Abstract
Recently, all applications of BACE1 inhibitors failed as therapeutical targets for Alzheimer´s disease (AD) due to severe side effects. Therefore, alternative ways for treatment development are a hot research topic. The present analysis investigates BACE1 protein–protein interaction networks and attempts to solve the absence of complete knowledge about pathways involving BACE1. A bioinformatics analysis matched the functions of the non-substrate interaction network with Voltage-gated potassium channels, which also appear as top priority protein nodes. Targeting BACE1 interactions with PS1 and GGA-s, blocking of BACE1 access to APP by BRI3 and RTN-s, activation of Wnt signaling and upregulation of β-catenin, and brain delivery of the extracellular domain of p75NTR, are the main alternatives to the use of BACE 1 inhibitors highlighted by the analysis. The pathway enrichment analysis also emphasized substrates and substrate candidates with essential biological functions, which cleavage must remain controlled. They include ephrin receptors, ROBO1, ROBO2, CNTN-s, CASPR-s, CD147, CypB, TTR, APLP1/APLP2, NRXN-s, and PTPR-s. The analysis of the interaction subnetwork of BACE1 functionally related to inflammation identified a connection to three cardiomyopathies, which supports the hypothesis of the common molecular mechanisms with AD. A lot of potential shows the regulation of BACE1 activity through post-translational modifications. The interaction network of BACE1 and its phosphorylation enzyme CSNK1D functionally match the Circadian clock, p53, and Hedgehog signaling pathways. The regulation of BACE1 glycosylation could be achieved through N-acetylglucosamine transferases, α-(1→6)-fucosyltransferase, β-galactoside α-(2→6)-sialyltransferases, galactosyltransferases, and mannosidases suggested by the interaction network analysis of BACE1-MGAT3. The present analysis proposes possibilities for the alternative control of AD pathology.
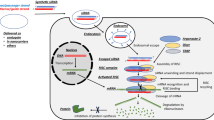
Graphic abstract






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files Table S1–S3.
Abbreviations
- BBB:
-
Blood–brain barrier
- CAM:
-
Cell adhesion molecule
- DRM:
-
Detergent-resistant membrane
- ER:
-
Endoplasmic reticulum
- ECM:
-
Extracellular matrix organization
- FN:
-
Fibronectin
- GB:
-
Glioblastoma
- GGA:
-
Golgi associated
- GPI:
-
Glycosylphosphatidylinositol
- GGA:
-
Golgi associated, gamma-adaptin ear-containing, ARF-binding protein
- HA:
-
Hyaluronan
- IGF:
-
Insulin-like growth factor
- IGFBP:
-
Insulin-like growth factor-binding protein
- KO:
-
Knockout
- L1CAM:
-
L1 cell adhesion molecule protein
- PTP:
-
Protein-tyrosine phosphatase
- PTPR:
-
Protein-tyrosine phosphatase, receptor-type
- PPIN:
-
Protein–protein interaction network
- RTN:
-
Reticulon
- MMP:
-
Matrix metalloproteinase
- NRX:
-
Neurexin
- TGN:
-
Trans-golgi network
References
Achilleos A, Huffman NT, Marcinkiewicyz E et al (2015) MBTPS1/SKI-1/S1P proprotein convertase is required for ECM signaling and axial elongation during somitogenesis and vertebral development. Hum Mol Genet 24:2884–2898. https://doi.org/10.1093/hmg/ddv050
Akiyama H, Kawamata T, Dedhar S, McGeer PL (1991) Immunohistochemical localization of vitronectin, its receptor and beta-3 integrin in Alzheimer brain tissue. J Neuroimmunol 32:19–28. https://doi.org/10.1016/0165-5728(91)90067-h
Alanis-Lobato G, Andrade-Navarro MA, Schaefer MH (2017) HIPPIE v2.0: Enhancing meaningfulness and reliability of protein-protein interaction networks. Nucleic Acids Res 45:D408–D414. https://doi.org/10.1093/nar/gkw985
Allen NJ, Bennett ML, Foo LC et al (2012) Astrocyte glypicans 4 and 6 promote formation of excitatory synapses via GluA1 AMPA receptors. Nature 486:410–414. https://doi.org/10.1038/nature11059
Andre B, Duterme C, Van Moer K et al (2011) Hyal2 is a glycosylphosphatidylinositol-anchored, lipid raft-associated hyaluronidase. Biochem Biophys Res Commun 411:175–179. https://doi.org/10.1016/j.bbrc.2011.06.125
Andreyeva A, Nieweg K, Horstmann K et al (2012) C-terminal fragment of N-cadherin accelerates synapse destabilization by amyloid-β. Brain 135:2140–2154. https://doi.org/10.1093/brain/aws120
Bader JS, Chaudhuri A, Rothberg JM, Chant J (2004) Gaining confidence in high-throughput protein interaction networks. Nat Biotechnol 22:78–85. https://doi.org/10.1038/nbt924
Banerji S, Ni J, Wang SX et al (1999) LYVE-1, a new homologue of the CD44 glycoprotein, is a lymph-specific receptor for hyaluronan. J Cell Biol 144:789–801. https://doi.org/10.1083/jcb.144.4.789
Bennett BD, Denis P, Haniu M et al (2000) A furin-like convertase mediates propeptide cleavage of BACE, the Alzheimer’s β-secretase. J Biol Chem 275:37712–37717. https://doi.org/10.1074/jbc.M005339200
Berglund EO, Murai KK, Fredette B et al (1999) Ataxia and abnormal cerebellar microorganization in mice with ablated contactin gene expression. Neuron 24:739–750. https://doi.org/10.1016/S0896-6273(00)81126-5
Bhat MA, Rios JC, Lu Y et al (2001) Axon-glia interactions and the domain organization of myelinated axons requires neurexin IV/caspr/paranodin. Neuron 30:369–383. https://doi.org/10.1016/S0896-6273(01)00294-X
Bosse KR, Raman P, Zhu Z et al (2017) Identification of GPC2 as an oncoprotein and candidate immunotherapeutic target in high-risk neuroblastoma. Cancer Cell 32:295-309.e12. https://doi.org/10.1016/j.ccell.2017.08.003
Bot N, Schweizer C, Ben HS, Fraering PC (2011) Processing of the synaptic cell adhesion molecule neurexin-3β by Alzheimer disease α- and γ-secretases. J Biol Chem 286:2762–2773. https://doi.org/10.1074/jbc.M110.142521
Bouyain S, Watkins DJ (2010) The protein tyrosine phosphatases PTPRZ and PTPRG bind to distinct members of the contactin family of neural recognition molecules. Proc Natl Acad Sci U S A 107:2443–2448. https://doi.org/10.1073/pnas.0911235107
Brady-Kalnay SM, Rimm DL, Tonks NK (1995) Receptor protein tyrosine phosphatase PTPμ associates with cadherins and catenins in vivo. J Cell Biol 130:977–986. https://doi.org/10.1083/jcb.130.4.977
Bras J, Djaldetti R, Alves AM et al (2016) Exome sequencing in a consanguineous family clinically diagnosed with early-onset Alzheimer’s disease identifies a homozygous CTSF mutation. Neurobiol Aging 46:236.e1-236.e6. https://doi.org/10.1016/j.neurobiolaging.2016.06.018
Brochmann Murray EJ, Murray SS, Simon R, Behnam K (2007) Recombinant expression, isolation, and proteolysis of extracellular matrix-secreted phosphoprotein-24 kDa. Connect Tissue Res 48:292–299. https://doi.org/10.1080/03008200701692404
