Abstract
Various branches of any living species are destined to specified identification. Henceforth, imposing of bioinformatics, including various supportive software along with wet lab experiments, has led to conclude genus and species of unidentified strains. Further, for the construction of phylogenetic trees, the principle of minimum evolution has been considered. Since few years, rapid sequencing of DNA has been adapted, and data on nucleotide sequences of genome in diverse organisms have accumulated at an accelerated pace. In the present study, molecular identification of the fungus, Beauveria bassiana, isolated from the soil of a tea cultivation area was endeavoured. Soil from Bogidhola tea garden of Golaghat District, Assam, India, has captured an attention as it holds some natural pesticidal properties. Since B. bassiana possesses substantial pesticidal property, an attempt was carried out to identify the presence of B. bassiana in that particular soil of tea garden, followed by its molecular identification. Sequencing of D2 subunit spanning 28S rDNA of isolated fungi had been completed for molecular identification imposing bioinformatic tools and among 15 isolates compared where a correction of 100% identification was observed with isolate of B. bassiana strain CCFC225554.
Similar content being viewed by others
Background
Tea is a substantial source of alkaloids and antioxidant which have arrested the increasing interest of a remarkable beverage and health drink. Also, India is the world’s largest tea producer country. The maximum production of tea in India is from north- Bengal, Assam and few parts of southern India (Rio et al., 2004). On the contrary, being susceptible to diseases caused by various insect pests, most tea garden owners protect their tea trees by using synthetic insecticides Being the sources of toxic chemicals, synthetic insecticides cause hormonal disorders, cancer, infertility, and Parkinson’s disease. Therefore, an attempt has been taken to find out natural pest-controlling methods in order to substitute synthetic insecticides (Hazarika et al., 2009).
It has been found that, irrespective of pesticides supplementation, tea plantation at Bogidhola Tea Estate in Golaghat district, Assam, are rarely affected by pests compared to other tea plantation areas of the district. Therefore, it can be inferred from the observation that the soil from tea plantation area of Bogidhola Tea Estate probably contains some biological pest control agents that protect tea plants from certain pests.
Apart from micro- and macroelements, microbes are important components of soil which play an important role in rendering nutrition and protection to tree against pests. Among soilborne microbes, fungi play an important role in controlling insect as they secrete mycotoxins that inhibit the growth of pests. Beauveria bassiana is one of the most common and prominent soilborne fungi that play a major role in controlling insect. The spores of B. bassiana get attached to insect cuticle. Afterwards, spores penetrate into haemocel followed by proliferation of hyphae under suitable conditions. During proliferation, they secrete mycotoxins which ultimately desiccate the hosts (Feng et al., 1994; da Costa et al., 2001). Therefore, it can be assumed that the soil from Bogidhola Tea Estate contains B. bassiana, which in turn control pests attack by destroying them.
In this study, an attempt was carried out to isolate B. bassiana strain from tea garden soil of Golaghat district, Assam, and identify it with the aid of 28S rDNA sequencing. Sequencing, which is a procedure to determine the precise order of nucleotides within a DNA, is one of the most secured methods for identifying an organism as it makes the comparison of DNA sequences among biological species easy to resolve their evolutionary relationships. It is demonstrated that 28S rDNA sequences (28S rDNA) have been used for phylogenetic studies and the relationships among distantly related taxa and also among closely related taxa (Ui and Won, 1999 and Ninet et al., 2003). DNA sequence that coded for ribosomal RNA was proposed to be ribosomal DNA (rDNA). rDNA of eukaryotes consists of a tandem repeat of an operon, composed of NTS (non-transcribed spacer), ETS (external transcribed spacer), 18S, ITS1 (Internal transcribed spacer 1), 5.8S, ITS2, and 28S tracts (Richard et al., 2008). Coding regions of rDNA are highly specific to species but ITS regions vary due to insertions, deletions, and point mutations. The different coding regions of the rDNA repeats are highly conserved and typically show distinct evolutionary rates (Fig. 1).
