Abstract
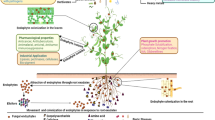
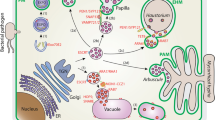
The plant pathogen, Bipolaris sorokiniana (teleomorph: Cochliobolus sativus), is of global concern as it attacks many economically important cereals and grasses. During the infection process, phytopathogenic fungi are known to secrete a variety of proteins collectively known as the secretome, analyzing which can help in deciphering the mechanism of fungal pathogenesis. In this study, we performed in silico secretome analysis of C. sativus strain ND90Pr using established secretome prediction pipeline involving software tools such as SignalP, TargetP, TMHMM, big-PI Fungal Predictor, ProtComp, and WoLF PSORT. Using these software and other prediction criteria, we identified 196 probable secretory proteins from the B. sorokiniana proteome. Characterization of the predicted secretome revealed proteins that may have probable functions in degradation of the plant cell wall, lipids, proteins, and nucleic acids, as well as in pathogenesis and metabolism. Further, the PHI-base analysis identified 38 proteins having a possible role in pathogenicity and virulence. This study helped to predict the composition of the secretome of B. sorokiniana and extrapolate its role in plant infection and pathogen survival. It may provide clues for developing new control strategies targeting the vital fungal secretory proteins.





Similar content being viewed by others
Data availability
The proteome of C. sativus strain ND90Pr (anamorph: B. sorokiniana) v 1.0 used in the study was downloaded from the JGI Mycocosm database (http://genome.jgi.doe.gov/pages/dynamicOrganismDownload.jsf?organism=Cocsa1). The analyzed data and the datasets supporting the conclusions of this article are included within the article and the associated Online Resources.
Abbreviations
- Aa:
-
Amino acid
- Avr:
-
Avirulence
- CHRD:
-
Chordin
- CWDEs:
-
Cell wall degrading enzymes
- ECM:
-
Extracellular matrix
- ETS:
-
Effector-triggered susceptibility
- FAD:
-
Flavin adenine dinucleotide
- GH:
-
Glycosyl hydrolase
- GMC:
-
Glucose methanol choline
- GPI:
-
Glycosylphosphatidylinositol
- LysM:
-
Lysine motif
- PHI:
-
Pathogen Host Interaction database
- pI:
-
Isoelectric point
- SCR:
-
Small and cysteine-rich
- START:
-
StAR related lipid transfer
- TM:
-
Transmembrane domain
- TMHMM:
-
Transmembrane Helices Hidden Markov Model
References
Acharya K, Dutta AK, Pradhan P (2011) Bipolaris sorokiniana (Sacc.) Shoem.: The most destructive wheat fungal pathogen in the warmer areas. Aust J Crop Sci 5:1064. https://doi.org/10.13080/z-a.2013.100.025
Ademark P, de Vries RP, Hagglund P, Stalbrand H, Visser J (2001) Cloning and characterization of Aspergillus niger genes encoding an alpha-galactosidase and a beta-mannosidase involved in galactomannan degradation. Eur J Biochem 268:2982–2990. https://doi.org/10.1046/j.1432-1327.2001.02188.x
Apoga D, Jansson H-B (2000) Visualization and characterization of the extracellular matrix of Bipolaris sorokiniana. Mycol Res 104:564–575. https://doi.org/10.1017/S0953756299001641
Apoga D, Jansson H-B, Tunlid A (2001) Adhesion of conidia and germlings of the plant pathogenic fungus Bipolaris sorokiniana to solid surfaces. Mycol Res 105:1251–1260. https://doi.org/10.1016/s0953-7562(08)61997-8
Aragunde H, Biarnes X, Planas A (2018) Substrate recognition and specificity of chitin deacetylases and related family 4 carbohydrate esterases. Int J Mol Sci 19:412. https://doi.org/10.3390/ijms19020412
Bell AA, Wheeler MH (1986) Biosynthesis and functions of fungal melanins. Ann Rev Phytopathol 24:411–451. https://doi.org/10.1146/annurev.py.24.090186.002211