Cai J, Qi X, Kociok N et al (2012) β-secretase (BACE1) inhibition causes retinal pathology by vascular dysregulation and accumulation of age pigment. EMBO Mol Med 4:980–991. https://doi.org/10.1002/emmm.201101084
Charlwood J, Dingwall C, Matico R et al (2001) Characterization of the glycosylation profiles of Alzheimer’s β-secretase protein Asp-2 expressed in a variety of cell lines. J Biol Chem 276:16739–16748. https://doi.org/10.1074/jbc.M009361200
Checler F, Afram E, Pardossi-Piquard R, Lauritzen I (2021) Is γ-secretase a beneficial inactivating enzyme of the toxic APP C-terminal fragment C99? J Biol Chem 296:100489–100508. https://doi.org/10.1016/j.jbc.2021.100489
Chen H, Li C, Zhou T, et al (2020) Secreted phosphoprotein 24 kD (Spp24) inhibits growth of human osteosarcoma through the BMP-2/Smad signaling pathway. 1–20. https://doi.org/10.21203/rs.3.rs-27515/v1
Chibnik LB, White CC, Mukherjee S et al (2018) Susceptibility to neurofibrillary tangles: role of the PTPRD locus and limited pleiotropy with other neuropathologies. Mol Psychiatry 23:1521–1529. https://doi.org/10.1038/mp.2017.20
Chin CH, Chen SH, Wu HH et al (2014) CytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol 8:1–7. https://doi.org/10.1186/1752-0509-8-S4-S11
Cline MS, Smoot M, Cerami E et al (2007) Integration of biological networks and gene expression data using Cytoscape. Nat Protoc 2:2366–2382. https://doi.org/10.1038/nprot.2007.324
Costantini C, Ko MH, Jonas MC, Puglielli L (2007) A reversible form of lysine acetylation in the ER and Golgi lumen controls the molecular stabilization of BACE1. Biochem J 407:383–395. https://doi.org/10.1042/BJ20070040
Cuccaro ML, Carney RM, Zhang Y et al (2016) SORL1 mutations in early- and late-onset Alzheimer disease. Neurol Genet 2:e116–e123. https://doi.org/10.1212/NXG.0000000000000116
Cui J, Xiao J, Tagliabracci VS et al (2015) A secretory kinase complex regulates extracellular protein phosphorylation. Elife 2015:1–18. https://doi.org/10.7554/eLife.06120
Cummings RD, Trowbridge IS, Kornfeld S (1982) A mouse lymphoma cell line resistant to the leukoagglutinating lectin from Phaseolus vulgaris is deficient in UDP-GlcNAc:α-D-mannoside β1,6 N-acetylglucosaminyltransferase. J Biol Chem 257:13421–13427. https://doi.org/10.1016/s0021-9258(18)33465-3
De Cat B, Muyldermans SY, Coomans C et al (2003) Processing by proprotein convertases is required for glypican-3 modulation of cell survival, Wnt signaling, and gastrulation movements. J Cell Biol 163:625–635. https://doi.org/10.1083/jcb.200302152
DeWit J, O’Sullivan ML, Savas JN et al (2013) Unbiased discovery of glypican as a receptor for LRRTM4 in regulating excitatory synapse development. Neuron 79:696–711. https://doi.org/10.1016/j.neuron.2013.06.049
Dislich B (2013) The global proteomic profiling of the BACE1 knockout mouse brain allows the identification of novel BACE1 candidate substrates in vivo. Dtsch Zent für Neurodegener Erkrankungen und Tech Univ München, Diss für Medizin
Dislich B, Wohlrab F, Bachhuber T et al (2015) Label-free quantitative proteomics of mouse cerebrospinal fluid detects β-site APP cleaving enzyme (BACE1) protease substrates in vivo. Mol Cell Proteomics 14:2550–2563. https://doi.org/10.1074/mcp.M114.041533
Dollt C, Becker K, Michel J et al (2017) The shedded ectodomain of Lyve-1 expressed on M2-like tumor-associated macrophages inhibits melanoma cell proliferation. Oncotarget 8:103682–103692. https://doi.org/10.18632/oncotarget.21771
Du Y, Grandis JR (2015) Receptor-type protein tyrosine phosphatases in cancer. Chin J Cancer 34:61–69. https://doi.org/10.5732/cjc.014.10146
Eggert S, Paliga K, Soba P et al (2004) The proteolytic processing of the amyloid precursor protein gene family members APLP-1 and APLP-2 involves α-, β-, γ-, and ε-like cleavages: Modulation of APLP-1 processing by N-glycosylation. J Biol Chem 279:18146–18156. https://doi.org/10.1074/jbc.M311601200
Endres K, Postina R, Schroeder A et al (2005) Shedding of the amyloid precursor protein-like protein APLP2 by disintegrin-metalloproteinases: retinoic acid-induced upregulation of substrate and proteinase ADAM10 during neuronal cell differentiation. FEBS J 272:5808–5820. https://doi.org/10.1111/j.1742-4658.2005.04976.x
Enright AJ, Van Dongen S, Ouzounis CA (2002) An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res 30:1575–1584. https://doi.org/10.1093/nar/30.7.1575
Eriksson PS, Perfilieva E, Björk-Eriksson T et al (1998) Neurogenesis in the adult human hippocampus. Nat Med 4:1313–1317. https://doi.org/10.1038/3305
Etchegaray J-P, Machida KK, Noton E et al (2009) Casein kinase 1 delta regulates the pace of the mammalian circadian clock. Mol Cell Biol 29:3853–3866. https://doi.org/10.1128/mcb.00338-09
Fan L, Xu D, Wang W et al (2013) Caspr interaction with amyloid precursor protein reduces amyloid-β generation in vitro. Neurosci Lett 548:255–260. https://doi.org/10.1016/j.neulet.2013.05.055
Fifield AL, Hanavan PD, Faigel DO et al (2020) Molecular inhibitor of QSOX1 suppresses tumor growth in vivo. Mol Cancer Ther 19:112–122. https://doi.org/10.1158/1535-7163.MCT-19-0233
Filser S, Ovsepian SV, Masana M et al (2015) Pharmacological inhibition of BACE1 impairs synaptic plasticity and cognitive functions. Biol Psychiatry 77:729–739. https://doi.org/10.1016/j.biopsych.2014.10.013
Fleck D, van Bebber F, Colombo A et al (2013) Dual cleavage of neuregulin 1 type III by BACE1 and ADAM17 liberates its EGF-like domain and allows paracrine signaling. J Neurosci 33:7856–7869. https://doi.org/10.1523/JNEUROSCI.3372-12.2013
Franz M, Rodriguez H, Lopes C et al (2018) GeneMANIA update 2018. Nucleic Acids Res 46:W60–W64. https://doi.org/10.1093/nar/gky311
Garton KJ, Gough PJ, Philalay J et al (2003) Stimulated shedding of vascular cell adhesion molecule 1 (VCAM-1) is mediated by tumor necrosis factor-α-converting enzyme (ADAM 17). J Biol Chem 278:37459–37464. https://doi.org/10.1074/jbc.M305877200
Gautam V, D’Avanzo C, Hebisch M et al (2014) BACE1 activity regulates cell surface contactin-2 levels. Mol Neurodegener 9:1–10. https://doi.org/10.1186/1750-1326-9-4
Geng Y, Xu C, Wang Y, Zhang L (2020) Quiescin sulfhydryl oxidase 1 regulates the proliferation, migration and invasion of human glioblastoma cells via PI3K/Akt pathway. Onco Targets Ther 13:5721–5729. https://doi.org/10.2147/OTT.S255941
