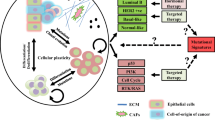
Abstract
Breast cancer is the most commonly diagnosed cancer in women in United States. From data of American Cancer Society from 2007 reported total of 178,480 women diagnosed with breast cancer. The death rate from breast cancer has decreased in North America over time, but still accounts for second highest cancer death, following lung cancer. Breast cancer is staged based on tumor size, nodal involvement, and distant metastasis like any other solid tumors. However clinical staging is not the only important factor in management of breast cancer. Various molecular features divides breast cancer into many subgroups – that act differently, and respond differently from therapy. Thus the focus of breast cancer treatment has evolved focusing on specific targets. The most important biologic markers in subtyping of breast cancer so far are hormone receptor positivity and HER2/neu protein expression. Five molecular subtypes using intrinsic gene set include Basal mRNA, HER2 + mRNA, Luminal AmRNA, Luminal B mRNA, and Normal-like mRNA. In addition, better understanding of genetic target of breast cancer has given us arsenal of personalized, and more effective treatment approach.
This review will focus on examples that highlight several mechanism of tumorigenesis, giving us not just understanding of gene pathways and the molecular biology, that could lead us to therapeutic target. Several important molecular targets have been investigated in preclinical and clinical trials, others are yet to be explored. We will also describe genetic mechanisms discovery related to overcoming resistance to current targeted therapies in breast cancer, including hormone receptor expression and HER 2- neu amplification. We will also review other exciting developments in understanding of breast cancer, the tumor microenvironment and cancer stem cells, and targeting agents in that area.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sjogren S, et al. Prognostic and predictive value of c-erbB-2 overexpression in primary breast cancer, alone and in combination with other prognostic markers. J Clin Oncol. 1998;16(2):462–9.
Slamon DJ, et al. Studies of the HER-2/neu proto-oncogene in human breast and ovarian cancer. Science. 1989;244(4905):707–12.
Graus-Porta D, et al. ErbB-2, the preferred heterodimerization partner of all ErbB receptors, is a mediator of lateral signaling. EMBO J. 1997;16(7):1647–55.
Dankort D, et al. Grb2 and Shc adapter proteins play distinct roles in Neu (ErbB-2)-induced mammary tumorigenesis: implications for human breast cancer. Mol Cell Biol. 2001;21(5):1540–51.
Slamon DJ, et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med. 2001;344(11):783–92.
Gajria D, Chandarlapaty S. HER2-Amplified breast cancer: mechanisms of trastuzumab resistance and novel targeted therapies. Expert Rev Anticancer Ther. 2011;11(2):263–75.
Romond EH, et al. Trastuzumab plus adjuvant chemotherapy for operable HER2-positive breast cancer. N Engl J Med. 2005;353(16):1673–84.
Piccart-Gebhart MJ, et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med. 2005;353(16):1659–72.
Bange J, Zwick E, Ullrich A. Molecular targets for breast cancer therapy and prevention. Nat Med. 2001;7(5):548–52.
Ritter CA, et al. Mechanisms of resistance development against trastuzumab (Herceptin) in an in vivo breast cancer model. Int J Clin Pharmacol Ther. 2004;42(11):642–3.
Lu Y, et al. Insulin-like growth factor-I receptor signaling and resistance to trastuzumab (Herceptin). J Natl Cancer Inst. 2001;93(24):1852–7.
Shattuck DL, et al. Met receptor contributes to trastuzumab resistance of Her2-overexpressing breast cancer cells. Cancer Res. 2008;68(5):1471–7.
Serra V, et al. PI3K Inhibition results in enhanced HER signaling and acquired ERK dependency in HER2-overexpressing breast cancer. Oncogene. 2011;30(22):2547–57.
Scaltriti M, et al. Expression of p95HER2, a truncated form of the HER2 receptor, and response to anti-HER2 therapies in breast cancer. J Natl Cancer Inst. 2007;99(8):628–38.
Berns K, et al. A functional genetic approach identifies the PI3K pathway as a major determinant of trastuzumab resistance in breast cancer. Cancer Cell. 2007;12(4):395–402.
Nagata Y, et al. PTEN activation contributes to tumor inhibition by trastuzumab, and loss of PTEN predicts trastuzumab resistance in patients. Cancer Cell. 2004;6(2):117–27.
Hynes NE, Lane HA. ERBB receptors and cancer: the complexity of targeted inhibitors. Nat Rev Cancer. 2005;5(5):341–54.
