Abstract
Multidrug resistance (MDR1) and breast cancer resistance protein (BCRP) play important roles in drug absorption and distribution. Computational prediction of substrates for both transporters can help reduce time in drug discovery. This study aimed to predict the efflux activity of MDR1 and BCRP using multiple machine learning approaches with molecular descriptors and graph convolutional networks (GCNs). In vitro efflux activity was determined using MDR1- and BCRP-expressing cells. Predictive performance was assessed using an in-house dataset with a chronological split and an external dataset. CatBoost and support vector regression showed the best predictive performance for MDR1 and BCRP efflux activities, respectively, of the 25 descriptor-based machine learning methods based on the coefficient of determination (R2). The single-task GCN showed a slightly lower performance than descriptor-based prediction in the in-house dataset. In both approaches, the percentage of compounds predicted within twofold of the observed values in the external dataset was lower than that in the in-house dataset. Multi-task GCN did not show any improvements, whereas multimodal GCN increased the predictive performance of BCRP efflux activity compared with single-task GCN. Furthermore, the ensemble approach of descriptor-based machine learning and GCN achieved the highest predictive performance with R2 values of 0.706 and 0.587 in MDR1 and BCRP, respectively, in time-split test sets. This result suggests that two different approaches to represent molecular structures complement each other in terms of molecular characteristics. Our study demonstrated that predictive models using advanced machine learning approaches are beneficial for identifying potential substrate liability of both MDR1 and BCRP.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multidrug resistance (MDR1) and breast cancer resistance protein (BCRP) highly contribute to drug absorption and distribution (1). In particular, these efflux transporters are expressed in the blood–brain barrier (BBB) and prevent the brain penetration of drugs (2,3,4,5,6). Therefore, in vitro screening to eliminate MDR1 and BCRP substrates is utilized to develop medicines for central nervous system (CNS) diseases.
Efflux activity prediction can help reduce the cost and time in drug discovery. MDR1 has a large binding pocket and recognizes various structurally diverse compounds (7). Therefore, the accurate prediction of substrates via molecular docking simulations using protein structural information remains challenging. Quantitative structure–activity relationships (QSAR) using machine learning techniques, a type of artificial intelligence (AI), have been used to improve absorption, distribution, metabolism, and excretion (ADME) properties in the drug discovery process (8,9,10,11,12,13,14,15). Several computational classification models have been reported to predict substrates and modulators of MDR1, BCRP, and other ABC transporters including various techniques (16,17,18,19,20,21,22,23,24,25,26). Few studies have presented regression models for predicting the efflux activity of MDR1 and BCRP (12, 27, 28).
Various machine learning algorithms are essential for exploring the best prediction model. The automated machine learning framework comprehensively investigates multiple algorithms and minimizes technical issues. This study used PyCaret, an open-source machine learning library in Python that automates the machine learning workflow with minimum coding, as a machine learning approach using molecular descriptors (29). In a recent study, the model for predicting the fraction of a drug unbound in plasma using PyCaret outperformed those using other automated frameworks (30). In contrast to descriptor-based approaches, graph convolutional networks (GCN) based on molecular graphs and convolutional neural networks (CNN) based on compound images have recently gained attention as a recent trend in machine learning (31,32,33).
The GCN for molecular properties is a powerful approach for implementing multi-task and multimodal learning. Multi-task learning attempts to learn multiple different tasks simultaneously and has been utilized for predicting ADME parameters (13, 33,34,35). Additionally, GCN can combine different types of information, such as the chemical structures of molecules and amino acid sequences of proteins, through multimodal learning. A recent study applied multimodal GCN to the classification of molecular properties (36). However, the effectiveness of multimodal GCN in predicting ADME properties, including transporter activity, remains unclear.
This study developed a predictive model for MDR1 and BCRP activities using descriptor-based machine learning and GCNs. The predictive performances of the GCN models, including multi-task and multimodal learning, were compared with those of multiple descriptor-based machine learning approaches. In addition, an ensemble of descriptor-based machine learning and GCNs was used to enhance predictivity.
Materials and Methods
Materials
The test compounds were prepared by Takeda Pharmaceutical Company (Fujisawa, Japan) to determine efflux activity in MDR1- and BCRP-expressing cells. All other reagents and solvents were of analytical grade or better and were commercially available.
