Abstract
The rapid pace of economic development has resulted in the release of several polycyclic aromatic hydrocarbons (PAHs) into the environment. Microbial degradation using white-rot fungi is a promising method for the removal of PAHs from the environment. In the present study, biodegradation of recalcitrant PAH by a white-rot fungus, Trametes maxima IIPLC-32, was investigated using pyrene. The pyrene concentration decreased by 79.80%, 65.37%, and 56.37% within 16 days from the initial levels of 10 mg L−1, 25 mg L−1, and 50 mg L−1, respectively. Gas chromatographic–mass spectrometric identification of prominent metabolites 1-hydroxypyrene, 2-methyl-1-naphthyl acetic acid, di-n-butyl phthalate, and diethyl phthalate helped in determining the pyrene degradation pathway. The presence of 81 extracellular proteins was revealed by secretome analysis. The identified proteins up-regulated in response to pyrene degradation were classified into detoxification proteins (6.12%), redox proteins (6.12%), stress proteins (4.08%), metabolic-related proteins (26.53%), translation and transcriptional proteins (49%), catalytic proteins (49%), and other proteins (8.16%). Knowledge of secretome analysis in pyrene degradation helped to understand the degradation mechanism of pyrene. Also, the study suggests that T. maxima IIPLC-32 has the potential to be used in the bioremediation of PAH contaminated aquatic environment.
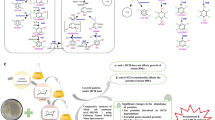
Graphical abstract






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
Change history
27 February 2023
A Correction to this paper has been published: https://doi.org/10.1007/s11356-023-26148-5
References
Acevedo F, Pizzul L, Castillo P et al (2011) Degradation of polycyclic aromatic hydrocarbons by the Chilean white-rot fungus Anthracophyllum discolor. J Hazard Mater 185:212–219. https://doi.org/10.1016/j.jhazmat.2010.09.020
Agrawal N, Shahi SK (2017) Degradation of polycyclic aromatic hydrocarbon (pyrene) using novel fungal strain Coriolopsis byrsina strain APC5. Int Biodeterior Biodegradation 122:69–81. https://doi.org/10.1016/j.ibiod.2017.04.024
Agrawal N, Verma P, Shahi SK (2018) Degradation of polycyclic aromatic hydrocarbons (phenanthrene and pyrene) by the ligninolytic fungi Ganoderma lucidum isolated from the hardwood stump. Bioresour Bioprocess 5:11. https://doi.org/10.1186/s40643-018-0197-5
Al-Hawash AB, Alkooranee JT, Zhang X, Ma F (2018) Fungal degradation of polycyclic aromatic hydrocarbons. Int J Pure Appl Biosci 6:8–24. https://doi.org/10.18782/2320-7051.6302
Al Farraj DA, Hadibarata T, Elshikh MS et al (2020) Biotransformation and degradation pathway of pyrene by filamentous soil fungus Trichoderma sp. F03. Water Air Soil Pollut 231:1–9. https://doi.org/10.1007/s11270-020-04514-0
Al-Hawash AB, Al-Qurnawi WS, Abbood HA et al (2020) Pyrene-degrading fungus ceriporia lacerata RF-7 from contaminated soil in Iraq. Polycycl Aromat Compd 1–9. https://doi.org/10.1080/10406638.2020.1713183
Baranger C, Pezron I, Lins L et al (2021) A compartmentalized microsystem helps understanding the uptake of benzo(a)pyrene by fungi during soil bioremediation processes. Sci Total Environ 784:147151. https://doi.org/10.1016/j.scitotenv.2021.147151
Barman SR, Banerjee P, Mukhopadhayay A, Das P (2017) Biodegradation of acenapthene and naphthalene by Pseudomonas mendocina: process optimization, and toxicity evaluation. J Environ Chem Eng 5:4803–4812. https://doi.org/10.1016/j.jece.2017.09.012