Hynes NE, MacDonald G. ErbB receptors and signaling pathways in cancer. Curr Opin Cell Biol. 2009;21(2):177–84.
Lee-Hoeflich ST, et al. A central role for HER3 in HER2-amplified breast cancer: implications for targeted therapy. Cancer Res. 2008;68(14):5878–87.
Sergina NV, et al. Escape from HER-family tyrosine kinase inhibitor therapy by the kinase-inactive HER3. Nature. 2007;445(7126):437–41.
Yao E, et al. Suppression of HER2/HER3-mediated growth of breast cancer cells with combinations of GDC-0941 PI3K inhibitor, trastuzumab, and pertuzumab. Clin Cancer Res. 2009;15(12):4147–56.
Sachdev JC, Jahanzeb M. Blockade of the HER family of receptors in the treatment of HER2-positive metastatic breast cancer. Clin Breast Cancer. 2011;12:19–29.
Sanchez-Martin M, Pandiella A. Differential action of small molecule HER kinase inhibitors on receptor heterodimerization: therapeutic implications. Int J Cancer. 2011;131:244–52.
Noonberg SB, Benz CC. Tyrosine kinase inhibitors targeted to the epidermal growth factor receptor subfamily: role as anticancer agents. Drugs. 2000;59(4):753–67.
Blackwell KL, et al. Randomized study of lapatinib alone or in combination with trastuzumab in women with ErbB2-positive, trastuzumab-refractory metastatic breast cancer. J Clin Oncol. 2010;28(7):1124–30.
Blackwell KL, et al. Single-agent lapatinib for HER2-overexpressing advanced or metastatic breast cancer that progressed on first- or second-line trastuzumab-containing regimens. Ann Oncol. 2009;20(6):1026–31.
Kaufman B, et al. Lapatinib monotherapy in patients with HER2-overexpressing relapsed or refractory inflammatory breast cancer: final results and survival of the expanded HER2+ cohort in EGF103009, a phase II study. Lancet Oncol. 2009;10(6):581–8.
ALTTO (Adjuvant Lapatinib and/or Trastuzumab Treatment Optimisation) study. BIG 2-06/N063D. NCT00490139.
Gupta M, et al. Clinical implications of pathophysiological and demographic covariates on the population pharmacokinetics of trastuzumab emtansine, a HER2-targeted antibody-drug conjugate, in patients with HER2-positive metastatic breast cancer. J Clin Pharmacol. 2012;52:691–703.
Baselga J, Swain SM. CLEOPATRA: a phase III evaluation of pertuzumab and trastuzumab for HER2-positive metastatic breast cancer. Clin Breast Cancer. 2010;10(6):489–91.
Verma S, et al. Trastuzumab emtansine for HER2-positive advanced breast cancer. N Engl J Med 367:1783-1791, doi:10.1056/NEJMoa1209124 (2012).
Kai S, Hara H. Allogeneic hematopoietic stem cell transplantation. Ther Apher Dial. 2003;7(3):285–91.
Kai T, Spradling A. An empty drosophila stem cell niche reactivates the proliferation of ectopic cells. Proc Natl Acad Sci USA. 2003;100(8):4633–8.
Tokuzawa Y, et al. Fbx15 is a novel target of Oct3/4 but is dispensable for embryonic stem cell self-renewal and mouse development. Mol Cell Biol. 2003;23(8):2699–708.
Fiebiger W, et al. Occurrence of a pulmonary carcinoid following allogeneic stem cell transplantation for chronic myelogenous leukemia: a case report. Ann Hematol. 2003;82(6):374–6.
Smalley M, Ashworth A. Stem cells and breast cancer: a field in transit. Nat Rev Cancer. 2003;3(11):832–44.
Zhang Y, et al. Elevated insulin-like growth factor 1 receptor signaling induces antiestrogen resistance through the MAPK/ERK and PI3K/Akt signaling routes. Breast Cancer Res. 2011;13(3):R52.
Study of BMS-754807 combined with letrozole or BMS-754807 alone in patients with hormone receptor-positive breast cancer and resistance to non-steroidal aromatase inhibitors. Clinical Trials.gov # NCT01225172.
Litzenburger BC, et al. High IGF-IR activity in triple-negative breast cancer cell lines and tumorgrafts correlates with sensitivity to anti-IGF-IR therapy. Clin Cancer Res. 2011;17(8):2314–27.