In Vitro Permeability MDR1 and BCRP-Expressing Cells
The efflux ratio (ER) was determined by previously described methods (12). Test compounds solubilized in dimethyl sulfoxide (DMSO) were added to transport buffer (Hanks’ balanced salt solution with 10 mM HEPES, pH 7.4) at a final concentration of 2 μM (DMSO < 1%) on either the apical or basolateral side of the transwell chamber with Madin–Darby canine kidney (MDCK)-MDR1 from NIH and MDCK-BCRP from Solvo Biotechnology (Szeged, Hungary).
The confluent cell monolayers on the transwell were incubated for 1 h at 37°C with 5% CO2. The test compounds were quantified by liquid chromatography–tandem mass spectrometry (LC-MS/MS; Applied Biosystems, Foster City, CA, USA). The ER and permeation of the test compounds from the apical to basolateral (A to B) or B to A direction were determined. The apparent permeability coefficient Papp (cm/s) was calculated using the following equation:
where dCr/dt is the cumulative concentration of the compound in the receiver chamber as a function of time (µM/s); Vr is the volume of the solution in the receiver chamber (0.075 mL on the apical side, 0.25 mL on the basolateral side); A is the surface area for transport, i.e., 0.0804 cm2 for the monolayer area; and C0 is the initial concentration in the donor chamber (µM).
The ER was calculated using the following equation:
Data Preparation
The ER for 9490 and 3440 compounds in MDR1- and BCRP-expressing cells, respectively, were used as the proprietary internal dataset. As data splitting, we conducted a time split. The entire dataset was chronologically divided into a training set (compounds used until May 2022) and a test set (from June 2022 to March 2023). As an external dataset, the ER for 46 and 47 compounds were determined in MDR1- and BCRP-expressing cells, respectively.
Descriptor-Based Approach
Various molecular descriptors were generated using alvaDesc (1.0.16) (Alvascience Srl, Lecco, Italy), which provides physicochemical properties such as lipophilicity, polarity, molar refractivity, and pharmacophore. The 3D descriptors and descriptors with N/A were removed. The generated 2D descriptors were applied to the feature selection.
The Boruta algorithm was applied as the feature selection method, a wrapper around the random forest algorithm to identify essential features for further analysis (37, 38). After the feature selection, PyCaret 2.3.6 was used for data splitting, model selection, and hyperparameter turning (29). Twenty-five machine learning algorithms, including linear regression (lr), lasso regression (lasso), ridge regression (ridge), elastic net (en), least angle regression (lar), lasso least angle regression (llar), orthogonal matching pursuit (omp), Bayesian ridge (br), automatic relevance determination (ard), passive aggressive regressor (par), random sample consensus (ransac), TheilSen regressor (tr), Huber regressor (huber), kernel ridge (kr), support vector machine (svm), K-nearest neighbors (knn), decision tree regressor (dt), random forest regressor (rf), extra trees regressor (et), AdaBoost regressor (ada), gradient boosting regressor (gbr), MLP regressor (mlp), extreme gradient boosting (xgboost), light gradient boosting machine (lightgbm), and CatBoost regressor (catboost), were employed for model building. For model selection, tenfold cross-validation was applied only to the training datasets. Model performance was assessed using the percentage within a twofold error, the coefficient of determination (R2), and root-mean-squar error (RMSE) values. The percentage within a twofold error is the percentage of predicted values within twofold of the observed values and is used to evaluate the prediction acceptability. R2 and RMSE were calculated using the following equations:
Hyperparameter tuning was performed for the model showing the best performance in tenfold cross-validation. The number of iterations in the grid search for hyperparameter tuning was set to 100. Finally, the model performance was evaluated using test and external datasets.
Graph-Based Approach
The training and test sets from the descriptor-based approach were used in the graph-based approach. We used a GCN comprising two graph convolutional layers and two linear transformation layers (39). We used the atom type, degree, hybridization, aromaticity, formal charge, number of implicit Hs on the atom, number of radical electrons of the atom, and total Hs on the atom as atom features. The percentage within a twofold error, R2, and RMSE were used to evaluate model performance. Integrated Gradients was used to interpret which chemical substructures influenced ER prediction (40).
The model was trained on 80% of the training dataset for 200 epochs, with a batch size of 1024. The mean absolute error (MAE) was used as the loss function. Adam was used as an optimization algorithm (41). Hyperparameters of Adam, including learning rate, exponential decay rates, epsilon (a parameter for numerical stability), and weight decay (L2 penalty), were optimized using Optuna to maximize the accuracy of the other 20% of the training dataset (42). The number of optimization trials was set to 100. We used the Captum library to perform Integrated Gradients (43).
