Abstract
Chromosomal structural changes can be induced by the addition of specific alien chromosomes called ‘gametocidal (Gc) chromosomes’. In the monosomic addition of the Gc chromosome to common wheat, chromosomal breaks occur in gametes receiving no Gc chromosome, and the broken ends heal and stabilize in the subsequent generations. Thus, by the so-called Gc system, deficiencies and translocations can be induced in common wheat and also in alien chromosome addition and substitution lines of common wheat. Deficiencies of wheat and alien chromosomes were cytologically identified by the chromosome banding and in situ hybridization techniques. The plants carrying deletions or wheat-alien translocations were established as new aneuploid lines of common wheat with sub-arm aneuploidy. Those for wheat chromosomes are called deletion stocks and those for alien chromosomes are called dissection lines. The new aneuploids have been used for cytological chromosome mapping and have corrected some mistakes in genetic mapping.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Aneuploids of Common Wheat
The hexaploid nature of common wheat (Triticum aestivum L., 2n = 6x = 42) enables us to produce aneuploids rather easily. Almost 60 year ago Sears (1954) first reported the systematic production of aneuploid lines in common wheat. These so-called conventional aneuploids comprise different series of aneuploid lines of Chinese Spring wheat, such as monosomics, nullisomic, nullisomic-tetrasomics and ditelosomics (Sears 1954; Sears and Sears 1978), which have been used extensively for the genetic and genomic studies of wheat. He allowed everybody to use those aneuploids freely. I believe that his generosity stimulated the subsequent development of not only genetic and cytogenetic studies but also molecular and genomics studies of wheat. Common wheat can also tolerate the addition of alien chromosomes from related wild and cultivated species, such as goatgrass, wheatgrass, rye and barley. In some cases, alien chromosomes substitute for wheat chromosomes very well to be established as common wheat cultivars, e.g., those carrying a 1R(1B) substitution.
The new aneuploids described here are not really new, but I would like to distinguish them from the conventional aneuploid lines that have aneuploidy of whole chromosomes or chromosome arms. I would like to define these new aneuploids as ‘aneuploids that have sub-arm aneuploidy’, which can be induced by two genetic mechanisms, as well as by artificial mutation. By the suppression of the pairing control (ph: pairing homoeologous) gene, homoeologous pairing is induced between wheat and alien chromosomes to generate wheat-alien recombinant chromosomes in alien substitution lines of common wheat. Also, unique chromosomes called gametocidal (Gc) chromosomes, which were introduced from specific wild species of the genus Aegilops, induce chromosomal breakage in common wheat to generate deficient chromosomes and wheat-alien recombinant chromosomes. I named the common wheat lines carrying deficient wheat chromosomes ‘deletion stocks’ (Endo and Gill 1996), and I proposed to call generically the wheat lines carrying deletions or translocations of alien chromosomes ‘dissection’ lines (Endo 2007).
Advanced Techniques to Check Aneuploids
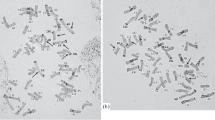
The addition of whole alien chromosomes or chromosome arms is generally detrimental to the performance of common wheat. The new aneuploids with dissected chromosomes at the sub-arm level are more useful not only for breeding purposes but also for genomic analysis. Two cytological techniques ‘chromosome banding’ and ‘fluorescence in situ hybridization’ are most useful for the identification of chromosomes and sub-arm aberrations. C-banding allows us to identify all common wheat chromosomes (Gill et al. 1991) and not only to verify the conventional wheat aneuploids in mitotic metaphase cells but also to detect deficient chromosomes. Fluorescence in situ hybridization (FISH) is useful for locating specific DNA sequences on chromosomes. The genomic in situ hybridization (GISH), which uses probes of total genomic DNA from alien species, is useful for the detection of alien chromosomes and chromosomal segments introduced into wheat. The combination of chromosome banding and FISH/GISH is an even more powerful cytological technique for identifying alien chromosomes in wheat because we can identify chromosomes and locate specific DNA sequences in the same chromosome preparation (Fig. 8.1).
A mitotic metaphase cell of the barley 6H addition line of common wheat depicted by the combination of C-banding and FISH/GISH. The C-banding pattern shows that this line has a chromosomal complement of common wheat, and FISH indicates the presence of rDNA sequences in NORs, and GISH confirmed the disomic addition of barley chromosome 6H
Gametocidal Mechanism
Some alien chromosomes called gametocidal (Gc) chromosomes ensure their existence in common wheat in a selfish manner. When the Gc chromosome exists in the monosomic condition, two types of gametophyte are produced, those carrying the Gc chromosome and those without the Gc chromosome, and chromosome breakage occurs only in the latter gametophytes (Fig. 8.2). Such Gc-induced chromosomal breakage leads to either the sterility of gametes or the production of fertile gametes carrying chromosomal mutations, and the induced chromosomal mutations become stabilized in subsequent generations (Endo 1990).
