Abstract
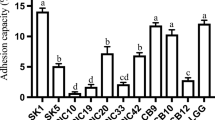
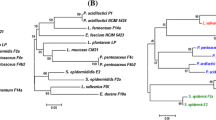
The objective of this study was to isolate, identify, and assess the safety and functionality in vitro of putative probiotic bacterial strains. Isolation procedures were based on standard methods using elective and selective media. The isolates were identified by comparative 16S rRNA sequencing analysis while their safety was determined according to the safety tests recommended by the FAO/WHO such as antibiotic resistance, hemolysin, and biogenic amine production. Most of the isolates did not pass the in vitro safety tests; therefore, only Lactiplantibacillus plantarum (from ant intestine and cheese), Lacticaseibacillus paracasei (from goat milk and kimchi), Enterococcus faecium (from chili doenjang and vegetables with kimchi ingredients), Limosilactobacillus fermentum (from saliva), and Companilactobacillus alimentarius (from kimchi) were identified and selected for further studies. The isolates were further differentiated by rep-PCR and identified to the strain level by genotypic (16S rRNA) and phenotypic (Gen III) approaches. Subsequently, the strain tolerance to acid and bile was evaluated resulting in good viability after simulated gastrointestinal tract passage. Adhesion to mucin in vitro and the presence of mub, mapA, and ef-tu genes confirmed the adhesive potential of the strains and the results of features associated with adhesion such as hydrophobicity and zeta potential extended the insights. This study reflects the importance of fermented and non-fermented food products as a promising source of lactic acid bacteria with potential probiotic properties. Additionally, it aims to highlight the challenges associated with the selection of safe strains, which often fail in the in vitro tests, thus hindering the possibilities of “uncovering” novel and safe probiotic strains.



Similar content being viewed by others
Data availability
All data generated related to this project is available upon request from the authors of the manuscript.
References
Alp D, Kuleaşan H (2019) Adhesion mechanisms of lactic acid bacteria: conventional and novel approaches for testing. World J Microbiol Biotechnol 35:156. https://doi.org/10.1007/s11274-019-2730-x
Amaral DMF, Silva LF, Casarotti SN et al (2017) Enterococcus faecium and Enterococcus durans isolated from cheese: survival in the presence of medications under simulated gastrointestinal conditions and adhesion properties. J Dairy Sci 100:933–949. https://doi.org/10.3168/jds.2016-11513
Arellano-Ayala K, Ascencio-Valle FJ, Gutiérrez-González P et al (2020) Hydrophobic and adhesive patterns of lactic acid bacteria and their antagonism against foodborne pathogens on tomato surface (Solanum lycopersicum L.). J Appl Microbiol 129:876–891. https://doi.org/10.1111/jam.14672
Audisio MC, Torres MJ, Sabaté DC et al (2011) Properties of different lactic acid bacteria isolated from Apis mellifera L. bee-gut. Microbiol Res 166:1–13. https://doi.org/10.1016/j.micres.2010.01.003
Aziz K, Tariq M, Zaidi A (2019) Biofilm development in L. fermentum under shear flow & sequential GIT digestion. FEMS Microbiol Lett 366:1–10. https://doi.org/10.1093/femsle/fnz064
Bhakta JN, Bhattacharya S, Lahiri S, Panigrahi AK (2023) Probiotic characterization of arsenic-resistant lactic acid bacteria for possible application as arsenic bioremediation tool in fish for safe fish food production. Probiot Antimicrob Proteins 15:889–902. https://doi.org/10.1007/s12602-022-09921-9
Bover-Cid S, Holzapfel WH (1999) Improved screening procedure for biogenic amine production by lactic acid bacteria. Int J Food Microbiol 53:33–41. https://doi.org/10.1016/s0168-1605(99)00152-x
Buntin N, de Vos W, Hongpattarakere T (2017) Variation of mucin adhesion, cell surface characteristics, and molecular mechanisms among Lactobacillus plantarum isolated from different habitats. Appl Microbiol Biotechnol 101:7663–7674. https://doi.org/10.1007/s00253-017-8482-3