Graf ER, Zhang X, Jin SX et al (2004) Neurexins induce differentiation of GABA and glutamate postsynaptic specializations via neuroligins. Cell 119:1013–1026. https://doi.org/10.1016/j.cell.2004.11.035
Gray-Owen SD, Blumberg RS (2006) CEACAM1: contact-dependent control of immunity. Nat Rev Immunol 6:433–446. https://doi.org/10.1038/nri1864
Greco S, Zaccagnini G, Fuschi P et al (2017) Increased BACE1-AS long noncoding RNA and β-amyloid levels in heart failure. Cardiovasc Res 113:453–463. https://doi.org/10.1093/cvr/cvx013
Hampel H, Vassar R, De Strooper B et al (2020a) The β-secretase BACE1 in Alzheimer’s disease. Biol Psychiatry 89:1–12. https://doi.org/10.1016/j.biopsych.2020.02.001
Hampel H, Vassar R, De Strooper B et al (2020b) The β-secretase BACE1 in Alzheimer’s disease. Biol Psychiatry 89:745–756. https://doi.org/10.1016/j.biopsych.2020.02.001
Hanavan PD, Borges CR, Katchman BA et al (2015) Ebselen inhibits QSOX1 enzymatic activity and suppresses invasion of pancreatic and renal cancer cell lines. Oncotarget 6:18418–18428. https://doi.org/10.18632/oncotarget.4099
Hanoulle X, Melchior A, Sibille N et al (2007) Structural and functional characterization of the interaction between cyclophilin B and a heparin-derived oligosaccharide. J Biol Chem 282:34148–34158. https://doi.org/10.1074/jbc.M706353200
Hartung H-P, Reiners K, Archelos JJ et al (1995) Circulating adhesion molecules and tumor necrosis factor receptor in multiple sclerosis: correlation with magnetic resonance imaging. Ann Neurol 38:186–193. https://doi.org/10.1002/ana.410380210
Hayward AN, Gordon WR (2018) Dystroglycan proteolysis is conformationally-regulated and disrupted by disease-associated mutations. BioRxiv. https://doi.org/10.1101/279315
He W, Lu Y, Qahwash I et al (2004) Reticulon family members modulate BACE1 activity and amyloid-beta peptide generation. Nat Med 10:959–965. https://doi.org/10.1038/nm1088
He X, Li F, Chang WP, Tang J (2005) GGA proteins mediate the recycling pathway of memapsin 2 (BACE). J Biol Chem 280:11696–11703. https://doi.org/10.1074/jbc.M411296200
He W, Hu J, Xia Y, Yan R (2014) β-site amyloid precursor protein cleaving enzyme 1(BACE1) regulates Notch signaling by controlling the cleavage of Jagged 1 (Jag1) and Jagged 2 (Jag2) proteins. J Biol Chem 289:20630–20637. https://doi.org/10.1074/jbc.m114.579862
Heber S, Herms J, Gajic V et al (2000) Mice with combined gene knock-outs reveal essential and partially redundant functions of amyloid precursor protein family members. J Neurosci 20:7951–7963. https://doi.org/10.1523/jneurosci.20-21-07951.2000
Hein MY, Hubner NC, Poser I et al (2015) A human interactome in three quantitative dimensions organized by stoichiometries and abundances. Cell 163:712–723. https://doi.org/10.1016/j.cell.2015.09.053
Hemming ML, Elias JE, Gygi SP, Selkoe DJ (2009) Identification of β-secretase (BACE1) substrates using quantitative proteomics. PLoS ONE 4:e8477–e8490. https://doi.org/10.1371/journal.pone.0008477
Hessler S, Zheng F, Hartmann S et al (2015) β-secretase BACE1 regulates hippocampal and reconstituted M-currents in a β-subunit-like fashion. J Neurosci 35:3298–3311. https://doi.org/10.1523/JNEUROSCI.3127-14.2015
Hitt B, Riordan SM, Kukreja L et al (2012) β-Site amyloid precursor protein (APP)-cleaving enzyme 1 (BACE1)-deficient mice exhibit a close homolog of L1 (CHL1) loss-of-function phenotype involving axon guidance defects. J Biol Chem 287:38408–38425. https://doi.org/10.1074/jbc.M112.415505
Hitt BD, Jaramillo TC, Chetkovich DM, Vassar R (2010) BACE1-/- mice exhibit seizure activity that does not correlate with sodium channel level or axonal localization. Mol Neurodegener 5:31–44. https://doi.org/10.1186/1750-1326-5-31
Hogl S, Kuhn PH, Colombo A, Lichtenthaler SF (2011) Determination of the proteolytic cleavage sites of the amyloid precursor-like protein 2 by the proteases ADAM10, BACE1 and γ-secretase. PLoS ONE 6:e21337–e22148. https://doi.org/10.1371/journal.pone.0021337
Holmes BB, DeVos SL, Kfoury N et al (2013) Heparan sulfate proteoglycans mediate internalization and propagation of specific proteopathic seeds. Proc Natl Acad Sci U S A 110:E3138–E3147. https://doi.org/10.1073/pnas.1301440110
Holt CE, Dickson BJ (2005) Sugar codes for axons? Neuron 46:169–172. https://doi.org/10.1016/j.neuron.2005.03.021
Hou H, Fan Q, He W et al (2017) BACE1 deficiency causes abnormal neuronal clustering in the dentate gyrus. Stem Cell Rep 9:217–230. https://doi.org/10.1016/j.stemcr.2017.05.030
Hsu LJ, Schultz L, Hong Q et al (2009) Transforming growth factor β1 signaling via interaction with cell surface hyal-2 and recruitment of WWOX/WOX1. J Biol Chem 284:16049–16059. https://doi.org/10.1074/jbc.M806688200
Hsu LJ, Chiang MF, Sze CI et al (2016) HYAL-2-WWOX-SMAD4 signaling in cell death and anticancer response. Front Cell Dev Biol 4:1–11. https://doi.org/10.3389/fcell.2016.00141
Hsu LJ, Hong Q, Chen ST et al (2017) Hyaluronan activates Hyal-2/WWOX/Smad4 signaling and causes bubbling cell death when the signaling complex is overexpressed. Oncotarget 8:19137–19155. https://doi.org/10.18632/oncotarget.13268
Hu X, Hicks CW, He W et al (2006) Bace1 modulates myelination in the central and peripheral nervous system. Nat Neurosci 9:1520–1525. https://doi.org/10.1038/nn1797
Hu X, He W, Diaconu C et al (2008) Genetic deletion of BACE1 in mice affects remyelination of sciatic nerves. FASEB J 22:2970–2980. https://doi.org/10.1096/fj.08-106666
Hu X, Zhou X, He W et al (2010) BACE1 deficiency causes altered neuronal activity and neurodegeneration. J Neurosci 30:8819–8829. https://doi.org/10.1523/JNEUROSCI.1334-10.2010
Hu X, He W, Luo X et al (2013) BACE1 regulates hippocampal astrogenesis via the Jagged1-Notch pathway. Cell Rep 4:40–49. https://doi.org/10.1016/j.celrep.2013.06.005
Hummel V, Kallmann BA, Wagner S et al (2001) Production of MMPs in human cerebral endothelial cells and their role in shedding adhesion molecules. J Neuropathol Exp Neurol 60:320–327. https://doi.org/10.1093/jnen/60.4.320
Hur JY, Teranishi Y, Kihara T et al (2012) Identification of novel γ-secretase-associated proteins in detergent-resistant membranes from brain. J Biol Chem 287:11991–12005. https://doi.org/10.1074/jbc.M111.246074
Huth T, Schmidt-Neuenfeldt K, Rittger A et al (2009) Non-proteolytic effect of beta-site APP-cleaving enzyme 1 (BACE1) on sodium channel function. Neurobiol Dis 33:282–289. https://doi.org/10.1016/j.nbd.2008.10.015
Huttlin EL, Ting L, Bruckner RJ et al (2015) The BioPlex network: a systematic exploration of the human interactome. Cell 162:425–440. https://doi.org/10.1016/j.cell.2015.06.043
