Abstract
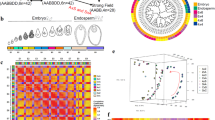
Heterosis specifies the superior performance of heterozygous individuals and although used in plant breeding the underlying molecular mechanisms still remain largely elusive. In this study, we demonstrate the manifestation of heterosis in hybrid maize embryo and endosperm tissue 6 days after fertilization in crosses of several inbred lines. We provide a comparative analysis of heterosis-associated gene expression in these tissues by a combined approach of suppression subtractive hybridization and microarray hybridizations. Non-additive expression pattern indicated a trans-regulatory mechanism to act early after fertilization in hybrid embryo and endosperm although the majority of genes showed mid-parental expression levels in embryo and dosage dependent expression levels in endosperm. The consistent expression pattern within both tissues and both inbred line genotype combinations of genes coding for chromatin related proteins pointed to heterosis-related epigenetic processes. These and genes involved in other biological processes, identified in this study, might provide entry points for the investigation of regulatory networks associated with the specification of heterosis.




Similar content being viewed by others
References
Achard P, Cheng H, De Grauwe L, Decat J, Schoutteten H, Moritz T, Van Der Straeten D, Peng J, Harberd NP (2006) Integration of plant responses to environmentally activated phytohormonal signals. Science 311:91–94
Achard P, Liao L, Jiang C, Desnos T, Bartlett J, Fu X, Harberd NP (2007) DELLAs contribute to plant photomorphogenesis. Plant Physiol 143:1163–1172
Adams KL, Cronn R, Percifield R, Wendel JF (2003) Genes duplicated by polyploidy shows unequal contributions to the transcriptome and organ-specific reciprocal silencing. PNAS 100:4649–4654
Alleman M, Doctor J (2000) Genomic imprinting in plants: observations and evolutionary implications. Plant Mol Biol 43:147–161
Auger DL, Gray AD, Ream TS, Kato A, Coe EH Jr, Birchler JA (2005) Non-additive gene expression in diploid and triploid hybrids of maize. Genetics 169:389–397
Baroux C, Spillane C, Grossniklaus U (2002) Genomic imprinting during seed development. Adv Genet 46:165–214
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc 57:289–300
Berger F (1999) Endosperm development. Curr Opin Plant Biol 2:28–32
Birchler JA (1993) Dosage analysis of maize endosperm development. Annu Rev Genet 27:181–204
Birchler JA, Auger DL, Riddle NC (2003) In search of the molecular basis of heterosis. Plant Cell 15:2236–2239
Brink RA, Cooper DC (1947) The endosperm in seed development. Bot Rev 13:423–541
Chandler V, Stam M (2004) Chromatin conversations: mechanisms and implications of paramutation. Nat Rev Gen 5:532–544
Chinnusamy V, Gong Z, Zhu JK (2008) Abscisic acid-mediated epigenetic processes in plant development and stress responses. J Integr Plant Biol 50:1187–1195
Costa LM, Gutièrrez-Marcos JF, Dickinson HG (2004) More than a yolk: the short life and complex times of the plant endosperm. Trends Plant Sci 9:507–514
Drews GN, Lee D, Christensen CA (1998) Genetic analysis of female gametophyte development and function. Plant Cell 10:5–17
Fransz PF, de Jong FH (2002) Chromatin dynamics in plants. Curr Opin Plant Biol 5:560–567
Fu X, Harberd NP (2003) Auxin promotes Arabidopsis root growth by modulating gibberellin response. Nature 421:740–743
Gao XH, Huang XZ, Xiao SL, Fu XD (2008) Evolutionarily conserved DELLA-mediated gibberellin signaling in plants. J Integr Plant Biol 50:825–834
Goldberg RB, de Paiva G, Yadegari R (1994) Plant embryogenesis: zygote to seed. Science 266:605–614
Grant-Downton R, Dickinson H (2004) Plants, pairing and phenotypes: two’s company? Trends Genet 20:188–195
Groenewald JH, Botha FC (2008) Down-regulation of pyrophosphate: fructose 6-phosphate 1-phosphotransferase (PFP) activity in sugarcane enhances sucrose accumulation in immature internodes. Transgenic Res 17:85–92
Guo M, Rupe MA, Danilevskaya ON, Yang X, Hu Z (2003) Genome-wide mRNA profiling reveals heterochronic allelic variation and a new imprinted gene in hybrid maize endosperm. Plant J 36:30–44
