Abstract
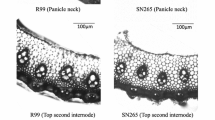
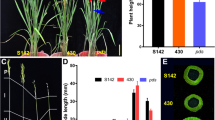
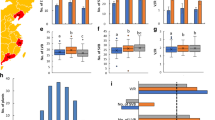
The quantitative trait locus controlling the number of primary rachis branches (PRBs) in rice was identified using backcrossed inbred lines of Sasanishiki/Habataki//Sasanishiki///Sasanishiki. The resultant gene was ABERRANT PANICLE ORGANIZATION 1 (APO1). Habataki-genotype segregated reciprocal recombinant lines for the APO1 locus increased both the number of PRB (12–13%) and the number of grains per panicle (9–12%), which increased the grain yield per plant (5–7%). Further recombination dividing this region revealed that different alleles regulated the number of PRB and the number of grains per panicle. The PRB1 allele, which includes the APO1 open reading frame (ORF) and the proximal promoter region, controlled only the number of PRB but not the number of grains per panicle. In contrast, the HI1 allele, which includes only the distal promoter region, increased the grain yield and harvest index in Habataki-genotype plants, nevertheless, the ORF expressed was Sasanishiki type. It also increased the number of large vascular bundles in the peduncle. APO1 expression occurred not only in developing panicles but also in the developing vascular bundle systems. In addition, Habataki plants displayed increased APO1 expression in comparison to Sasanishiki plants. It suggests that APO1 enhances the formation of vascular bundle systems which, consequently, promote carbohydrate translocation to panicles. The HI1 allele is suggested to regulate the amount of APO1 expression, and thereby control the development of vascular bundle systems. These findings may be useful to improve grain yield as well as quality through the improvement of translocation efficiency.










Similar content being viewed by others
References
Ando T, Yamamoto T, Shimizu T, Ma XF, Shomura A, Takeuchi Y, Lin SY, Yano M (2008) Genetic dissection and pyramiding of quantitative traits for panicle architecture by using chromosomal segment substitution lines in rice. Theor Appl Genet 116:881–890
Ashikari M, Matsuoka M (2006) Identification, isolation and pyramiding of quantitative trait loci for rice breeding. Trends Plant Sci 11:344–350
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A, Angeles ER, Qian Q, Kitano H, Matsuoka M (2005) Cytokinin oxidase regulates rice grain production. Science 309:741–745
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Durfee T, Roe JL, Sessions RA, Inouye C, Serikawa K, Feldmann KA, Weigel D, Zambryski PC (2003) The F-box-containing protein UFO and AGAMOUS participate in antagonistic pathways governing early petal development in Arabidopsis. Proc Natl Acad Sci 100:8571–8576
Fukuoka H, Ogawa T, Mitsuhara I, Iwai T, Isuzugawa K, Nishizawa Y, Gotoh Y, Nishizawa Y, Tagiri A, Ugaki M, Ohshima M, Yano H, Murai N, Niwa Y, Hibi T, Ohashi Y (2000) Agrobacterium-mediated transformation of monocot and dicot plants using the NCR promoter derived from soybean chlorotic mottle virus. Plant Cell Rep 19:815–820
Fukushima A, Akita S (1997) Varietal differences of the course and differentiation time of large vascular bundles in the rachis of rice. Jpn J Crop Sci 66:24–28
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300
Hirose T, Terao T (2004) A comprehensive expression analysis of the starch synthase gene family in rice (Oryza sativa L.). Planta 220:9–16
Hirose T, Takano M, Terao T (2002) Cell wall invertase in developing rice caryopsis: molecular cloning of OsCIN1 and analysis of its expression in relation to its role in grain filling. Plant Cell Physiol 43:452–459
Hirose T, Ohdan T, Nakamura Y, Terao T (2006) Expression profiling of genes related to starch synthesis in rice leaf sheaths during the heading period. Physiol Plant 128:425–435
Huang X, Qian Q, Liu Z, Sun H, He S, Luo D, Xia G, Chu C, Li J, Fu X (2009) Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet 41:494–497