Capozzi V, Russo P, Ladero V et al (2012) Biogenic amines degradation by Lactobacillus plantarum: toward a potential application in wine. Front Microbiol 3:12. https://doi.org/10.3389/fmicb.2012.00122
Cervantes-Elizarrarás A, Cruz-Cansino N, Ramírez-Moreno E et al (2019) In vitro probiotic potential of lactic acid bacteria isolated from aguamiel and pulque and antibacterial activity against pathogens. Appl Sci 9:601. https://doi.org/10.3390/app9030601
Cheon MJ, Lim SM, Lee NK et al (2020) Probiotic properties and neuroprotective effects of Lactobacillus buchneri KU200793 isolated from Korean fermented foods. Int J Mol Sci 21:1227. https://doi.org/10.3390/ijms21041227
Corry JEL, Curtis GDW, Baird RM, Adams M, Moss MO, Ogden I (2011) Handbook of culture media for food and water microbiology. Corry JEL, Curtis GDW, Baird RM (Eds). Perlego Ltd, London, UK. Retrieved from https://www.perlego.com/book/787652/handbook-of-culture-media-for-food-and-water-microbiology-pdf (Original work published 2011)
Cowan MM, Van der Mei HC, Stokroos I et al (1992) Heterogeneity of surfaces of subgingival bacteria as detected by zeta potential measurements. J Dent Res 71:1803–1806. https://doi.org/10.1177/00220345920710110701
De Castilho NPA, Nero LA, Todorov SD (2019) Molecular screening of beneficial and safety determinants from bacteriocinogenic lactic acid bacteria isolated from Brazilian artisanal calabresa. Lett Appl Microbiol 69:204–211. https://doi.org/10.1111/lam.13194
De la Fuente-Salcido NM, Castañeda-Ramírez JC, García-Almendárez BE et al (2015) Isolation and characterization of bacteriocinogenic lactic bacteria from M-Tuba and Tepache, two traditional fermented beverages in México. Food Sci Nutr 3:434–442. https://doi.org/10.1002/fsn3.236
De Wouters T, Jans C, Niederberger T et al (2015) Adhesion potential of intestinal microbes predicted by physico-chemical characterization methods. PLoS ONE 10:e0136437. https://doi.org/10.1371/journal.pone.0136437
dos Santos KMO, De Matos CR, Salles HO et al (2020) Exploring beneficial/virulence properties of two dairy-related strains of Streptococcus infantarius subsp. infantarius. Probiotics Antimicrob Proteins 12:1524–1541. https://doi.org/10.1007/s12602-020-09637-8
EFSA (2012) Guidance on the assessment of bacterial susceptibility to antimicrobials of human and veterinary importance. EFSA J 10:2740. https://doi.org/10.2903/j.efsa.2012.2740
Escobar-Ramírez MC, Jaimez-Ordaz J, Escorza-Iglesias VA et al (2020) Lactobacillus pentosus ABHEAU-05: An in vitro digestion resistant lactic acid bacterium isolated from a traditional fermented Mexican beverage. Rev Argent Microbiol 52(305–314):10. https://doi.org/10.1016/j.ram.2019.10.005
FAO/WHO, Food and Agriculture Organization and World Health Organization (2002) Health and nutritional properties of probiotics in food including powder milk with live lactic acid bacteria. Report of a joint FAO/WHO expert consultation on evaluation of health and nutritional properties of probiotics in food including powder milk with live lactic acid bacteria. Cordoba, Argentina, Available from: https://www.iqb.es/digestivo/pdfs/probioticos.pdf. [Last accessed: January 29, 2023]
Garcia-Gonzalez N, Prete R, Battista N et al (2018) Adhesion properties of food-associated Lactobacillus plantarum strains on human intestinal epithelial cells and modulation of IL-8 release. Front Microbiol 9:2392. https://doi.org/10.3389/fmicb.2018.02392
Georghiou PR, Doggett AM, Kielhofner MA et al (1994) Molecular fingerprinting of Legionella species by repetitive element PCR. J Clin Microbiol 32:2989–2994. https://doi.org/10.1128/jcm.32.12.2989-2994.1994
Gevers D, Huys G, Swings J (2001) Applicability of rep-PCR fingerprinting for identification of Lactobacillus species. FEMS Microbiol Lett 205:31–36. https://doi.org/10.1111/j.1574-6968.2001.tb10921.x
Gheziel C, Russo P, Arena MP et al (2019) Evaluating the probiotic potential of Lactobacillus plantarum strains from Algerian infant feces: towards the design of probiotic starter cultures tailored for developing countries. Probiotics Antimicrob Proteins 11:113–123. https://doi.org/10.1007/s12602-018-9396-9
