Abstract
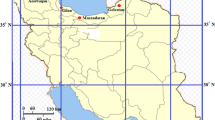
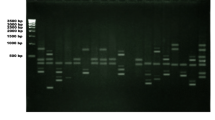
Hordem murinum is a widespread weedy/wild species, growing in different ecological conditions in Iran. Populations of H. murinum subsp. glaucum (2n = 2x = 14) and subsp. leporinum (2n = 4x, 6x = 28, 42) are found in this region. Inter-retroelement amplified polymorphism (IRAP) markers were used to analyse the genetic diversity of 57 accessions of H. murinum from different regions of Iran, and to examine patterns of diversity related to the taxonomy and geography. Eight IRAP primer combinations amplified a total of 241 distinct DNA fragments sized 150-1400 bp, from which 236 (97.9%) were polymorphic. On average, each primer combination amplified about 30.12 fragments (ranged from 23 to 34) in PCR. The patterns of genetic diversity were closely correlated with taxonomic groups, ploidy levels and geographic origin. Along with the high genetic diversity, three geographic sub-genepools were evident, 1: in the North-Northeast region along the Alborz Mountains, 2: in the West-Northwest region along the Zagros Mountains, and 3: in the Central — Southern region. The genetic diversity in diploids was higher than polyploids. Also genetic diversity in W-NW region along the Zagros Mountains was considerably higher than that of the other regions.
Similar content being viewed by others
References
Amirouche R. & Misset M.T. 2003. Hordein polymorphism in diploid and tetraploid Mediterranean populations of the Hordeum murinum L. complex. Plant Syst. Evol. 242: 83–99.
Baum B.R. & Bailey L.G. 1984a. Taxonomic studies in wall barley (Hordeum murinum) and sea barley (Hordeum marinum). I. Character investigation: assessment of new and traditional characters. Can. J. Bot. 62: 753–762.
Baum B.R. & Bailey L.G. 1984b. Taxonomic studies in wall barley (Hordeum murinum) and sea barley (Hordeum marinum). II. Multivariate morphometrics. Can. J. Bot. 62: 2754–2764.
Baum B.R. & Bailey L.G. 1990. Key and synopsis of North American Hordeum species. Can. J. Bot. 68: 2433–2442.
Blattner F.R. 2004. Phylogenetic analysis of Hordeum (Poaceae) as inferred by nuclear rDNA ITS sequences. Mol. Phylogenet. Evol. 33: 289–299.
Blattner F.R. 2009. Progress in phylogenetic analysis and a new infrageneric classification of the barley genus Hordeum (Poaceae: Triticeae). Breeding Sci. 59: 471–480.
Booth T.A. & Richards A.J. 1978. Studies in the Hordeum murinum L. aggregate: disc electrophoresis of seed proteins. Bot. J. Linn. Soc. 76: 115–125.
Bor N.L. 1970. Graminae in: Rechinger K.H. (ed), Flora Iranica, Akademische Druck-u. Verlagsanstalt, Graz. v. 70: 232–244.
Boronnikova S.V. & Kalendar R.N. 2010. Using IRAP markers for analysis of genetic variability in populations of resource and rare species of plants. Russ. J. Genet. 46: 36–42.
Bothmer R. von, Flink J. & Landström T. 1987. Meiosis in interspecific Hordeum hybrids. II. Triploid combinations. Evol. Trends Plants 1: 41–50.
Bothmer R. von, Flink J. & Landström T. 1988. Meiosis in interspecific Hordeum hybrids. 6. Tetraploid (4x - 4x) hybrids. Genome 30: 479–485.
Bothmer R. von, Jacobsen N., Baden C., Jørgensen R.B. & Linde-Laursen I. 1995. An ecogeographical study of the genus Hordeum, 2nd ed, International Plant Genetic Resources Institute, Rom.
Boyko E., Kalendar R., Korzun V., Gill B. & Schulman A.H. 2002. Combined mapping of Aegilops tauschii by retrotransposon, microsatellite, and gene markers. Plant Mol. Biol. 48: 767–790.