Breddam K (1986) Serine carboxypeptidases. A review. Carlsberg Res Commun 51:83. https://doi.org/10.1007/bf02907561
Brown NA, Antoniw J, Hammond-Kosack KE (2012) The predicted secretome of the plant pathogenic fungus Fusarium graminearum: A refined comparative analysis. PLoS One 7:e33731. https://doi.org/10.1371/journal.pone.0033731
Carresi L, Pantera B, Zoppi C et al (2006) Cerato-platanin, a phytotoxic protein from Ceratocystis fimbriata: expression in Pichia pastoris, purification and characterization. Protein Expr Purif 49:159–167. https://doi.org/10.1016/j.pep.2006.07.006
Chand R, Kumar M, Kushwaha C, Shah K, Joshi AK (2014) Role of melanin in release of extracellular enzymes and selection of aggressive isolates of Bipolaris sorokiniana in barley. Curr Microbiol 69:202–211. https://doi.org/10.1007/s00284-014-0559-y
Choi J, Park J, Kim D et al (2010) Fungal secretome database: integrated platform for annotation of fungal secretomes. BMC Genom 11:105. https://doi.org/10.1186/1471-2164-11-105
Commenil P, Belingheri L, Bauw G, Dehorter B (1999) Molecular characterization of a lipase induced in Botrytis cinerea by components of grape berry cuticle. Physiol Mol Plant Pathol 55:37–43. https://doi.org/10.1006/pmpp.1999.0206
Condon BJ, Leng Y, Wu D et al (2013) Comparative genome structure, secondary metabolite, and effector coding capacity across Cochliobolus pathogens. PLoS Genet 9:e1003233. https://doi.org/10.1371/journal.pgen.1003233
Conesa A, Gotz S, Garcia-Gomez JM et al (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676. https://doi.org/10.1093/bioinformatics/bti610
Cortazar AR, Aransay AM, Alfaro M, Oguiza JA, Lavin JL (2014) SECRETOOL: integrated secretome analysis tool for fungi. Amino Acids 46:471–473. https://doi.org/10.1007/s00726-013-1649-z
Cosgrove DJ (2005) Growth of the plant cell wall. Nat Rev Mol Cell Biol 6:850. https://doi.org/10.1038/nrm1746
de Vries RP, Visser J (2001) Aspergillus enzymes involved in degradation of plant cell wall polysaccharides. Microbiol Mol Biol Rev 65:497–522. https://doi.org/10.1128/mmbr.65.4.497-522.2001
do Amaral AM, Antoniw J, Rudd JJ, Hammond-Kosack KE (2012) Defining the predicted protein secretome of the fungal wheat leaf pathogen Mycosphaerella graminicola. PLoS One 7:e49904. https://doi.org/10.1371/journal.pone.0049904
Dodds PN, Lawrence GJ, Catanzariti A-M, Ayliffe MA, Ellis JG (2004) The Melampsora lini AvrL567 avirulence genes are expressed in haustoria and their products are recognized inside plant cells. Plant Cell 16:755–768. https://doi.org/10.1105/tpc.020040
Eisenhaber B, Schneider G, Wildpaner M, Eisenhaber F (2004) A sensitive predictor for potential GPI lipid modification sites in fungal protein sequences and its application to genome-wide studies for Aspergillus nidulans, Candida albicans, Neurospora crassa, Saccharomyces cerevisiae and Schizosaccharomyces pombe. J Mol Biol 337:243–253. https://doi.org/10.1016/j.jmb.2004.01.025
El-Gebali S, Mistry J, Bateman A et al (2019) The Pfam protein families database in 2019. Nucleic Acids Res 47:D427–D432. https://doi.org/10.1093/nar/28.1.263
Emanuelsson O, Nielsen H, Brunak S, Von Heijne G (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300:1005–1016. https://doi.org/10.1006/jmbi.2000.3903
Emanuelsson O, Brunak S, Von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2:953. https://doi.org/10.1038/nprot.2007.131
Gao JX, Gao SG, Li YQ, Chen J (2015) Genome-wide prediction and analysis of the classical secreted proteins of Curvularia lunata. J Plant Prot Res 42:869–876. https://doi.org/10.13802/j.cnki.zwbhxb.2015.06.002
