Abstract
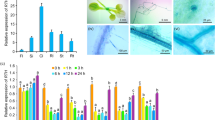
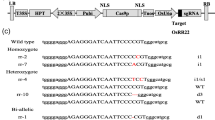
The group 5 LEA (late embryogenesis abundant) proteins are an atypical LEA protein group, which is associated with resistance to multiple stresses. In this study, OsLea14-A gene was isolated from Oryza sativa L., which encodes a 5C LEA protein with 151 amino acids. The qPCR analysis showed that OsLea14-A expressed in all tissues and organs at all times. The expression of OsLea14-A in the panicles of plumping stage were dramatically increased. The heterologous expression of OsLea14-A in Escherichia coli improved its growth performance under salinity, desiccation, high temperature, and freeze-thaw stresses. The purified OsLea14-A protein can protect LDH activity from freeze-thaw-, heat-, and desiccation-induced inactivation. The overexpression of OsLea14-A in rice improved tolerance to dehydration, high salinity, CuSO4, and HgCl2, but excluding K2Cr2O7. The analysis of metal contents showed that the accumulation of OsLea14-A protein in transgenic rice could increase the accumulation of Hg, but could not increase the accumulation of Na, Cr, and Cu after HgCl2, NaCl, K2Cr2O7, and CuSO4 treatment, respectively. These results suggested that OsLea14-A conferred multiple stress tolerance and Hg accumulation, which made it a possible gene in genetic improvement for plants to acclimatize itself to multiple stresses and remediate Hg-contaminated soil.








Similar content being viewed by others
References
Amara I, Capellades M, Ludevid MD, Pagès M, Goday A (2013) Enhanced water stress tolerance of transgenic maize plants over-expressing LEA Rab28 gene. J Plant Physiol 170:864–873
Battaglia M, Olvera-Carrillo Y, Garciarrubio A, Campos F, Covarrubias AA (2008) The enigmatic LEA proteins and other hydrophilins. Plant Physiol 148:6–24
Boucher V, Buitink J, Lin X, Boudet J, Hoekstra FA, Hundertmark M, Renard D, Leprince O (2010) MtPM25 is an atypical hydrophobic late embryogenesis-abundant protein that dissociates cold and desiccation-aggregated proteins. Plant Cell Environ 33:418–430
Bradford NM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cao Y, Xiang X, Geng M, You Q, Huang X (2017) Effect of HbDHN1 and HbDHN2 genes on abiotic stress responses in Arabidopsis. Front Plant Sci 8:470
Ciccarelli FD, Bork P (2005) The WHy domain mediates the response to desiccation in plants and bacteria. Bioinformatics 21:1304–1307
Drira M, Saibi W, Brini F, Gargouri A, Masmoudi K, Hanin M (2013) The K-segments of the wheat dehydrin DHN-5 are essential for the protection of lactate dehydrogenase and β-glucosidase activities in vitro. Mol Biotechnol 54:643–650
French-Pacheco L, Cuevas-Velazquez CL, Rivillas-Acevedo L, Covarrubias AA, Amero C (2018) Metal-binding polymorphism in late embryogenesis abundant protein AtLEA4-5, an intrinsically disordered protein. Peer J 6:e4930
Fuxreiter M, Simon I, Friedrich P, Tompa P (2004) Preformed structural elements feature in partner recognition by intrinsically unstructured proteins. J Mol Biol 338:1015–1026
Gao J, Lan T (2016) Functional characterization of the late embryogenesis abundant (LEA) protein gene family from Pinus tabuliformis (Pinaceae) in Escherichia coli. Sci Rep 6:19467
Hara M, Monna S, Murata T, Nakano T, Amano S, Nachbar M, Wätzig H (2016) The Arabidopsis KS-type dehydrin recovers lactate dehydrogenase activity inhibited by copper with the contribution of His residues. Plant Sci 245:135–142
He S, Tan L, Hu Z, Chen G, Wang G, Hu T (2012) Molecular characterization and functional analysis by heterologous expression in E. coli under diverse abiotic stresses for OsLEA5, the atypical hydrophobic LEA protein from Oryza sativa L. Mol Gen Genomics 287:39–54