Kleinsmith LJ, Pierce Jr GB. Multipotentiality of single embryonal carcinoma cells. Cancer Res. 1964;24:1544–51.
Nishimura EK, et al. Dominant role of the niche in melanocyte stem-cell fate determination. Nature. 2002;416(6883):854–60.
Folkman J. Tumor angiogenesis in women with node-positive breast cancer. Cancer J Sci Am. 1995;1(2):106–8.
Folkman J. Angiogenesis in cancer, vascular, rheumatoid and other disease. Nat Med. 1995;1(1):27–31.
Dayan F, et al. A dialogue between the hypoxia-inducible factor and the tumor microenvironment. Cancer Microenviron. 2008;1(1):53–68.
Fujita Y, Abe R, Shimizu H. Clinical approaches toward tumor angiogenesis: past, present and future. Curr Pharm Des. 2008;14(36):3820–34.
Poon RT, Fan ST, Wong J. Clinical implications of circulating angiogenic factors in cancer patients. J Clin Oncol. 2001;19(4):1207–25.
Baeriswyl V, Christofori G. The angiogenic switch in carcinogenesis. Semin Cancer Biol. 2009;19(5):329–37.
Zhao J, et al. Correlation between serum vascular endothelial growth factor and endostatin levels in patients with breast cancer. Cancer Lett. 2004;204(1):87–95.
Weidner N, et al. Tumor angiogenesis and metastasis–correlation in invasive breast carcinoma. N Engl J Med. 1991;324(1):1–8.
Weidner N, et al. Tumor angiogenesis correlates with metastasis in invasive prostate carcinoma. Am J Pathol. 1993;143(2):401–9.
Semenza GL. Involvement of oxygen-sensing pathways in physiologic and pathologic erythropoiesis. Blood. 2009;114(10):2015–9.
Semenza GL. Regulation of oxygen homeostasis by hypoxia-inducible factor 1. Physiology (Bethesda). 2009;24:97–106.
Simon MC, Keith B. The role of oxygen availability in embryonic development and stem cell function. Nat Rev Mol Cell Biol. 2008;9(4):285–96.
Tang N, et al. Loss of HIF-1alpha in endothelial cells disrupts a hypoxia-driven VEGF autocrine loop necessary for tumorigenesis. Cancer Cell. 2004;6(5):485–95.
Rosen LS. Clinical experience with angiogenesis signaling inhibitors: focus on vascular endothelial growth factor (VEGF) blockers. Cancer Control. 2002;9(2 Suppl):36–44.
LeCouter J, et al. Identification of an angiogenic mitogen selective for endocrine gland endothelium. Nature. 2001;412(6850):877–84.
LeCouter J, Ferrara N. EG-VEGF and the concept of tissue-specific angiogenic growth factors. Semin Cell Dev Biol. 2002;13(1):3–8.
Ferrara N, et al. EG-VEGF and Bv8: a novel family of tissue-restricted angiogenic factors. Biochim Biophys Acta. 2004;1654(1):69–78.
Ferrara N, et al. Differential expression of the angiogenic factor genes vascular endothelial growth factor (VEGF) and endocrine gland-derived VEGF in normal and polycystic human ovaries. Am J Pathol. 2003;162(6):1881–93.
Kamiyama M, et al. VEGF receptor antisense therapy inhibits angiogenesis and peritoneal dissemination of human gastric cancer in nude mice. Cancer Gene Ther. 2002;9(2):197–201.
Mathur SP, et al. Serum vascular endothelial growth factor C (VEGF-C) as a specific biomarker for advanced cervical cancer: relationship to insulin-like growth factor II (IGF-II), IGF binding protein 3 (IGF-BP3) and VEGF-A [corrected]. Gynecol Oncol. 2005;98(3):467–83.
Mathur RS, Mathur SP. Vascular endothelial growth factor (VEGF) up-regulates epidermal growth factor receptor (EGF-R) in cervical cancer in vitro: this action is mediated through HPV-E6 in HPV-positive cancers. Gynecol Oncol. 2005;97(1):206–13.
Bachtiary B, et al. Serum VEGF levels in patients undergoing primary radiotherapy for cervical cancer: impact on progression-free survival. Cancer Lett. 2002;179(2):197–203.
Cao X, et al. Upregulation of VEGF-A and CD24 gene expression by the tGLI1 transcription factor contributes to the aggressive behavior of breast cancer cells. Oncogene. 2012;31(1):104–15.
Liu L, et al. VEGF 936C>T polymorphism and breast cancer risk: evidence needed further clarification. Breast Cancer Res Treat. 2011;127(2):569–71.