Multi-task learning was applied to build a single model to predict MDR1 and BCRP activities for the same compound. The model architecture was the same as that of the single-task GCN, and the output layer size was changed to two. The loss function, optimization algorithm, and method of hyperparameter tuning were the same as in the single-task GCN.
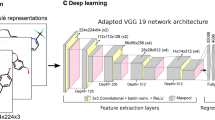
Multimodal learning utilizes the chemical structures of drug molecules and amino acid sequences of the two transporters. Chemical structures were encoded using two graph convolutional layers into 128-dimensional feature vectors. The amino acid sequences of MDR1 and BCRP were also encoded with one one-dimensional convolutional layer and one linear transformation layer into 32-dimensional feature vectors. After concatenating the two types of feature vectors, two linear transformation layers predicted the ER from the feature vectors. The training and hyperparameter tuning methods were the same as those for the single-task GCN.
For graph-based approaches, PyTorch (44), Deep Graph Library (45), and DGL-LifeSci (46) were used. We used the GCN Predictor implementation in DGL-LifeSci with default parameters as the GCN architecture. To illustrate the molecules, we used RDKit, an open-source cheminformatics software (47).
Results
Dataset Analysis
The ER histogram did not follow a normal distribution, and the ER was log-transformed to reduce the skewness of the measurement variable (Fig. 1). ER distribution was comparable between the time-split training and test sets. The distribution pattern of the BCRP ER in the external dataset differed from that in the in-house dataset (Fig. S1).
Distribution of MDR1 and BCRP ER and log(ER) values in the training set (solid column) and test set (open column). Values represent the relative frequency of MDR1 ER a and log(ER) b in MDR1 and ER c and log(ER) d in BCRP. The relative frequency was calculated by dividing a frequency count by the sum of all frequencies
Descriptor-Based Approach
The predictive performance of the 25 machine learning models was compared using tenfold cross-validation for each training set. The model performance was evaluated using the percentage within a twofold error, R2, and RMSE. The CatBoost showed the best performance in the time-split training set (Table I). The ER predictivity based on CatBoost was evaluated using an independent test set from the training set. The R2 and RMSE values were 0.673 and 0.424, respectively, and 59.6% of the predicted ER values were within twofold of the observed values (Table II). Tree-based models enable the calculation of the feature importance, and the top 10 important features of the MDR1 ER prediction model were investigated (Table S1).
Support vector regression produced the best results for BCRP ER prediction in the time-split training set (Table I). The R2 and RMSE values were 0.536 and 0.333, respectively, in an independent time-split test set, and 63.9% of the predicted ER values were within twofold of the observed values (Table II).
Compared with the in-house time-split test set, the predictive performance in the external dataset was poor. In the external MDR1 and BCRP datasets, the percentages within the twofold error were 41.3% and 36.2%, respectively (Fig. S3).
Graph-Based Approach
The predictive performance of the GCN is shown in Table II. The model performance was evaluated using an independent test set from the training set. The R2 values of the MDR1 and BCRP ER predictions for the test set were 0.651 and 0.484, respectively. In the external MDR1 and BCRP dataset, the predictive performances were poorer than that in the in-house time-split test set, and the percentage within twofold error was 47.8% and 55.3%, respectively (Fig. S2). The chemical substructure contributing to the predicted ER was visualized using Integrated Gradients in the external dataset (Fig. S4).
The results of multi-task and multimodal learning are shown in Table II. Multi-task learning did not improve ER prediction accuracy. Meanwhile, based on R2 and RMSE, multimodal learning improved the predictive performance of BCRP ER.
Ensemble Approach
The ensemble model was developed to utilize the advantages of both the descriptor-based and graph-based approaches. The average values of the predicted ER with descriptor-based machine learning and the GCN were used for the final predicted ER. In both tasks, the accuracy was higher than that of the descriptor- and graph-based approaches (Table II). The R2 values of the MDR1 and BCRP ER predictions were 0.706 and 0.587, respectively, in the time-split test set. The predictive performance in the external dataset was lower than that in the in-house time-split test set, and 50.0% and 55.3% of the predicted MDR1 ER and BCRP ER values, respectively, were within twofold of the observed values (Figs. 2 and 3).
Discussion
The physiologically based pharmacokinetic (PBPK) model has been used for in vivo intestinal absorption and brain permeability and integrates quantitative ER values in each process. In particular, we reported that unbound brain-to-plasma partitioning (Kp,uu,brain) could be predicted using the ER in MDR1 and BCRP. An accurate ER can predict Kp,uu,brain by incorporating it into the PBPK model.
