Abstract
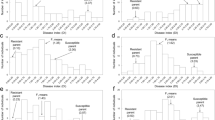
Resistance against a Ralstonia solanacearum race 3-phylotype II strain JT516 was assessed in a F2:3 and a population of inbred lines (RIL), both derived from a cross between L. esculentum cv. Hawaii 7996 (partially resistant) and L. pimpinellifolium WVa700 (susceptible). Resistance criteria used were the percentage of wilted plants to calculate the AUDPC value, and bacterial colonization scores in roots and stem (hypocotyl and epicotyl) assessed in two independent greenhouse experiments conducted during the cool and hot seasons in Réunion Island, France. Symptoms were more severe during the cool season trials. Heritability estimates in individual seasons ranged from 0.82 to 0.88, depending on resistance criterion. A set of 76 molecular markers was used for quantitative trait loci (QTL) mapping using the single- and composite- interval mapping methods, as well as ANOVA. Four QTLs, named Bwr- followed by a number indicating their map location, were identified. They explained from 3.2 to 29.8% of the phenotypic variation, depending on the resistance criterion and the season. A major QTL, Bwr-6, and a minor one, Bwr-3, were detected in each season for all resistance criteria. Both QTLs showed stronger effects in the hot season than in the cool one. Their role in resistance to R. solanacearum race 3-phylotype II was subsequently confirmed in the RIL population derived from the same cross. Two other QTLs, Bwr-4 and Bwr-8, with intermediate and minor effects, respectively, were only detected in the hot season, demonstrating that environmental factors may strongly influence the expression of resistance against the race 3-phylotype II strain JT516. These QTLs were compared with those detected in the RIL population against race 1-phylotype I strain JT519 as well as those detected in other previous studies in the same genetic background against other race 1-phylotype I and II strains. This comparison revealed the possible occurrence of some phylotype-specific resistance QTLs in Hawaii 7996.


Similar content being viewed by others
References
Acosta JC, Gilbert JC, Quinon VL (1964) Heritability of bacterial wilt resistance in tomato. P Am Soc Hortic Sci 84:455–462
Anaïs G (1986) Utilisation de la résistance variétale dans la lutte contre le flétrissement bactérien de la tomate dû au Pseudomonas solanacearum. In: Smith EF (ed)INRA Bulletin Technique d’Information. Ministère de l’Agriculture, Paris, pp 409–411, 449–453
Anand N, Sadashiva AT, Tikoo SK, Ramkishun M, Reddy K (1993) Resistance to bacterial wilt in tomato: gene dosage effects. In: Hartman GL and Hayward AC (eds) Bacterial wilt. ACIAR Proceedings 45. Kaoshiung, Taiwan, pp 142–148
Basten CJ, Weir BS, Zeng ZB (2000) QTL cartographer. A reference manual and tutorial for QTL mapping. Department of Statistics, North Carolina State University, Raleigh, North Carolina
Buddenhagen IW (1986) Bacterial wilt revisited. In: Persley GJ (ed) Bacterial wilt in Asia and the South Pacific. ACIAR Proceedings, ACIAR, Camberra, pp 126–139
Buddenhagen IW, Kelman A (1964) Biological and physiological aspects of bacterial wilt caused by Pseudomonas solanacearum. Annu Rev Phytopathol 2:203–230
Buddenhagen I, Sequeira L, Kelman A (1962) Designation of races in Pseudomonas solanacearum. Phytopathology 52:726
Carmeille A, Prior P, Kodja H, Chiroleu F, Luisetti J, Besse P (2006) Evaluation of resistance to race 3 biovar 2 of Ralstonia solanacearum in tomato germplasm. J Phytopathology 154:1–5
Cook D, Sequeira L (1991) Genetic and biochemical characterization of a Pseudomonas solanacearum gene cluster required for extracellular polysaccharide production and virulence. J Bacteriol 173:1654–1662