Bianco F, Race M, Papirio S, Esposito G (2022) Phenanthrene biodegradation in a fed–batch reactor treating a spent sediment washing solution: Techno–economic implications for the recovery of ethanol as extracting agent. Chemosphere 286:131361. https://doi.org/10.1016/j.chemosphere.2021.131361
Bishnoi NR, Mehta U, Sain U, Pandit GG (2005) Quantification of polycyclic aromatic hydrocarbons in tea and coffee samples of Mumbai city (India) by high performance liquid chromatography. Environ Monit Assess 107:399–406. https://doi.org/10.1007/s10661-005-3547-7
Chan SMN, Luan T, Wong MH, Tam NFY (2006) Removal and biodegradation of polycyclic aromatic hydrocarbons by Selenastrum capricornutum. Environ Toxicol Chem 25:1772–1779. https://doi.org/10.1897/05-354R.1
Das N, Chandran P (2011) Microbial degradation of petroleum hydrocarbon contaminants: an overview. Biotechnol Res Int 941810. https://doi.org/10.4061/2011/941810
Ferhan M, Leao AL, de Melo IS et al (2012) Ligninases production and partial purification of mnp from brazilian fungal isolate in submerged fermentation. Ferment Technol 1:106. https://doi.org/10.4172/2167-7972.1000106
Gabriele I, Race M, Papirio S, Esposito G (2021) Phytoremediation of pyrene-contaminated soils: A critical review of the key factors affecting the fate of pyrene. J Environ Manag 293:112805. https://doi.org/10.1016/j.jenvman.2021.112805
Ghosal D, Ghosh S, Dutta TK, Ahn Y (2016) Current state of knowledge in microbial degradation of polycyclic aromatic hydrocarbons (PAHs): a review. Front Microbiol 7:1369. https://doi.org/10.3389/fmicb.2016.01369
Ghosh D, Nagpal S, Bhat MA et al (2015) Gel-free proteomics reveals neoplastic potential in endometrium of infertile patients with stage IV ovarian endometriosis. J Reprod Health Med 1:83–95. https://doi.org/10.1016/j.jrhm.2015.06.003
Hadibarata T, Ayu R (2013) Biodegradation and metabolite transformation of pyrene by basidiomycetes fungal isolate Armillaria sp. F022. Bioprocess Biosyst Eng 36:461–468. https://doi.org/10.1007/s00449-012-0803-4
Hernández-Vega JC, Cady B, Kayanja G et al (2017) Detoxification of polycyclic aromatic hydrocarbons (PAHs) in Arabidopsis thaliana involves a putative flavonol synthase. J Hazard Mater 321:268–280. https://doi.org/10.1016/j.jhazmat.2016.08.058
Hidayat A, Yanto DHY (2018) Biodegradation and metabolic pathway of phenanthrene by a new tropical fungus, Trametes hirsuta D7. J Environ Chem Eng 6:2454–2460. https://doi.org/10.1016/j.jece.2018.03.051
Houshani M, Salehi-Lisar SY, Motafakkerazad R, Movafeghi A (2019) Uptake and distribution of phenanthrene and pyrene in roots and shoots of maize (Zea mays L.). Environ Sci Pollut Res 26:9938–9944. https://doi.org/10.1007/s11356-019-04371-3
Isaac P, Lozada M, Dionisi HM et al (2015) Differential expression of the catabolic nahAc gene and its effect on PAH degradation in Pseudomonas strains isolated from contaminated Patagonian coasts. Int Biodeterior Biodegradation 105:1–6. https://doi.org/10.1016/j.ibiod.2015.08.011
Juhasz AL, Naidu R (2000) Bioremediation of high molecular weight polycyclic aromatic hydrocarbons: a review of the microbial degradation of benzo(a)pyrene. Int Biodeterior Biodegradation 45:57–88. https://doi.org/10.1016/S0964-8305(00)00052-4
Kashyap N, Moholkar VS (2021) Intensification of pyrene degradation by native Candida tropicalis MTCC 184 with sonication: Kinetic and mechanistic investigation. Chem Eng Process Intensif 164:108415. https://doi.org/10.1016/j.cep.2021.108415