The automated machine learning framework using PyCaret minimizes technical hurdles. In the descriptor-based prediction of the MDR1 ER, CatBoost, an open-source gradient boosting library developed by Yandex (48), showed the highest predictivity in the time-split test set. Researchers from various fields have successfully utilized CatBoost for machine learning using big data since 2017 (7). However, there are no reports on the prediction of ADME-Tox parameters using CatBoost. In contrast, in the descriptor-based prediction of the BCRP ER, support vector regression, a popular machine learning algorithm widely used for the classification and regression of ADME properties in several studies (49, 50), showed the highest predictivity in the time-split test set. The comprehensive automated machine learning framework can help to select the best algorithm from traditional and contemporary algorithms.
Previous research has demonstrated that the graph-based approach is superior to descriptor-based prediction for multiple ADME parameters (34). However, the MDR1 and BCRP ER predictive performance of GCN was poorer than that of the descriptor-based approaches in our study. We used a simple GCN architecture comprising two graph convolutional layers and two linear transformation layers. Meanwhile, recent sophisticated models, such as graph attention networks (GATs) (51) and message passing neural networks (MPNNs) (52), can enhance predictive performance. Additionally, many features are available for each atom in a molecule that captures both the electrons’ properties and the bonds in which the atom participates. Modifying atomic features and introducing bond features may improve the performance of graph-based approaches.
Previous research has demonstrated that the multi-task GCN is superior to the single-task GCN for predicting intrinsic clearance and solubility (31). However, herein, the multi-task GCN showed poorer performance than the single-task GCN, presumably due to the imbalance in the dataset size between the two tasks, which produces many missing values, particularly in the smaller dataset (i.e., the BCRP dataset in this study). The possibility that many missing values lower the performance of multi-task learning has been reported in previous research (53). To improve the performance of multi-task learning, datasets with few missing values may be required.
In contrast, multimodal learning may help overcome the challenge of imbalanced ADME datasets. Although the improvement of predictive performance using multimodal learning was not observed in MDR1 ER prediction, multimodal learning improved the predictivity of the BCRP ER (Table II and Fig. S2). This may be because multimodal learning overcomes the limited data problem by combining the information from two modalities. In our case, the BCRP dataset was smaller than that of MDR1, and multimodal learning solved the difficulty of the limited BCRP dataset. Meanwhile, the study’s multimodal learning approach is constrained by the number of protein sequence types. This study only used the protein sequences of two transporters, indicating that the constructed model has only been categorically informed that the transporters are different, and the model’s ability to recognize similarities and disparities between the protein sequences of diverse transporters has not been tested. Thus, further studies using datasets of efflux activities, including more diverse transporters, are required to investigate the utility of multimodal learning.
In both descriptor-based machine learning and molecular graph-based approaches, the predictive performances in the external dataset were lower than those in the in-house time-split set, indicating that the coverage of chemical space differed between the two datasets. In practice, descriptor-based PCA plots demonstrated that some compounds in the external dataset did not fit the distribution observed in the in-house training set (Fig. 4). The applicability domain must be considered to estimate the model’s reliability and coverage.
Furthermore, we applied Integrated Gradients to the GCN models to interpret which compound substructures contributed to the predicted ER. Several studies have reported substructures that affect the MDR1 and BCRP ERs using a fingerprint-based machine learning approach (54,55,56,57). Herein, using GCN, the primary amine moiety was frequently recognized to increase the ER, whereas hydroxyl groups (especially secondary alcohols) and halogens (Cl and F) were occasionally recognized to decrease the ER (Fig. S4). While hydroxyl groups and halogens are common in MDR1 non-substrates (55, 57), the role of primary amines in efflux activity has not been reported in the context of machine learning. QSAR studies have reported that substructures of inhibitors, such as amino groups, fluorine, and chlorine, can be recognized by ABC transporters (58), although the action of the inhibitors on the transporters may not be the same as that of the substrates. Additionally, some contributing substructures differ between the fingerprint-based machine learning approach in previous studies and the graph-based approach in this study; for example, nitrile and thial groups frequently occur in non-substrates (54). These findings suggest that combining multiple approaches, including fingerprint-, descriptor-, and graph-based approaches, would strengthen knowledge about the chemical structure contributing to ER. GCN enables structure-based interpretation, thereby improving the efficiency of structural optimization in the drug discovery process.