El-Rabey H.A., Badr A., Schäfer-Pregl R., Martin W. & Salamini F. 2002. Speciation and species separation in Hordeum L. (Poaceae) resolved by discontinuous molecular markers. Plant Biol. 4: 567–575.
Flavell A.J., Smith D.B. & Kumar A. 1992. Extreme heterogeneity of Ty1-copia group retrotransposons in plants. Mol. Gen. Genet. 231: 233–242.
Gawel N.J. & Jarret R.L. 1991. A modified CTAB extraction procedure for Musa and Ipomoea. Plant Mol. Biol. Rep. 9(3): 262–266.
Giles B.E. & Lefkovitch L.P. 1986. A taxonomic investigation of the Hordeum murinum complex (Poaceae). Plant Syst. Evol. 153: 181–197.
Giles B.E. 1984. A comparison between quantitative and biochemical variation in the wild barley Hordeum murinum. Evolution 38: 34–41.
Graner A., Bjørnstad Å., Konishi T. & Ordon F. 2003. Molecular diversity of the barley genome, in: Bothmer R. Von, Van Hintum T.J.L., Knüpffer H. & Sato K., (eds), Diversity in barley (Hordeum vulgare), Elsevier, Amsterdam, the Netherlands.
Heslop-Harrison J.S., Brandes A., Taketa S., Schmidt T., Vershinin A.V., Alkhimova E.G., Kamm A., Doudrick R.L., Schwarzacher T., Katsiotis A., Kubis S., Kumar A., Pearce S.R., Flavell A.J. & Harrison G.E. 1997. The chromosomal distributions of Ty1-copia group retrotransposable elements in higher plants and their implications for genome evolution. Genetica 100: 197–204.
Jaaska V. 1992. Isoenzyme variation in barley (Hordeum L.). Hereditas 116: 29–35.
Jaccard P. 1908. Nouvelles recherches sur la distribution florale. Bulletin de la Société Vaudoise des Sciences Naturelles 44: 223–270. (cited and implemented in NTSYSpc program, version 2.02e).
Jacobsen N. & Bothmer R. von 1995. Taxonomy in the Hordeum murinum complex (Poaceae). Nord. J. Bot. 15: 449–458.
Jakob S.S. & Blattner F.R. 2010. Two extinct diploid progenitors were involved in allopolyploid formation in the Hordeum murinum (Poaceae: Triticeae) taxon complex. Mol. Phylogenet. Evol. 55: 650–659.
Kakeda K., Taketa S. & Komatsuda T. 2009. Molecular phylogeny of the genus Hordeum using thioredoxin-like gene sequences. Breeding Sci.59: 595–601.
Kalendar R., Grab T., Regina M., Suoniemi A. & Schulman A.H. 1999. IRAP and REMAP: two new retrotransposon-based DNA fingerprinting techniques. Theor. Appl. Genet. 98: 704–711.
Kalendar R., Tanskanen J., Immonen S., Nevo E. & Schulman A.H. 2000. Genome evolution of wild barley (Hordeum spontaneum) by BARE-1 retrotransposon dynamics in response to sharp microclimatic divergence. Proc. Nat. Acad. Sci. USA 97: 6603–6607.
Kubis S., Schmidt T. & Heslop-Harrison J.S. 1998. Repetitive DNA elements as a major component of plant genomes. Ann. Bot. 82(supp. A): 45–55.
Li W., Zhang P., Fellers J.P., Friebe B. & Gill B.S. 2004. Sequence composition, organization, and evolution of the core Triticeae genome. Plant J. 40: 500–511.
Linde-Laursen I., Bothmer R. von & Jacobsen N. 1989. Giemsa C-banded karyotypes of Hordeum marinum and H. murinum. Genome 32: 629–639.
Manninen O., Kalendar R., Robinson J. & Schulman A.H. 2000. Application of BAR/M retrotransposon markers to the mapping of a major resistance gene for net blotch in barley. Mol. Gen. Genet. 264: 325–334.
