Summary
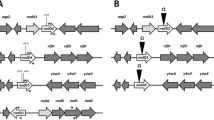
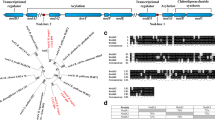
The nucleotide sequence of the nod box locus n4 in Rhizobium meliloti was determined and revealed six genes organized in a single transcriptional unit, which are induced in response to a plant signal such as luteolin. Mutations in these genes influence the early steps of nodule development on Medicago, but have no detectable effect on Melilotus, another host for R. meliloti. Based on sequence homology, the first open reading frame (ORF) corresponds to the nodM gene and the last to the nodN gene of Rhizobium leguminosarum. The others do not exhibit similarity to any genes sequenced so far, so we designated them as nolF, nolG, nolH and nolI, respectively. We found that the n4 locus, and especially the nodM and nodN genes, are involved in the production of the root hair deformation (Had) factor. NodM exhibits homology to amidotransferases, primarily to the d-glucosamine synthetase encoded by the glmS gene of Escherichia coli. We demonstrated that in E. coli the regulatory gene nodD together with luteolin can activate nod genes. On this basis we showed that nodM complemented an E. coli glmS −mutation, indicating that nodM can be considered as a glmS gene under plant signal control. Moreover, exogenously supplied d-glucosamine restored nodulation of Medicago by nodM mutants. Our data suggest that in addition to the housekeeping glmS gene of R. meliloti, nodM as a second glmS copy provides glucosamine in sufficient amounts for the synthesis of the Had factor.
Similar content being viewed by others
References
Aguilar OM, Reilander H, Arnold W, Pühler A (1987) Rhizobium meliloti nifN (fixF) gene is part of an operon regulated by a nifA dependent promoter and codes for a polypeptide homologous to the nifK gene product. J Bacteriol 169:5393–5400
Banfalvi Z, Kondorosi A (1989) Production of root hair deformation factors by Rhizobium meliloti nodulation genes in Escherichia coli:hsnD (nodH) is involved in the plant host specific modification of the nodABC factor. Plant Mol Biol 13:1–12
Banfalvi Z, Randhawa GS, Kondorosi E, Kiss A, Kondorosi A (1983) Construction and characterization of R prime plasmids carrying symbiotic genes of Rhizobium meliloti. Mol Gen Genet 189:129–135
Berg DE, Weiss A, Crossland L (1980) Polarity of Tn5 insertion mutations in Escherichia coli. J Bacteriol 142:439–446
Beringer JE, Beynon JL, Buchanan-Wollaston AV, Johnston AWB (1978) Transfer of the drug resistance transposon Tn5 to Rhizobium. Nature 276:633–634
Casadaban MJ (1975) Fusion of the Escherichia coli lac genes to the ara promoter: A general technique using bacteriophage Mu-1 insertions. Proc Natl Acad Sci USA 72:809–813
Casadaban MJ (1976) Transposition and fusion of the lac genes to selected promoters in Escherichia coli using bacteriophage lambda and Mu. J Mol Biol 104:541–555
Cervantes E, Sharma SB, Maillet F, Vasse J, Truchet G, Rosenberg C (1989) The Rhizobium meliloti host range nodQ gene encodes a protein which shares homology with translation elongation and initiation factors. Mol Microbiol 3:745–755
Chang ACY, Cohen SN (1978) Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the p15A cryptic miniplasmid. J Bacteriol 134:1141–1156
Corbin D, Barran L, Ditta G (1983) Organization and expression of Rhizobium meliloti nitrogen fixation genes. Proc Natl Acad Sci USA 80:3005–3009
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res 16:10881–10890
Debellé F, Rosenberg C, Vasse J, Maillet F, Martinez E, Dénarié J, Truchet G (1986) Assignment of symbiotic developmental phenotypes to common and specific nodulation (nod) genetic loci of Rhizobium meliloti. J Bacteriol 168:1075–1086
de Bruijn FJ, Lupski JR (1984) The use of transposon Tn5 mutagenesis in the rapid generation of correlated physical and genetic map of DNA segments cloned into multicopy plasmids — a review. Gene 27:131–149