In this study, the ensemble model achieved higher predictive performance than the descriptor- and graph-based approaches (Table II). Adding molecular descriptors to a graph convolutional model improves the predictive performance for molecular property prediction (59). The molecular graph-based approach learns the relationship between the chemical structures and efflux activity. In contrast, the descriptor-based approach uses many kinds of molecular descriptors, such as constitutional, topological, and pharmacophore, in addition to chemical structures, including functional groups and fragment counts. Among the top 10 features extracted from the MDR1 ER prediction model, molecular descriptors directly expressing chemical structures were limited, such as the frequency of C–N at topological distance 1 and the number of double bonds (Table S1). The ensemble approach can consider both types of features, leading to high predictive performance.
Conclusions
This study successfully developed prediction models for ER in MDR1- and BCRP-expressing cells using molecular descriptor-based machine learning and graph-based approaches. Multimodal learning outperformed the other GCN approaches in predicting the BCRP ER. Because GCN enables the visualization of atomic contributions to the prediction result, this information would be useful for clarifying the structure spot that increases ER and improving ER in the drug optimization process. Finally, ensemble approaches combined with descriptor-based machine learning and GCN improved the prediction of both MDR1 and BCRP ERs, enabling early decision-making in compound prioritization.
Data Availability
We have provided data that can be disclosed to supplemental material.
References
Yamamoto S, Kosugi Y, Hirabayashi H, Moriwaki T. Impact of P-glycoprotein on intestinal absorption of an inhibitor of apoptosis protein antagonist in rats: mechanisms of nonlinear pharmacokinetics and food effects. Pharm Res. 2018;35(10):190. https://doi.org/10.1007/s11095-018-2470-x.
Breedveld P, Pluim D, Cipriani G, Wielinga P, van Tellingen O, Schinkel AH, et al. The effect of Bcrp1 (Abcg2) on the in vivo pharmacokinetics and brain penetration of imatinib mesylate (Gleevec): implications for the use of breast cancer resistance protein and P-glycoprotein inhibitors to enable the brain penetration of imatinib in patients. Cancer Res. 2005;65(7):2577–82. https://doi.org/10.1158/0008-5472.CAN-04-2416.
Sakata A, Tamai I, Kawazu K, Deguchi Y, Ohnishi T, Saheki A, et al. In vivo evidence for ATP-dependent and P-glycoprotein-mediated transport of cyclosporin A at the blood-brain barrier. Biochem Pharmacol. 1994;48(10):1989–92. https://doi.org/10.1016/0006-2952(94)90601-7.
Schinkel AH, Smit JJ, van Tellingen O, Beijnen JH, Wagenaar E, van Deemter L, et al. Disruption of the mouse mdr1a P-glycoprotein gene leads to a deficiency in the blood-brain barrier and to increased sensitivity to drugs. Cell. 1994;77(4):491–502. https://doi.org/10.1016/0092-8674(94)90212-7.
Kodaira H, Kusuhara H, Ushiki J, Fuse E, Sugiyama Y. Kinetic analysis of the cooperation of P-glycoprotein (P-gp/Abcb1) and breast cancer resistance protein (Bcrp/Abcg2) in limiting the brain and testis penetration of erlotinib, flavopiridol, and mitoxantrone. J Pharmacol Exp Ther. 2010;333(3):788–96. https://doi.org/10.1124/jpet.109.162321.
Enokizono J, Kusuhara H, Ose A, Schinkel AH, Sugiyama Y. Quantitative investigation of the role of breast cancer resistance protein (Bcrp/Abcg2) in limiting brain and testis penetration of xenobiotic compounds. Drug Metab Dispos. 2008;36(6):995–1002. https://doi.org/10.1124/dmd.107.019257.
Hancock JT, Khoshgoftaar TM. CatBoost for big data: an interdisciplinary review. J Big Data. 2020;7(1):94. https://doi.org/10.1186/s40537-020-00369-8.
Kosugi Y, Hosea N. Direct comparison of total clearance prediction: computational machine learning model versus bottom-up approach using in vitro assay. Mol Pharm. 2020;17(7):2299–309. https://doi.org/10.1021/acs.molpharmaceut.9b01294.
Kosugi Y, Hosea N. Prediction of oral pharmacokinetics using a combination of in silico descriptors and in vitro ADME properties. Mol Pharm. 2021;18(3):1071–9. https://doi.org/10.1021/acs.molpharmaceut.0c01009.
Korotcov A, Tkachenko V, Russo DP, Ekins S. Comparison of deep learning with multiple machine learning methods and metrics using diverse drug discovery data sets. Mol Pharm. 2017;14(12):4462–75. https://doi.org/10.1021/acs.molpharmaceut.7b00578.