Kim S, Kweon O, Jones RC et al (2007) Complete and integrated pyrene degradation pathway in Mycobacterium vanbaalenii PYR-1 Based on Systems Biology. J Bacteriol 189:464–472. https://doi.org/10.1128/JB.01310-06
Kumari M, Ghosh P, Thakur IS (2020) Evaluation of a biosurfactant producing bacterial strain Pseudomonas sp. ISTPY2 for efficient pyrene degradation and landfill soil bioremediation through soil microcosm and proteomic studies. Bioresour Technol Rep 12:100607. https://doi.org/10.1016/j.biteb.2020.100607
Lazim ZM, Hadibarata T (2016) Ligninolytic fungus Polyporus sp. S133 mediated metabolic degradation of fluorene. Braz J Microbiol 47:610–616. https://doi.org/10.1016/j.bjm.2016.04.015
Lee H, Jang Y, Lee YM et al (2015) Enhanced removal of PAHs by Peniophora incarnata and ascertainment of its novel ligninolytic enzyme genes. J Environ Manag 164:10–18. https://doi.org/10.1016/j.jenvman.2015.08.036
Li Z, Cabana H, Lecka J et al (2021) Efficiencies of selected biotreatments for the remediation of PAH in diluted bitumen contaminated soil microcosms. Biodegradation 32:563–576. https://doi.org/10.1007/s10532-021-09952-z
Ling J, Zhang G, Sun H et al (2011) Isolation and characterization of a novel pyrene-degrading Bacillus vallismortis strain JY3A. Sci Total Environ 409:1994–2000. https://doi.org/10.1016/j.scitotenv.2011.02.020
Lyon F (1994) IARC monographs on the evaluation of carcinogenic risks to humans. Some Ind Chem 60:389–433
Nzila A, Ramirez CO, Musa MM et al (2018) Pyrene biodegradation and proteomic analysis in Achromobacter xylosoxidans, PY4 strain. Int Biodeterior Biodegradation 130:40–47. https://doi.org/10.1016/j.ibiod.2018.03.014
Oluseyi T, Olayinka K, Alo B, Smith RM (2011) Comparison of extraction and clean-up techniques for the determination of polycyclic aromatic hydrocarbons in contaminated soil samples. Afr J Environ Sci Technol 5:482–493
Park H, Min B, Jang Y et al (2019) Comprehensive genomic and transcriptomic analysis of polycyclic aromatic hydrocarbon degradation by a mycoremediation fungus, Dentipellis sp. KUC8613. Appl Microbiol Biotechnol 103:8145–8155. https://doi.org/10.1007/s00253-019-10089-6
Pozdnyakova NN (2012) Involvement of the ligninolytic system of white-rot and litter-decomposing fungi in the degradation of polycyclic aromatic hydrocarbons. Biotechnol Res Int. 243217. https://doi.org/10.1155/2012/243217
Ravelet C, Krivobok S, Sage L, Steiman R (2000) Biodegradation of pyrene by sediment fungi. Chemosphere 40:557–563. https://doi.org/10.1016/S0045-6535(99)00320-3
Saraswathy A, Hallberg R (2005) Mycelial pellet formation by Penicillium ochrochloron species due to exposure to pyrene. Microbiol Res 160:375–383. https://doi.org/10.1016/j.micres.2005.03.001
Saraswathy A, Hallberg R (2002) Degradation of pyrene by indigenous fungi from a former gasworks site. FEMS Microbiol Lett 210:227–232. https://doi.org/10.1111/j.1574-6968.2002.tb11185.x
Suman SK, Khatri M, Dhawaria M et al (2018) Potential of Trametes maxima IIPLC-32 derived laccase for the detoxification of phenolic inhibitors in lignocellulosic biomass prehydrolysate. Int Biodeterior Biodegradation 133:1–8. https://doi.org/10.1016/j.ibiod.2018.05.009
Vasconcelos MRS, Vieira GAL, Otero IVR et al (2019) Pyrene degradation by marine-derived ascomycete: process optimization, toxicity, and metabolic analyses. Environ Sci Pollut Res 26:12412–12424. https://doi.org/10.1007/s11356-019-04518-2
Velmurugan N, Lee H, Cha H, Lee Y (2017) Proteomic analysis of the marine-derived fungus Paecilomyces sp. strain SF-8 in response to polycyclic aromatic hydrocarbons. Bot Mar 60:381–392. https://doi.org/10.1515/bot-2016-0101