Gao M, Yin X, Yang W et al (2017) GDSL lipases modulate immunity through lipid homeostasis in rice. PLoS Pathog 13:e1006724. https://doi.org/10.1371/journal.ppat.1006724
Gasteiger E, Hoogland C, Gattiker A et al (2005) Protein identification and analysis tools on the ExPASy server. In: The Proteomics Protocols Handbook. Springer, Berlin, pp 571–607. https://doi.org/10.1385/1-59259-890-0:571
Godfrey D, Bohlenius H, Pedersen C et al (2010) Powdery mildew fungal effector candidates share N-terminal Y/F/WxC-motif. BMC Genom 11:317. https://doi.org/10.1186/1471-2164-11-317
Grant CE, Bailey TL, Noble WS (2011) FIMO: scanning for occurrences of a given motif. Bioinformatics 27:1017–1018. https://doi.org/10.1093/bioinformatics/btr064
Gupta PK, Vasistha NK, Aggarwal R, Joshi AK (2018) Biology of B. sorokiniana (syn. Cochliobolus sativus) in genomics era. J Plant Biochem Biotechnol 27:123–138. https://doi.org/10.1007/s13562-017-0426-6
Harris PV, Welner D, McFarland KC et al (2010) Stimulation of lignocellulosic biomass hydrolysis by proteins of glycoside hydrolase family 61: structure and function of a large, enigmatic family. Biochemistry 49:3305–3316. https://doi.org/10.1021/bi100009p
Heard S, Brown NA, Hammond-Kosack K (2015) An interspecies comparative analysis of the predicted secretomes of the necrotrophic plant pathogens Sclerotinia sclerotiorum and Botrytis cinerea. PLoS One 10:e0130534. https://doi.org/10.1371/journal.pone.0130534
Henrissat B, Bairoch A (1993) New families in the classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 293:781–788. https://doi.org/10.1042/bj2930781
Horton P, Park K-J, Obayashi T et al (2007) WoLF PSORT: protein localization predictor. Nucleic acids research 35:W585-W587. https://doi.org/10.1093/nar/gkm259
Horton P, Park K-J, Obayashi T, Nakai K (2006) Protein subcellular localization prediction with WoLF PSORT. In: Proceedings of the 4th Asia-Pacific Bioinformatics Conference. World Scientific, pp 39–48. https://doi.org/10.1142/9781860947292_0007
Jedrzejas MJ (2000) Structure, function, and evolution of phosphoglycerate mutases: comparison with fructose-2, 6-bisphosphatase, acid phosphatase, and alkaline phosphatase. Prog Biophys Mol Biol 73:263. https://doi.org/10.1016/s0079-6107(00)00007-9
Jeong JS, Mitchell TK, Dean RA (2007) The Magnaporthe grisea snodprot1 homolog, MSP1, is required for virulence. FEMS Microbiol Lett 273:157–165. https://doi.org/10.1105/tpc.107.051219
Jiang RHY, Tripathy S, Govers F, Tyler BM (2008) RXLR effector reservoir in two Phytophthora species is dominated by a single rapidly evolving superfamily with more than 700 members. Proc Natl Acad Sci 105:4874–4879. https://doi.org/10.1073/pnas.0709303105
Joly DL, Feau N, Tanguay P, Hamelin RC (2010) Comparative analysis of secreted protein evolution using expressed sequence tags from four poplar leaf rusts (Melampsora spp.). BMC Genom 11:422. https://doi.org/10.1186/1471-2164-11-422
Kleemann J, Rincon-Rivera LJ, Takahara H et al (2012) Sequential delivery of host-induced virulence effectors by appressoria and intracellular hyphae of the phytopathogen Colletotrichum higginsianum. PLoS Pathog 8:e1002643. https://doi.org/10.1371/journal.ppat.1002643
Krogh A, Larsson B, Von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a Hidden Markov Model: application to complete genomes. J Mol Biol 305:567–580. https://doi.org/10.1006/jmbi.2000.4315
Kumar J, Schafer P, Huckelhoven R et al (2002) Bipolaris sorokiniana, a cereal pathogen of global concern: cytological and molecular approaches towards better control. Mol Plant Pathol 3:185–195. https://doi.org/10.1046/j.1364-3703.2002.00120.x
Lannou C (2012) Variation and selection of quantitative traits in plant pathogens. Ann Rev Phytopathol 50. https://doi.org/10.1146/annurev-phyto-081211-173031
