Abstract
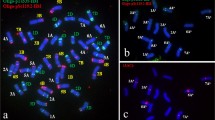
Fluorescence in situ hybridization (FISH) can reveal minor structural differences of chromosomes. The karyotype of common wheat (Triticum aestivum L.) based on FISH pattern is seldom reported. In this study, non-denaturing FISH (ND-FISH) using Oligo-pSc119.2-1, Oligo-pTa535-1 and (AAG)6 as probes was used to investigate the chromosomal structure of 85 common wheat including 83 wheat-rye 1RS.1BL translocation cultivars/lines, a wheatrye 1RS.1AL translocation cultivar Amigo and Chinese Spring (CS). Two, three, two, three, six, three and four structural types respectively for 1A, 2A, 3A, 4A, 5A, 6A and 7A chromosomes were observed. Two, eight, two, two, four and six types of chromosome for 2B, 3B, 4B, 5B, 6B and 7B were respectively detected. The structure of 1B chromosomes in Amigo and CS is different. Five, two, two and two types of chromosomal structure respectively for 1D, 2D, 3D and 5D were distinguished. Polymorphisms of 1RS.1BL, 4D, 6D and 7D chromosomes were not detected. Chromosomes 1AI, 2AI, 3AI, 4AI, 5AIII, 6AI, 7AIII, 2BI, 3BV, 4BI, 5BII, 6BIII, 7BI, 1DIV, 2DI, 3DI and 5DII appeared in these 85 wheat cultivars/lines at high frequency. Each of the 85 wheat cultivars/lines has a unique karyotype. Amigo is a complex translocation cultivar. The FISH karyotype of wheat chromosomes built in this study provide a reference for the future analyzing wheat genetic stocks and help to learn structural variations of wheat chromosomes. In addition, the results in this study indicate that oligonucleotide probes and ND-FISH technology can be used to identify individual wheat cultivar.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Badaeva, E.D., Dedkova, O.S., Gay, G., Pukhalskyi, V.A., Zelenin, A.V., Bernard, S., Bernard, M. 2007. Chromosomal rearrangements in wheat: their types and distribution. Genome 50:907–926.
Badaeva, E.D., Amosova, A.V., Goncharov, N.P., Macas, J., Ruban, A.S., Grechishnikova, I.V., Zoshchuk, S.A., Houben, A. 2015. A set of cytogenetic markers allows the precise identification of all A-genome chromosomes in diploid and polyploidy wheat. Cytogenet. Genome Res. 146:71–79.
Cabo, S., Carvalho, A., Martin, A., Lima-Brito, J. 2014. Structural rearrangements detected in newly-formed hexaploid tritordeum after three sequential FISH experiments with repetitive DNA sequences. J. Genet. 93: 183–188. doi:10.1007/s12041-014-0328-5.
Carmona, A., Friero, E., Bustos, A.D., Jouve, N., Cuadrado, A. 2013. Cytogenetic diversity of SSR motifs within and between Hordeum species carrying the H genome: H. vulgare L. and H. bulbosum L. Theor. Appl. Genet. 126:949–961.
Carvalho, A., Guedes-Pinto, H., Lima-Brito, J. 2013. Polymorphism of the simple sequence repeat (AAC)5 in the nucleolar chromosomes of old Portuguese wheat cultivars. J. Genet. 92:583–586.
Cuadrado, Á., Golczyk, H., Jouve, N. 2009. A novel, simple and rapid nondenaturing FISH (ND-FISH) technique for the detection of plant telomeres. Potential used and possible target structures detected. Chromosome Res. 17:755–762.
Cuadrado, Á., Jouve, N. 2010. Chromosomal detection of simple sequence repeats (SSRs) using nondenaturing FISH (ND-FISH). Chromosoma 119:495–503.
Cuadrado, Á., Carmona, A., Jouve, N. 2013. Chromosomal characterization of the three subgenomes in the polyploids of Hordeum murinum L.: New insight into the evolution of this complex. PLoS ONE 8:e81385.
Delgado, A., Carvalho, A., Martín, C., Martín, A., Lima-Brito, J. 2017. Genomic reshuffling in advanced lines of hexaploid tritordeum. Genet. Resour. Crop Evol. 64:1331–1353.
Du, P., Zhuang, L.F., Wang, Y.Z., Yuan, L., Wang, Q., Wan, D.R., Dawadondup, Tan, L.J., Shen, J., Xu, H.B., Zhao, H., Chu, C.G., Qi, Z.J. 2017. Development of oligonucleotides and multiplex probes for quick and accurate identification of wheat and Thinopyrum bessarabicum chromosomes. Genome 60:93–103.
Friebe, B., Bill, B.S. 1994. C-band polymorphism and structural rearrangements detected in common wheat (Triticum aestivum). Euphytica 78:1–5.
Fu, S.L., Chen, L., Wang, Y.Y., Li, M., Yang, Z.J., Qiu, L., Yan, B.J., Ren, Z.L., Tang, Z.X. 2015. Oligonucleotide probes for ND-FISH analysis to identify rye and wheat chromosomes. Sci. Rep. 5:10552.