Ditta G, Stanfield S, Corbin D, Helinski DR (1980) Broad host range DNA cloning system for gram-negative bacteria: construction of a gene bank of Rhizobium meliloti. Proc Natl Acad Sci USA 77:7347–7351
Downie JA (1989) The nodL gene from Rhizobium leguminosarum is homologous to the acetyl transferases encoded by lacA and cysE. Mol Microbiol 3:1649–1651
Economou A, Hamilton WDO, Johnston AWB, Downie JA (1990) The Rhizobium nodulation gene nodO encodes a Ca2+-binding protein that is exported without N-terminal cleavage and is homologous to haemolysin and related proteins. EMBO J 9:349–354
Faucher C, Maillet F, Vasse J, Rosenberg C, Van Brussel AAN, Truchet G, Dénarié J (1988) Rhizobium meliloti host range nodes gene determines production of an alfalfa-specific extracellular signal. J Bacteriol 170:5489–5499
Fickett JW (1982) Recognition of protein coding regions in DNA sequences. Nucleic Acids Res 10:5302–5318
Figurski DH, Helinski DR (1979) Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci USA 76:1648–1652
Fisher F, Egelhoff TT, Mulligan JT, Long SR (1988) Specific binding of proteins from Rhizobium meliloti cell-free extracts containing nodD to DNA sequences upstream of inducible nodulation genes. Genes Dev 2:282–293
Gerhold D, Stacey G, Kondorosi A (1989) Use of a promoter-specific probe to identify two loci from the Rhizobium meliloti nodulation regulon. Plant Mol Biol 12:181–188
Ghosh S, Blumental HJ, Davidson E, Roseman S (1960) Glucosamine metabolism V Enzymatic synthesis of glucosamine-6-phosphate. J Biol Chem 235:1265–1273
Henikoff S (1987) Unidirectional digestion with Exonuclease III in DNA sequence analysis. Methods Enzymol 155:156–165
Hong GF, Burn JE, Johnston AWB (1987) Evidence that DNA involved in the expression of nodulation (nod) genes in Rhizobium binds to the product of the regulatory gene nodD. Nucleic Acids Res 15:9677–9689
Horvath B, Kondorosi E, John M, Schmidt J, Török I, Györgypal Z, Barabas I, Wieneke U, Schell J, Kondorosi A (1986) Organization, structure and symbiotic function of Rhizobium meliloti nodulation genes determining host specificity for alfalfa. Cell 46:335–343
Horvath B, Bachem C, Schell J, Kondorosi A (1987) Host specific regulation of nodulation genes in Rhizobium is mediated by a plant signal, interacting with the nodD gene product. EMBO J 6:841–848
Kondorosi A, Svab Z, Kiss GB, Dixon RA (1977) Ammonia assimilation and nitrogen fixation in Rhizobium meliloti. Mol Gen Genet 151:221–226
Kondorosi E, Banfalvi Z, Kondorosi A (1984) Physical and genetic analysis of a symbiotic region of Rhizobium meliloti: Identification of nodulation genes. Mol Gen Genet 193:445–452
Kondorosi E, Gyuris J, Schmidt J, John M, Duda E, Hoffmann B, Schell J, Kondorosi A (1989) Positive and negative control of nod gene expression in Rhizobium meliloti is required for optimal nodulation. EMBO J 8:1331–1340
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lerouge P, Roche P, Faucher C, Maillet F, Truchet G, Prome JC, Dénarié J (1990) Symbiotic host-specificity of Rhizobium meliloti is determined by a sulphated and acylated oligosaccharide signal. Nature 344:781–784
Long SR (1989) Rhizobium-legume nodulation: Life together in the underground. Cell 56:203–214
Maentsaelae P, Zalkin H (1984) Glutamine nucleotide sequence of Saccharomyces cerevisiae ADE4 encoding phosphoribosyl-pyrophosphate amidotransferase. J Biol Chem 259:8478–8484
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: A laboratory manual, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Mount DW, Conrad B (1984) Microcomputer programs for graphic analysis of nucleic acid and protein sequences. Nucleic Acids Res 12:811–817
Mulligan JT, Long SR (1985) Induction of Rhizobium meliloti nodC expression by plant exudate requires nodD. Proc Natl Acad Sci USA 82:6609–6613