Deletion Stocks of Common Wheat
Two Gc chromosomes have been used to generate deletion and dissection lines. One is chromosome 2C derived from Aegilops cylindirica Host. and the other is chromosome 3C derived from Aegilops triuncialis L. (Endo 1988). For the production of the deletion stocks, I started cytological selection in the progeny of the Gc (mainly 2C) monosomic addition line crossed with euploid Chinese Spring. Deletions took place in the heterozygous condition and often multiple deletions occurred in single plants as shown in Fig. 8.3. I further selected the selfed progeny of such plants to obtain single-deletion homozygotes. In some cases I ended up getting multiple-deletion homozygous lines and also failed to obtain homozygotes for specific chromosome arms like the short arm of chromosome 4B. Table 8.1 summarizes the distribution of deletion breakpoints among different chromosome arms. I examined about 500 primary progeny, and in the end I identified 436 deletions and established deletion-homozygous lines for about 350 (ca. 80 %) of the deletions. This means that almost one deletion per plant occurred on average. In other words, we can expect to obtain one deletion for a specific chromosome in 4 ~ 5 % of plants we examine. The breakpoints of the deletions seem to be randomly distributed, but it is difficult to tell whether or not there are hotspots of Gc-induced chromosomal breaks.
Undoubtedly the deletion stocks are useful for chromosome mapping, especially for deletion mapping of molecular markers. Werner et al. (1992) conducted deletion mapping of RFLP using the deletion stocks for the first time. Since then a series of papers on wheat chromosome deletion or bin mapping of various DNA markers using deletion stocks have been published (e.g. Qi et al. 2004). These deletion-homozygous stocks are distributed from National BioResource Project-Wheat (http://www.shigen.nig.ac.jp/wheat/komugi/).
Dissection of Alien Chromosomes
The Gc system can also be used to dissect alien (A) chromosomes introduced into common wheat. The cross scheme is simple: First, make a cross between the Gc disomic addition and an alien disomic addition; then, backcross the hybrid to the alien addition to produce plants disomic for the alien chromosome and monosomic for the Gc chromosome (42 + A” + Gc’) (Endo 2007). In the progeny of this plant, we can find plants carrying structurally changed alien chromosomes, as well as aberrant wheat chromosomes.
By FISH/GISH we can undoubtedly identify deletions and translocations of alien chromosomes and can tell the exact breakpoint of a translocation between alien and wheat chromosomes (Fig. 8.4). The Gc system was successfully applied to produce dissection lines of various alien chromosomes in common wheat, such as barley chromosomes (2H, Joshi et al. 2011; 3H, Sakai et al. 2009; 4H, Sakata et al. 2010; 5H, Ashida et al.2007; 7H, Masoudi-Nejad et al. 2005), rye 1R chromosomes (Tsuchida et al. 2008; Gyawali et al. 2009, 2010) and rye B chromosomes (Endo et al. 2008). All possible kinds of rearrangements of alien chromosomes have been obtained (Fig. 8.5).
Examples of alien chromosome aberrations induced by the Gc system. The green FISH signals indicate the subtelomeric repeats HvT01 for the barley 5H and 7H chromosomes derived from a cultivar Betzes, the pSc200 repeats for the rye 1R chromosome from a cultivar Imperial, and the E1100 repeats for the rye B chromosome from a Siberian cultivar. The GISH signals are shown in red for all chromosomes
PCR-Based Mass Selection of Gc-Induced Deletions for Specific Chromosomes
The chromosomal deficiencies in the deletion stocks of common wheat were cytologically identified and characterized by chromosome banding. Then the breakpoints of the deleted chromosomes were analyzed with DNA markers. Once the positions of the DNA markers are located on a chromosome, more deficiencies of the chromosome can be identified using chromosome-specific DNA markers and appropriate aneuploids. For example, using PCR-based 6B-specific markers, a search was made for deficiencies of chromosome 6B among the progeny from a cross between nullisomic 6B-tetrasomic 6A and a monosomic 2C addition line as the pollen parent (Fig. 8.6). Any deficiencies occurring in the 6B chromosome within the two markers could be detected by PCR because no 6B homologue was transmitted from the female parent. Thus, 102 (5.0 %) of the 2,041 hybrid plants were found to have a deficiency in either or both chromosome arms (unpublished data). In a similar way, Gc-induced deletions of alien chromosomes in common wheat can be identified by PCR. This PCR selection should find deletions overlooked by cytology and moreover probably detect structural changes that do not exist in the root tips but exist in aerial parts of plants. Joshi et al. (2013) conducted PCR analysis in 81 plants carrying a cytologically normal-appearing 2H chromosome in root tips and detected 2H aberration in the leaves of 6 of them. This fact implied the ongoing production of aberrations after fertilization.