Wei K, Yin H, Peng H et al (2017) Characteristics and proteomic analysis of pyrene degradation by Brevibacillus brevis in liquid medium. Chemosphere 178:80–87. https://doi.org/10.1016/j.chemosphere.2017.03.049
Wen J, Gao D, Zhang B, Liang H (2011) Co-metabolic degradation of pyrene by indigenous white-rot fungus Pseudotrametes gibbosa from the northeast China. Int Biodeterior Biodegradation 65:600–604. https://doi.org/10.1016/j.ibiod.2011.03.003
Wu M, Guo X, Wu J, Chen K (2020) Effect of compost amendment and bioaugmentation on PAH degradation and microbial community shifting in petroleum-contaminated soil. Chemosphere 256:126998. https://doi.org/10.1016/j.chemosphere.2020.126998
Yuan H, Yao J, Masakorala K, Wang F (2014) Isolation and characterization of a newly isolated pyrene-degrading Acinetobacter strain USTB-X. Environ Sci Pollut Res 21:2724–2732. https://doi.org/10.1007/s11356-013-2221-9
Zeneli A, Kastanaki E, Simantiraki F, Gidarakos E (2019) Monitoring the biodegradation of TPH and PAHs in refinery solid waste by biostimulation and bioaugmentation. J Environ Chem Eng 7:103054. https://doi.org/10.1016/j.jece.2019.103054
Zhang H, Zhang S, He F et al (2016) Characterization of a manganese peroxidase from white-rot fungus Trametes sp.48424 with strong ability of degrading different types of dyes and polycyclic aromatic hydrocarbons. J Hazard Mater 320:265–277. https://doi.org/10.1016/j.jhazmat.2016.07.065
Zhou H, Li X, Hu B et al (2021) Assembly of fungal mycelium-carbon nanotube composites and their application in pyrene removal. J Hazard Mater 415:125743. https://doi.org/10.1016/j.jhazmat.2021.125743
Zhu Y, Chen K, Ding Y et al (2019) Metabolic and proteomic mechanism of benzo(a)pyrene degradation by Brevibacillus brevis. Ecotoxicol Environ Saf 172:1–10. https://doi.org/10.1016/j.ecoenv.2019.01.044
Acknowledgements
The authors are highly thankful to Dr. Vijay Tak, Scientist at Defence Research and Development Establishment (Gwalior, India), and Mr. Saurabh Nagpal, Agilent Technologies (Gurugram, India) for LC-MSn analysis.
Funding
This study was supported by the Council of Scientific and Industrial Research (CSIR), New Delhi, India, under the project OLP-1094; INSPIRE program, Department of Science and Technology (New Delhi, India) and Academy of Scientific and Innovative Research (Ghaziabad, India) provided fellowship.
Author information
Authors and Affiliations
Contributions
AI contributed to conceptualization, writing—original draft, and formal analysis; SKS contributed to writing—reviewing and editing, methodology, and validation; BPV and DT performed formal analysis; AR was involved in conceptualization, visualization, and funding acquisition; PKK contributed to conceptualization, resources, investigation, writing—reviewing and editing, and supervision. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
Not applicable
Consent to participate
Not applicable
Consent for publication
All authors have given consent for publication
Competing interests
No competing interest
Additional information
Responsible Editor: Robert Duran
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Imam, A., Suman, S.K., Vempatapu, B.P. et al. Pyrene remediation by Trametes maxima: an insight into secretome response and degradation pathway. Environ Sci Pollut Res 29, 44135–44147 (2022). https://doi.org/10.1007/s11356-022-18888-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-18888-7