Brueggeman RS, Edwards MC, Friesen TL (2015) Necrotrophic effector-triggered susceptibility (NETS) underlies the barley-Pyrenophora teres f. teres interaction specific to chromosome 6H. Mol Plant Pathol 16:188–200. https://doi.org/10.1111/mpp.12172
Ma Z, Lapitan NL, Steffenson B (2004) QTL mapping of net blotch resistance genes in a doubled-haploid population of six-rowed barley. Euphytica 137:291–296
Manninen OM, Jalli M, Kalendar R, Schulman A, Afanasenko O, Robinson J (2006) Mapping of major spot-type and net-type net-blotch resistance genes in the Ethiopian barley line CI 9819. Genome 49:1564–1571. https://doi.org/10.1139/g06-119
Martin A, Platz GJ, de Klerk D, Fowler RA, Smit F, Potgieter FG, Prins R (2018) Identification and mapping of net form of net blotch resistance in South African barley. Mol Breed 38:53. https://doi.org/10.1007/s11032-018-0814-1
Mascher M et al (2017) A chromosome conformation capture ordered sequence of the barley genome. Nature 544:427–433. https://doi.org/10.1038/nature22043
Massman J et al (2010) Genome-wide association mapping of Fusarium head blight resistance in contemporary barley breeding germplasm. Mol Breed 27:439–454. https://doi.org/10.1007/s11032-010-9442-0
Mathre D (1997) Compendium of Barley Diseases. The American Phytopathological Society, St. Paul, MN
Maurer A et al (2015) Modelling the genetic architecture of flowering time control in barley through nested association mapping. BMC Genom 16:290
Mitterbauer E et al (2017) Growth response of 98 barley (Hordeum vulgare L.) genotypes to elevated CO2 and identification of related quantitative trait loci using genome-wide association studies. Plant Breed 136:483–497. https://doi.org/10.1111/pbr.12501
Mohammadi M et al (2015) A genome-wide association study of malting quality across eight US barley breeding programs. Theor Appl Genet 128:705–721. https://doi.org/10.1007/s00122-015-2465-5
Moll E, Flath K, Tessenow I (2010) Assessment of resistance in cereal cultivars Design and analysis of experiments using the SAS-application RESI 2 Berichte aus dem Julius Kühn-Institut: 154
Muqaddasi QH, Reif JC, Li Z, Basnet BR, Dreisigacker S, Röder MS (2017) Genome-wide association mapping and genome-wide prediction of anther extrusion in CIMMYT spring wheat. Euphytica 213:73. https://doi.org/10.1007/s10681-017-1863-y
Murray GM, Brennan JP (2009) The current and potential costs from diseases of barley in Australia. Grains Research and Development Corporation, Barton
Nordborg M et al (2002) The extent of linkage disequilibrium in Arabidopsis thaliana. Nat Genet 30:190–193. https://doi.org/10.1038/ng813
O’Boyle P et al (2014) Mapping net blotch resistance in ‘Nomini’and CIho 2291 barley. Crop Sci 54:2596–2602
Poland JA, Brown PJ, Sorrells ME, Jannink JL (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS ONE 7:e32253. https://doi.org/10.1371/journal.pone.0032253
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rafalski A, Morgante M (2004) Corn and humans: recombination and linkage disequilibrium in two genomes of similar size. Trends Genetics 20:103–111
Raman H, Platz G, Chalmers K, Raman R, Read B, Barr A, Moody D (2003) Mapping of genomic regions associated with net form of netblotch resistance in barley. Aust J Agric Res 54:1359–1367
Reif JC, Melchinger AE, Frisch M (2005) Genetical and mathematical properties of similarity and dissimilarity coefficients applied in plant breeding and seed bank management. Crop Sci 45:1–7
Remington DL et al (2001) Structure of linkage disequilibrium and phenotypic associations in the maize genome. Proc Natl Acad Sci 98:11479–11484
Richards J, Chao S, Friesen T, Brueggeman R (2016) Fine mapping of the barley chromosome 6H net form net blotch susceptibility locus. G3 (Bethesda) 6:1809–1818. https://doi.org/10.1534/g3.116.028902
Richards JK, Friesen TL, Brueggeman RS (2017) Association mapping utilizing diverse barley lines reveals net form net blotch seedling resistance/susceptibility loci. Theor Appl Genet 130:915–927. https://doi.org/10.1007/s00122-017-2860-1
Richter K, Schondelmaier J, Jung C (1998) Mapping of quantitative trait loci affecting Drechslera teres resistance in barley with molecular markers. Theor Appl Genet 97:1225–1234
Robinson J, Jalli M (1997) Quantitative resistance to Pyrenophora teres in six Nordic spring barley accessions. Euphytica 94:201–208
Rode J, Ahlemeyer J, Friedt W, Ordon F (2011) Identification of marker-trait associations in the German winter barley breeding gene pool (Hordeum vulgare L.). Mol Breed 30:831–843. https://doi.org/10.1007/s11032-011-9667-6
Saari E, Prescott J (1975) Scale for appraising the foliar intensity of wheat diseases. Plant Disease Reporter 59:377–380
Sannemann W, Huang BE, Mathew B, Léon J (2015) Multi-parent advanced generation inter-cross in barley: high-resolution quantitative trait locus mapping for flowering time as a proof of concept. Mol Breed 35:86