Ishiguro H, Okubo T, Kuwabara Y et al (2017) NOTCH1 activates the Wnt/β-catenin signaling pathway in colon cancer. Oncotarget 8:60378–60389. https://doi.org/10.18632/oncotarget.19534
Jin SM, Cho HJ, Kim YW et al (2012) Aβ-induced Ca2+ influx regulates astrocytic BACE1 expression via calcineurin/NFAT4 signals. Biochem Biophys Res Commun 425:649–655. https://doi.org/10.1016/j.bbrc.2012.07.123
Jonas MC, Costantini C, Puglielli L (2008) PCSK9 is required for the disposal of non-acetylated intermediates of the nascent membrane protein BACE1. EMBO Rep 9:916–922. https://doi.org/10.1038/embor.2008.132
Kapoor A, Nation DA (2021) Role of Notch signaling in neurovascular aging and Alzheimer’s disease. Semin Cell Dev Biol 116:90–97. https://doi.org/10.1016/j.semcdb.2020.12.011
Katchman BA, Antwi K, Hostetter G et al (2011) Quiescin sulfhydryl oxidase 1 promotes invasion of pancreatic tumor cells mediated by matrix metalloproteinases. Mol Cancer Res 9:1621–1631. https://doi.org/10.1158/1541-7786.MCR-11-0018
Katchman BA, Ocal IT, Cunliffe HE et al (2013) Expression of quiescin sulfhydryl oxidase 1 is associated with a highly invasive phenotype and correlates with a poor prognosis in luminal B breast cancer. Breast Cancer Res 15:R28–R41. https://doi.org/10.1186/bcr3407
Kawamura Y, Kikuchi A, Takada R et al (2001) Inhibitory effect of a presenilin 1 mutation on the Wnt signalling pathway by enhancement of β-catenin phosphorylation. Eur J Biochem 268:3036–3041. https://doi.org/10.1046/j.1432-1327.2001.02197.x
Kerrien S, Alam-Faruque Y, Aranda B et al (2007) IntAct—open source resource for molecular interaction data. Nucleic Acids Res 35:561–565. https://doi.org/10.1093/nar/gkl958
Kilger E, Buehler A, Woelfing H et al (2011) BRI2 protein regulates β-amyloid degradation by increasing levels of secreted insulin-degrading enzyme (IDE). J Biol Chem 286:37446–37457. https://doi.org/10.1074/jbc.M111.288373
Kim DY, Carey BW, Wang H et al (2007) BACE1 regulates voltage-gated sodium channels and neuronal activity. Nat Cell Biol 9:755–764. https://doi.org/10.1038/ncb1602
Kim WH, Watanabe H, Lomoio S, Tesco G (2021) Spatiotemporal processing of neural cell adhesion molecules 1 and 2 by BACE1 in vivo. J Biol Chem 296:100372. https://doi.org/10.1016/j.jbc.2021.100372
Kitazume S, Tachida Y, Oka R et al (2001) Alzheimer’s beta-secretase, beta-site amyloid precursor protein-cleaving enzyme, is responsible for cleavage secretion of a golgi-resident sialyltransferase. Proc Natl Acad Sci U S A 98:13554–13559. https://doi.org/10.1073/pnas.241509198
Kizuka Y, Nakano M, Kitazume S et al (2016) Bisecting GlcNAc modification stabilizes BACE1 protein under oxidative stress conditions. Biochem J 473:21–30. https://doi.org/10.1042/BJ20150607
Kizuka Y, Kitazume S, Fujinawa R et al (2015) An aberrant sugar modification of BACE 1 blocks its lysosomal targeting in Alzheimer’s disease. EMBO Mol Med 7:175–189. https://doi.org/10.15252/emmm.201404438
Kondratova AA, Kondratov RV (2012) The circadian clock and pathology of the ageing brain. Nat Rev Neurosci 13:325–335. https://doi.org/10.1038/nrn3208
Kuhn PH, Marjaux E, Imhof A et al (2007) Regulated intramembrane proteolysis of the interleukin-1 receptor II by α-, β-, and γ-secretase. J Biol Chem 282:11982–11995. https://doi.org/10.1074/jbc.M700356200
Kuhn PH, Koroniak K, Hogl S et al (2012) Secretome protein enrichment identifies physiological BACE1 protease substrates in neurons. EMBO J 31:3157–3168. https://doi.org/10.1038/emboj.2012.173
Kume H, Murayama KS, Araki W (2009) The two-hydrophobic domain tertiary structure of reticulon proteins is critical for modulation of β-secretase BACE1. J Neurosci Res 87:2963–2972. https://doi.org/10.1002/jnr.22112
Kuzuya A, Uemura K, Kitagawa N et al (2007) Presenilin 1 is involved in the maturation of beta-site amyloid precursor protein-cleaving enzyme 1 (BACE1). J Neurosci Res 85:153–165. https://doi.org/10.1002/jnr.21104
La Marca R, Cerri F, Horiuchi K et al (2011) TACE (ADAM17) inhibits Schwann cell myelination. Nat Neurosci 14:857–865. https://doi.org/10.1038/nn.2849
Laird FM, Cai H, Savonenko AV et al (2005) BACE1, a major determinant of selective vulnerability of the brain to amyloid-beta amyloidogenesis, is essential for cognitive, emotional, and synaptic functions. J Neurosci 25:11693–11709. https://doi.org/10.1523/JNEUROSCI.2766-05.2005
Lake DF, Faigel DO (2014) The emerging role of qsox1 in cancer. Antioxid Redox Signal 21:485–496. https://doi.org/10.1089/ars.2013.5572
Lao L, Shen J, Tian H et al (2016) Secreted phosphoprotein 24 kD inhibits growth of human prostate cancer cells stimulated by BMP-2. Anticancer Res 36:5773–5780. https://doi.org/10.21873/anticanres.11161
Leavesley DI, Kashyap AS, Croll T et al (2013) Vitronectin - master controller or micromanager? IUBMB Life 65:807–818. https://doi.org/10.1002/iub.1203
Lee KB, Murray SS, Duarte MEL et al (2011) Effects of the bone morphogenetic protein binding protein spp24 (secreted phosphoprotein 24 kD) on the growth of human lung cancer cells. J Orthop Res 29:1712–1718. https://doi.org/10.1002/jor.21383
Li B, Niswander LA (2020) TMEM132A, a novel Wnt signaling pathway regulator through Wntless (WLS) interaction. Front Cell Dev Biol 8:1–14. https://doi.org/10.3389/fcell.2020.599890
Li C-S, Tian H, Zou M et al (2015) Secreted phosphoprotein 24 kD (Spp24) inhibits growth of human pancreatic cancer cells caused by BMP-2. Biochem Biophys Res Commun 466:167–172. https://doi.org/10.1016/j.bbrc.2015.08.124
Li N, Fu H, Hewitt SM et al (2017) Therapeutically targeting glypican-2 via single-domain antibody-based chimeric antigen receptors and immunotoxins in neuroblastoma. Proc Natl Acad Sci U S A 114:E6623–E6631. https://doi.org/10.1073/pnas.1706055114
Lichtenthaler SF, Dominguez DI, Westmeyer GG et al (2003) The cell adhesion protein P-selectin glycoprotein ligand-1 is a substrate for the aspartyl protease BACE1. J Biol Chem 278:48713–48719. https://doi.org/10.1074/jbc.M303861200
Linneberg C, Toft CLF, Kjaer-Sorensen K, Laursen LS (2019) L1cam-mediated developmental processes of the nervous system are differentially regulated by proteolytic processing. Sci Rep 9:1–14. https://doi.org/10.1038/s41598-019-39884-x
Lopes CT, Franz M, Kazi F et al (2010) Cytoscape Web: an interactive web-based network browser. Bioinformatics 26:2347–2348. https://doi.org/10.1093/bioinformatics/btq430