Wang Y, Liu H, Fan Y, Chen X, Yang Y, Zhu L, et al. In silico prediction of human intravenous pharmacokinetic parameters with improved accuracy. J Chem Inf Model. 2019;59(9):3968–80. https://doi.org/10.1021/acs.jcim.9b00300.
Kosugi Y, Mizuno K, Santos C, Sato S, Hosea N, Zientek M. Direct comparison of the prediction of the unbound brain-to-plasma partitioning utilizing machine learning approach and mechanistic neuropharmacokinetic model. AAPS J. 2021;23(4):72. https://doi.org/10.1208/s12248-021-00604-x.
Liu K, Sun X, Jia L, Ma J, Xing H, Wu J, et al. Chemi-Net: A Molecular Graph Convolutional Network for Accurate Drug Property Prediction. Int J Mol Sci. 2019;20(14):3389. https://doi.org/10.3390/ijms20143389.
Kamiya Y, Handa K, Miura T, Yanagi M, Shigeta K, Hina S, et al. In silico prediction of input parameters for simplified physiologically based pharmacokinetic models for estimating plasma, liver, and kidney exposures in rats after oral doses of 246 disparate chemicals. Chem Res Toxicol. 2021;34(2):507–13. https://doi.org/10.1021/acs.chemrestox.0c00336.
Ekins S, Waller CL, Swaan PW, Cruciani G, Wrighton SA, Wikel JH. Progress in predicting human ADME parameters in silico. J Pharmacol Toxicol Methods. 2000;44(1):251–72. https://doi.org/10.1016/s1056-8719(00)00109-x.
Eric S, Kalinic M, Ilic K, Zloh M. Computational classification models for predicting the interaction of drugs with P-glycoprotein and breast cancer resistance protein. SAR QSAR Environ Res. 2014;25(12):939–66. https://doi.org/10.1080/1062936X.2014.976265.
Hazai E, Hazai I, Ragueneau-Majlessi I, Chung SP, Bikadi Z, Mao Q. Predicting substrates of the human breast cancer resistance protein using a support vector machine method. BMC Bioinformatics. 2013;14:130. https://doi.org/10.1186/1471-2105-14-130.
Zhong L, Ma CY, Zhang H, Yang LJ, Wan HL, Xie QQ, et al. A prediction model of substrates and non-substrates of breast cancer resistance protein (BCRP) developed by GA-CG-SVM method. Comput Biol Med. 2011;41(11):1006–13. https://doi.org/10.1016/j.compbiomed.2011.08.009.
Gantner ME, Peroni RN, Morales JF, Villalba ML, Ruiz ME, Talevi A. Development and validation of a computational model ensemble for the early detection of BCRP/ABCG2 substrates during the drug design stage. J Chem Inf Model. 2017;57(8):1868–80. https://doi.org/10.1021/acs.jcim.7b00016.
Sedykh A, Fourches D, Duan J, Hucke O, Garneau M, Zhu H, et al. Human intestinal transporter database: QSAR modeling and virtual profiling of drug uptake, efflux and interactions. Pharm Res. 2013;30(4):996–1007. https://doi.org/10.1007/s11095-012-0935-x.
Garg P, Dhakne R, Belekar V. Role of breast cancer resistance protein (BCRP) as active efflux transporter on blood-brain barrier (BBB) permeability. Mol Divers. 2015;19(1):163–72. https://doi.org/10.1007/s11030-014-9562-2.
Ose A, Toshimoto K, Ikeda K, Maeda K, Yoshida S, Yamashita F, et al. Development of a support vector machine-based system to predict whether a compound is a substrate of a given drug transporter using its chemical structure. J Pharm Sci. 2016;105(7):2222–30. https://doi.org/10.1016/j.xphs.2016.04.023.
Namasivayam V, Stefan K, Gorecki L, Korabecny J, Soukup O, Jansson PJ, et al. Physicochemistry shapes bioactivity landscape of pan-ABC transporter modulators: anchor point for innovative Alzheimer’s disease therapeutics. Int J Biol Macromol. 2022;217:775–91. https://doi.org/10.1016/j.ijbiomac.2022.07.062.
Namasivayam V, Stefan K, Pahnke J, Stefan SM. Binding mode analysis of ABCA7 for the prediction of novel Alzheimer’s disease therapeutics. Comput Struct Biotechnol J. 2021;19:6490–504. https://doi.org/10.1016/j.csbj.2021.11.035.
Namasivayam V, Stefan K, Silbermann K, Pahnke J, Wiese M, Stefan SM. Structural feature-driven pattern analysis for multitarget modulator landscapes. Bioinformatics. 2022;38(5):1385–92. https://doi.org/10.1093/bioinformatics/btab832.
