Abstract
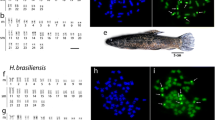
Conventional and molecular cytogenetic analyses were performed in specimens of the Neotropical Crenuchus spilurus freshwater fish species from a single location (Caeté River, Brazil). All specimens presented diploid values of 2n = 38 chromosomes (12 m + 4sm + 2st + 20a), the lowest reported for family Crenuchidae up to now. A single pair of nucleolar organizing regions (NORs) was detected in the subtelocentric chromosome pair no. 9 by silver-staining and fluorescence in situ hybridization (FISH) with 18S rDNA sequence-specific probe. Two pairs of 5S rRNA gene clusters were found either interstitial or terminally located in the long arms of the acrocentric chromosome pairs nos. 10 and 13. Heterochromatic regions were clearly observed in the short arms of the NOR-bearing chromosome pair and weakly-positive to the pericentromeric regions of most acrocentric chromosomes. Additionally, no sex chromosomes were identified in the surveyed specimens. Crenuchidae have signals of several mechanisms involved in karyotype diversification within this family: differential location of heterochromatin-rich regions, multiplication, and translocation of rDNA clusters, presence/absence of sex chromosomes, macrostructural changes in morphology and number of chromosomes. This variety of karyotype patterns reveals the importance of widening cytogenetic studies to more taxa for better know the chromosomal evolution occurred in this group.



Similar content being viewed by others
References
Alves-Costa FA, Wasko AP, Oliveira C, Foresti F, Martins C (2006) Genomic organization and evolution of the 5S ribossomal DNA in Tilapiini fishes. Genetica 127:243–252. https://doi.org/10.1007/s10709-005-4013-8
Amaro-Ghilardi RC, Silva MJJ, Rodrigues MT, Yonenaga-Yassuda Y (2008) Chromosomal studies in four species of genus Chaunus (Bufonidae, Anura): localization of telomeric and ribosomal sequences after fluorescence in situ hybridization (FISH). Genetica 134:159–168. https://doi.org/10.1007/s10709-007-9218-6
Arai R (2011) Fish karyotypes – a check list. Springer, Tokio
Centofante L, Bertollo LAC, Moreira-Filho O (2001) Comparative cytogenetics among sympatric species of Characidium (Pisces, Characiformes). Diversity analysis with the description of ZW sex chromosome system and natural triploidy. Caryologia 54:253–260. https://doi.org/10.1080/00087114.2001.10589233
Centofante L, Bertollo LAC, Buckup PA, Moreira-Filho O (2003) Chromosomal divergence and maintenance of sympatric Characidium fish species (Crenuchidae, Characidiinae). Hereditas 138:213–218. https://doi.org/10.1034/j.1601-5223.2003.01714.x
Cioffi MB, Bertollo LAC (2012) Chromosomal distribution and evolution of repetitive DNAs in fish. In: Garrido-Ramos M (ed) Genome dynamics, 1st edn. Karger, Basel, pp 197–221
Cioffi MB, Molina WFM, Artoni RF, Bertollo LAC (2012) Chromosomes as tools for discovering biodiversity – the case of Erythrinidae fish family. In: Tirunilai P (ed) Recent trends in cytogenetic studies - methodologies and applications, 1st edn. In Tech, Shanghai, pp 125–146
Foresti F, Almeida-Toledo LF, Toledo SA (1981) Polymorphic nature of nucleolus organizer regions in fishes. Cytogenet Cell Genet 31:137–144. https://doi.org/10.1159/000131639
Gold JR, Li CY, Shipley NS, Powers PK (1990) Improved methods for working with fish chromosomes with a review of metaphase chromosome banding. J Fish Biol 37:563–575. https://doi.org/10.1111/j.1095-8649.1990.tb05889.x
Hatanaka T, Galetti PM (2004) Mapping of the 18S and 5S ribosomal RNA genes in the fish Prochilodus argenteus Agassiz, 1829 (Characiformes, Prochilodontidae). Genetica 122:239–244. https://doi.org/10.1007/s10709-004-2039-y
Howell WM, Black DA (1980) Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia 36:1014–1015. https://doi.org/10.1007/BF01953855
Levan A, Fredga K, Sandberg A (1964) Nomenclature for centromeric positions on chromosomes. Hereditas 52:201–220. https://doi.org/10.1111/j.1601-5223.1964.tb01953.x
Lucchini S, Nardi I, Barsacchi G, Batistoni R, Andronico F (1993) Molecular cytogenetics of the ribosomal (18S + 28S and 5S) DNA loci in primitive and advanced urodele amphibians. Genome 36:762–773. https://doi.org/10.1139/g93-101
Machado TC, Pansonato-Alves JC, Pucci MB, Nogaroto V, Almeida MC, Oliveira C, Foresti F, Bertollo LAC, Moreira-Filho O, Artoni RF, Vicari MR (2011) Chromosomal painting and ZW sex chromosomes differentiation in Characidium (Characiformes, Crenuchidae). BMC Genet 12:65. https://doi.org/10.1186/1471-2156-12-65
Maistro EL, Mata EP, Oliveira C, Foresti F (1998) Unusual occurrence of a ZZ/ZW sex-chromosome system and supernumerary chromosomes in Characidium Cf. fasciatum (Pisces, Characiformes, Characidiinae). Genetica 104:1–7. https://doi.org/10.1023/A:1003242020259
Maistro EL, Jesus CM, Oliveira C, Moreira-Filho O, Foresti F (2004) Cytogenetic analysis of A-B chromosomes and ZZ-ZW sex chromosomes of Characidium gomesi (Teleostei, Characiformes, Crenuchidae). Cytologia 69:181–186. https://doi.org/10.1508/cytologia.69.181
