Abstract
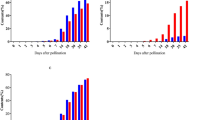
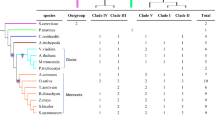
To elucidate the roles of the isogenes encoding starch synthase (EC 2.4.1.21) in rice (Oryza sativa L.), a comprehensive expression analysis of the gene family was conducted. Extensive searches for starch synthase genes were done in the databases of both the whole genome and full-length cDNAs of rice, and ten genes were revealed to comprise the starch synthase gene family. Multi-sequence alignment analysis of the starch synthase proteins from rice and other plant species suggested that they were grouped into five classes, soluble starch synthase I (SSI), SSII, SSIII, SSIV and granule-bound starch synthase (GBSS). In rice, there was one gene for SSI, three for SSII and two each for SSIII, IV and GBSS. The expression pattern of the ten genes in the developing caryopsis was examined by semi-quantitative RT–PCR analysis. Based on the temporal expression patterns, the ten genes could be divided into three groups: (i) early expressers (SSII-2, III-1, GBSSII), which are expressed in the early stage of grain filling; (ii) late expressers (SSII-3, III-2, GBSSI), which are expressed in the mid to later stage of grain filling; and (iii) steady expressers (SSI, II-1, IV-1, IV-2), which are expressed relatively constantly during grain filling. Within a caryopsis, the three gene groups spatially share their expression, i.e. “early expressers” in the pericarp, the “late expressers” in the endosperm” and the “steady expressers” in both tissues. In addition, this grouping was reflected in the expression pattern of various rice tissues: expression in non-endosperm, endosperm or all tissues examined. The implications in this spatio-temporal work sharing of starch synthesis isogenes are discussed.






Similar content being viewed by others
Abbreviations
- DAF :
-
Days after flowering
- GBSS :
-
Granule-bound starch synthase
- SS :
-
Soluble starch synthase
References
Cao H, Imparl-Radosevich J, Guan H, Keeling PL, James MG, Myers AM (1999) Identification of the soluble starch synthase activities of maize endosperm. Plant Physiol 120:205–215
Chang S, Puryear J, Cairney J (1993) A simplified method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116
Craig J, Lloyd JR, Tomlinson K, Barber L, Edwards A, Wang TL, Martin C, Hedley CL, Smith AM (1998) Mutations in the gene encoding starch synthase II profoundly alter amylopectin structure in pea embryos. Plant Cell 10:413–426
Gao M, Fisher DK, Kim KN, Shannon JC, Guiltinan MJ (1997) Independent genetic control of maize starch-branching enzymes IIa and IIb. Plant Physiol 114:69–78
Gao M, Wanat J, Stinard PS, James MG, Myers AM (1998) Characterization of dull1, a maize gene coding for a novel starch synthase. Plant Cell 10:399–412
Gubler F, Li Z, Fieg S, Jacobsen JV, Morell MK (2000) Cloning and characterization of a starch synthase I gene (Accession No. AF234163) from barley (PGR00-047). Plant Physiol 122:1459
Harn C, Knight M, Ramakrishnan A, Guan H, Keeling PL, Wasserman BP (1998) Isolation and characterization of the zSSIIa and zSSIIb starch synthase cDNA clones from maize endosperm. Plant Mol Biol 37:639–649
Hirose T, Takano M, Terao T (2002) Cell wall invertase in developing rice caryopsis: molecular cloning of OsCIN1 and analysis of its expression in relation to its role in grain filling. Plant Cell Physiol 43:452–459
James MG, Denyer K, Myers AM (2003) Starch synthesis in the cereal endosperm. Curr Opin Plant Biol 6:215–22
Li Z, Sun F, Xu S, Chu X, Mukai Y, Yamamoto M, Ali S, Rampling L, Kosar-Hashemi, Rahman S, Morell MK (2003) The structural organisation of the gene encoding class II starch synthase of wheat and barley and the evolution of the genes encoding starch synthase in plants. Funct Integr Genomics 3:76–85
Matsuoka M (1990) Classification and characterization of cDNA that encodes the light-harvesting chlorophyll a/b binding protein of photosystem II from rice. Plant Cell Physiol 31:519–526
Mizuno K, Kobayashi E, Tachibana M, Kawasaki T, Fujiwara T, Funane K, Kobayashi M, Baba T (2001) Characterization of an isoform of rice starch branching enzyme, RBE4, in developing seeds. Plant Cell Physiol 42:349–357
Nakamura T, Vrinten P, Hayakawa K, Ikeda J (1998) Characterization of a granule-bound starch synthase isoform found in the pericarp of wheat. Plant Physiol 118:451–459
Nakamura Y (2002) Towards a better understanding of the metabolic system for amylopectin biosynthesis in plants: rice endosperm as a model tissue. Plant Cell Physiol 43:718–725
Patron N, Smith AM, Fahy BF, Hylton CM, Naldrett MJ, Rossnagel BC, Denyer K (2002) The altered pattern of amylose accumulation in the endosperm of low-amylose barley cultivars is attributable to a single mutant allele of granule-bound starch synthase I with a deletion in the 5′-non-coding region. Plant Physiol 130:190–198
Preiss J, Sivak MN (1996) Starch synthesis in sinks and sources. In: Zamski E, Schaffer AA (eds) Photoassimilate distribution in plants and crops. Dekker, New York, pp 63–96
Sato K (1984) Starch granules in tissues of rice plants and their changes in relation to plant growth. JARQ 18:78–86
Smith AM, Denyer K, Martin C (1997) The synthesis of the starch granule. Annu Rev Plant Physiol Plant Mol Biol 48:67–87
Takaiwa F, Oono K, Wing D, Kato A (1991) Sequence of three members and expression of a new major superfamily of glutelin genes from rice. Plant Mol Biol 17:875–885
Tomlinson KL, Lloyd JR, Smith AM (1997) Importance of isoforms of starch-branching enzyme in determining the structure of starch in pea leaves. Plant J 11:31–43
Tomlinson K, Craig J, Smith AM (1998) Major differences in isoform composition of starch synthase between leaves and embryos of pea (Pisum sativum L.). Planta 204:86–92
UmemotoT, Yano M, Satoh H, Shomura A, Nakamura Y (2002) Mapping of a gene responsible for the difference in amylopectin structure between japonica-type and indica-type rice varieties. Theor Appl Genet 104:1–8
Vrinten PL, Nakamura T (2000) Wheat granule-bound starch synthase I and II are encoded by separate genes that are expressed in different tissues. Plant Physiol 122:255–263
Wang YJ, White P, Pollak L, Jane JL (1993) Characterization of starch structures of 17 maize endosperm mutant genotypes with Oh43 inbred line background. Cereal Chem 70:171–179
Zhu T, Budworth P, Chen W, Provart N, Chang HS, Guimil S, Su W, Estes B, Zou G, Wang X (2003) Transcriptional control of nutrient partitioning during rice grain filling. Plant Biotechnol J 1:59–70
Acknowledgement
The authors thank Kiiko Takatsuto for expert technical assistances, and Dr. Naohiro Aoki for helpful discussion. This work is supported in part by grants from the Ministry of Agriculture, Forestry and Fisheries of Japan (Pioneer Research Project Fund, PRPF-1206) to T.H. and T.T., and from the Ministry of Education, Culture, Sports, Science and Technology of Japan (No. 15580016) to T.H.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hirose, T., Terao, T. A comprehensive expression analysis of the starch synthase gene family in rice (Oryza sativa L.). Planta 220, 9–16 (2004). https://doi.org/10.1007/s00425-004-1314-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-004-1314-6