Silbermann K, Stefan SM, Elshawadfy R, Namasivayam V, Wiese M. Identification of Thienopyrimidine Scaffold as an inhibitor of the ABC transport protein ABCC1 (MRP1) and related transporters using a combined virtual screening approach. J Med Chem. 2019;62(9):4383–400. https://doi.org/10.1021/acs.jmedchem.8b01821.
Ohashi R, Watanabe R, Esaki T, Taniguchi T, Torimoto-Katori N, Watanabe T, et al. Development of simplified in vitro P-Glycoprotein substrate assay and in silico prediction models to evaluate transport potential of P-Glycoprotein. Mol Pharm. 2019;16(5):1851–63. https://doi.org/10.1021/acs.molpharmaceut.8b01143.
Fang C, Wang Y, Grater R, Kapadnis S, Black C, Trapa P, et al. Prospective validation of machine learning algorithms for absorption, distribution, metabolism, and excretion prediction: an industrial perspective. J Chem Inf Model. 2023;63(11):3263–74. https://doi.org/10.1021/acs.jcim.3c00160.
Ali M. PyCaret: An open source, low-code machine learning library in Python. https://pycaret.org/. Accessed 2 Jul 2023.
Mulpuru V, Mishra N. In Silico prediction of fraction unbound in human plasma from chemical fingerprint using automated machine learning. ACS Omega. 2021;6(10):6791–7. https://doi.org/10.1021/acsomega.0c05846.
Broccatelli F, Trager R, Reutlinger M, Karypis G, Li M. Benchmarking accuracy and generalizability of four graph neural networks using large in vitro ADME datasets from different chemical spaces. Mol Inform. 2022;41(8):2100321. https://doi.org/10.1002/minf.202100321.
Grebner C, Matter H, Kofink D, Wenzel J, Schmidt F, Hessler G. Application of deep neural network models in drug discovery programs. ChemMedChem. 2021;16(24):3772–86. https://doi.org/10.1002/cmdc.202100418.
Martinez Mora A, Subramanian V, Miljkovic F. Multi-task convolutional neural networks for predicting in vitro clearance endpoints from molecular images. J Comput Aided Mol Des. 2022;36:443–57. https://doi.org/10.1007/s10822-022-00458-1.
Feinberg EN, Joshi E, Pande VS, Cheng AC. Improvement in ADMET prediction with multitask deep featurization. J Med Chem. 2020;63(16):8835–48. https://doi.org/10.1021/acs.jmedchem.9b02187.
Ramsundar B, Liu B, Wu Z, Verras A, Tudor M, Sheridan RP, et al. Is multitask deep learning practical for pharma? J Chem Inf Model. 2017;57(8):2068–76. https://doi.org/10.1021/acs.jcim.7b00146.
Kojima R, Ishida S, Ohta M, Iwata H, Honma T, Okuno Y. kGCN: a graph-based deep learning framework for chemical structures. J Cheminform. 2020;12(1):32. https://doi.org/10.1186/s13321-020-00435-6.
Pfeifer B, Holzinger A, Schimek MG. Robust random forest-based all-relevant feature ranks for trustworthy AI. Stud Health Technol Inform. 2022;294:137–8. https://doi.org/10.3233/SHTI220418.
Lilhore UK, Poongodi M, Kaur A, Simaiya S, Algarni AD, Elmannai H, et al. Hybrid model for detection of cervical cancer using causal analysis and machine learning techniques. Comput Math Methods Med. 2022;2022:4688327. https://doi.org/10.1155/2022/4688327.
Kipf TN, Welling M. Semi-supervised classification with graph convolutional networks. Preprint at https://arxiv.org/abs/1609.02907 (2016).
Sundararajan M, Taly A, Yan Q. Axiomatic attribution for deep networks. Proceedings of the 34th International Conference on Machine Learning, PMLR. 2017;70:3319–28.
Kingma DP, Ba J. Adam: A method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Akiba T, Sano S, Yanase T, Ohta T, Koyama M. Optuna: A Next-generation Hyperparameter Optimization Framework. Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining. 2019;2623–31.
Kokhlikyan N, Miglani V, Martin M, Wang E, Alsallakh B, Reynolds J, et al. Captum: A unified and generic model interpretability library for pytorch. Preprint at https://arxiv.org/abs/2009.07896 (2020).
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, et al. Pytorch: An imperative style, high-performance deep learning library. Advances in neural information processing systems. 2019;32:8024–35.
