Abstract
Chordoma is a rare and aggressive bone tumor. An accurate investigation of tumor heterogeneity is necessary for the development of effective therapeutic strategies. This study aims to assess the poorly understood tumor heterogeneity of chordomas and identify potential therapeutic targets. Single-cell RNA sequencing was performed to delineate the transcriptomic landscape of chordomas. Six tumor samples of chordomas were obtained, and 33,737 cells passed the quality control test and were analyzed. The main cellular populations identified with specific markers were as follows: chordoma cells (16,052 [47.6%]), fibroblasts (6945 [20.6%]), mononuclear phagocytes (4734 [14.0%]), and T/natural killer (NK) cells (3944 [11.7%]). Downstream analysis of each cell type was performed. Six subclusters of chordomas exhibited properties of an epithelial-like extracellular matrix, stem cells, and immunosuppressive activity. Although few immune checkpoints were detected on cytotoxic immune cells such as T and NK cells, a strong immunosuppressive effect was exerted on the Tregs and M2 macrophages. In addition, the cellular interactions were indicative of enhancement of the TGF-β signaling pathway being the main mechanism for tumor progression, invasion, and immunosuppression. These findings, especially from the analysis of molecular targeted therapy and tumor immune microenvironment, may help in the identification of therapeutic targets in chordomas.






Similar content being viewed by others
Code availability
All scripts used are available at https://github.com/restore1997/chordoma.
References
McMaster ML et al (2001) Chordoma: incidence and survival patterns in the United States, 1973–1995. Cancer Causes Control 12(1):1–11
Walcott BP et al (2012) Chordoma: current concepts, management, and future directions. Lancet Oncol 13(2):e69-76
Tarpey PS et al (2017) The driver landscape of sporadic chordoma. Nat Commun 8(1):890
Thanindratarn P et al (2019) Advances in immune checkpoint inhibitors for bone sarcoma therapy. J Bone Oncol 15:100221
Stacchiotti S, Sommer J, Chordoma G (2015) Global consensus, Building a global consensus approach to chordoma: a position paper from the medical and patient community. Lancet Oncol 16(2):71–83
Dagogo-Jack I, Shaw AT (2018) Tumour heterogeneity and resistance to cancer therapies. Nat Rev Clin Oncol 15(2):81–94
Meng T et al (2019) Molecular targeted therapy in the treatment of chordoma: a systematic review. Front Oncol 9:30
Bompas E et al (2015) Sorafenib in patients with locally advanced and metastatic chordomas: a phase II trial of the French Sarcoma Group (GSF/GETO). Ann Oncol 26(10):2168–2173
Tawbi HA et al (2017) Pembrolizumab in advanced soft-tissue sarcoma and bone sarcoma (SARC028): a multicentre, two-cohort, single-arm, open-label, phase 2 trial. Lancet Oncol 18(11):1493–1501
Guo M et al (2019) Epigenetic heterogeneity in cancer. Biomark Res 7:23
Potter SS (2018) Single-cell RNA sequencing for the study of development, physiology and disease. Nat Rev Nephrol 14(8):479–492
Zou MX et al (2019) Clinical impact of the immune microenvironment in spinal chordoma: immunoscore as an independent favorable prognostic factor. Neurosurgery 84(6):E318-e333
Morimoto Y et al (2019) Prognostic significance of VEGF receptors expression on the tumor cells in skull base chordoma. J Neuro-oncol 144(1):65–77
Wang Y et al (2020) Changing technologies of RNA sequencing and their applications in clinical oncology. Front Oncol 10:447
Zhang Y, Zhang Z (2020) The history and advances in cancer immunotherapy: understanding the characteristics of tumor-infiltrating immune cells and their therapeutic implications. Cell Mol Immunol 17(8):807–821
Neftel C et al (2019) An integrative model of cellular states, plasticity, and genetics for glioblastoma. Cell 178(4):835-849.e21
Zhang M et al (2020) Single-cell transcriptomic architecture and intercellular crosstalk of human intrahepatic cholangiocarcinoma. J Hepatol 73(5):1118–1130
Maynard A et al (2020) Therapy-induced evolution of human lung cancer revealed by single-cell RNA sequencing. Cell 182(5):1232-1251.e22