Cook D, Sequeira L (1994) Strain differentiation of Pseudomonas solanacearum by molecular genetics methods. In: Hayward AC and Hartman GL (eds) Bacterial wilt: the disease and its causative agent, Pseudomonas solanacearum. CAB International, Wallingford, UK, pp 77–93
Cook D, Barlow E, Sequeira L (1989) Genetic diversity of Pseudomonas solanacearum: detection of restriction fragment length polymorphism with DNA that specify virulence and the hypersensitive response. Mol Plant Microbe Interact 2:113–121
Cook D, Barlow E, Sequeira L (1991) DNA probes as tools for the study of host-pathogen evolution: the example of Pseudomonas solanacearum. In: Hennecke H, Verma DPS (eds) Advances in molecular genetics of plant microbe interactions. Kluwer, Dordrecht, pp 103–108
Danesh D, Aarons S, McGill GE, Young ND (1994) Genetic dissection of oligogenic resistance to bacterial wilt in tomato. Mol Plant Microbe Interact 7:464–471
Fegan M, Prior P (2005) How complex is the " Ralstonia solanacearum species complex"? In: Allen C, Prior P and Hayward AC (eds) Bacterial wilt: The disease and Ralstonia solanacearum species complex. APS Press, St Paul, pp 449–461
Fegan M, Taghavi L, Sly LI, Hayward AC (1998) Phylogeny, diversity and molecular diagnostics of Ralstonia solanacearum. In: Prior P, Allen C, Elphinstone J (eds) Bacterial wilt disease: molecular and ecological aspects. Springer, Berlin Heidelberg New York, pp 19–33
French ER (1986) Interaction between strains of Pseudomonas solanacearum, its hosts and the environment. In: Persley GJ (ed) Bacterial wilt in Asia and South Pacific, ACIAR proceedings, ACIAR, Camberra, pp 99–104
Granada GA, Sequeira L (1983) Survival of Pseudomonas solanacearum in soil, rhizosphere and plant roots. Can J Microbiol 29:433–440
Grimault V, Prior P (1993) Bacterial wilt resistance in tomato associated with tolerance of vascular tissues to Pseudomonas solanacearum. Plant Pathol 42:589–594
Grimault V, Anaïs G, Prior P (1994) Distribution of Pseudomonas solanacearum in the stem tissues of tomato plants with different levels of resistance to bacterial wilt. Plant Pathol 43:663–668
Haldane J (1919) The combination of linkage values, and the calculation of distance between loci of linked factors. J Genet 8:299–309
Hayward AC (1964) Characteristics of Pseudomonas solanacearum. J Appl Bacteriol 27:265–277
Hayward AC (1991) Biology and epidemiology of bacterial wilt caused by Pseudomonas solanacearum. Annu Rev Phytopathol 29:65–87
Hayward AC (1994) The hosts of Pseudomonas solanacearum. In: Hayward AC, Hartman GL (eds) Bacterial wilt: the disease and its causative agent, Pseudomonas solanacearum. CAB International, Wallingford, UK, pp 9–25
He LY, Sequeira L, Kelman A (1983) Characteristics of strains of Pseudomonas solanacearum from China. Plant Dis 67:1357–1361
Janse JD (1996) Potato brown rot in Western Europe—history, present occurence and some remarks on possible origin, epidemiology and control strategies. EPPO Bull 26:679–695
Kelman A, Sequeira L (1965) Root-to-root spread of Pseudomonas solanacearum. Phytopathology 55:304–309
Kim SH, Olson TN, Schaad NW, Moorman GW (2003) Ralstonia solanacearum race 3, biovar 2, the causal agent of brown rot of potato, identified in geraniums in Pennsylvania, Delaware, and Connecticut. Plant Dis 87:450
Knapp S, Stroup W, Ross W (1985) Exact confidence intervals for heritabiility on a progeny mean basis. Crop Sci 9:257–262
Lander ES, Botstein D (1989) Mapping mendelian factors underlying quantitative trait using RFLP linkage maps. Genetics 121:185–199
Lander ES, Geen P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) Mapmaker: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Lefebvre V, Palloix A, Caranta C, Pochard E (1995) Construction of an interspecific integrated linkage map of pepper using molecular markers and doubled haploid progenies. Genome 38:112–121
Monma S, Sakata Y (1993) Inheritance of resistance to bacterial wilt in tomato. In: Hartman GL, Hayward AC (eds) Bacterial wilt. ACIAR proceedings, Kaoshiung, Taiwan, pp 149–153