Gill, B.S., Friebe, B., Endo, T.R. 1991. Standard karyotype and nomenclature system for description of chromosome bands and structural aberrations in wheat (Triticum aestivum). Genome 34:830–839.
Han, F.P., Lamb, J.C., Birchler, A. 2006. High frequency of centromere inactivation resulting in stable dicentric chromosomes of maize. Pro. Natl. Acad. Sci. U.S.A. 103:3238–3243.
Jiang, J., Friebe, B., Gill, B.S. 1994. Chromosome painting of Amigo wheat. Theor. Appl. Genet. 89:811–813.
Kirov, I.V., Kiseleva, A.V., Laere K.V., Roy, N.V., Khrustaleva, L.I. 2017. Tandem repeats of Allium fistulosum associated with major chromosomal landmarks. Mol. Genet. Genomics 292:453–464.
Komuro, S., Endo, R., Shikata, K., Kato, A. 2013. Genomic and chromosomal distribution patterns of various repeated DNA sequences in wheat revealed by a fluorescence in situ hybridization procedure. Genome 56:131–137.
Landjeva, S., Korzun, V., Börner, A. 2007. Molecular markers: actual and potential contributions to wheat genome characterization and breeding. Euphytica 156:271–296.
Li, G.R., Gao, D., Zhang, H.G., Li, J.B., Wang, H.J., La, S.X., Ma, J.W., Yang, Z.J. 2016. Molecular cytogenetic characterization of Dasypyrum breviaristatum chromosomes in wheat background revealing the genomic divergence between Dasypyrum species. Mol. Cytogenet. 9:6.
Mirzaghaderi, G., Houben, A., Badaeva, E. 2014. Molecular-cytogenetic analysis of Aegilops triuncialis and identification of its chromosomes in the background of wheat. Mol. Cytogenet. 7:91.
Mukai, Y., Nakahara, Y., Yamamoto, M. 1993. Simultaneous discrimination of the three genomes in hexaploid wheat by multicolour fluorescence in situ hybridization using total genomic and highly repeated DNA probes. Genome 36:489–494.
Pavia, I., Carvalho, A., Rocha, L., Gaspar M.J., Lima-Brito, J. 2014. Physical location of SSR regions and cytogenetic instabilities in Pinus sylvestris chromosomes revealed by ND-FISH. J. Genet. 93:567–571.
Pedersen, C., Langridge, P. 1997. Identification of the entire chromosome complement of bread wheat by two-colour FISH. Genome 40:598–593.
Schneider, A., Linc, G., Molnár-Láng, M. 2003. Fluorescence in situ hybridization polymorphism using two repetitive DNA clones in different cultivars of wheat. Plant Breed. 122: 396–400.
Sharma, S.K., Yamamoto, M., Mukai, Y. 2016. Molecular cytogenetic approaches in exploration of important chromosomal landmarks in plants. In: Rajpal, R., Rao, S.R., Raina, S.N. (eds), Sustainable Development and Biodiversity. Springer Press. Geneva, Switzerland, pp. 127–148.
Tang, S.Y., Qiu, L., Xiao, Z.Q., Fu, S.L., Tang Z.X. 2016. New oligonucleotide probes for ND-FISH analysis to identify barley chromosomes and to investigate polymorphisms of wheat chromosomes. Genes 7:118.
Tang, Z.X., Li, M., Chen, L., Wang, Y.Y., Ren, Z.L., Fu, S.L. 2014a. New types of wheat chromosomal structural variations in derivatives of wheat-rye hybrids. PloS ONE 9:e110282.
Tang, Z.X., Yang, Z.J., Fu, S.L. 2014b. Oligonucleotides replacing the roles of repetitive sequences pAs1, pSc119.2, pTa-535, pTa71, CCS1, and pAWRC.1 for FISH analysis. J. Appl. Genetics 55:313–318.
Xiao, Z.Q., Tang, S.Y., Qiu, L., Tang, Z.X., Fu, S.L. 2017. Oligonucleotides and ND-FISH displaying different arrangements of tandem repeats and identification of Dasypyrum villosum chromosomes in wheat backgrounds. Molecules 22:973.
Zhu, M.Q., Du, P., Zhuang, L.F., Chu, C.G., Zhao, H., Qi, Z.J. 2017. A simple and efficient non-denaturing FISH method for maize chromosome differentiation using single-strand oligonucleotide probes. Genome online May 4:1–8.
Acknowledgement
This project was supported by the National Natural Science Foundation of China (No. 31471498).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by M. Molnár-Láng
Electronic Supplementary Material (ESM)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Jiang, M., Xaio, Z.Q., Fu, S.L. et al. FISH Karyotype of 85 Common Wheat Cultivars/Lines Displayed by ND-FISH Using Oligonucleotide Probes. CEREAL RESEARCH COMMUNICATIONS 45, 549–563 (2017). https://doi.org/10.1556/0806.45.2017.049
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1556/0806.45.2017.049