This DNA provides phylogenetic information of species belonging to wide systematic levels (Hillis and Dixon, 1991). The D2 domain consists generally a 600-bp nucleotide domain, located at the 5′ end of a large subunit of rDNA. Kurtzman and Robnett (1995) and Kwiatkowski et al. (2012) showed that with the help of sequence divergence of the D2 domain, most species of yeast and fungi could be identified.
The objectives of this work were to (1) identify the B. bassiana strain isolated from the soil of Bogidhola tea estate, Assam, and (2) find out the supporting reason that reinforce the presence of B. bassiana in that particular tea garden soil.
Materials and methods
Field sites and collection of soil sample
The field of study was conducted from soil sample of Bogidhola Tea Estate, Golaghat district, Assam, India. From the site, four soil samples were collected within 100 m to each other and labelled (S1, S2, S3, and S4). Five hundred grams of each soil sample was taken from the top 10 cm of the soil surface. Afterwards, the samples were air dried at room temperature for 7 days followed by passing through 0.71-mm mesh sieve and were subsequently packed in plastic bags (Suzuki et al., 1999 and Safavi, 2010).
Screening of microorganisms present in the collected soil samples
Thirty grams of Sabouraud’s dextrose agar (SDAY) media, a tailor-made media composed of dextrose (20 kg/m3), mycological peptone (10 kg/m3), agar (kg/m3), and yeast extract (1%), was prepared by dissolving into 1-L de-ionised water followed by pouring in four separate petri dishes (P1, P2, P3, and P4) and left to set. The pH of the media was maintained at 5.8 during the course of procedure. Next, 1 g of soil sample was taken from each of the S1, S2, S3, and S4 and inoculated in SDAY media of P1, P2, P3, and P4, respectively. Subsequently, all petri dishes were kept in total darkness at 298 K for 5 days. P1, P2, and P4 were selected for subculture and isolation of B. bassiana with profused conidia growth (Safavi, 2010 and Mondal et al., 2015).
Primary screening: isolation of entomopathogenic fungi
Isolation of pure spore culture of entomopathogenic fungi is crucial for developing pure mycelia of B. bassiana. Another tailor-made media (PDAD media), consisting of potato dextrose agar supplemented with 1 g/L yeast extract, 0.0004 kg chloramphenicol, 0.00004 kg streptomycin, 4 units of penicillin, 0.05 g/L gentamicin, and 0.004% active ingredient dodine, were prepared in order to endorse the growth of entomopathogenic fungi only. Commercial fungicide ‘Syllit’ (65% active ingredient) was used as the source of dodine. The pH of PDAD media was maintained at 6.9 throughout the study. While chloromphenicol, being a broad spectrum antibiotic, inhibits the growth of a wide-range Gram-positive and Gram-negative bacteria, the combination of streptomycin, gentamycin, and penicillin inhibits the bacterial and yeast growth. After preparing, PDAD media were poured in a petri dish (P5) and left to set. Afterwards, a subculture was done on the P5 with the inocula taken from each of the mycelia developed on P1, P2, and P4. Prolific growths of entomopathogenic mycelia were observed in the subcultured media (P5) after 5-day incubation at 298 K temperature in total darkness. A control strain of B. bassiana (strain ARSEF502) was also grown under similar incubation conditions on the PDAD media. Identification of B. bassiana mycelia grown in P5 was achieved by comparing the growth morphology with those mycelia growth of B. bassiana control strain (Fernandes et al., 2010).