Mhiri C., Morel J.-B., Vernhettes S., Casacuberta J.M., Lucas H. & Grandbastien M.A. 1997. The promoter of the tobacco Tnt1 retrotransposon is induced by wounding and by abiotic stress. Plant Mol. Biol. 33: 257–266.
Morrell P.L. & Clegg M.T. 2007. Genetic evidence for a second domestication of barley (Hordeum vulgare) east of the Fertile Crescent. Proc. Nat. Acad. Sci. USA 104: 3289–3294.
Nei M. & Li W.H. 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Nat. Acad. Sci. USA 76: 5269–5273. (cited and implemented in NTSYSpc program, version 2.02e).
Nevo E., Beiles A., Kaplan D., Storch N. & Zohary D. 1986. Genetic diversity and environmental association of wild barley, Hordeum spontaneum (Poaceae), in Iran. Plant Syst. Evol. 153: 141–146.
Ourari M., Ainouche A., Coriton O., Huteau V., Brown S., Misset M-T., Ainouche M. & Amirouche R. 2011. Diversity and evolution of the Hordeum murinum polyploid complex in Algeria. Genome 54: 639–654
Parsa A. 1950. Flore de l’Iran. Publication du Ministere de l’Education Museum d’Histoire Naturelle de Tehran, Tehran.
Peakall R. & Smouse P.E. 2006. GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol. Notes 6: 288–295.
Rajhathy T. & Morrison J.W. 1962. Cytogenetic studies in the genus Hordeum L. VI. The murinum complex. Can. J. Genet. Cytol. 4: 240–247.
Saeidi H., Rahiminejad M.R. & Heslop-Harrison J.S. 2008. Retroelement insertional polymorphisms, diversity and phylogeography within diploid, D-genome Aegilops tauschii (Triticeae, Poaceae) sub-taxa in Iran. Ann. Bot. 101: 855–861.
Saeidi H., Rahiminejad M.R., Vallian S. & Heslop-Harrison J.S. 2006. Biodiversity of diploid D-genome Aegilops tauschii Coss. in Iran measured using microsatellites. Genet. Resour. Crop Evol. 53: 1477–1484.
Takeda S., Sugimoto K., Otsuki H. & Hirochika H. 1998. Transcriptional activation of the tobacco retrotransposon Tto1 by wounding and methyl jasmonate. Plant Mol. Biol.36: 365–376.
Taketa S., Takeda K. & Bothmer R. von 1999. Detection of Hordeum marinum genome in three polyploid Hordeum species and cytotypes by genomic in situ hybridization. Hereditas 130: 185–188.
Taketa S., Takeda K. & Bothmer R. von 2001. Physical locations of 5s and 18s-25s rDNA in Asian and American diploid Hordeum species with the I genome. Heredity 86: 522–530.
Tanno K., Bothmer R., Yamane K., Takeda K. & Komastuda T. 2010. Analysis of DNA sequence polymorphism at the cMWG699 locus reveals phylogenetic relationships and allopolyploidy within Hordeum murinum subspecies. Hereditas 147: 34–42.
Teo C.H., Tan S.H., Ho C.L., Faridah Q.Z., Othman Y.R., Heslop-Harrison J.S., Kalendar R. & Schulman A.H. 2005. Genome constitution and classification using retrotransposon-based markers in the orphan crop banana. J. Plant Biol. 48: 96–105.
Vanderwiel P.L., Voytas D.F. & Wendel F.J. 1993. Copia-like retrotransposable element evolution in diploid and polyploid cotton (Gossypium L.). J. Mol. Evol. 36: 429–447.
Voytas D.F., Cummings M.P., Konieczny A., Ausubel F.M. & Rodermel S.R. 1992. Copia-like retrotransposons are ubiquitous among plants. Proc. Nat. Acad. Sci. USA 89: 7124–7128.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sharifi-Rigi, P., Saeidi, H. & Rahiminejad, M.R. Genetic diversity and geographic distribution of variation of Hordeum murinum as revealed by retroelement insertional polymorphisms in Iran. Biologia 69, 469–477 (2014). https://doi.org/10.2478/s11756-014-0340-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-014-0340-5