López-Bendito G, Flames N, Ma L et al (2007) Robo1 and Robo2 cooperate to control the guidance of major axonal tracts in the mammalian forebrain. J Neurosci 27:3395–3407. https://doi.org/10.1523/JNEUROSCI.4605-06.2007
Lorente-Gea L, Garcia B, Martin C et al (2020) Heparan sulfate proteoglycans undergo differential expression alterations in Alzheimer disease brains. J Neuropathol Exp Neurol 79:474–483. https://doi.org/10.1093/jnen/nlaa016
Ma QH, Futagawa T, Yang WL et al (2008) A TAG1-APP signalling pathway through Fe65 negatively modulates neurogenesis. Nat Cell Biol 10:283–294. https://doi.org/10.1038/ncb1690
Ma JH, Qin L, Li X (2020) Role of STAT3 signaling pathway in breast cancer. Cell Commun Signal 18:1–13. https://doi.org/10.1186/s12964-020-0527-z
Maness PF, Schachner M (2007) Neural recognition molecules of the immunoglobulin superfamily: signaling transducers of axon guidance and neuronal migration. Nat Neurosci 10:19–26. https://doi.org/10.1038/nn1827
Marambaud P, Wen PH, Dutt A et al (2003) A CBP binding transcriptional repressor produced by the PS1/ε-cleavage of N-cadherin is inhibited by PS1 FAD mutations. Cell 114:635–645. https://doi.org/10.1016/j.cell.2003.08.008
Masliah E, Xie F, Dayan S et al (2010) Genetic deletion of Nogo/Rtn4 ameliorates behavioral and neuropathological outcomes in amyloid precursor protein transgenic mice. Neuroscience 169:488–494. https://doi.org/10.1016/j.neuroscience.2010.04.045
Matsuda S, Giliberto L, Matsuda Y et al (2005) The familial dementia BRI2 gene binds the Alzheimer gene amyloid-β precursor protein and inhibits amyloid-β production. J Biol Chem 280:28912–28916. https://doi.org/10.1074/jbc.C500217200
Matsuda S, Matsuda Y, D’Adamio L (2009) BRI3 inhibits amyloid precursor protein processing in a mechanistically distinct manner from its homologue dementia gene BRI2. J Biol Chem 284:15816–15825. https://doi.org/10.1074/jbc.M109.006403
McConlogue L, Buttini M, Anderson JP et al (2007) Partial reduction of BACE1 has dramatic effects on Alzheimer plaque and synaptic pathology in APP transgenic mice. J Biol Chem 282:26326–26334. https://doi.org/10.1074/jbc.M611687200
McEver RP, Cummings RD (1997) Role of PSGL-1 binding to selectins in leukocyte recruitment. J Clin Invest 100:485–492. https://doi.org/10.1172/jci119556
Merico D, Isserlin R, Stueker O et al (2010) Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS ONE 5:e13984–e13995. https://doi.org/10.1371/journal.pone.0013984
Mi HK, Puglielli L (2009) Two endoplasmic reticulum (ER)/ER golgi intermediate compartment-based lysine acetyltransferases post-translationally regulate BACE1 levels. J Biol Chem 284:2482–2492. https://doi.org/10.1074/jbc.M804901200
Moore SA, Saito F, Chen J et al (2002) Deletion of brain dystroglycan recapitulates aspects of congenital muscular dystrophy. Nature 418:422–425. https://doi.org/10.1038/nature00838
Morel C, Adami P, Musard J-F et al (2007) Involvement of sulfhydryl oxidase QSOX1 in the protection of cells against oxidative stress-induced apoptosis. Exp Cell Res 313:3971–3982. https://doi.org/10.1016/j.yexcr.2007.09.003
Munir J, Van Ngu T, Na Ayudthaya PD, Ryu S (2020) Downregulation of glypican-4 facilitates breast cancer progression by inducing cell migration and proliferation. Biochem Biophys Res Commun 526:91–97. https://doi.org/10.1016/j.bbrc.2020.03.064
Nagae M, Yamaguchi Y, Taniguchi N, Kizuka Y (2020) 3D structure and function of glycosyltransferases involved in N-glycan maturation. Int J Mol Sci 21:1–23. https://doi.org/10.3390/ijms21020437
Nahalkova J, Volkmann I, Aoki M et al (2010) CD147, a γ-secretase associated protein is upregulated in Alzheimer’s disease brain and its cellular trafficking is affected by presenilin-2. Neurochem Int 56:67–76. https://doi.org/10.1016/j.neuint.2009.09.003
Nahálková J (2016) The protein interaction network with functional roles in tumorigenesis, neurodegeneration, and ageing. Mol Cell Biochem 423:187–196. https://doi.org/10.1007/s11010-016-2836-5
Nahálková J (2020a) The molecular mechanisms associated with PIN7, a protein-protein interaction network of seven pleiotropic proteins. J Theor Biol 487:110124–110131. https://doi.org/10.1016/j.jtbi.2019.110124
Nahálková J (2020b) Linking TPPII to the protein interaction and signalling networks. Comput Biol Chem 87:107291–107399. https://doi.org/10.1016/j.compbiolchem.2020.107291
Nahálková J (2020c) Exploring the sirtuin functionality in ageing through human protein interaction networks. SN Comput Sci 1:183–195. https://doi.org/10.1007/s42979-020-00192-1
Nahálková J (2022) Focus on molecular functions of anti-aging deacetylase SIRT3. Biochemistry (Moscow) 87:21–34. https://doi.org/10.1134/S0006297922010035
Nalbant D, Youn H, Nalbant SI et al (2005) FAM20: an evolutionarily conserved family of secreted proteins expressed in hematopoietic cells. BMC Genomics 6:1–21. https://doi.org/10.1186/1471-2164-6-11
Needham B, Wlodek M, Ciccotosto G et al (2008) Identification of the Alzheimer’s disease amyloid precursor protein (APP) and its homologue APLP2 as essential modulators of glucose and insulin homeostasis and growth. J Pathol 215:155–163. https://doi.org/10.1002/path.2343
Nelson WJ, Nusse R (2004) Convergence of Wnt, β-catenin, and cadherin pathways. Science 303:1483–1487. https://doi.org/10.1126/science.1094291
Nishida-Fukuda H, Araki R, Shudou M et al (2016) Ectodomain shedding of lymphatic vessel endothelial hyaluronan receptor 1 (LYVE-1) is induced by vascular endothelial growth factor A (VEGF-A). J Biol Chem 291:10490–10500. https://doi.org/10.1074/jbc.M115.683201
Oh Y, Kim EY, Kim Y et al (2011) Neuroprotective effects of overexpressed cyclophilin B against Aβ-induced neurotoxicity in PC12 cells. Free Radic Biol Med 51:905–920. https://doi.org/10.1016/j.freeradbiomed.2011.05.036
Oh-hashi K, Naruse Y, Amaya F et al (2003) Cloning and characterization of a novel GRP78-binding protein in the rat brain. J Biol Chem 278:10531–10537. https://doi.org/10.1074/jbc.M212083200
Oh-Hashi K, Koga H, Nagase T et al (2012) Characterization of the expression and cell-surface localization of transmembrane protein 132A. Mol Cell Biochem 370:23–33. https://doi.org/10.1007/s11010-012-1394-8
Oikari LE, Yu C, Okolicsanyi RK et al (2020) HSPGs glypican-1 and glypican-4 are human neuronal proteins characteristic of different neural phenotypes. J Neurosci Res 98:1619–1645. https://doi.org/10.1002/jnr.24666