Development of pure B. bassiana mycelia
Mycelia of B. bassiana were further harvested from the mycelia subculture developed in P5 by a microbiological loop and immediately suspended in a 0.01% tween 80 solution, following vortexes for 30s. Next, 20-μL aliquot of mycelia suspension was transferred to a petri dish (P6) containing PDAD media, following incubation for 5 days under 298 K at total darkness. A pure mycelia growth of B. bassiana was observed on the surface of the P6 and hence confirmed by comparing to the mycelia growth of B. bassiana control strain. Afterwards, mycelia of B. bassiana isolate, developed in the P6, were removed from the agar surface with a sterile scalpel and transferred to a sterile petri dish. Subsequently, mycelium was cut into small pieces (less than 0·5 mm) with a sterile scalpel and was transferred in 100-ml Potato Dextrose broth. After 1 week, the dried pure mycelia of the fungal isolate were harvested by centrifugation and stored in a freezer. Additionally, lactophenol cotton blue staining was done in order to identify the nature of the fungus grown in P6 containing PDAD (Al-Samarrai and Schmid, 2000; Castrillo et al., 2003 and Fernandes et al., 2010).
A purified mycelium culture of isolated B. bassiana strain, developed over P6, is illustrated in Fig. 2.
Fungal DNA extraction
In a 1.5-ml centrifuge tube, 0.005 g of dried pure mycelia was suspended in a solution composed of 150 μl of 10%(w/v) sodium dodecyl sulfate, 500 μl of Tris buffer (100 mm Tris HCL, pH 8.0, 40 mM ethylene diamine tetra acetic acid [EDTA]), and 300 μl of Benzyl Chloride and consequently incubated at 55 °C for 30 min. Subsequent treatment with RNase (1 mg/ml) and phenol:chloroform:isoamylalcohol (25:24:1) was undertaken in order to remove proteins. Afterwards, the tube was vortexed at 10000×g for 1 min, and next, the supernatant was separated. After that, DNA was precipitated from supernatant by adding 2.5 volumes 100% ice-cold ethanol. Then, the DNA pellet was washed carefully with 2 volumes of 70% ethanol and then re-suspended with distilled water (Urtz and Rice, 1997 and Park et al., 2001).
PCR amplification and sequencing of the D2 region of the large subunit (28S) rDNA gene
The nuclear rDNA segment corresponding to the D2 region of fungal 28S ribosomal RNA (28S-D2 rDNA) was amplified through polymerase chain reaction (PCR), carried out using a Perkin-Elmer GeneAmp PCR System 2400. LF402Fmix1 [sequence: GTGAAATTGYTRAAAGGGAA] and LF402Fmix3 [sequence: GTGAAATTGTCAAAAGGGAA] were used as forward primers whereas, TW13 [sequence: GGTCCGTGTTTCAAGACG] and LR3 [sequence: GGTCCGTGTTTCAAGAC] were used as reverse primers with the aim of amplifying D2 region of 28S rDNA only (Porter and Golding, 2012 and Tedersoo et al., 2015). PCR procedure was initialized with the denaturation of DNA template strands at 95 °C for 5 min, followed by 30 cycles of amplification (denaturation at 95 °C for 1 min succeeding annealing at 45 °C for 1 min and next extension for 1 min at 72 °C) and then final extension for 5mins at 72 °C. Purification of amplified DNA product, following isolation, was done in a mandatory basis prior to sequencing of that amplified DNA. The amplified DNA was excised in × 0.5 TBE buffer by gel electrophoresis in a 1.6% agarose gel supplemented with the ethidium bromide with the purpose of isolating and purifying it, and a single discrete PCR amplicon band of 700 bp was observed in agarose gel (Fig. 3).
The isolated and purified DNA was extracted from agarose gel, using a QIAGEN gel elution kit (Qiagen, Wartworth, CA). Afterwards, direct sequencing of the PCR products was done, according to the manufacturer’s protocol, by a ABI 3730xl Genetic Analyser (ThermoFisher scientific) with a BDT v3.1 Cycle sequencing kit (ThermoFisher scientific). LF402Fmix1 primer and TW13 primer were used for sense and antisense sequencing, respectively (Park et al., 2001 and Park et al., 2002, and Ninet et al., 2003).