Guo M, Rupe MA, Yang X, Crasta O, Zinselmeier C, Smith OS, Bowen B (2006) Genome-wide transcript analysis of maize hybrids: allelic additive gene expression and yield heterosis. TAG 113:831–845
Hämmerle B, Ferrus R (2003) Expression of enhancers is altered in Drosophila melanogaster hybrids. Evol Dev 5:221–230
Henikoff S, Ahmad K (2005) Assembly of variant histones into chromatin. Annu Rev Cell Dev Biol 21:133–153
Hochholdinger F, Hoecker N (2007) Towards the molecular basis of heterosis. Trends Plant Sci 12:427–432
Hoecker N, Keller B, Piepho H-P, Hochholdinger F (2005) Manifestation of heterosis during early maize (Zea mays L.) root development. TAG 12:421–429
Hoecker N, Keller B, Muthreich N, Chollet D, Descombes P, Piepho HP, Hochholdinger F (2008) Comparison of maize (Zea mays L.) F1-hybrid and parental inbred line primary root transcriptomes suggests organ-specific patterns of nonadditive gene expression and conserved expression trends. Genetics 179:1275–1283
Hong SK, Kitano H, Satoh H, Nagato Y (1996) How is the embryo size genetically regulated in rice? Development 122:2051–2058
Huang Y, Zhang L, Zhang J, Yuan D, Xu C, Li X, Zhou D, Wang S, Zhang Q (2006) Heterosis and polymorphisms of gene expression in an elite rice hybrid as revealed by microarray analysis of 9198 unique ESTs. Plant Mol Biol 62:579–591
Laux T, Jürgens G (1997) Embryogenesis: a new start in life. Plant Cell 9:989–1000
Le Q, Gutiérrez-Marcos JF, Costa LM, Meyer S, Dickinson HG, Lörz H, Kranz E, Scholten S (2005) Construction and screening of subtracted cDNA libraries from limited populations of plant cells: a comparative analysis of gene expression between maize egg cells and central cells. Plant J 44:167–178
Lin BY (1984) Ploidy barrier to endosperm development in maize. Genetics 107:103–115
Lopes MA, Larkins BA (1993) Endosperm origin, development, and function. Plant Cell 5:1383–1399
Melchinger AE (1999) Genetic diversity and heterosis. In: International symposium on genetics and exploitation of heterosis in crop plants, Mexico City, 17–22 August 1997, pp 99–118
Mertens E (1991) Pyrophosphate-dependent phosphofructokinase, an anaerobic glycolytic enzyme? FEBS Lett 285:1–5
Meyer S, Pospisil H, Scholten S (2007) Heterosis associated gene expression in maize embryo six days after fertilization exhibits additive, dominant and overdominant pattern. Plant Mol Biol 63:381–391
Michalak P, Noor MAF (2003) Genome-wide patterns of expression in Drosophila pure species and hybrid males. Mol Biol Evol 20:1070–1076
Okamoto T, Scholten S, Lörz H, Kranz E (2005) Identification of genes that are up- or down-regulated in the apical or basal cell of maize two-celled embryos and monitoring their expression during zygote development by a cell manipulation- and PCR-based approach. Plant Cell Physiol 46:332–833
Olsen OA (1998) Endosperm development. Plant Cell 10:485–488
Olsen OA (2001) Endosperm development: cellularization and cell fate specification. Annu Rev Plant Physiol Plant Mol Biol 52:233–267
Olsen OA (2004) Nuclear endosperm development in cereals and Arabidopsis thaliana. Plant Cell 16:214–227
Pang SZ, DeBoer DL, Wan Y, Ye G, Layton JG, Neher MK, Armstrong CL, Fry JE, Hinchee MA, Fromm ME (1996) An improved green fluorescent protein gene as a vital marker in plants. Plant Physiol 112:893–900
Peng J, Carol P, Richards DE, King KE, Cowling RJ, Murphy GP, Harberd NP (1997) The Arabidopsis GAI gene defines a signalling pathway that negatively regulates gibberellin response. Genes Dev 11:3194–3205
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ, Worland AJ, Pelica F, Sudhakar D, Christou P, Snape JW, Gale MD, Harberd NP (1999) ‘Green revolution’ genes encode mutant gibberellin response modulators. Nature 400:256–261
Piepho H-P (2005) Optimal allocation in designs for assessing heterosis from cDNA gene expression data. Genetics 171:359–364
Rhoades MM, Dempsey E (1966) Induction of chromosome doubling at meiosis by the elongate gene in maize. Genetics 54:505–522