Liu L, et al. Expression of connective tissue growth factor is in agreement with the expression of VEGF, VEGF-C, -D and associated with shorter survival in gastric cancer. Pathol Int. 2007;57(11):712–8.
Nicolini A, et al. Vascular endothelial growth factor (VEGF) and other common tissue prognostic indicators in breast cancer: a case-control study. Int J Biol Markers. 2004;19(4):275–81.
Kostopoulos I, et al. Evaluation of the prognostic value of HER-2 and VEGF in breast cancer patients participating in a randomized study with dose-dense sequential adjuvant chemotherapy. Breast Cancer Res Treat. 2006;96(3):251–61.
Gadducci A, et al. Serum preoperative vascular endothelial growth factor (VEGF) in epithelial ovarian cancer: relationship with prognostic variables and clinical outcome. Anticancer Res. 1999;19(2B):1401–5.
Des Guetz G, et al. Microvessel density and VEGF expression are prognostic factors in colorectal cancer. Meta-analysis of the literature. Br J Cancer. 2006;94(12):1823–32.
Bremnes RM, Camps C, Sirera R. Angiogenesis in non-small cell lung cancer: the prognostic impact of neoangiogenesis and the cytokines VEGF and bFGF in tumours and blood. Lung Cancer. 2006;51(2):143–58.
Balbi G, et al. Vascular endothelial growth factor (VEGF): can we use it as prognostic factor in endometrial cancer? Minerva Ginecol. 2006;58(5):411–5.
Jain RK. Normalization of tumor vasculature: an emerging concept in antiangiogenic therapy. Science. 2005;307(5706):58–62.
Hughes CC. Endothelial-stromal interactions in angiogenesis. Curr Opin Hematol. 2008;15(3):204–9.
Xian X, et al. Pericytes limit tumor cell metastasis. J Clin Invest. 2006;116(3):642–51.
Lindblom P, et al. Endothelial PDGF-B retention is required for proper investment of pericytes in the microvessel wall. Genes Dev. 2003;17(15):1835–40.
Augustin HG. Translating angiogenesis research into the clinic: the challenges ahead. Br J Radiol. 2003;76(Spec No 1):S3–10.
Ferrara N, Gerber HP, LeCouter J. The biology of VEGF and its receptors. Nat Med. 2003;9(6):669–76.
Gerber HP, Ferrara N. The role of VEGF in normal and neoplastic hematopoiesis. J Mol Med (Berl). 2003;81(1):20–31.
D’Agostino Sr RB. Changing end points in breast-cancer drug approval – the avastin story. N Engl J Med. 2011;365(2):e2.
Baselga J. Targeting the phosphoinositide-3 (PI3) kinase pathway in breast cancer. Oncologist. 2011;16(Suppl 1):12–9.
Alvarez RH, Valero V, Hortobagyi GN. Emerging targeted therapies for breast cancer. J Clin Oncol. 2010;28(20):3366–79.
O’Regan RM, Khuri FR. Farnesyl transferase inhibitors: the next targeted therapies for breast cancer? Endocr Relat Cancer. 2004;11(2):191–205.
Eckert LB, et al. Involvement of Ras activation in human breast cancer cell signaling, invasion, and anoikis. Cancer Res. 2004;64(13):4585–92.
Li T, et al. Phase II trial of the farnesyltransferase inhibitor tipifarnib plus fulvestrant in hormone receptor-positive metastatic breast cancer: New York Cancer Consortium Trial P6205. Ann Oncol. 2009;20(4):642–7.
Johnston SR, et al. A phase II, randomized, blinded study of the farnesyltransferase inhibitor tipifarnib combined with letrozole in the treatment of advanced breast cancer after antiestrogen therapy. Breast Cancer Res Treat. 2008;110(2):327–35.
Stambolic V, et al. Utility of metformin in breast cancer treatment, is neoangiogenesis a risk factor? Breast Cancer Res Treat. 2009;114(2):387–9.
Dowling RJ, et al. Metformin inhibits mammalian target of rapamycin-dependent translation initiation in breast cancer cells. Cancer Res. 2007;67(22):10804–12.
Huang HY, et al. Inhibition of human breast cancer cell invasion by siRNA against urokinase-type plasminogen activator. Cancer Invest. 2010;28(7):689–97.
Choudhary C, et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science. 2009;325(5942):834–40.
Bolden JE, Peart MJ, Johnstone RW. Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov. 2006;5(9):769–84.