Granato D, Bergonzelli GE, Pridmore RD et al (2004) Cell surface-associated elongation factor Tu mediates the attachment of Lactobacillus johnsonii NCC533 (La1) to human intestinal cells and mucins. Infect Immun 72:2160–2169. https://doi.org/10.1128/IAI.72.4.2160-2169.2004
Guo XH, Kim JM, Nam HM, Park SY, Kim JM (2010) Screening lactic acid bacteria from swine origins for multistrain probiotics based on in vitro functional properties. Anaerobe 16:321–326. https://doi.org/10.1016/j.anaerobe.2010.03.006
Gutiérrez-Sarmiento W, Peña-Ocaña BA, Lam-Gutiérrez A et al (2022) Microbial community structure, physicochemical characteristics and predictive functionalities of the Mexican tepache fermented beverage. Microb Res 260:127045. https://doi.org/10.1016/j.micres.2022.127045
Halasz A, Baráth Á, Holzapfel WH (1999) The influence of starter culture selection on sauerkraut fermentation. Z Lebensm Unters Forsch 208:434–438. https://doi.org/10.1007/S002170050443
Halebian S, Harris B, Finegold SM et al (1981) Rapid method that aids in distinguishing gram-positive from gram-negative anaerobic bacteria. J Clin Microbiol 13:444–448. https://doi.org/10.1128/jcm.13.3.444-448.1981
Han Q, Kong B, Chen Q et al (2017) In vitro comparison of probiotic properties of lactic acid bacteria isolated from Harbin dry sausages and selected probiotics. J Funct Foods 32:391–400. https://doi.org/10.1016/j.jff.2017.03.020
Ji Y, Kim H, Park H et al (2013) Functionality and safety of lactic bacterial strains from Korean kimchi. Food Control 31:467–473. https://doi.org/10.1016/j.foodcont.2012.10.034
Jung MJ, Kim J, Lee SH et al (2022) Role of combinated lactic acid bacteria in bacterial, viral, and metabolite dynamics during fermentation of vegetable food, kimchi. Food Res Int 157:111261. https://doi.org/10.1016/j.foodres.2022.111261
Kim E, Won JE, Yang SM et al (2022) Diversity of a lactic acid bacterial community during fermentation of gajami-sikhae, a traditional Korean fermented fish, as determined by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Foods 11:909. https://doi.org/10.3390/foods11070909
Kim MJ, Kwak HS, Jung HY et al (2016) Microbial communities related to sensory attributes in Korean fermented soybean paste (doenjang). Food Res Int 89(Pt 1):724–732. https://doi.org/10.1016/j.foodres.2016.09.032
Kamarinou CS, Papadopoulou OS, Doulgeraki AI et al (2022) Mapping the key technological and functional characteristics of indigenous lactic acid bacteria isolated from Greek traditional dairy products. Microorganisms 10:246. https://doi.org/10.3390/microorganisms10020246
Klare I, Konstabel C, Müller-Bertling S et al (2005) Evaluation of new broth media for microdilution antibiotic susceptibility testing of lactobacilli, pediococci, lactococci, and bifidobacteria. Appl Environ Microbiol 71:8982–8986. https://doi.org/10.1128/AEM.71.12.8982-8986.2005
Klingberg TD, Axelsson L, Naterstad K et al (2005) Identification of potential probiotic starter cultures for Scandinavian-type fermented sausages. Int J Food Microbiol 105:419–431. https://doi.org/10.1016/j.ijfoodmicro.2005.03.020
Ladero V, Fernández M, Calles-Enríquez M et al (2012) Is the production of the biogenic amines tyramine and putrescine a species-level trait in enterococci? Food Microbiol 30:132–138. https://doi.org/10.1016/j.fm.2011.12.016
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, USA, pp 115–175
Laparra JM, Sanz Y (2009) Comparison of in vitro models to study bacterial adhesion to the intestinal epithelium. Lett Appl Microbiol 49:695–701. https://doi.org/10.1111/j.1472-765X.2009.02729.x
Lee J, Kim S, Kang CH (2022) Screening and probiotic properties of lactic acid bacteria with potential immunostimulatory activity isolated from kimchi. Fermentation 9:4. https://doi.org/10.3390/fermentation9010004
