Abstract
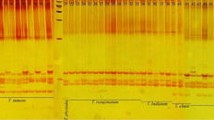
No information is available on the transferability and amplification quality of microsatellite (SSR) markers of the public domain inBrassica carinata A. Braun. The objective of the presented research was to study the amplification of a set of 73 SSRs fromB. nigra (L.) Koch andB. napus L. inB. carinata, and to compare the results with those obtained in the amplification of the same markers in otherBrassica species of the U triangle. This set of SSRs fromB. nigra (B genome) andB. napus (AC genome) allows the identification of the 3 basic genomes of theBrassica species tested. 94.3% of the SSR markers fromB. nigra and 97.4% of those fromB. napus amplified SSR-specific products inB. carinata. Very high-quality amplification with a strong signal and easy scoring inB. carinata was recorded for 52.8% of the specific loci fromB. nigra SSRs and 59.3% of the specific loci fromB. napus SSRs, compared to 66.7% inB. nigra and 62.8% inB. napus. Genome specificity and amplification quality ofB. nigra andB. napus SSR markers in the 6 species under study is reported. High-quality transferable SSR markers provide an efficient and cost-effective platform to advance in molecular research inB. carinata.
Similar content being viewed by others
References
Bornet B, Branchard M, 2004. Use of ISSR fingerprints to detect microsatellites and genetic diversity in several relatedBrassica taxa andArabidopsis thaliana. Hereditas 140: 245–248.
Batley J, Hopkins CJ, Cogan NOI, Hand M, Jewell E, Kaur J, et al. 2007. Identification and characterization of simple sequence repeat markers fromBrassica napus expressed sequences. 7: 886–889.
Berry ST, Leon AJ, Hanfrey CC, Challis P, Burkholz SR, Barnes GK, et al. 1995. Molecular markers analysis ofHelianthus annuus L. 2. Construction of aRFLP linkage map for cultivated sunflower. Theor Appl Genet 91: 195–199.
Burgess B, Mountford H, Hopkins CJ, Love C, Ling A, Spangenberg G, et al. 2006. Identification and characterization of simple sequence repeat (SSR) markers derivedin silico fromBrassica oleracea genome shotgun sequences.Mol Ecol Notes 6: 1191–1194.
Dice LR, 1945. Measures of the amount of ecologic association between species. Ecology 26: 297–302.
Genet T, Viljoen CD, Labuschagne MT, 2005. Genetic analysis of Ethiopian mustard genotypes using amplified fragment length polymorphism (AFLP) markers. Afr J Biotechnol 4: 891–897.
Gutierrez MV, Vaz Patto MC, Huguet T, Cubero JI, Moreno MT, Torres AM, 2005. Cross-species amplification ofMedicago truncatula microsatellites across three major pulse crops. Theor Appl Genet 110: 1210–1217.
Hasan M, Seyis F, Badani AG, Pons-Kühnemann J, Friedt W, Lühs W, et al. 2006. Analysis of genetic diversity in theBrassica napus L. gene pool using SSR markers. Genet Res Crop Evol 53: 793–802.
Hopkins CJ, Cogan NOI, Hand M, Jewell E, Kaur J, Li X, et al. 2007. Sixteen new simple sequence repeat markers fromBrassica juncea expressed sequences and their cross-species amplification. 7: 697–700.
Lowe AJ, Jones AE, Raybould AF, Trick M, Moule CL, Edwards KJ, 2002. Transferability and genome specificity of a new set of microsatellite primers amongBrassica species of the U triangle. Mol Ecol Not 2: 711.
Lowe AJ, Moule C, Trick M, Edwards KJ, 2004. Efficient large-scale development of microsatellites for marker and mapping applications inBrassica crop species. Theor Appl Genet 108: 1103–1112.
Mantel NA, 1967. The detection of disease clustering and a generalized regression approach. Cancer Res 27: 209–220.
Plieske J, Struss D, 2001. Microsatellite markers for genome analysis inBrassica. I. Development inBrassica napus and abundance inBrassicaceae species. Theor Appl Genet 102: 689–694.
Pradhan AK, Prakash S, Mukhopadhyay A, Pental D, 1992. Phylogeny ofBrassica and allied genera based on variation in chloroplast and mitochondrial DNA patterns: molecular and taxonomic classifications are incongruous. Theor Appl Genet 85: 331–340.
Rohlf FJ, 1998. NTSYS-pc numerical taxonomy and multivariate analysis system, version 2.02. Exeter Software, Setauket, New York, USA: Exeter Software.
Suwabe K, Iketani H, Nunome T, Kage T, Hirai M, 2002. Isolation and characterization of microsatellites inBrassica rapa L. Theor Appl Genet 93: 534–538.
Teklewold A, Becker HC, 2006. Geographic pattern of genetic diversity among 43 Ethiopian mustardBrassica carinata (A. Braun) accessions as revealed by RAPD analysis. Genet Res Crop Evol 53: 1173–1185.
Tsunoda S, 1980. Eco-physiology of wild and cultivated forms inBrassica and allied genera. In: Tsunoda S, Hinata K, Gómez-Campo C, eds.Brassica crops and wild allies, Tokyo: Japan Scientific Societies Press: 109–120.
U N, 1935. Genomic analysis ofBrassica with special reference to the experimental formation ofB. napus and its peculiar mode of fertilization. Jpn J Bot 7: 389–452.
Warwick SI, Black LD, 1991. Molecular systematics ofBrassica and allied genera (subtribe Brassicinae, Brassiceae) chloroplast genome and cytodeme congruence. Theor Appl Genet 82: 81–92.
Warwick SI, Gugel RK, McDonald T, Falk KC, 2006. Genetic variation of Ethiopian mustard (Brassica carinata A. Braun) germplasm in western Canada. Genet Res Crop Evol 53: 297–312.
Westman AL, Kresovich S, 1998. The potential for cross-taxa simple-sequence repeat (SSR) amplification betweenArabidopsis thaliana L. and crop brassicas. Theor Appl Genet 96: 272–281.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Márquez-Lema, A., Velasco, L. & Pérez-Vich, B. Transferability, amplification quality, and genome specificity of microsatellites inBrassica carinata and related species. J Appl Genet 51, 123–131 (2010). https://doi.org/10.1007/BF03195720
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF03195720