Oliveira SM, Ribeiro CA, Cardoso I, Saraiva MJ (2011) Gender-dependent transthyretin modulation of brain amyloid-β Levels: evidence from a mouse model of Alzheimer’s disease. J Alzheimer’s Dis 27:429–439. https://doi.org/10.3233/JAD-2011-110488
Orchard S, Ammari M, Aranda B et al (2014) The MIntAct project—IntAct as a common curation platform for 11 molecular interaction databases. Nucleic Acids Res 42:D358–D363. https://doi.org/10.1093/nar/gkt1115
Orentas RJ, Yang JJ, Wen X et al (2012) Identification of cell surface proteins as potential immunotherapy targets in 12 pediatric cancers. Front Oncol 2:1–16. https://doi.org/10.3389/fonc.2012.00194
Osborn L, Hession C, Tizard R et al (1989) Direct expression cloning of vascular cell adhesion molecule 1, a cytokine-induced endothelial protein that binds to lymphocytes. Cell 59:1203–1211. https://doi.org/10.1016/0092-8674(89)90775-7
Östman A, Hellberg C, Böhmer FD (2006) Protein-tyrosine phosphatases and cancer. Nat Rev Cancer 6:307–320. https://doi.org/10.1038/nrc1837
Oti M (2006) Predicting disease genes using protein-protein interactions. J Med Genet 43:691–698. https://doi.org/10.1136/jmg.2006.041376
Pantazopoulos H, Katsel P, Haroutunian V et al (2020) Molecular signature of extracellular matrix pathology in schizophrenia. Eur J Neurosci. https://doi.org/10.1111/ejn.15009
Park J-J, Yi JY, Jin YB et al (2012) Sialylation of epidermal growth factor receptor regulates receptor activity and chemosensitivity to gefitinib in colon cancer cells. Biochem Pharmacol 83:849–857. https://doi.org/10.1016/j.bcp.2012.01.007
Parr C, Mirzaei N, Christian M, Sastre M (2015) Activation of the Wnt/β-catenin pathway represses the transcription of the β-amyloid precursor protein cleaving enzyme (BACE1) via binding of T-cell factor-4 to BACE1 promoter. FASEB J 29:623–635. https://doi.org/10.1096/fj.14-253211
Paumier JM, Thinakaran G (2019) Matrix metalloproteinase 13, a new target for therapy in Alzheimer’s disease. Genes Dis 6:1–2. https://doi.org/10.1016/j.gendis.2019.02.004
Peer E, Aichberger SK, Vilotic F et al (2021) Casein kinase 1d encodes a novel drug target in hedgehog–GLI-driven cancers and tumor-initiating cells resistant to Smo inhibition. Cancers 13:4227–4243. https://doi.org/10.3390/cancers13164227
Peles E, Nativ M, Lustig M et al (1997) Identification of a novel contactin-associated transmembrane receptor with multiple domains implicated in protein-protein interactions. EMBO J 16:978–988. https://doi.org/10.1093/emboj/16.5.978
Penzkofer S (2011) Screen for kinases affecting amyloidogenic cleavage by BACE1. Dissertation, University of Konstanz
Perez RG, Zheng H, Van Der Ploeg LHT, Koo EH (1997) The β-amyloid precursor protein of Alzheimer’s disease enhances neuron viability and modulates neuronal polarity. J Neurosci 17:9407–9414. https://doi.org/10.1523/jneurosci.17-24-09407.1997
Pernodet N, Hermetet F, Adami P et al (2012) High expression of QSOX1 reduces tumorigenesis, and is associated with a better outcome for breast cancer patients. Breast Cancer Res 14:1–15. https://doi.org/10.1186/bcr3341
Pike CJ, Carroll JC, Rosario ER, Barron AM (2009) Protective actions of sex steroid hormones in Alzheimer’s disease. Front Neuroendocrinol 30:239–258. https://doi.org/10.1016/j.yfrne.2009.04.015
Poliak S, Salomon D, Elhanany H et al (2003) Juxtaparanodal clustering of Shaker-like K+ channels in myelinated axons depends on Caspr2 and TAG-1. J Cell Biol 162:1149–1160. https://doi.org/10.1083/jcb.200305018
Pratte M, Rougon G, Schachner M, Jamon M (2003) Mice deficient for the close homologue of the neural adhesion cell L1 (CHL1) display alterations in emotional reactivity and motor coordination. Behav Brain Res 147:31–39. https://doi.org/10.1016/S0166-4328(03)00114-1
Priller C, Bauer T, Mitteregger G et al (2006) Synapse formation and function is modulated by the amyloid precursor protein. J Neurosci 26:7212–7221. https://doi.org/10.1523/JNEUROSCI.1450-06.2006
Pruessmeyer J, Martin C, Hess FM et al (2010) A Disintegrin and metalloproteinase 17 (ADAM17) mediates inflammation-induced shedding of syndecan-1 and -4 by lung epithelial cells. J Biol Chem 285:555–564. https://doi.org/10.1074/jbc.M109.059394
Rajapaksha TW, Eimer WA, Bozza TC, Vassar R (2011) The Alzheimer’s -secretase enzyme BACE1 is required for accurate axon guidance of olfactory sensory neurons and normal glomerulus formation in the olfactory bulb. Mol Neurodegener 6:88–96. https://doi.org/10.1186/1750-1326-6-88
Ramamoorthy S, Pereira AC (2020) Neurotoxic astrocytes secreted glypican-4 drives Alzheimer’s tau pathology. SSRN Electron J. https://doi.org/10.2139/ssrn.3716976
Ray WJ, Yao M, Nowotny P et al (1999) Evidence for a physical interaction between presenilin and Notch. Proc Natl Acad Sci U S A 96:3263–3268. https://doi.org/10.1073/pnas.96.6.3263
Rida PCG, Syed MI, Aneja R (2019) Time will tell: Circadian clock dysregulation in triple negative breast cancer. Front Biosci 11:178–192. https://doi.org/10.2741/S533
Rogaeva E, Meng Y, Lee JH et al (2007) The neuronal sortilin-related receptor SORL1 is genetically associated with Alzheimer disease. Nat Genet 39:168–177. https://doi.org/10.1038/ng1943
Rudan Njavro J, Klotz J, Dislich B et al (2020) Mouse brain proteomics establishes MDGA1 and CACHD1 as in vivo substrates of the Alzheimer protease BACE1. FASEB J 34:2465–2482. https://doi.org/10.1096/fj.201902347R
Sachse CC, Kim YH, Agsten M et al (2013) BACE1 and presenilin/γ-secretase regulate proteolytic processing of KCNE1 and 2, auxiliary subunits of voltage-gated potassium channels. FASEB J 27:2458–2467. https://doi.org/10.1096/fj.12-214056
Sanchez-Pulido L, Ponting CP (2018) TMEM132: An ancient architecture of cohesin and immunoglobulin domains define a new family of neural adhesion molecules. Bioinformatics 34:721–724. https://doi.org/10.1093/bioinformatics/btx689
Sato M, Takahashi K, Nagayama K et al (2005) Identification of chromosome arm 9p as the most frequent target of homozygous deletions in lung cancer. Genes, Chromosom Cancer 44:405–414. https://doi.org/10.1002/gcc.20253
Satoh J, Tabunoki H, Ishida T et al (2013) Accumulation of a repulsive axonal guidance molecule RGMa in amyloid plaques: a possible hallmark of regenerative failure in Alzheimer’s disease brains. Neuropathol Appl Neurobiol 39:109–120. https://doi.org/10.1111/j.1365-2990.2012.01281.x
Savonenko AV, Melnikova T, Laird FM et al (2008) Alteration of BACE1-dependent NRG1/ErbB4 signaling and schizophrenia-like phenotypes in BACE1-null mice. Proc Natl Acad Sci U.S.A. 105:5585–5590. https://doi.org/10.1073/pnas.0710373105