Serenius M (2006) Population structure of Pyrenophora teres, the causal agent of net blotch of barley (Doctoral Dissertation). Agrifood Research Reports 78:60
Shjerve RA, Faris JD, Brueggeman RS, Yan C, Zhu Y, Koladia V, Friesen TL (2014) Evaluation of a Pyrenophora teres f teres mapping population reveals multiple independent interactions with a region of barley chromosome 6H. Fungal Genet Biol 70:104–112. https://doi.org/10.1016/j.fgb.2014.07.012
Silvar C et al (2010) Screening the Spanish barley core collection for disease resistance. Plant Breed 129:45–52
Simmonds J et al (2014) Identification and independent validation of a stable yield and thousand grain weight QTL on chromosome 6A of hexaploid wheat (Triticum aestivum L.). BMC Plant Biol 14:191
Smedegård-Petersen V (1971) Pyrenophora teres f. maculata f. nov. and Pyrenophora teres f. teres on barley in Denmark. Yearb R Vet Agric Univ (Copenhagen) 1971:124–144
Smedegård-Petersen V (1976) Pathogenesis and genetics of net-spot blotch and leaf stripe of barley caused by Pyrenophora teres and Pyrenophora graminea (Doctoral Dissertation) Copenhagen, p 176
Steffenson BJ, Webster R (1992) Quantitative resistance to Pyrenophora teres f. teres in barley. Phytopathology 82:407–411
Steffenson B, Hayes P, Kleinhofs A (1996) Genetics of seedling and adult plant resistance to net blotch (Pyrenophora teres f. teres) and spot blotch (Cochliobolus sativus) in barley. Theor Appl Genet 92:552–558
Stein N, Herren G, Keller B (2001) A new DNA extraction method for high-throughput marker analysis in a large-genome species such as Triticum aestivum. Plant Breed 120:354–356
Storey JD, Tibshirani R (2003) Statistical significance for genomewide studies. Proc Natl Acad Sci 100:9440–9445
Tamang P, Neupane A, Mamidi S, Friesen T, Brueggeman R (2015) Association mapping of seedling resistance to spot form net blotch in a worldwide collection of barley. Phytopathology 105:500–508. https://doi.org/10.1094/PHYTO-04-14-0106-R
Tekauz A (1985) A numerical scale to classify reactions of barley to Pyrenophora teres. Can J Plant Path 7:181–183
Tekauz A (1990) Characterization and distribution of pathogenic variation in Pyrenophora teres f. teres and P. teres f. maculata from western Canada. Can J Plant Pathol 12:141–148
Trofimovskaya AY, Afanasenko O, Levitin MM (1983) Sources of barley resistance to the causal agent of net blotch (Drechslera teres). Rep Acad Agricu Sci 3:19–21
Vatter T, Maurer A, Kopahnke D, Perovic D, Ordon F, Pillen K (2017) A nested association mapping population identifies multiple small effect QTL conferring resistance against net blotch (Pyrenophora teres f. teres) in wild barley. PLoS ONE 12:e0186803. https://doi.org/10.1371/journal.pone.0186803
Wallwork H, Butt M, Capio E (2016) Pathogen diversity and screening for minor gene resistance to Pyrenophora teres f. teres in barley and its use for plant breeding. Australas Plant Pathol 45:527–531
Wang H, Smith KP, Combs E, Blake T, Horsley RD, Muehlbauer GJ (2012) Effect of population size and unbalanced data sets on QTL detection using genome-wide association mapping in barley breeding germplasm. Theor Appl Genet 124:111–124. https://doi.org/10.1007/s00122-011-1691-8
Wehner GG, Balko CC, Enders MM, Humbeck KK, Ordon FF (2015) Identification of genomic regions involved in tolerance to drought stress and drought stress induced leaf senescence in juvenile barley. BMC Plant Biol 15:125
Wonneberger R, Ficke A, Lillemo M (2017a) Identification of quantitative trait loci associated with resistance to net form net blotch in a collection of Nordic barley germplasm. Theor Appl Genet 130:2025–2043. https://doi.org/10.1007/s00122-017-2940-2
Wonneberger R, Ficke A, Lillemo M (2017b) Mapping of quantitative trait loci associated with resistance to net form net blotch (Pyrenophora teres f teres) in a doubled haploid Norwegian barley population. PLoS ONE 12:e0175773. https://doi.org/10.1371/journal.pone.0175773
Yan J, Shah T, Warburton ML, Buckler ES, McMullen MD, Crouch JJ (2009) Genetic characterization and linkage disequilibrium estimation of a global maize collection using SNP markers. PLoS ONE 4:e8451
Zhang Z et al (2010) Mixed linear model approach adapted for genome-wide association studies. Nat Genet 42:355
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome J 1:5–20. https://doi.org/10.3835/plantgenome2008.02.0089
Acknowledgements
This research was supported by the German Research Society (DFG) (OR 72/11-1) and the Russian Foundation for Basic Research (RFBR) (No 15-54-12365 NNIO_a).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Kevin Smith.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Novakazi, F., Afanasenko, O., Anisimova, A. et al. Genetic analysis of a worldwide barley collection for resistance to net form of net blotch disease (Pyrenophora teres f. teres). Theor Appl Genet 132, 2633–2650 (2019). https://doi.org/10.1007/s00122-019-03378-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-019-03378-1