Wang M, Zheng D, Ye Z, Gan Q, Li M, Song X, et al. Deep graph library: A graph-centric, highly-performant package for graph neural networks. Preprint at https://arxiv.org/abs/1909.01315 (2019).
Li M, Zhou J, Hu J, Fan W, Zhang Y, Gu Y, et al. DGL-LifeSci: an open-source toolkit for deep learning on graphs in life science. ACS Omega. 2021;6(41):27233–8. https://doi.org/10.1021/acsomega.1c04017.
Landrum G. RDKit: Open-source cheminformatics. https://www.rdkit.org.
Dorogush AV, Ershov V, Gulin A. CatBoost: gradient boosting with categorical features support. Preprint at https://arxiv.org/abs/1810.11363 (2018).
Hou T, Wang J, Li Y. ADME evaluation in drug discovery. 8. The prediction of human intestinal absorption by a support vector machine. J Chem Inf Model. 2007;47(6):2408–15. https://doi.org/10.1021/ci7002076.
Gombar VK, Hall SD. Quantitative structure-activity relationship models of clinical pharmacokinetics: clearance and volume of distribution. J Chem Inf Model. 2013;53(4):948–57. https://doi.org/10.1021/ci400001u.
Veličković P, Cucurull G, Casanova A, Romero A, Lio P, Bengio Y. Graph attention networks. Preprint at https://arxiv.org/abs/1710.10903 (2017).
Gilmer J, Schoenholz SS, Riley PF, Vinyals O, Dahl GE, editors. Neural message passing for quantum chemistry. Proceedings of the 34th International Conference on Machine Learning, PMLR. 2017;70:1263–72.
Kuenneth C, Rajan AC, Tran H, Chen L, Kim C, Ramprasad R. Polymer informatics with multi-task learning. Patterns. 2021;2(4):100238. https://doi.org/10.1016/j.patter.2021.100238.
Wang Z, Chen Y, Liang H, Bender A, Glen RC, Yan A. P-glycoprotein substrate models using support vector machines based on a comprehensive data set. J Chem Inf Model. 2011;51(6):1447–56. https://doi.org/10.1021/ci2001583.
Poongavanam V, Haider N, Ecker GF. Fingerprint-based in silico models for the prediction of P-glycoprotein substrates and inhibitors. Bioorg Med Chem. 2012;20(18):5388–95. https://doi.org/10.1016/j.bmc.2012.03.045.
Li D, Chen L, Li Y, Tian S, Sun H, Hou T. ADMET evaluation in drug discovery. 13. Development of in silico prediction models for P-glycoprotein substrates. Mol Pharm. 2014;11(3):716–26. https://doi.org/10.1021/mp400450m.
Shaikh N, Sharma M, Garg P. Selective fusion of heterogeneous classifiers for predicting substrates of membrane transporters. J Chem Inf Model. 2017;57(3):594–607. https://doi.org/10.1021/acs.jcim.6b00508.
Namasivayam V, Silbermann K, Wiese M, Pahnke J, Stefan SM. C@PA: computer-aided pattern analysis to predict multitarget abc transporter inhibitors. J Med Chem. 2021;64(6):3350–66. https://doi.org/10.1021/acs.jmedchem.0c02199.
Yang K, Swanson K, Jin W, Coley C, Eiden P, Gao H, et al. Analyzing learned molecular representations for property prediction. J Chem Inf Model. 2019;59(8):3370–88. https://doi.org/10.1021/acs.jcim.9b00237.
Acknowledgements
We thank the research members at Takeda Pharmaceutical Company Ltd. for their experimental and technical contributions and Hiroko Hishinuma for technical assistance with PyCaret coding.
Funding
The study used internal budget of Takeda Pharmaceutical Company Limited.
Author information
Authors and Affiliations
Contributions
Asahi Adachi: Substantial contributions to conduct the machine learning and interpretation of data for the work.; drafting the work manuscript.
Tomoki Yamashita: Interpretation of data for the work; review the manuscript.
Shigehiko Kanaya: Review the manuscript; final approval of the version to be published.
Yohei Kosugi: Substantial contributions to the conception and design of the work; interpretation of data for the work; review the manuscript; final approval of the version to be published.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing financial interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Adachi, A., Yamashita, T., Kanaya, S. et al. Ensemble Machine Learning Approaches Based on Molecular Descriptors and Graph Convolutional Networks for Predicting the Efflux Activities of MDR1 and BCRP Transporters. AAPS J 25, 88 (2023). https://doi.org/10.1208/s12248-023-00853-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-023-00853-y