Weichert W, et al. Expression of class I histone deacetylases indicates poor prognosis in endometrioid subtypes of ovarian and endometrial carcinomas. Neoplasia. 2008;10(9):1021–7.
Weichert W, et al. Association of patterns of class I histone deacetylase expression with patient prognosis in gastric cancer: a retrospective analysis. Lancet Oncol. 2008;9(2):139–48.
Weichert W, et al. Histone deacetylases 1, 2 and 3 are highly expressed in prostate cancer and HDAC2 expression is associated with shorter PSA relapse time after radical prostatectomy. Br J Cancer. 2008;98(3):604–10.
Weichert W, et al. Class I histone deacetylase expression has independent prognostic impact in human colorectal cancer: specific role of class I histone deacetylases in vitro and in vivo. Clin Cancer Res. 2008;14(6):1669–77.
Cebrian A, et al. Genetic variants in epigenetic genes and breast cancer risk. Carcinogenesis. 2006;27(8):1661–9.
Cress WD, Seto E. Histone deacetylases, transcriptional control, and cancer. J Cell Physiol. 2000;184(1):1–16.
Ozdag H, et al. Differential expression of selected histone modifier genes in human solid cancers. BMC Genomics. 2006;7:90.
Qian DZ, et al. Class II histone deacetylases are associated with VHL-independent regulation of hypoxia-inducible factor 1 alpha. Cancer Res. 2006;66(17):8814–21.
Kim SH, et al. Regulation of the HIF-1alpha stability by histone deacetylases. Oncol Rep. 2007;17(3):647–51.
Peart MJ, et al. Novel mechanisms of apoptosis induced by histone deacetylase inhibitors. Cancer Res. 2003;63(15):4460–71.
Rodriguez-Gonzalez A, et al. Role of the aggresome pathway in cancer: targeting histone deacetylase 6-dependent protein degradation. Cancer Res. 2008;68(8):2557–60.
Vousden KH, Lane DP. p53 in health and disease. Nat Rev Mol Cell Biol. 2007;8(4):275–83.
Vervoorts J, Luscher-Firzlaff J, Luscher B. The ins and outs of MYC regulation by posttranslational mechanisms. J Biol Chem. 2006;281(46):34725–9.
Spange S, et al. Acetylation of non-histone proteins modulates cellular signalling at multiple levels. Int J Biochem Cell Biol. 2009;41(1):185–98.
Crabb SJ, et al. Characterisation of the in vitro activity of the depsipeptide histone deacetylase inhibitor spiruchostatin A. Biochem Pharmacol. 2008;76(4):463–75.
Moore PS, et al. Gene expression profiling after treatment with the histone deacetylase inhibitor trichostatin A reveals altered expression of both pro- and anti-apoptotic genes in pancreatic adenocarcinoma cells. Biochim Biophys Acta. 2004;1693(3):167–76.
Glaser KB, et al. Differential protein acetylation induced by novel histone deacetylase inhibitors. Biochem Biophys Res Commun. 2004;325(3):683–90.
Frew AJ, et al. Combination therapy of established cancer using a histone deacetylase inhibitor and a TRAIL receptor agonist. Proc Natl Acad Sci USA. 2008;105(32):11317–22.
Lindemann RK, Gabrielli B, Johnstone RW. Histone-deacetylase inhibitors for the treatment of cancer. Cell Cycle. 2004;3(6):779–88.
Minucci S, Pelicci PG. Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nat Rev Cancer. 2006;6(1):38–51.
Munster PN, et al. A phase II study of the histone deacetylase inhibitor vorinostat combined with tamoxifen for the treatment of patients with hormone therapy-resistant breast cancer. Br J Cancer. 2011;104(12):1828–35.
Schneider BJ, et al. Phase I study of vorinostat (suberoylanilide hydroxamic acid, NSC 701852) in combination with docetaxel in patients with advanced and relapsed solid malignancies. Invest New Drugs. 2012;30:249–57.
Grant C, et al. Romidepsin: a new therapy for cutaneous T-cell lymphoma and a potential therapy for solid tumors. Expert Rev Anticancer Ther. 2010;10(7):997–1008.
Chumsri S, et al. Aromatase inhibitors and xenograft studies. Steroids. 2011;76(8):730–5.
Chen S, et al. The HDAC inhibitor LBH589 (panobinostat) is an inhibitory modulator of aromatase gene expression. Proc Natl Acad Sci USA. 2010;107(24):11032–7.