Levin DE (2005) Cell wall integrity signaling in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 69:262–291. https://doi.org/10.1128/mmbr.69.2.262-291.2005
Lewis TE, Sillitoe I, Dawson N et al (2018) Gene3D: Extensive prediction of globular domains in proteins. Nucleic Acids Res 46:D435–D439. https://doi.org/10.1093/nar/gkx1187
Li W, Jaroszewski L, Godzik A (2001) Clustering of highly homologous sequences to reduce the size of large protein databases. Bioinformatics 17:282–283. https://doi.org/10.1093/bioinformatics/17.3.282
Lum G, Min XJ (2011) FunSecKB: the fungal secretome knowledgebase. Database 2011. https://doi.org/10.1093/database/bar001
Marshall R, Kombrink A, Motteram J et al (2011) Analysis of two in planta expressed LysM effector homologues from the fungus Mycosphaerella graminicola reveals novel functional properties and varying contributions to virulence on wheat. Plant Physiol 156:756–769. https://doi.org/10.1104/pp.111.176347
Martinez D, Berka RM, Henrissat B et al (2008) Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nat Biotechnol 26:553. https://doi.org/10.1038/nbt1403
Mayer AM, Harel E (1979) Polyphenol oxidases in plants. Phytochemistry 18:193–215. https://doi.org/10.1016/0031-9422(79)80057-6
Meinken J, Asch DK, Neizer-Ashun KA et al (2014) FunSecKB2: a fungal protein subcellular location knowledgebase. Comput Mol Biol 4. https://doi.org/10.5376/cmb.2014.04.0007
Mitchell AL, Attwood TK, Babbitt PC et al (2019) InterPro in 2019: improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res 47:D351–D360. https://doi.org/10.1093/nar/gky1100
Motteram J, Kufner I, Deller S et al (2009) Molecular characterization and functional analysis of MgNLP, the sole NPP1 domain-containing protein, from the fungal wheat leaf pathogen Mycosphaerella graminicola. Mol Plant Microbe Interact 22:790–799. https://doi.org/10.1094/mpmi-22-7-0790
Mueller O, Kahmann R, Aguilar G et al (2008) The secretome of the maize pathogen Ustilago maydis. Fungal Genet Biol 45:S63–S70. https://doi.org/10.1016/j.fgb.2008.03.012
Ohm RA, Feau N, Henrissat B et al (2012) Diverse lifestyles and strategies of plant pathogenesis encoded in the genomes of eighteen Dothideomycetes fungi. PLoS Pathog 8:e1003037. https://doi.org/10.1371/journal.ppat.1003037
Paper JM, Scott-Craig JS, Adhikari ND, Cuomo CA, Walton JD (2007) Comparative proteomics of extracellular proteins in vitro and in planta from the pathogenic fungus Fusarium graminearum. Proteomics 7:3171–3183. https://doi.org/10.1002/pmic.200700184
Persson BL, Lagerstedt JO, Pratt JR et al (2003) Regulation of phosphate acquisition in Saccharomyces cerevisiae. Curr Genet 43:225–244. https://doi.org/10.1007/s00294-003-0400-9
Polizeli M, Rizzatti ACS, Monti R et al (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67:577–591. https://doi.org/10.1007/s00253-005-1904-7
Pollet A, Delcour JA, Courtin CM (2010) Structural determinants of the substrate specificities of xylanases from different glycoside hydrolase families. Crit Rev Biotechnol 30:176–191. https://doi.org/10.3109/07388551003645599
Rao MB, Tanksale AM, Ghatge MS, Deshpande VV (1998) Molecular and biotechnological aspects of microbial proteases. Microbiol Mol Biol Rev 62:597–635. https://doi.org/10.1128/mmbr.62.3.597-635.1998
Rodrigues ML, Nosanchuk JD, Schrank A et al (2011) Vesicular transport systems in fungi. Future Microbiol 6:1371–1381. https://doi.org/10.2217/fmb.11.112
Schilling B, Lerch K (1995) Amine oxidases from Aspergillus niger: identification of a novel flavin-dependent enzyme. Biochimica et Biophysica Acta (BBA)-General Subjects 1243:529–537. https://doi.org/10.1016/0304-4165(94)00183-x