Hirayama T, Shinozaki K (2010) Research on plant abiotic stress responses in the post-genome era: past, present and future. Plant J 61:1041–1052
Hofgen R, Willmitzer L (1988) Storage of competent cells for Agrobacterium transformation. Nucleic Acids Res 16:9877
Hu TZ (2008) OsLEA3, a late embryogenesis abundant protein gene from rice, confers tolerance to water deficit and salt stress to transgenic rice. Russ J Plant Physiol 55:530–537
Hunault G, Jaspard E (2010) LEAPdb: a database for the late embryogenesis abundant proteins. BMC Genomics 11:221
Hundertmark M, Hincha DK (2008) LEA (late embryogenesis abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genomics 9:118
Jaspard E, Hunault G (2014) Comparison of amino acids physico-chemical properties and usage of late embryogenesis abundant proteins, hydrophilins and WHy domain. PLoS One 9:e109570
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202
Kim HS, Lee JH, Kim JJ, Kim CH, Jun SS, Hong YN (2005) Molecular and functional characterization of CaLEA6, the gene for a hydrophobic LEA protein from Capsicum annuum. Gene 344:115–123
Kovacs D, Kalmar E, Torok Z, Tompa P (2008) Chaperone activity of ERD10 and ERD14, two disordered stress-related plant proteins. Plant Physiol 147:381–390
Krüger C, Berkowitz O, Stephan UW, Hell R (2002) A metal-binding member of the late embryogenesis abundant protein family transports iron in the phloem of Ricinus communis L. J Biol Chem 277:25062–25069
Kyte J, Dolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132
Liang J, Zhou M, Zhou X, Jin Y, Xu M, Lin J (2013) JcLEA, a novel LEA-like protein from Jatropha curcas, confers a high level of tolerance to dehydration and salinity in Arabidopsis thaliana. PLoS One 8:e83056
Liang Y, Xiong Z, Zheng J, Xu D, Zhu Z, Xiang J, Gan J, Raboanatahiry N, Yin Y, Li M (2016) Genome-wide identification, structural analysis and new insights into late embryogenesis abundant (LEA) gene family formation pattern in Brassica napus. Sci Rep 6:24265
Linding R, Jensen LJ, Diella F, Bork P, Gibson TJ, Russell RB (2003) Protein disorder prediction: implications for structural proteomics. Structure 11:1453–1459
Liu G, Xu H, Zhang L, Zheng Y (2011) Fe binding properties of two soybean (Glycine max L.) LEA4 proteins associated with antioxidant activity. Plant Cell Physiol 52:994–1002
Liu Y, Wang L, Xing X, Sun L, Pan J, Kong X, Zhang M, Li D (2013) ZmLEA3, a multifunctional group 3 LEA protein from maize (Zea mays L.), is involved in biotic and abiotic stresses. Plant Cell Physiol 54:944–959
Liu Y, Wang L, Jiang S, Pan J, Cai G, Li D (2014) Group 5 LEA protein, ZmLEA5C, enhance tolerance to osmotic and low temperature stresses in transgenic tobacco and yeast. Plant Physiol Biochem 84:22–31
Liu Y, Xie L, Liang X, Zhang S (2015) CpLEA5, the late embryogenesis abundant protein gene from Chimonanthus praecox, possesses low temperature and osmotic resistances in prokaryote and eukaryotes. Int J Mol Sci 16:26978–26990
Magwanga RO, Lu P, Kirungu JN, Lu H, Wang X, Cai X, Zhou Z, Zhang Z, Salih H, Wang K, Liu F (2018) Characterization of the late embryogenesis abundant (LEA) proteins family and their role in drought stress tolerance in upland cotton. BMC Genet 19:6
Mertens J, Aliyu H, Cowan DA (2018) LEA proteins and the evolution of the WHy domain. Appl Environ Microbiol 84:e00539–e00518
Mowla SB, Cuypers A, Driscoll SP, Kiddle G, Thomson J, Foyer CH, Theodoulou FL (2006) Yeast complementation reveals a role for an Arabidopsis thaliana late embryogenesis abundant (LEA)-like protein in oxidative stress tolerance. Plant J 48:743–756