Mah JH, Park Y, Jin Y et al (2019) Bacterial production and control of biogenic amines in Asian fermented soybean foods. Foods 8:85. https://doi.org/10.3390/foods8020085
Maikranz E, Spengler C, Thewes N et al (2020) Different binding mechanisms of Staphylococcus aureus to hydrophobic and hydrophilic surfaces. Nanoscale 12:19267–19275. https://doi.org/10.1039/D0NR03134H
Mendling W (2016) Vaginal microbiota. In: Schwiertz A (ed) Microbiota of the human body. Advances in experimental medicine and biology, vol 902. Springer, Cham. https://doi.org/10.1007/978-3-319-31248-4_6
Mulaw G, Sisay Tessema T, Muleta D et al (2019) In vitro evaluation of probiotic properties of lactic acid bacteria isolated from some traditionally fermented Ethiopian food products. Int J Microb 2019:Article ID 7179514. https://doi.org/10.1155/2019/7179514
Neuhaus FC, Baddiley J (2003) A continuum of anionic charge: structures and functions of D-alanyl-teichoic acids in gram-positive bacteria. Microbiol Mol Biol Rev 67(4):686–723. https://doi.org/10.1128/MMBR.67.4.686-723.2003
Ngalimat MS, Raja Abd Rahman RNZ, Yusof MT et al (2019) Characterization of bacteria isolated from the stingless bee, Heterotrigona itama, honey, bee bread and propolis. Peer J 7:e7478. https://doi.org/10.7717/peerj.7478
Nordström EA, Teixeira C, Montelius C et al (2021) Lactiplantibacillus plantarum 299v (LP299V®): Three decades of research. Benef Microbes 12:441–465. https://doi.org/10.3920/BM2020.0191
Nunziata L, Brasca M, Morandi S et al (2022) Antibiotic resistance in wild and commercial non-enterococcal lactic acid bacteria and bifidobacteria strains of dairy origin: an update. Food Microbiol 104:103999. https://doi.org/10.1016/j.fm.2022.103999
Ovalle-Marmolejo XY, Redondo-Solano M, Granados-Chinchilla F et al (2023) Effect of stress factors on the production of biogenic amines by lactic acid bacteria isolated from fermented Mexican foods (cheese and beer). Food Control 146:09553. https://doi.org/10.1016/j.foodcont.2022.109553
Patra JK, Das GI, Paramithiotis S et al (2016) Kimchi and other widely consumed traditional fermented foods of Korea: A Review. Front Microbiol 7:1493. https://doi.org/10.3389/fmicb.2016.01493
Pavli A, Maltezou HC, Papadakis A et al (2016) Respiratory infections and gastrointestinal illness on a cruise ship: a three-year prospective study. Travel Med Infect Dis 14:389–397. https://doi.org/10.1016/j.tmaid.2016.05.019
Pelletier C, Bouley C, Cayuela C et al (1997) Cell surface characteristics of Lactobacillus casei subsp. casei, Lactobacillus paracasei subsp. paracasei, and Lactobacillus rhamnosus strains. Appl Environ Microbiol 63:1725–1731. https://doi.org/10.1128/aem.63.5.1725-1731.1997
Pineiro M, Stanton C (2007) Probiotic bacteria: legislative framework - requirements to evidence basis. J Nutr 137:850S-S853. https://doi.org/10.1093/jn/137.3.850S
Pontieri E (2018) The staphylococcal hemolysins. In: Savini V (ed) Pet-to-man travelling staphylococci. Academic Press. Massachusetts, USA, pp 103–116
Ribière C, Hegarty C, Stephenson H et al (2019) Gut and whole- body microbiota of the honeybee separate thriving and thriving hives. Microb Ecol 78:195–205. https://doi.org/10.1007/s00248-018-1287-9
Rocha-Mendoza D, Kosmerl E, Miyagusuku-Cruzado G et al (2020) Growth of lactic acid bacteria in milk phospholipids enhances their adhesion to Caco-2 cells. J Dairy Sci 103:7707–7718. https://doi.org/10.3168/jds.2020-18271
Rubio R, Jofré A, Martín B et al (2014) Characterization of lactic acid bacteria isolated from infant feces as potential probiotic starter cultures for fermented sausages. Food Microbiol 38:303–311. https://doi.org/10.1016/j.fm.2013.07.015
Saarela M, Mogensen G, Fondén R et al (2000) Probiotic bacteria: safety, functional and technological properties. J Biotechnol 84:197–215. https://doi.org/10.1016/s0168-1656(00)00375-8
Son SH, Jeon HL, Yang SJ et al (2017) Probiotic lactic acid bacteria isolated from traditional Korean fermented foods based on β-glucosidase activity. Food Sci Biotechnol 27:123–129. https://doi.org/10.1007/s10068-017-0212-1