Scholefield Z, Yates EA, Wayne G et al (2003) Heparan sulfate regulates amyloid precursor protein processing by BACE1, the Alzheimer’s β-secretase. J Cell Biol 163:97–107. https://doi.org/10.1083/jcb.200303059
Schrötter A, Pfeiffer K, El Magraoui F et al (2012) The amyloid precursor protein (APP) family members are key players in S-adenosylmethionine formation by MAT2A and modify BACE1 and PSEN1 gene expression-relevance for Alzheimer’s disease. Mol Cell Proteomics 11:1274–1288. https://doi.org/10.1074/mcp.M112.019364
Schwarzman AL, Gregori L, Vitek MP et al (1994) Transthyretin sequesters amyloid β protein and prevents amyloid formation. Proc Natl Acad Sci U S A 91:8368–8372. https://doi.org/10.1073/pnas.91.18.8368
Seales EC, Shaikh FM, Woodard-Grice AV et al (2005) A protein kinase C/Ras/ERK signaling pathway activates myeloid fibronectin receptors by altering β1 integrin sialylation. J Biol Chem 280:37610–37615. https://doi.org/10.1074/jbc.M508476200
Shen LL, Li WW, Xu YL et al (2019) Neurotrophin receptor p75 mediates amyloid β-induced tau pathology. Neurobiol Dis 132:104567–104578. https://doi.org/10.1016/j.nbd.2019.104567
Shi CY, Fan Y, Liu B, Lou WH (2013) HIF1 contributes to hypoxia-induced pancreatic cancer cells invasion via promoting QSOX1 expression. Cell Physiol Biochem 32:561–568. https://doi.org/10.1159/000354460
Shin TM, Isas JM, Hsieh CL et al (2008) Formation of soluble amyloid oligomers and amyloid fibrils by the multifunctional protein vitronectin. Mol Neurodegener 3:1–12. https://doi.org/10.1186/1750-1326-3-16
Singh J, Itahana Y, Knight-Krajewski S et al (2004) Proteolytic enzymes and altered glycosylation modulate dystroglycan function in carcinoma cells. Cancer Res 64:6152–6159. https://doi.org/10.1158/0008-5472.CAN-04-1638
Sjödin S, Brinkmalm G, Öhrfelt A et al (2019) Endo-lysosomal proteins and ubiquitin CSF concentrations in Alzheimer’s and Parkinson’s disease. Alzheimer’s Res Ther 11:117–133. https://doi.org/10.1186/s13195-019-0533-9
Stark C (2006) BioGRID: a general repository for interaction datasets. Nucleic Acids Res 34:D535–D539. https://doi.org/10.1093/nar/gkj109
Straub V, Campbell KP (1997) Muscular dystrophies and the dystrophin-glycoprotein complex. Curr Opin Neurol 10:168–175. https://doi.org/10.1097/00019052-199704000-00016
Stützer I, Selevsek N, Esterházy D et al (2013) Systematic proteomic analysis identifies β-site amyloid precursor protein cleaving enzyme 2 and 1 (BACE2 and BACE1) substrates in pancreatic β-cells. J Biol Chem 288:10536–10547. https://doi.org/10.1074/jbc.M112.444703
Sun X, Tong Y, Qing H et al (2006) Increased BACE1 maturation contributes to the pathogenesis of Alzheimer’s disease in Down syndrome. FASEB J 20:1361–1368. https://doi.org/10.1096/fj.05-5628com
Szklarczyk D, Morris JH, Cook H et al (2017) The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res 45:D362–D368. https://doi.org/10.1093/nar/gkw937
Takahashi M, Kizuka Y, Ohtsubo K et al (2016) Disease-associated glycans on cell surface proteins. Mol Aspects Med 51:56–70. https://doi.org/10.1016/j.mam.2016.04.008
Tamura K, Chiu YW, Shiohara A et al (2020) EphA4 regulates Aβ production via BACE1 expression in neurons. FASEB J 34:16383–16396. https://doi.org/10.1096/fj.202001510R
Tian H, Bi X, Li CS et al (2013) Secreted phosphoprotein 24 kD (Spp24) and Spp14 affect TGF-β induced bone formation differently. PLoS ONE 8:1–10. https://doi.org/10.1371/journal.pone.0072645
Truong PH, Ciccotosto GD, Merson TD et al (2019) Amyloid precursor protein and amyloid precursor-like protein 2 have distinct roles in modulating myelination, demyelination, and remyelination of axons. Glia 67:525–538. https://doi.org/10.1002/glia.23561
Uetani N (2000) Impaired learning with enhanced hippocampal long-term potentiation in PTPdelta-deficient mice. EMBO J 19:2775–2785. https://doi.org/10.1093/emboj/19.12.2775
Usenovic M, Suon S, Gretzula CA, Parmentier-Batteur S (2018) P3–192: novel targets for blocking the uptake of tau oligomers in hipsc neurons. Alzheimer’s Dement 14:P1140–P1141. https://doi.org/10.1016/j.jalz.2018.06.1550
Vassar R (2019) Editorial: Implications for BACE1 inhibitor clinical trials: adult conditional BACE1 knockout mice exhibit axonal organization defects in the hippocampus. J Prev Alzheimer’s Dis 6:78–84. https://doi.org/10.14283/jpad.2019.3
Vetrivel KS, Zhang X, Meckler X et al (2008) Evidence that CD147 modulation of β-amyloid (Aβ) levels is mediated by extracellular degradation of secreted Aβ. J Biol Chem 283:19489–19498. https://doi.org/10.1074/jbc.M801037200
Vogel P, Hansen GM, Read RW et al (2012) Amelogenesis imperfecta and other biomineralization defects in Fam20a and Fam20c null mice. Vet Pathol 49:998–1017. https://doi.org/10.1177/0300985812453177
Von Arnim CAF, Kinoshita A, Peltan ID et al (2005) The low density lipoprotein receptor-related protein (LRP) is a novel β-secretase (BACE1) substrate. J Biol Chem 280:17777–17785. https://doi.org/10.1074/jbc.M414248200
Von Einem B, Wahler A, Schips T et al (2015) The Golgi-localized γ-ear-containing ARF-binding (GGA) proteins alter amyloid-β precursor protein (APP) processing through interaction of their GAE domain with the beta-Site APP cleaving enzyme 1 (BACE1). PLoS ONE 10:e0129047–e0129068. https://doi.org/10.1371/journal.pone.0129047
Walter J, Fluhrer R, Hartung B et al (2001) Phosphorylation regulates intracellular trafficking of β-secretase. J Biol Chem 276:14634–14641. https://doi.org/10.1074/jbc.M011116200
Wang H, Song L, Laird F et al (2008) BACE1 knock-outs display deficits in activity-dependent potentiation of synaptic transmission at mossy fiber to CA3 synapses in the hippocampus. J Neurosci 28:8677–8681. https://doi.org/10.1523/JNEUROSCI.2440-08.2008
Wang YC, Juan HC, Wong YH et al (2013) Protogenin prevents premature apoptosis of rostral cephalic neural crest cells by activating the α5βl1-integrin. Cell Death Dis 4:e651–e663. https://doi.org/10.1038/cddis.2013.177
Wang H, Megill A, Wong PC et al (2014) Postsynaptic target specific synaptic dysfunctions in the CA3 area of BACE1 knockout mice. PLoS ONE 9:e92279–e92288. https://doi.org/10.1371/journal.pone.0092279
Wang X, Jiang W, Luo S, et al (2020) The C. elegans homolog of TMEM132D, a human panic-disorder and anxiety risk gene, modulates neuronal morphogenesis through the WAVE-regulatory complex. BioRxiv. https://doi.org/10.1101/2020.11.16.385906