Park SC, Kim YH, Jeong JC, Kim CY, Lee H-S, Bang JW, Kwas SS (2011) Sweetpotato late embryogenesis abundant 14 (IbLEA14) gene influences lignification and increases osmotic- and salt stress-tolerance of transgenic calli. Planta 233:621–634
Patil A, Nakamura H (2006) Disordered domains and high surface charge confer hubs with the ability to interact with multiple proteins in interaction networks. FEBS Lett 580:2041–2045
Rodriguez-Salazar J, Moreno S, Espín G (2017) LEA proteins are involved in cyst desiccation resistance and other abiotic stresses in Azotobacter vinelandii. Cell Stress Chaperones 22:397–408
Saitu N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic tress. Mol Biol Evol 4:406–425
Sasaki K, Christov NK, Tsuda S, Imai R (2014) Identification of a novel LEA protein involved in freezing tolerance in wheat. Plant Cell Physiol 55:136–147
Sharma A, Kumar D, Kumar S, Rampuria S, Reddy AR, Kirti PB (2016) Ectopic expression of an atypical hydrophobic group 5 LEA protein from wild peanut, Arachis diogoi confers abiotic stress tolerance in tobacco. PLoS One 11:e0150609
Shi J, Liu M, Chen Y, Wang J, Lu C (2016) Heterologous expression of the dehydrin-like protein gene AmCIP from Ammopiptanthus mongolicus enhances viability of Escherichia coli and tobacco under cold stress. Plant Growth Regul 79:71–80
Singh S, Cornilescu CC, Tyler RC, Cornilescu G, Tonelli M, Lee MS, Markley JL (2005) Solution structure of a late embryogenesis abundant protein (LEA14) from Arabidopsis thaliana, a cellular stress-related protein. Protein Sci 14:2601–2609
Tunnacliffe A, Wise MJ (2007) The continuing conundrum of the LEA proteins. Naturwissenschaften 94:791–812
Wang M, Li P, Li C, Pan Y, Jiang X, Zhu D, Zhao Q, Yu J (2014) SiLEA14, a novel atypical LEA protein, confers abiotic stress resistance in foxtail millet. BMC Plant Biol 14:290
Wang X, Zhang L, Zhang Y, Bai Z, Liu H, Zhang D (2017) Triticum aestivum WRAB18 functions in plastids and confers abiotic stress tolerance when overexpressed in Escherichia coli and Nicotiania benthamiana. PLoS One 12:e0171340
Yancey PH (2005) Organic osmolytes as compatible, metabolic and counteracting cytoprotectants in high osmolarity and other stresses. J Exp Biol 208:2819–2830
Yang W, Zhang L, Lv H, Li H, Zhang Y, Xu Y, Yu J (2015) The K-segments of wheat dehydrin WZY2 are essential for its protective functions under temperature stress. Front Plant Sci 6:406
Zhang X, Lu S, Jiang C, Wang Y, Lv B, Shen J, Ming F (2014) RcLEA, a late embryogenesis abundant protein gene isolated from Rosa chinensis, confers tolerance to Escherichia coli and Arabidopsis thaliana and stabilizes enzyme activity under diverse stresses. Plant Mol Biol 85:333–347
Zhou Y, He P, Xu Y, Liu Q, Yang Y, Liu S (2017) Overexpression of CsLEA11, a Y3SK2-type dehydrin gene from cucumber (Cucumis sativus), enhances tolerance to heat and cold in Escherichia coli. AMB Express 7:182
Funding
This work was funded by the Basic and Advanced Research Project of Chongqing CSTC (cstc2018jcyjAX0665) and the Natural Science Foundation Project of Wanzhou District (201403063).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Gangrong Shi
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hu, T., Liu, Y., Zhu, S. et al. Overexpression of OsLea14-A improves the tolerance of rice and increases Hg accumulation under diverse stresses. Environ Sci Pollut Res 26, 10537–10551 (2019). https://doi.org/10.1007/s11356-019-04464-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-019-04464-z