Schneiter R, Di Pietro A (2013) The CAP protein superfamily: function in sterol export and fungal virulence. Biomolecular Concepts 4:519–525. https://doi.org/10.1515/bmc-2013-0021
Setlow P (1975) Identification and localization of the major proteins degraded during germination of Bacillus megaterium spores. J Biol Chem 250:8159–8167
Sperschneider J, Dodds PN, Gardiner DM et al (2015) Advances and challenges in computational prediction of effectors from plant pathogenic fungi. PLoS Pathog 11:e1004806. https://doi.org/10.1371/journal.ppat.1004806
Steinberg G (2015) Cell biology of Zymoseptoria tritici: Pathogen cell organization and wheat infection. Fungal Genet Biol 79:17–23. https://doi.org/10.1016/j.fgb.2015.04.002
Stergiopoulos I, van den Burg HA, Okmen B et al (2010) Tomato Cf resistance proteins mediate recognition of cognate homologous effectors from fungi pathogenic on dicots and monocots. Proc Natl Acad Sci 107:7610–7615. https://doi.org/10.1073/pnas.1002910107
Stergiopoulos I, Kourmpetis YAI, Slot JC et al (2012) In silico characterization and molecular evolutionary analysis of a novel superfamily of fungal effector proteins. Mol Biol Evol 29:3371–3384. https://doi.org/10.1093/molbev/mss143
Sutzl L, Foley G, Gillam EMJ, Boden M, Haltrich D (2019) The GMC superfamily of oxidoreductases revisited: analysis and evolution of fungal GMC oxidoreductases. Biotechnol Biofuels 12:118. https://doi.org/10.1186/s13068-019-1457-0
Tjalsma H, Bolhuis A, Jongbloed JDH, Bron S, van Dijl JM (2000) Signal peptide-dependent protein transport in Bacillus subtilis: a genome-based survey of the secretome. Microbiol Mol Biol Rev 64:515–547. https://doi.org/10.1128/mmbr.64.3.515-547.2000
Tsujishita Y, Hurley JH (2000) Structure and lipid transport mechanism of a StAR-related domain. Nat Struct Mol Biol 7:408. https://doi.org/10.1038/75192
van den Brink J, de Vries RP (2011) Fungal enzyme sets for plant polysaccharide degradation. Appl Microbiol Biotechnol 91:1477–1492. https://doi.org/10.1007/s00253-011-3473-2
van den Burg HA, Harrison SJ, Joosten MHAJ, Vervoort J, de Wit PJGM (2006) Cladosporium fulvum Avr4 protects fungal cell walls against hydrolysis by plant chitinases accumulating during infection. Mol Plant Microbe Interact 19:1420–1430. https://doi.org/10.1094/mpmi-19-1420
Vincent D, Bedon F (2013) Secretomics of plant-fungus associations: more secrets to unravel. J Plant Biochem Physiol 1:e117. https://doi.org/10.4172/2329-9029.1000e117
Vivek-Ananth RP, Mohanraj K, Vandanashree M et al (2018) Comparative systems analysis of the secretome of the opportunistic pathogen Aspergillus fumigatus and other Aspergillus species. Sci Rep 8:1–16. https://doi.org/10.1038/s41598-018-25016-4
Vogel JP, Raab TK, Schiff C, Somerville SC (2002) PMR6, a pectate lyase-like gene required for powdery mildew susceptibility in Arabidopsis. Plant Cell 14:2095–2106. https://doi.org/10.1105/tpc.003509
Vu AL, Dee MM, Gwinn KD, Ownley BH (2011) First report of spot blotch and common root rot caused by Bipolaris sorokiniana on switchgrass in Tennessee. Plant Dis 95:1195–1195. https://doi.org/10.1094/pdis-12-10-0880
Wessels JGH (1996) Hydrophobins: proteins that change the nature of the fungal surface. In: Advances in Microbial Physiology, vol 38. Elsevier, Amsterdam, pp 1–45. https://doi.org/10.1016/s0065-2911(08)60154-x
Winnenburg R, Baldwin TK, Urban M et al (2006) PHI-base: a new database for pathogen host interactions. Nucleic Acids Res 34:D459–D464. https://doi.org/10.1093/nar/gkj047
Wu S-C, Kauffmann S, Darvill AG, Albersheim P (1995) Purification, cloning and characterization of two xylanases from Magnaporthe grisea, the rice blast fungus. Mol Plant Microbe Interact 8:506–514. https://doi.org/10.1094/mpmi-8-0506