Warde-Farley D, Donaldson SL, Comes O et al (2010) The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res 38:214–220. https://doi.org/10.1093/nar/gkq537
Weyer SW, Klevanski M, Delekate A et al (2011) APP and APLP2 are essential at PNS and CNS synapses for transmission, spatial learning and LTP. EMBO J 30:2266–2280. https://doi.org/10.1038/emboj.2011.119
Willem M, Garratt AN, Novak B et al (2006) Control of peripheral nerve myelination by the β-secretase BACE1. Science 314:664–666. https://doi.org/10.1126/science.1132341
Wong HK, Sakurai T, Oyama F et al (2005) β subunits of voltage-gated sodium channels are novel substrates of β-site amyloid precursor protein-cleaving enzyme (BACE1) and γ-secretase. J Biol Chem 280:23009–23017. https://doi.org/10.1074/jbc.M414648200
Wong YH, Lu AC, Wang YC et al (2010) Protogenin defines a transition stage during embryonic neurogenesis and prevents precocious neuronal differentiation. J Neurosci 30:4428–4439. https://doi.org/10.1523/JNEUROSCI.0473-10.2010
Woo J, Kwon SK, Choi S et al (2009) Trans-synaptic adhesion between NGL-3 and LAR regulates the formation of excitatory synapses. Nat Neurosci 12:428–437. https://doi.org/10.1038/nn.2279
Woodard-Grice AV, McBrayer AC, Wakefield JK et al (2008) Proteolytic shedding of ST6Gal-I by BACE1 regulates the glycosylation and function of α4β1 integrins. J Biol Chem 283:26364–26373. https://doi.org/10.1074/jbc.M800836200
Wu M, Huang C, Gan K et al (2006) LRRC4, a putative tumor suppressor gene, requires a functional leucine-rich repeat cassette domain to inhibit proliferation of glioma cells in vitro by modulating the extracellular signal-regulated kinase/protein kinase B/nuclear factor-kappaB pathway. Mol Biol Cell 17:3534–3542. https://doi.org/10.1091/mbc.e05-11-1082
Wu T, Li Y, Chen B (2020) B4GALT3 promotes cell proliferation and invasion in glioblastoma. Neurol Res 42:463–470. https://doi.org/10.1080/01616412.2020.1740465
Yamagata M, Weiner JA, Sanes JR (2002) Sidekicks: synaptic adhesion molecules that promote lamina-specific connectivity in the retina. Cell 110:649–660. https://doi.org/10.1016/S0092-8674(02)00910-8
Yao XQ, Jiao SS, Saadipour K et al (2015) P75NTR ectodomain is a physiological neuroprotective molecule against amyloid-beta toxicity in the brain of Alzheimer’s disease. Mol Psychiatry 20:1301–1310. https://doi.org/10.1038/mp.2015.49
Young-Pearse TL, Bai J, Chang R et al (2007) A critical function for β-amyloid precursor protein in neuronal migration revealed by in utero RNA interference. J Neurosci 27:14459–14469. https://doi.org/10.1523/JNEUROSCI.4701-07.2007
Yurchenko V, Constant S, Eisenmesser E, Bukrinsky M (2010) Cyclophilin-CD147 interactions: a new target for anti-inflammatory therapeutics. Clin Exp Immunol 160:305–317. https://doi.org/10.1111/j.1365-2249.2010.04115.x
Yurchenko V, O’Connor M, Dai WW et al (2001) CD147 is a signaling receptor for cyclophilin B. Biochem Biophys Res Commun 288:786–788. https://doi.org/10.1006/bbrc.2001.5847
Zhang C, Wang X, Cui J et al (2016) Synthetic analogues of betulinic acid as potent inhibitors of PS1/BACE1 interaction to reduce Aβ generation. Chin J Chem 35:103–112. https://doi.org/10.1002/cjoc.201600611
Zhang F, Wei J, Li X et al (2018) Early candidate urine biomarkers for detecting Alzheimer’s disease before amyloid-β plaque deposition in an APP (swe)/PSEN1 dE9 transgenic mouse model. J Alzheimer’s Dis 66:613–637. https://doi.org/10.3233/JAD-180412
Zhang XF, Wang J, Jia HL et al (2019) Core fucosylated glycan-dependent inhibitory effect of QSOX1-S on invasion and metastasis of hepatocellular carcinoma. Cell Death Discov 5:84–95. https://doi.org/10.1038/s41420-019-0164-8
Zhao YY, Takahashi M, Gu JG et al (2008) Functional roles of N-glycans in cell signaling and cell adhesion in cancer. Cancer Sci 99:1304–1310. https://doi.org/10.1111/j.1349-7006.2008.00839.x
Zheng H, Jiang M, Trumbauer ME et al (1995) β-amyloid precursor protein-deficient mice show reactive gliosis and decreased locomotor activity. Cell 81:525–531. https://doi.org/10.1016/0092-8674(95)90073-X
Zhou L, Barão S, Laga M et al (2012) The neural cell adhesion molecules L1 and CHL1 are cleaved by BACE1 protease in vivo. J Biol Chem 287:25927–25940. https://doi.org/10.1074/jbc.M112.377465
Zuberi K, Franz M, Rodriguez H et al (2013) GeneMANIA prediction server 2013 update. Nucleic Acids Res 41:115–122. https://doi.org/10.1093/nar/gkt533
Zuhl AM, Nolan CE, Brodney MA et al (2016) Chemoproteomic profiling reveals that cathepsin D off-target activity drives ocular toxicity of β-secretase inhibitors. Nat Commun 7:1–14. https://doi.org/10.1038/ncomms13042
Acknowledgements
The research was conducted from the financial resources of Biochemworld co., Uppsala County, Sweden.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict to declare.
Ethical approval
Not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
232_2022_225_MOESM1_ESM.xlsx
Supplementary file1 (XLSX 70 KB) The full result of the pathway enrichment and gene function prediction analysis of the BACE1 non-substrate PPIN. The network was constructed by GeneMania (3.5.2) application run under Cytoscape (3.9.0) environment utilizing the H. sapiens database (update 13-07-2017) and including all types of interaction networks. The table also includes the input protein list (column Gene 7-35)
232_2022_225_MOESM2_ESM.xlsx
Supplementary file2 (XLSX 373 KB) The full result of the pathway enrichment and gene function analysis of the BACE1 substrate PPIN. The network was constructed by applying the GeneMania (3.5.2) application run under Cytoscape (3.7.2) environment utilizing the H. sapiens database (update 13-07-2017) and including all types of interaction networks. The table also includes the input protein list (column Gene 7-226)
232_2022_225_MOESM3_ESM.xlsx
Supplementary file3 (XLSX 67 KB) The full result of the pathway enrichment analysis and gene function prediction analysis of the interaction network of BACE1 and its substrates PRTG, VCAM1, ST6GAL1, and the integrins glycosylated by ST6GAL1 (ITGA5, ITGA4, ITGB1). The network was constructed by applying the GeneMania (3.5.2) application run under Cytoscape (3.7.2) environment utilizing the H. sapiens database (update 13-07-2017) and including all types of the interaction networks
Rights and permissions
About this article
Cite this article
Nahálková, J. Finding New Ways How to Control BACE1. J Membrane Biol 255, 293–318 (2022). https://doi.org/10.1007/s00232-022-00225-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-022-00225-1