Martins C, Galetti PM (2001) Organization of 5S rDNA in species of the fish Leporinus: two different genomic locations are characterized by distinct nontranscribed spacers. Genome 44:903–910. https://doi.org/10.1139/gen-44-5-903
Miyazawa CS, Galetti PM (1994) First cytogenetical studies in Characidium species (Pisces: Characiformes, Characidiinae). Cytologia 59:73–79. https://doi.org/10.1508/cytologia.59.73
Noleto RB, Amorin AP, Vicari MR, Artoni RF, Cestari MM (2009) An unusual ZZ/ZW sex chromosome system in Characidium fishes (Crenuchidae, Characiformes) with the presence of rDNA sites. J Fish Biol 75:448–453. https://doi.org/10.1111/j.1095-8649.2009.02342.x
Oliveira C, Wright JM (1998) Molecular cytogenetic analysis of heterochromatin in the chromosomes of tilapia, Oreochromis niloticus (Teleostei: Cichlidae). Chromosom Res 6:205–211. https://doi.org/10.1023/A:1009211701829
Oliveira C, Foresti F, Hilsdorf AWS (2009) Genetics of neotropical fish: from chromosomes to populations. Fish Physiol Biochem 35:81–100. https://doi.org/10.1007/s10695-008-9250-1
Oliveira C, Avelino GS, Abe KT et al (2011) Phylogenetic relationships within the speciose family Characidae (Teleostei: Ostariophysi: Characiformes) based on multilocus analysis and extensive ingroup sampling. BMC Evol Biol 11:275. https://doi.org/10.1186/1471-2148-11-275
Pansonato-Alves JC, Paiva LRS, Oliveira C, Foresti F (2010) Interspecific chromosomal divergences in the genus Characidium (Teleostei: Characiformes: Crenuchidae). Neotrop Ichthyol 8:77–86. https://doi.org/10.1590/S1679-62252010000100010
Pansonato-Alves JC, Vicari MR, Oliveira C, Foresti F (2011a) Chromosomal diversification in populations of Characidium Cf. gomesi (Teleostei: Crenuchidae). J Fish Biol 78:183–194. https://doi.org/10.1111/j.1095-8649.2010.02847.x
Pansonato-Alves JC, Oliveira C, Foresti F (2011b) Karyotypic conservatism in samples of Characidium Cf. zebra (Teleostei, Characiformes, Crenuchidae): physical mapping of ribosomal genes and natural triploidy. Genet Mol Biol 34:208–213. https://doi.org/10.1590/S1415-47572011005000005
Pazian MF, Shimabukuro-Dias CK, Pansonato-Alves JC, Oliveira C, Foresti F (2013) Chromosome painting of Z and W sex chromosomes in Characidium (Characiformes, Crenuchidae). Genetica 141:1–9. https://doi.org/10.1007/s10709-013-9701-1
Pinhal D, Yoshimura T, Araki C, Martins C (2011) The 5S rDNA family evolves through concerted and birth-and-death evolution in fish genomes: an example from freshwater stingrays. BMC Evol Biol 11:151. https://doi.org/10.1186/1471-2148-11-151
Pinkel D, Straume T, Gray JW (1986) Cytogenetic analysis using quantitative, high sensitivity, fluorescence hybridization. Proc Natl Acad Sci U S A 83:2934–2938. https://doi.org/10.1073/pnas.83.9.2934
Sajdak SL, Reed KM, Phillips RB (1998) Intraindividual and interspecies variation in the 5S rDNA of coregonid fish. J Mol Evol 46:680–688. https://doi.org/10.1007/PL00006348
Silva AR, Maistro EL (2006) Cytogenetic divergence between two sympatric species of Characidium (Teleostei, Characiformes, Crenuchidae) from the Machado River, Minas Gerais, Brazil. Genet Mol Biol 29:459–463. https://doi.org/10.1590/S1415-47572006000300010
Singh M, Kumar M, Nagpure NS, Kushwaha B, Gond I, Lakra WS (2009) Chromosomal localization of 18S and 5S rDNA using FISH in the genus Tor (Pisces, Cyprinidae). Genetica 137:245–252. https://doi.org/10.1007/s10709-009-9367-x
Sumner AT (1972) A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75:304–306. https://doi.org/10.1016/0014-4827(72)90558-7
Suzuki H, Sakurai S, Matsuda Y (1996) Rat rDNA spacer sequences and chromosomal assignment of the genes to the extreme terminal region of chromosome 19. Cytogenet Cell Genet 72:1–4. https://doi.org/10.1159/000134149
Vicari MR, Artoni RF, Moreira-Filho O, Bertollo LAC (2008) Diversification of a ZZ/ZW sex chromosome system in Characidium fish (Crenuchidae, Characiformes). Genetica 134:311–317. https://doi.org/10.1007/s10709-007-9238-2
Weiler KS, Wakimoto BT (1995) Heterochromatin and gene expression in Drosophila. Annu Rev Genet 29:577–605. https://doi.org/10.1146/annurev.ge.29.120195.003045
Acknowledgments
We are grateful to Ricardo Britzke for helping with the fish sampling. We would like to thank Carla Pereira who reviewed and greatly improved the version in English. The financial support for this study was provided by FAPESP (2009/50952-2). We also thank the anonymous reviewers for the suggestions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pazian, M.F., Oliveira, C. & Foresti, F. Cytogenetics characterization of Crenuchus spilurus (Günther, 1863): a remarkable low diploid value within family Crenuchidae (Characiformes). Biologia 73, 77–81 (2018). https://doi.org/10.2478/s11756-018-0006-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-018-0006-9