Abstract
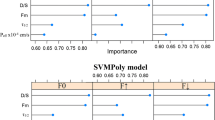
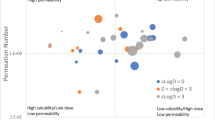
In this study, observed food effects of 473 drugs were categorized into positive, negative, or no effects and compared with the predictions made by machine learning (ML), the Biopharmaceutics Classification System (BCS) and refined Developability Classification System (rDCS). All methods used primarily in silico estimates for prediction, and for ML, four algorithms were evaluated using nested cross-validation to select important information from 371 features calculated based on the chemical structure. Approximately 18 features, including estimated solubility in biorelevant media, were selected as important, and the random forest classifier was the best among four algorithms with 36.6% error rate (ER) and 10.8% opposite prediction rate (OPR). The prediction by rDCS utilizing solubility in a biorelevant medium was somewhat inferior, but not by much; 41.0% ER and 11.4% OPR. Compared with these two methods, the prediction by BCS was inferior; 54.5% ER and 21.4% OPR. ER was improved modestly by using measured features instead of in silico estimates when BCS was applied to a subset of 151 drugs (46.4% from 55.0%). ML and rDCS predicted the food effects of the same subset using in silico estimates with ERs of 37.7% and 42.4%, respectively, suggesting that the predictions by ML and rDCS using in silico features are similar or more accurate than those by BCS using measured features. These results suggest that ML was useful in revealing essential features from complex information and, together with rDCS, is effective in predicting food effects during drug development, including early drug discovery.
Graphical abstract






Similar content being viewed by others
References
FDA. Assessing the Effects of Food on Drugs in INDs and NDAs – Clinical Pharmacology Considerations: Guidance for industry. 2019
Marroum PJ, Nuthalapati S, Parikh A, Shebley M, Hoffman D, Zha J, et al. Industry Perspective on Standardizing Food-Effect Studies for New Drug Development. Clin Pharmacokinet. 2018;57(8):901–9.
Riedmaier AE, DeMent K, Huckle J, Bransford P, Stillhart C, Lloyd R, et al. Use of Physiologically Based Pharmacokinetic (PBPK) Modeling for Predicting Drug-Food Interactions: an Industry Perspective. AAPS J. 2020;22(6):123.
O’Shea JP, Holm R, O’Driscoll CM, Griffin BT. Food for thought: formulating away the food effect - a PEARRL review. J Pharm Pharmacol. 2019;71(4):510–35.
Fleisher D, Li C, Zhou Y, Pao LH. Karim A. Drug, meal and formulation interactions influencing drug absorption after oral administration. Clinical implications. Clin Pharmacokinet. 1999;36(3):233–54.
Singh BN. A quantitative approach to probe the dependence and correlation of food-effect with aqueous solubility, dose/solubility ratio, and partition coefficient (Log P) for orally active drugs administered as immediate-release formulations. Drug Dev Res. 2005;65(2):55–75.
Gu CH, Li H, Levons J, Lentz K, Gandhi RB, Raghavan K, et al. Predicting effect of food on extent of drug absorption based on physicochemical properties. Pharm Res. 2007;24(6):1118–30.
Amidon GL, Lennernäs H, Shah VP, Crison JR. A theoretical basis for a biopharmaceutic drug classification: the correlation of in vitro drug product dissolution and in vivo bioavailability. Pharm Res. 1995;12(3):413–20.
ICH. Biopharmaceutics classification system-based biowaivers M9. 2019.
Butler JM, Dressman JB. The developability classification system: application of biopharmaceutics concepts to formulation development. J Pharm Sci. 2010;99(12):4940–54.
Rosenberger J, Butler J, Dressman J. A Refined Developability Classification System. J Pharm Sci. 2018;107(8):2020–32.
Rosenberger J, Butler J, Muenster U, Dressman J. Application of a Refined Developability Classification System. J Pharm Sci. 2019;108(3):1090–100.
Kosugi Y, Hosea N. Direct Comparison of Total Clearance Prediction: Computational Machine Learning Model versus Bottom-Up Approach Using In Vitro Assay. Mol Pharm. 2020;17(7):2299–309.
Watanabe R, Ohashi R, Esaki T, Kawashima H, Natsume-Kitatani Y, Nagao C, et al. Development of an in silico prediction system of human renal excretion and clearance from chemical structure information incorporating fraction unbound in plasma as a descriptor. Sci Rep. 2019;9(1):18782.
Wang Y, Liu H, Fan Y, Chen X, Yang Y, Zhu L, et al. In Silico Prediction of Human Intravenous Pharmacokinetic Parameters with Improved Accuracy. J Chem Inf Model. 2019;59(9):3968–80.