Wang Y, et al. MicroRNA: past and present. Front Biosci. 2007;12:2316–29.
Iorio MV, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 2005;65(16):7065–70.
Wang ZX, et al. MicroRNA-21 modulates chemosensitivity of breast cancer cells to doxorubicin by targeting PTEN. Arch Med Res. 2011;42(4):281–90.
Mertens-Talcott SU, et al. The oncogenic microRNA-27a targets genes that regulate specificity protein transcription factors and the G2-M checkpoint in MDA-MB-231 breast cancer cells. Cancer Res. 2007;67(22):11001–11.
Liang Z, et al. Blockade of invasion and metastasis of breast cancer cells via targeting CXCR4 with an artificial microRNA. Biochem Biophys Res Commun. 2007;363(3):542–6.
Shay JW. Telomerase in human development and cancer. J Cell Physiol. 1997;173(2):266–70.
Shay JW, Bacchetti S. A survey of telomerase activity in human cancer. Eur J Cancer. 1997;33(5):787–91.
Harley CB. Telomerase and cancer therapeutics. Nat Rev Cancer. 2008;8(3):167–79.
Joseph I, et al. The telomerase inhibitor imetelstat depletes cancer stem cells in breast and pancreatic cancer cell lines. Cancer Res. 2010;70(22):9494–504.
Imetelstat in combination with paclitaxel (with or without bevacizumab) in patients with locally recurrent or metastatic breast cancer. Clinical Trials.gov NCT01256762.
Gagne JP, et al. The expanding role of poly(ADP-ribose) metabolism: current challenges and new perspectives. Curr Opin Cell Biol. 2006;18(2):145–51.
Ame JC, Spenlehauer C, de Murcia G. The PARP superfamily. Bioessays. 2004;26(8):882–93.
Hartman AR, Ford JM. BRCA1 and p53: compensatory roles in DNA repair. J Mol Med (Berl). 2003;81(11):700–7.
Hartman AR, Ford JM. BRCA1 Induces DNA damage recognition factors and enhances nucleotide excision repair. Nat Genet. 2002;32(1):180–4.
Ford JM. Regulation of DNA damage recognition and nucleotide excision repair: another role for p53. Mutat Res. 2005;577(1–2):195–202.
Boulton S, Kyle S, Durkacz BW. Interactive effects of inhibitors of poly(ADP-ribose) polymerase and DNA-dependent protein kinase on cellular responses to DNA damage. Carcinogenesis. 1999;20(2):199–203.
McCabe N, et al. Deficiency in the repair of DNA damage by homologous recombination and sensitivity to poly(ADP-ribose) polymerase inhibition. Cancer Res. 2006;66(16):8109–15.
Gudmundsdottir K, Ashworth A. The roles of BRCA1 and BRCA2 and associated proteins in the maintenance of genomic stability. Oncogene. 2006;25(43):5864–74.
Alli E, et al. Defective repair of oxidative DNA damage in triple-negative breast cancer confers sensitivity to inhibition of poly(ADP-ribose) polymerase. Cancer Res. 2009;69(8):3589–96.
Fong PC, et al. Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N Engl J Med. 2009;361(2):123–34.
Fong PC, et al. Poly(ADP)-ribose polymerase inhibition: frequent durable responses in BRCA carrier ovarian cancer correlating with platinum-free interval. J Clin Oncol. 2010;28(15):2512–9.
Aguilar H, et al. Biological reprogramming in acquired resistance to endocrine therapy of breast cancer. Oncogene. 2010;29(45):6071–83.
Holst F, et al. Estrogen receptor alpha (ESR1) gene amplification is frequent in breast cancer. Nat Genet. 2007;39(5):655–60.
Kirkegaard T, et al. AKT activation predicts outcome in breast cancer patients treated with tamoxifen. J Pathol. 2005;207(2):139–46.
Turner N, et al. FGFR1 Amplification drives endocrine therapy resistance and is a therapeutic target in breast cancer. Cancer Res. 2010;70(5):2085–94.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Lim, B., Cream, L.V., Harvey, H.A. (2013). Update on Clinical Trials: Genetic Targets in Breast Cancer. In: El-Deiry, W. (eds) Impact of Genetic Targets on Cancer Therapy. Advances in Experimental Medicine and Biology, vol 779. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-6176-0_2
Download citation
DOI: https://doi.org/10.1007/978-1-4614-6176-0_2
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-6175-3
Online ISBN: 978-1-4614-6176-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)