Ikeda K, Nagasawa N, Natago Y (2001) ABERRANT PANICLE ORGANIZATION 1 temporally regulates meristem identity in rice. Dev Biol 282:349–360
Ikeda K, Ito M, Nagasawa N, Kyozuka J, Nagato Y (2007) Rice ABERRANT PANICLE ORGANIZATION 1, encoding an F-box protein, regulates meristem fate. Plant J 51:1030–1040
Ikeda-Kawakatsu K, Yasuno N, Oikawa T, Iida S, Nagato Y, Maekawa M, Kyozuka J (2009) Expression level of ABERRANT PANICLE ORGANIZATION 1 determines rice inflorescence form through control of cell proliferation in the meristem. Plant Physiol 150:736–747
Kobayashi S, Fukuta Y, Yagi T, Sato T, Osaki M, Khush GS (2004) Identification and characterization of quantitative trait loci affecting spikelet number per panicle in rice (Oryza sativa L.). Field Crops Res 89:253–262
Komatsu K, Maekawa M, Ujiie S, Satake Y, Furutani I, Okamoto H, Shimamoto K, Kyozuka J (2003a) LAX and SPA: major regulators of shoot branching in rice. Proc Natl Acad Sci 100:11765–11770
Komatsu M, Chujo A, Nagato Y, Shimamoto K, Kyozuka J (2003b) FRIZZY PANICLE is required to prevent the formation of axillary meristems and to establish floral meristem identity in rice spikelets. Development 130:3841–3850
Lanceras JC, Pantuwan G, Jongdee B, Toojinda T (2004) Quantitative trait loci associated with drought tolerance at reproductive stage in rice. Plant Physiol 135:384–399
Levin JZ, Meyerowitz EM (1995) UFO: an Arabidopsis gene involved in both floral meristem and floral organ development. Plant Cell 7:529–548
Mao B-B, Cai W-J, Zhang Z-H, Hu Z-L, Li P, Zhu L-H, Zhu Y-G (2003) Characterization of QTLs for harvest index and source-sink characters in a DH population of rice (Oryza sativa L.). Acta Genet Sin 30:1118–1126
Matsushima S (1957) Analysis of developmental factors determining yield and yield prediction in lowland rice. Bull Natl Inst Agric Sci A 5:1–271
Mei HW, Luo LJ, Ying CS, Wang YP, Yu XQ, Guo LB, Paterson AH, Li ZK (2003) Gene actions of QTLs affecting several agronomic traits resolved in a recombinant inbred rice population and two testcross populations. Theor Appl Genet 107:89–101
Miki D, Shimamoto K (2004) Simple RNAi vectors for stable and transient suppression of gene function in rice. Plant Cell Physiol 45:490–495
Moritoh S, Miki D, Akiyama M, Kawahara M, Izawa T, Maki H, Shimamoto K (2005) RNAi-mediated silencing of OsGEN-L(OsGEN-like), a new member of the RAD2/XPG nuclease family, causes male sterility by defect of microspore development in rice. Plant Cell Physiol 46:699–715
Murai M, Nakamura K, Saito M, Nagayama A, Ise K (2005) Yield-increasing effect of a major gene, Ur1(Undulate rachis-1) on different genetic backgrounds in rice. Breed Sci 55:279–285
Nagao S, Takahashi M, Kinoshita T (1958) Inheritance on a certain ear type in rice. Genetical studies on plant, XXIII. Mem Fac Agric Hokkaido Univ 3:38–47
Nagata K, Fukuta Y, Shimizu H, Yagi T, Terao T (2002) Quantitative trait loci for sink size and ripening traits in rice (Oryza sativa L.). Breed Sci 52:259–273
Nakagawa M, Shimamoto K, Kyozuka J (2002) Overexpression of RCN1 and RCN2, rice TERMINAL FLOWER 1/CENTRORADIALIS homologs, confers delay of phase transition and altered panicle morphology in rice. Plant J 29:743–750
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45
Pouteau S, Nicholls D, Tooke F, Coen E, Battey N (1998) Transcription pattern of a FIM homologue in Impatiens during floral development and reversion. Plant J 14:235–246
Sasahara H, Fukuta Y, Fukuyama T (1999) Mapping of QTLs for vascular bundle system and spike morphology in rice, Oryza sativa L. Breed Sci 49:75–81
Sasaki T, Matsumoto T, Yamamoto K et al (2002) The genome sequence and structure of rice chromosome 1. Nature 420:312–316
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M (2008) Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet 40:1023–1028