Pan XW et al (2020) Identification of a novel cancer stem cell subpopulation that promotes progression of human fatal renal cell carcinoma by single-cell RNA-seq analysis. Int J Biol Sci 16(16):3149–3162
Puram SV et al (2017) Single-cell transcriptomic analysis of primary and metastatic tumor ecosystems in head and neck cancer. Cell 171(7):1611-1624.e24
Jo VY, Fletcher CD (2014) WHO classification of soft tissue tumours: an update based on the 2013 (4th) edition. Pathology 46(2):95–104
Wang LM et al (2020) A novel isocitrate dehydrogenase 1 G131D mutation in glioblastoma. Chin Med J (Engl) 134(4):486–488
Stuart T et al (2019) Comprehensive integration of single-cell data. Cell 177(7):1888-1902.e21
Aran D et al (2019) Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol 20(2):163–172
Zhang X et al (2019) Cell Marker: a manually curated resource of cell markers in human and mouse. Nucleic Acids Res 47(D1):D721–D728
Yu G et al (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287
Trapnell C et al (2014) The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol 32(4):381–386
Jin S, Guerrero-Juarez CF (2021) Inference and analysis of cell-cell communication using Cell Chat. Nat Commun 12(1):1088
Zheng W et al (2019) Application of nomograms to predict overall and cancer-specific survival in patients with chordoma. J Bone Oncol 18:100247
Zhang R et al (2020) STMN1 upregulation mediates hepatocellular carcinoma and hepatic stellate cell crosstalk to aggravate cancer by triggering the MET pathway. Cancer Sci 111(2):406–417
Obayashi S et al (2017) Stathmin1 expression is associated with aggressive phenotypes and cancer stem cell marker expression in breast cancer patients. Int J Oncol 51(3):781–790
Shuliang S et al (2013) Involvement of ubiquitin-conjugating enzyme E2C in proliferation and invasion of prostate carcinoma cells. Oncol Res 21(3):121–127
Parte S et al (2019) PTTG1: a unique regulator of stem/cancer stem cells in the ovary and ovarian cancer. Stem Cell Rev Rep 15(6):866–879
Huang S et al (2018) Interleukin-6/signal transducer and activator of transcription 3 promotes prostate cancer resistance to androgen deprivation therapy via regulating pituitary tumor transforming gene 1 expression. Cancer Sci 109(3):678–687
Cottone L et al (2020) Frequent alterations in p16/CDKN2A identified by immunohistochemistry and FISH in chordoma. J Pathol Clin Res 6(2):113–123
Arora A et al (2020) Pan-cancer identification of clinically relevant genomic subtypes using outcome-weighted integrative clustering. Genome Med 12(1):110
Hung YP et al (2020) Dedifferentiated chordoma: clinicopathologic and molecular characteristics with integrative analysis. Am J Surg Pathol 44(9):1213–1223
Brandt D, Hedrich CM (2018) TCRαβ(+)CD3(+)CD4(-)CD8(-) (double negative) T cells in autoimmunity. Autoimmun Rev 17(4):422–430
Hwang I et al (2020) Tumor-associated macrophage, angiogenesis and lymphangiogenesis markers predict prognosis of non-small cell lung cancer patients. J Transl Med 18(1):443
Heilmann RM et al (2019) Mucosal expression of S100A12 (calgranulin C) and S100A8/A9 (calprotectin) and correlation with serum and fecal concentrations in dogs with chronic inflammatory enteropathy. Vet Immunol Immunopathol 211:64–74
Nüchel J et al (2018) TGFB1 is secreted through an unconventional pathway dependent on the autophagic machinery and cytoskeletal regulators. Autophagy 14(3):465–486
Derynck R, Turley SJ, Akhurst RJ (2021) TGFβ biology in cancer progression and immunotherapy. Nat Rev Clin Oncol 18(1):9–34
Takeuchi M et al (1997) On the mechanisms by which transforming growth factor-beta 2 alters antigen-presenting abilities of macrophages on T cell activation. Eur J Immunol 27(7):1648–1656
Kobie JJ et al (2003) Transforming growth factor beta inhibits the antigen-presenting functions and antitumor activity of dendritic cell vaccines. Cancer Res 63(8):1860–1864
Lamouille S, Xu J, Derynck R (2014) Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol 15(3):178–196
Miettinen PJ et al (1994) TGF-beta induced transdifferentiation of mammary epithelial cells to mesenchymal cells: involvement of type I receptors. J Cell Biol 127(6 Pt 2):2021–2036
Kornberg TB (2017) Distributing signaling proteins in space and time: the province of cytonemes. Curr Opin Genet Dev 45:22–27