Varma S, Simon R. Bias in error estimation when using cross-validation for model selection. BMC Bioinformatics. 2006;7:91.
Deng J, Zhu X, Chen Z, Fan CH, Kwan HS, Wong CH, et al. A Review of Food-Drug Interactions on Oral Drug Absorption. Drugs. 2017;77(17):1833–55.
Cappelli CI, Manganelli S, Lombardo A, Gissi A, Benfenati E. Validation of quantitative structure-activity relationship models to predict water-solubility of organic compounds. Sci Total Environ. 2013;463–464:781–9.
Kursa MB, Rudnicki WR. Feature Selection with the Boruta Package. J Stat Softw. 2010;36(11):1–13.
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, et al. Scikit-learn: Machine Learning in Python. JMLR. 2011;12:2825–30.
Anaconda Software Distribution. Computer software. Vers. 2–2.4.0. Anaconda, Nov. 2016. Web. <https://anaconda.com>.
Müller AC, Guido S. Introduction to Machine Learning with Python. O'Reilly Media, Inc. 2016.
Dahan A, Lennernäs H, Amidon GL. The fraction dose absorbed, in humans, and high jejunal human permeability relationship. Mol Pharm. 2012;9(6):1847–51.
Larregieu CA, Benet LZ. Distinguishing between the permeability relationships with absorption and metabolism to improve BCS and BDDCS predictions in early drug discovery. Mol Pharm. 2014;11(4):1335–44.
Lindenberg M, Kopp S, Dressman JB. Classification of orally administered drugs on the World Health Organization Model list of Essential Medicines according to the biopharmaceutics classification system. Eur J Pharm Biopharm. 2004;58(2):265–78.
Varma MV, Gardner I, Steyn SJ, Nkansah P, Rotter CJ, Whitney-Pickett C, et al. pH-Dependent solubility and permeability criteria for provisional biopharmaceutics classification (BCS and BDDCS) in early drug discovery. Mol Pharm. 2012;9(5):1199–212.
Kawai Y, Fujii Y, Tabata F, Ito J, Metsugi Y, Kameda A, et al. Profiling and trend analysis of food effects on oral drug absorption considering micelle interaction and solubilization by bile micelles. Drug Metab Pharmacokinet. 2011;26(2):180–91.
Sugano K, Kataoka M, Mathews Cda C, Yamashita S. Prediction of food effect by bile micelles on oral drug absorption considering free fraction in intestinal fluid. Eur J Pharm Sci. 2010;40(2):118–24.
Damle B, Ravandi F, Kaul S, Sonnichsen D, Ferreira I, Brooks D, et al. Effect of food on the oral bioavailability of UFT and leucovorin in cancer patients. Clin Cancer Res. 2001;7(3):517–23.
Yeh KC, Deutsch PJ, Haddix H, Hesney M, Hoagland V, Ju WD, et al. Single-dose pharmacokinetics of indinavir and the effect of food. Antimicrob Agents Chemother. 1998;42(2):332–8.
Daneshmend TK, Roberts CJ. The influence of food on the oral and intravenous pharmacokinetics of a high clearance drug: a study with labetalol. Br J Clin Pharmacol. 1982;14(1):73–8.
Xiao J, Tran D, Zhang X, Zhang T, Seo S, Zhu H, Zou P. Biliary Excretion-Mediated Food Effects and Prediction. AAPS J. 2020;22(6):124.
Li M, Zhao P, Pan Y, Wagner C. Predictive Performance of Physiologically Based Pharmacokinetic Models for the Effect of Food on Oral Drug Absorption: Current Status. CPT Pharmacometrics Syst Pharmacol. 2018;7(2):82–9.
Acknowledgements
This work was supported by AMED under Grant Number JP20be0304203.
Funding
This work was supported by AMED under Grant Number JP20be0304203.
Author information
Authors and Affiliations
Contributions
Substantial contributions to the conception or design of the work, Y.H., H.Y, and A.H.; Drafting the work or revising it critically for important intellectual content, Y.H. and A.H.; Final approval of the version to be published, Y.H., H.Y, and A.H., Agreement to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved, Y.H., H.Y, and A.H.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest associated with this manuscript. Yusuke Hoshino is an employee of Zeria Pharmaceutical Co., Ltd. However, Zeria Pharmaceutical Co., Ltd. did not contribute toward the analysis in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hoshino, Y., Yoshioka, H. & Hisaka, A. Comparison of Predictions by BCS, rDCS and Machine Learning for the Effect of Food on Oral Drug Absorption Based on Features Calculated In silico. AAPS J 24, 10 (2022). https://doi.org/10.1208/s12248-021-00664-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1208/s12248-021-00664-z