Wyss M, Brugger R, Kronenberger A et al (1999) Biochemical characterization of fungal phytases (myo-inositol hexakisphosphate phosphohydrolases): catalytic properties. Appl Environ Microbiol 65:367–373. https://doi.org/10.1128/aem.65.2.367-373.1999
Xue C, Park G, Choi W et al (2002) Two novel fungal virulence genes specifically expressed in appressoria of the rice blast fungus. Plant Cell 14:2107–2119. https://doi.org/10.1105/tpc.003426
Yamano H, Gannon J, Hunt T (1996) The role of proteolysis in cell cycle progression in Schizosaccharomyces pombe. EMBO J 15:5268–5279. https://doi.org/10.1002/j.1460-2075.1996.tb00912.x
Yashiroda H, Tanaka K (2003) But1 and But2 proteins bind to Uba3, a catalytic subunit of E1 for neddylation, in fission yeast. Biochem Biophys Res Commun 311:691–695. https://doi.org/10.1016/j.bbrc.2003.10.058
Zalucki YM, Jennings MP (2017) Signal peptidase I processed secretory signal sequences: selection for and against specific amino acids at the second position of mature protein. Biochem Biophys Res Commun 483:972–977. https://doi.org/10.1016/j.bbrc.2017.01.044
Zeng R, Gao S, Xu L, Liu X, Dai F (2018) Prediction of pathogenesis-related secreted proteins from Stemphylium lycopersici. BMC Microbiol 18:191. https://doi.org/10.1186/s12866-018-1329-y
Zhang S, Xu J-R (2014) Effectors and effector delivery in Magnaporthe oryzae. PLoS Pathog 10:e1003826. https://doi.org/10.1371/journal.ppat.1003826
Zhong R, Ye Z-H (2014) Secondary cell walls: biosynthesis, patterned deposition and transcriptional regulation. Plant Cell Physiol 56:195–214. https://doi.org/10.1093/pcp/pcu140
Zhong S, Ali S, Leng Y, Wang R, Garvin DF (2015) Brachypodium distachyon-Cochliobolus sativus pathosystem is a new model for studying plant-fungal interactions in cereal crops. Phytopathology 105:482–489. https://doi.org/10.1094/phyto-08-14-0214-r
Acknowledgements
The authors thank Ms. Pooja Sethiya (CSIR-NCL, Pune, India) and Ms. Meenakshi Kapase (CSIR-NCL, Pune, India) for initial technical support. GMP acknowledges the Academy of Scientific and Innovative Research (AcSIR), Ghaziabad, Uttar Pradesh, India for the Ph.D. research program, and thanks the Department of Science and Technology (DST), India for the INSPIRE research fellowship.
Funding
NYK acknowledges the Council of Scientific and Industrial Research (CSIR; https://www.csir.res.in/), India (grant number: BSC 0117), and the Department of Biotechnology (DBT; http://www.dbtindia.gov.in/), India (grant number: GAP 319026) for financial support. The funders had no role in study design, data collection, and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: Gauri M. Pathak and Narendra Y. Kadoo; Methodology: Gauri M. Pathak and Gayatri S. Gurjar; Formal analysis and investigation: Gauri M. Pathak; Writing - original draft preparation: Gauri M. Pathak; Writing - review and editing: Gauri M. Pathak, Gayatri S. Gurjar and Narendra Y. Kadoo; Funding acquisition: Narendra Y. Kadoo; Resources: Narendra Y. Kadoo; Supervision: Narendra Y. Kadoo.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
Not applicable.
Consent for publication
Not applicable.
Consent to participate
Not applicable.
Code availability
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(XLSX 949 kb)
Rights and permissions
About this article
Cite this article
Pathak, G.M., Gurjar, G.S. & Kadoo, N.Y. Insights of Bipolaris sorokiniana secretome - an in silico approach. Biologia 75, 2367–2381 (2020). https://doi.org/10.2478/s11756-020-00537-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-020-00537-4