Gulluoglu S et al (2016) The molecular aspects of chordoma. Neurosurg Rev 39(2):185–96
Tuysuz EC et al (2019) Distinctive role of dysregulated miRNAs in chordoma cancer stem-like cell maintenance. Exp Cell Res 380(1):9–19
Zou MX et al (2018) Clinicopathologic implications of CD8(+)/Foxp3(+) ratio and miR-574-3p/PD-L1 axis in spinal chordoma patients. Cancer Immunol Immunother 67(2):209–224
Zou MX et al (2019) The relationship between tumor-stroma ratio, the immune microenvironment, and survival in patients with spinal chordoma. Neurosurgery 85(6):E1095-e1110
Hildenbrand R et al (1998) Transforming growth factor-beta stimulates urokinase expression in tumor-associated macrophages of the breast. Lab Invest 78(1):59–71
Wei Y et al (2017) Fibroblast-specific inhibition of TGF-β1 signaling attenuates lung and tumor fibrosis. J Clin Invest 127(10):3675–3688
Correction: (2019) Tumor-secreted LOXL2 activates fibroblasts through FAK signaling. Mol Cancer Res, 17(10): 2141
Scheel C, Weinberg RA (2012) Cancer stem cells and epithelial-mesenchymal transition: concepts and molecular links. Semin Cancer Biol 22(5–6):396–403
Ramachandran A et al. (2018) TGF-β uses a novel mode of receptor activation to phosphorylate SMAD1/5 and induce epithelial-to-mesenchymal transition. 7
Stadhouders R, Lubberts E, Hendriks RW (2018) A cellular and molecular view of T helper 17 cell plasticity in autoimmunity. J Autoimmun 87:1–15
Acknowledgements
We acknowledge the contributions of specific colleagues, institutions, or agencies that aided the authors' efforts.
Funding
CAMS/PUMC Research Project #201920200501, Human Brain Tissue Bank Platform for Neurological Diseases.
Author information
Authors and Affiliations
Contributions
ZC involved in conception and design; WD and BZ took part in collection and assembly of data; WD and XL involved in data analysis and interpretation; all authors involved in manuscript writing and approval of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
262_2022_3152_MOESM1_ESM.pdf
Fig. S1 Immumohistochemical staining, A Hematoxylin-eosin staining results of chordoma tissues, B Immunohistochemistry of Brachyury staining of chordoma tissues (PDF 891 kb)
262_2022_3152_MOESM2_ESM.pdf
Fig. S2 Quality control process A Violin plots showing cell infiltration using nFeature_RNA, nCount_RNA, and percent_mt, B Dot plots showing the feature-feature relationships (PDF 507 kb)
262_2022_3152_MOESM3_ESM.pdf
Fig. S3 Visualization of all the cells based on sample and site, A UMAP plots of all the cells colored by samples, B UMAP plots of all the cells colored by site, C Bar plots showing the proportion of the identified cell types in various locations (PDF 1186 kb)
262_2022_3152_MOESM4_ESM.pdf
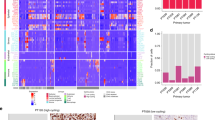
Fig. S4 Supplementary transcriptomic heterogeneity of tumor cells, A UMAP plot of tumor cells colored by samples, B UMAP plot of tumor cells colored by site, C Violin plots showing the expression of the immune checkpoints, D Bar plots showing the functional enrichment analysis of GO terms for each cluster of tumor cells, E Violin plots showing the expression of CDKN2A, F The ordering of tumor cells along pseudotime in a two-dimensional state space (colored by pseudotime), GO, Gene Ontology (PDF 1660 kb)
262_2022_3152_MOESM5_ESM.pdf
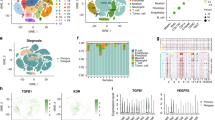
Fig. S5 Clustering and subtype analysis of supplementary T and NK cells, A UMAP plots of T and NK cells colored by samples, B UMAP plots of T and NK cells colored by site (PDF 5298 kb)
262_2022_3152_MOESM6_ESM.pdf
Fig. S6 The significant signaling pathways in the interplay between distinct cells, A The alluvial plot showing the outgoing (left) and incoming (right) signaling patterns of distinct cells, B Circle plot showing the significant inferred signaling networks (PDF 3809 kb)
Rights and permissions
About this article
Cite this article
Duan, W., Zhang, B., Li, X. et al. Single-cell transcriptome profiling reveals intra-tumoral heterogeneity in human chordomas. Cancer Immunol Immunother 71, 2185–2195 (2022). https://doi.org/10.1007/s00262-022-03152-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-022-03152-1