Swain MR, Anandharaj M, Ray RC et al (2014) Fermented fruits and vegetables of Asia: a potential source of probiotics. Biotechnol Res Int 2014:Article ID 250424. https://doi.org/10.1155/2014/250424
Tırıs G, Yanıkoğlu RS, Ceylan B et al (2023) A review of the currently developed analytical methods for the determination of biogenic amines in food products. Food Chem 398:133919. https://doi.org/10.1016/j.foodchem.2022.133919
Terpou A, Papadaki A, Lappa IK et al (2019) Probiotics in food systems: significance and emerging strategies towards improved viability and delivery of enhanced beneficial value. Nutrients 11:1591. https://doi.org/10.3390/nu11071591
Tsekouras G, Tryfinopoulou P, Panagou EZ (2022) Detection and identification of lactic acid bacteria in semi-finished beer products using molecular techniques. In Biology and Life Sciences Forum 6:122. https://doi.org/10.3390/Foods2021-11046
Todorov SD, Penna ALB, Venema K, Holzapfel WH, Chikindas ML (2024) Recommendations for the use of standardized abbreviations for the former Lactobacillus genera, reclassified in the year 2020 (opinion). Benef Microb ahead of print https://doi.org/10.1163/18762891-20230114
Tulini FL, Hymery N, Haertlé T et al (2016) Screening for antimicrobial and proteolytic activities of lactic acid bacteria isolated from cow, buffalo and goat milk and cheeses marketed in the southeast region of Brazil. J Dairy Res 83:115–124. https://doi.org/10.1017/S0022029915000606
Valledor SJD, Bucheli JEV, Holzapfel WH et al (2020) Exploring beneficial properties of the bacteriocinogenic Enterococcus faecium ST10Bz strain isolated from boza, a Bulgarian cereal-based beverage. Microorganisms 8:1474. https://doi.org/10.3390/microorganisms8101474
Vélez MP, De Keersmaecker SC, Vanderleyden J (2007) Adherence factors of Lactobacillus in the human gastrointestinal tract. FEMS Microbiol Lett 276:140–148. https://doi.org/10.1111/j.1574-6968.2007.00908.x
Wu YC, Lin ST, Guu JR et al (2020) Vagococcus silage sp. nov., isolated from brewer’s grain used to make silage in Taiwan. Int J Syst Evol Microbiol 70:1953–1960. https://doi.org/10.1099/ijsem.0.003999
Zheng J, Wittouck S, Salvetti E et al (2020) A taxonomic note on the genus Lactobacillus: description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int J Syst Evol Microbiol 70:2782–2858. https://doi.org/10.1099/ijsem.0.004107
Zoumpopoulou G, Tzouvanou A, Mavrogonatou E et al (2018) Probiotic features of lactic acid bacteria isolated from a diverse pool of traditional Greek dairy products regarding strain- host interactions. Probiotics Antimicrob Proteins 10:313–322. https://doi.org/10.1007/s12602-017-9311-9
Zuo F, Yu R, Feng X et al (2016) Characterization and in vitro properties of potential probiotic Bifidobacterium strains isolated from breast-fed infant feces. Ann Microbiol 66:1027–1037. https://doi.org/10.1007/s13213-015-1187-x
Funding
This work was supported by HEM Pharma, Pohang, Republic of Korea. The National Council on Science and Technology (CONACYT), Mexico, DF, Mexico provided financial support to KAA (reference 293853/472094). FAPESP (grant 2013/07914–8), University of Sao Paulo, Sao Paulo, SP, Brazil. Centre for Research and Development in Agrifood Systems and Sustainability, funded by FCT (UIDB/05937/2020 and UIDP/05937/2020), Fundação para a Ciência e a Tecnologia, Portugal. National Research Foundation (NRF) funded by the Ministry of Science and ICT (NRF-2018M3A9F3021964), Seoul, Republic of Korea.
Author information
Authors and Affiliations
Contributions
WHH, STD, and KAA: conceptualization. KAA: writing—original draft, formal analysis, and writing—review and editing. KAA, JL, JEVB, SDT, and HP: methodology (supporting). WHH and SDT: review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Arellano, K., Lim, J., Bucheli, J.E.V. et al. Identification of safe putative probiotics from various food products. Folia Microbiol (2024). https://doi.org/10.1007/s12223-024-01142-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12223-024-01142-7