Römisch-Margl L, Spielbauer G, Schützenmeister A, Schwab W, Piepho H-P, Genschel U, Gierl A (2010) Heterotoc patterns of sugar and amino acid components in developing maize kernels. TAG (this issue)
Rood SB, Buzzell RI, Mander LN, Pearce D, Pharis RP (1988) Gibberellins: a phytohormonal basis for heterosis in maize. Science 241:1216–1218
Russel SD (1992) Double fertilization. Int Rev Cytol 140:357–390
Sabelli PA, Larkins BA (2009) The development of endosperm in grasses. Plant Physiol 149:14–26
Scott RJ, Spielman M, Bailey J et al (1998) Parent-of-origin effects on seed development in Arabidopsis thaliana. Development 125:3329–3341
Shull GH (1908) The composition of a field of maize. Proc Am Breeders Assoc 4:296–301
Shull GH (1952) Beginning of the heterosis concept. In: Gowen JW (ed) Heterosis. Iowa State College Press, Ames, pp 14–48
Smith OS, Smith JSC, Bowen SL, Tenborg RA, Wall SJ (1990) Similarities among a group of elite maize inbreds as measured by pedigree, F1 grain yield, heterosis, and RFLPs. TAG 80:833–840
Song R, Messing J (2003) Gene expression of a gene family in maize based on noncollinear haplotypes. PNAS 100:9055–9060
Springer NM, Stupar RM (2007) Allelic variation and heterosis in maize: how do two halves make more than a whole? Genome Res 17:264–275
Stupar RM, Springer NM (2006) Cis-transcriptional variation in maize inbred lines B73 and Mo17 leads to additive expression patterns in the F1 hybrid. Genetics 173:2199–2210
Stupar RM, Hermanson PJ, Springer NM (2007) Nonadditive expression and parent-of-origin effects identified by microarray and allele-specific expression profiling of maize endosperm. Plant Physiol 145:411–425
Stupar RM, Gardiner JM, Oldre AG, Haun WJ, Chandler VL, Springer NM (2008) Gene expression analyses in maize inbreds and hybrids with varying levels of heterosis. BMC Plant Biol 8:33
Swanson-Wagner RA, Jia Y, DeCook R, Borsuk LA, Nettelton D, Schnable PS (2006) All possible modes of gene action are observed in a global comparison of gene expression in a maize F1 hybrid and its inbred parents. PNAS 103:6805–6810
Uzarowska A, Keller B, Piepho H-P, Schwarz G, Ingvardsen C, Wenzel G, Lübberstedt T (2007) Comparative expression profiling in meristems of inbred-hybrid triplets of maize based on morphological investigations of heterosis for plant height. Plant Mol Biol 63:21–34
Vuylsteke M, van Eeuwijk F, Van Hummelen P, Kuiper M, Zabeau M (2005) Genetic analysis of variation in gene expression in Arabidopsis thaliana. Genetics 171:1267–1275
Wang FH (1947) Embryological development of inbred and hybrid Zea mays L. Am J Bot 34:113–125
Acknowledgments
For providing seeds of the inbred lines used in this study we thank Prof. Dr. A. Melchinger (University of Hohenheim) and his co-workers. We thank T. Okamoto for allocating cDNAs of zygotes and two-celled embryos and André Schützenmeister for data processing for GEO submission. Furthermore we are especially grateful to Petra von Wiegen, Marlis Nissen and Sabina Miaskowska for excellent technical help with embryo and endosperm isolation and microarray hybridization. For assisting the glass house work we are thankful to Bärbel Hagemann. We thank Prof. Dr. Erhard Kranz and Prof. Dr. Horst Lörz for providing working space and support. This work was financially supported by the Deutsche Forschungsgemeinschaft (DFG) grant SCHO746/2 within the framework program “heterosis in plants” and by the Eiselen-Foundation (Ulm).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by T. Luebberstedt.
Contribution to the special issue “Heterosis in Plants”.
The microarray data of this study were deposited in Gene Expression Omnibus (GEO) at the National Centre for Biotechnology Information (NCBI) with the series entry GSE17754.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jahnke, S., Sarholz, B., Thiemann, A. et al. Heterosis in early seed development: a comparative study of F1 embryo and endosperm tissues 6 days after fertilization. Theor Appl Genet 120, 389–400 (2010). https://doi.org/10.1007/s00122-009-1207-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-1207-y