Simon R, Carpenter R, Doyle S, Coen E (1994) Fimbriata controls flower development by mediating between meristem and organ identity genes. Cell 78:99–107
Snyder FW, Carlson GE (1984) Selecting for partitioning of photosynthetic products in crops. Adv Agron 37:47–72
Song X-J, Huang W, Shi M, Zhu M-Z, Lin H-X (2007) A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet 39:623–630
Taylor S, Hofer J, Murfet I (2001) Stamina pistilloida, the pea ortholog of Fim and UFO, is required for normal development of flowers, inflorescences, and leaves. Plant Cell 13:31–46
Tian F, Zhu Z, Zhang B, Tan L, Fu Y, Wang X, Sun CQ (2006) Fine mapping of a quantitative trait locus for grain number per panicle from wild rice (Oryza rufipogon Griff.). Theor Appl Genet 113:619–629
Toki S, Hara N, Ono K, Onodera H, Tagiri A, Oka S, Tanaka H (2006) Early infection of scutellum tissue with Agrobacterium allows high-speed transformation of rice. Plant J 47:969–976
Tsugane K, Maekawa M, Takagi K, Takahara H, Qian Q, Eun CH, Iida S (2006) An active DNA transposon nDart causing leaf variegation and mutable dwarfism and its related elements in rice. Plant J 45:46–57
Wang E, Wand J, Zhu X, Hao W, Wang L, Li Q, Zhang L, He W, Lu B, Lin H, Ma H, Zhang G, He Z (2008) Control of rice grain-filling and yield by a gene with a potential signature of domestication. Nat Genet 40:1370–1374
Wilkinson MD, Haughn GW (1995) UNUSUAL FLORAL ORGANS Controls meristem identity and organ primordia fate in Arabidopsis. Plant Cell 7:1485–1499
Xiao J, Li J, Grandillo S, Ahn SN, Yuan L, Tanksley SD, McCouch SR (1998) Identification of trait-improving quantitative trait loci alleles from a wild rice relative, Oryza rufipogon. Genetics 150:899–909
Yagi T, Nagata K, Fukuta Y, Tamura K, Ashikawa I, Terao T (2001) QTL mapping of spikelet number in rice (Oryza sativa L.). Breed Sci 51:53–56
Yamanouchi U, Yano M, Lin H, Ashikari M, Yamada K (2002) A rice spotted leaf gene, Spl7, encodes a heat stress transcription factor protein. Proc Natl Acad Sci 99:7530–7535
Yoon D-B, Kang K-H, Kim H-J, Ju H-G, Kwon S-J, Suh J-P, Jeong O-Y, Ahn S-N (2006) Mapping quantitative trait loci for yield components and morphological traits in an advanced backcross population between Oryza grandiglumis and the O. sativa japonica cultivar Hwaseongbyeo. Theor Appl Genet 112:1052–1062
Zhang W-H, Kokubun M (2004) Historical changes in grain yield and photosynthetic rate of rice cultivars released in the 20th century in Tohoku region. Plant Prod Sci 7:36–44
Zhang S, Sandal N, Plolwick PL, Stiller J, Stougaard J, Fobert PR (2003) Proliferating Floral Organs (Pfo), a Lotus japonicus gene required for specifying floral meristem determinacy and organ identity, encodes an F-box protein. Plant J 33:607–619
Zhuang J-Y, Lin H-X, Lu J, Qian H-R, Hittalmani S, Huang N, Zheng K-L (1997) Analysis of QTL × environment interaction for yield components and plant height in rice. Theor Appl Genet 95:799–808
Acknowledgments
This work was supported by a Grant-in-aid from the Ministry of Agriculture, Forestry and Fishery (MAFF) of Japan (Bio-design project 1998–2007). SAS submission was performed with the assistance of the Computer Center for Agriculture, Forestry and Fisheries Research, MAFF, Japan. We thank Prof. K Shimamoto of Nara Inst. Sci Tec. for providing p2K1+ vector. We are also grateful to Mr. Tokuya Genba, Mr. Tuyoshi Kotake, Mr. Tetsuya Aota, Mr. Masanori Ichihashi, Mr. Shinobu Yuminamochi, Ms. Keiko Nozaki, Ms. Kiiko Takatsuto and Ms. Setsuko Hayashi for their excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Y. Xu.
Rights and permissions
About this article
Cite this article
Terao, T., Nagata, K., Morino, K. et al. A gene controlling the number of primary rachis branches also controls the vascular bundle formation and hence is responsible to increase the harvest index and grain yield in rice. Theor Appl Genet 120, 875–893 (2010). https://doi.org/10.1007/s00122-009-1218-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1218-8