Petersen GB, Stockwell PA, Hill DF (1988) Messenger RNA recognition in Escherichia coli: a possible second site of interaction with 16S ribosomal RNA. EMBO J 7:3957–3962
Putnoky P, Kondorosi A (1986) Two gene clusters of Rhizobium meliloti code for early essential nodulation functions and a third influences nodulation efficiency. J Bacteriol 167:881–887
Ratet P, Schell J, de Bruijn FJ (1988) Mini-Mu lac transposons with broad-host-range origins of conjugal transfer and replication designed for gene regulation studies in Rhizobiaceae. Gene 63:41–52
Rose RE (1988) The nucleotide sequence of pACYC184. Nucleic Acids Res 16:355
Rostas K, Kondorosi E, Horvath B, Simoncsits A, Kondorosi A (1986) Conservation of extended promoter regions of nodulation genes in Rhizobium. Proc Natl Acad Sci USA 83:1757–1761
Ruvkun GB, Ausubel FM (1981) A general method for site-directed mutagenesis in procaryotes. Nature 289:85–88
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Schmidt J, John M, Kondorosi E, Kondorosi A, Wieneke U, Schröder G, Schröder J, Schell J (1984) Mapping of the proteincoding regions of Rhizobium meliloti common nodulation genes. EMBO J 3:1705–1711
Schmidt J, Wingender R, John M, Wieneke U, Schell J (1988) Rhizobium meliloti nodA and nodB genes are involved in generating compounds that stimulate mitosis of plant cells. Proc Natl Acad Sci USA 8:8578–8582
Schwedock J, Long SR (1990) ATP sulphurylase activity of the nodP and nodQ gene products of Rhizobium meliloti. Nature 348:644–647
Shearman CA, Rossen L, Johnston AWB, Downie JA (1986) The Rhizobium leguminosarum nodulation gene nodF encodes a polypeptide similar to acyl-carrier protein and its regulated by nodD plus a factor in pea root exudate. EMBO J 5:647–652
Shine J, Dalgarno L (1974) The 3′-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci USA 71:1341–1346
Spaink HP, Wijffelman CA, Pees E, Okker RJH, Lugtenberg BJJ (1987) Rhizobium nodulation gene nodD as a determinant of host specificity. Nature 328:337–340
Surin BP, Downie JA (1988) Characterization of the Rhizobium leguminosarum genes nodLMN involved in efficient host-specific nodulation. Mol Microbiol 2:173–183
Tso JY, Salkin H, pan Cleemput M, Yanofsky C, Smith JM (1982) Nucleotide sequence of Escherichia coli purF and deduced amino acid sequence of glutamine phosphoribosylpyrophosphate amidotransferase. J Biol Chem 257:3525–3531
Vollmer SJ, Switzer RL, Hermodson MA, Bower SG, Zalkin H (1983) The glutamine-utilizing site of Bacillus subtilis glutamine phosphoribosylpyrophosphate amidotransferase. J Biol Chem 258:10582–10585
Von Heijne G (1986) A new method for predicting signal sequence cleavage sites. Nucleic Acids Res 14:4683–4690
Walker JF, Gay NJ, Saraste M, Eberle AN (1984) DNA sequence around the Escherichia coli unc operon. Biochem J 224:799–815
Wu HC, Wu TC (1971) Isolation and characterization of a glucosamine-requiring mutant of Escherichia coli K12 defective in glucosamine-6-phosphate synthetase. J Bacteriol 105:455–466
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequence of the M13mp18 and pUC19 vectors. Gene 33:103–119
Zaat SAJ, Van Brussel AAN, Tak T, Pees E, Lugtenberg BJJ (1987) Induction of the promoter of Rhizobium leguminosarum sym plasmid pRL1JI by plant flavanones and flavones. J Bacteriol 169:3388–3391
Author information
Authors and Affiliations
Additional information
Communicated by J. Schell
Rights and permissions
About this article
Cite this article
Baev, N., Endre, G., Petrovics, G. et al. Six nodulation genes of nod box locus 4 in Rhizobium meliloti are involved in nodulation signal production: nodM codes for d-glucosamine synthetase. Molec. Gen. Genet. 228, 113–124 (1991). https://doi.org/10.1007/BF00282455
Issue Date:
DOI: https://doi.org/10.1007/BF00282455