Epilogue
Thanks to the next-generation sequencing technologies, sequencing the entire genomes of wheat, barley and rye has become a reality. Several chromosomal landmarks will be needed to assemble contigs into supercontigs or even into chromosomes. The Gc system will help provide a virtually limitless number of such landmarks, namely breakpoints of deleted or translocated chromosomes of the new aneuploids of common wheat.
Lastly, I wish to express my gratitude to Prof. Bikram S. Gill for his encouragement to establish deletion stocks common wheat by using them for his wheat chromosome mapping studies, otherwise I would not have done these works mentioned above.
References
Ashida T, Nasuda S, Sato K, Endo TR (2007) Dissection of barley chromosome 5H in common wheat. Genes Genet Syst 82:123–133
Endo TR (1988) Induction of chromosomal structural changes by a chromosome of Aegilops cylindrica L. in common wheat. J Hered 79:366–370
Endo TR (1990) Gametocidal chromosomes and their induction of chromosome mutations in wheat. Jpn J Genet 65:135–152
Endo TR (2007) The gametocidal chromosome as a tool for chromosome manipulation in wheat. Chromosom Res 15:67–75
Endo TR, Gill BS (1996) The deletion stocks of common wheat. J Hered 87:295–307
Endo TR, Nasuda S, Jones N et al (2008) Dissection of rye B chromosomes, and nondisjunction properties of the dissected segments in a common wheat background. Genes Genet Syst 83:23–30
Gill BS, Friebe B, Endo TR (1991) Standard karyotype and nomenclature system for description of chromosome bands and structural aberrations in wheat (Triticum aestivum). Genome 34:830–839
Gyawali YP, Nasuda S, Endo TR (2009) Cytological dissection and molecular characterization of chromosome 1R derived from ‘Burgas 2’ common wheat. Genes Genet Syst 84:407–416
Gyawali YP, Nasuda S, Endo TR (2010) A cytological map of the short arm of rye chromosome 1R constructed with 1R dissection stocks of common wheat and PCR-based markers. Cytogenet Genome Res 129:224–233
Joshi GP, Nasuda S, Endo TR (2011) Dissection and cytological mapping of barley chromosome 2H in the genetic background of common wheat. Genes Genet Syst 86:231–248
Joshi GP, Endo TR, Nasuda S (2013) PCR and sequence analysis of barley chromosome 2H subjected to the gametocidal action of chromosome 2C. Theor Appl Genet 126:2381–2390
Masoudi-Nejad A, Nasuda S, Bihoreau M-T et al (2005) An alternative to radiation hybrid mapping for large-scale genome analysis in barley. Mol Genet Genomics 274:589–594
Qi LL, Echalier B, Chao S et al (2004) A chromosome bin map of 16,000 expressed sequence tag loci and distribution of genes among the three genomes of polyploidy wheat. Genetics 168:701–712
Sakai K, Nasuda S, Sato K, Endo TR (2009) Dissection of barley chromosome 3H in common wheat and comparison of 3H physical and genetic maps. Genes Genet Syst 84:25–34
Sakata M, Nasuda S, Endo TR (2010) Dissection of barley chromosome 4H in common wheat by the gametocidal system and cytological mapping of chromosome 4H with EST markers. Genes Genet Syst 85:19–29
Sears ER (1954) The aneuploids of common wheat. Missouri Agr Expt Sta Res Bull 572:1–59
Sears ER, Sears LMS (1978) The telocentric chromosomes of common wheat. In: Proceedings of 5th international wheat genetics symposium, New Delhi, pp 389–407
Tsuchida M, Fukushima T, Nasuda S et al (2008) Dissection of rye chromosome 1R in common wheat. Genes Genet Syst 83:43–53
Werner JE, Endo TR, Gill BS (1992) Toward a cytogenetically based physical map of the wheat genome. Proc Natl Acad Sci U S A 89:11307–11311
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution Noncommercial License, which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Copyright information
© 2015 The Author(s)
About this paper
Cite this paper
Endo, T.R. (2015). New Aneuploids of Common Wheat. In: Ogihara, Y., Takumi, S., Handa, H. (eds) Advances in Wheat Genetics: From Genome to Field. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55675-6_8
Download citation
DOI: https://doi.org/10.1007/978-4-431-55675-6_8
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55674-9
Online ISBN: 978-4-431-55675-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)