Nicole J-F (1995) Influence de facteurs biotiques ou abiotiques sur la dynamique des populations de Pseudomonas solanacearum au cours de l’infection et sur le développement de la maladie. PhD Thesis, University of Nantes, France, pp 93
Poussier S, Vandewalle P, Luisetti J (1999) Genetic diversity of african and world wide strains of Ralstonia solanacearum as determined by PCR-Restriction fragment length polymorphism analysis of the hrp gene region. Appl Environ Microbiol 65:2184–2194
Poussier S, Prior P, Luisetti J, Hayward AC, Fegan M (2000a) Partial sequencing of the hrpB and endoglucanase genes confirms and expands the known diversity within the Ralstonia solanacearum species complex. Syst Appl Microbiol 23:479–486
Poussier S, Trigalet-Demery D, Vandewalle P, Goffinet B, Luisetti J, Trigalet A (2000b) Genetic diversity of Ralstonia solanacearum as assessed by PCR-RFLP of the hrp gene region, AFLP and 16S rRNA sequence analysis, and identification of an African subdivision. Microbiol 146:1679–1692
Prior P, Allen C, Elphinstone J (1998) Bacterial wilt disease: molecular and ecological aspects. Springer, Berlin Heidelberg New York, pp 447
Schmit J (1978) Microscopic study of early stages of infection by Pseudomonas solanacearum E.F.S. on "in vitro" grown tomato seedlings. In: Proceedings of the IVth international conference on plant pathogenic bacteria. INRA, Angers, pp 841–856
Shapiro SS, Wilk MB (1965) An analysis of variance for normality (complete samples). Biometrika 52:591–611
Stuber CW, Edwards MD, Wendel JF (1987) Molecular marker facilitated investigations of quantitative trait loci in maize. II. Factors influencing yield and its component traits. Crop Sci 27:639–648
Thoquet P, Olivier J, Sperisen C, Rogowsky P, Laterrot H, Grimsley N (1996a) Quantitative trait loci determining resistance to bacterial wilt in tomato cultivar Hawaii 7996. Mol Plant Microbe Interact 9:826–836
Thoquet P, Olivier J, Sperisen C, Rogowsky P, Prior P, Anaïs G, Mangin B, Bazin B, Nazer R, Grimsley N (1996b) Polygenic resistance of tomato plants to bacterial wilt in the French West Indies. Mol Plant Microbe Interact 9:837–842
Vasse J, Frey P, Trigalet A (1995) Microscopic studies of intercellular infection and protoxylem invasion of tomato roots by Pseudomonas solanacearum. Mol Plant Microbe Interact 8:241–251
Wang J-F, Olivier J, Thoquet P, Mangin B, Sauviac L, Grimsley NH (2000) Resistance of tomato line Hawaii7996 to Ralstonia solanacearum Pss4 in Taiwan is controlled mainly by a major strain-specific locus. Mol Plant Microbe Interact 13:6–13
Williamson L, Nakaho K, Hudelson B, Allen C (2002) Ralstonia solanacearum Race 3, Biovar 2 strains isolated from geranium are pathogenic on potato. Plant Dis 86:987–991
Zeng ZB (1994) Precision mapping of quantitative trait loci. Genetics 136:1457–1468
Acknowledgments
The authors thank F. Chiroleu and J. Veslot (CIRAD Réunion) for help in statistical analyses, J.-J. Chéron (CIRAD Réunion) for technical assistance in the greenhouse trials, SCEA Bassin Plat for maintaining plant cuttings, and H. Kodja (Université Réunion) for his interest in this work. We also thank N. Grimsley (INRA Toulouse) and J-F. Wang (AVRDC Taiwan) for providing seeds of the mapping populations and the genotypic molecular data. This work was partially supported by funds from the Conseil Général of Réunion Island.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I. Paran
Rights and permissions
About this article
Cite this article
Carmeille, A., Caranta, C., Dintinger, J. et al. Identification of QTLs for Ralstonia solanacearum race 3-phylotype II resistance in tomato. Theor Appl Genet 113, 110–121 (2006). https://doi.org/10.1007/s00122-006-0277-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-006-0277-3