Data analysis
A consensus sequence of rDNA portion corresponding to D2 region of 28S rRNA of isolated Beauveria strain was found, using aligner software, with the help of sense and antisense sequencing. The consensus sequence found was further used to carry out BLAST in order to find out organisms from NCBI database having the exactly same sequence spanning the D2 region of 28S rDNA. Based on a maximum identity score achieved from BLAST, strains, having exactly same or nearly same sequence across the D2 region in 28S rDNA, were selected. The inferred rDNA sequence has been deposited in the NCBI database. A phylogenetic tree was constructed by neighbour-joining method using MEGA version 5.0 with an aim to comprehend the evolutionary relationship among the isolated Beauveria strain and other closely related taxa, and the strength of the internal branches of the consensus tree were statistically tested from 1000 bootstrap replication analysis. Kimura 2-parameter model was chosen for neighbour-joining-based phylogenetic tree reconstruction and evolutionary distance computation in the units of the number of base substitutions per site (Park et al., 2001).
Presence of Beauveria bassiana in the soil sample of other tea gardens
In order to discover whether the presence of B. bassiana is mandatory for insect pest control or not, an additional study was conducted with the aim of determining the presence of the tested fungi in the soil of other tea gardens situated at Golaghat district. Soil samples were collected from each of the tea gardens situated at Golaghat district, followed by attempting isolation of B. bassiana from their respective soil samples. From collection of soil samples to isolation of B. bassiana have been completed according to the same procedure followed during the fungal isolation from Bogidhola Tea Estate soil.
Results and discussion
Sequence alignment
Only a single band of 700 bp was achieved by gel electrophoresis of PCR products (Fig. 3). Therefore, it can be inferred that a particular segment of template rDNA was amplified and isolated by agarose gel electrophoresis. Additionally, direct sequencing procedure also confirmed the results.
With the help of direct sequencing procedure, a sequence of 570 bp (FS1) (5′-AACCAACAGGGATTGCCCCAGTAACGGCGAGTGAAGCGGCAACAGCTCAAAT TTGAAATCTGGCCCTCGGGTCCGAG TTGTAATTTGTAGAGGATGCTTTTGGCGAGGTGCCTTCCGAGTTCCCTGAAACGGGACGCCATAGAGGGTGAGAGCCCCGTCTGGTCGGACACCGAGCCTCTGTAAAGCTCCTTCAACGAGTCGAGTAGTTTGGGAATGCTGCTCAAAATGGGAGGTATATGTCTTCTAAAGCTAAATATTGGCCAGAGACCGATAGCGCACAAGTAGAGTGATCGAAAGATGAAAAGCACTTTGAAAAGAGGGTTAAAAAGTACGTGAAATTGTTGAAAGGGAAGCGCCTATGACCAGACTTGCGCCCGGTGAATCATCCAGCGTTCTCGCTGGTGCACTTTGCCGGGCACAGGCCAGCATCAGTTTGGCGCGGGGGAAAAAGGCTTCGGGAATGTGGCTCCCTCGGGAGTGTTATAGCCCGCTGCGTAATACCCTGCGCCGGACTGAGGTACGCGCATCGCAAGGATGCTGGCGTAATGGTCATCAGCGACCCGTCTTGAA) was found by the sense primer LF402Fmix1 and a reverse sequence of 570 bp (RS2) (5′-CAGCATCCTTGCGATGCGCGTACCTCAGTCCGGCGCAGGGTATTACGCAGCGGGCTATAACACTCCCGAGGGAGCCACATTCCCGAAGCCTTTTTCCCCCGCGCCAAACTGATGCTGGCCTGTGCCCGGCAAAGTGCACCAGCGAGAACGCTGGATGATTCACCGGGCGCAAGTCTGGTCATAGGCGCTTCCCTTTCAACAATTTCACGTACTTTTTAACCCTCTTTTCAAAGTGCTTTTCATCTTTCGATCACTCTACTTGTGCGCTATCGGTCTCTGGCCAATATTTAGCTTTAGAAGACATATACCTCCCATTTTGAGCAGCATTCCCAAACTACTCGACTCGTTGAAGGAGCTTTACAGAGGCTCGGTGTCCGACCAGACGGGGCTCTCACCCTCTATGGCGTCCCGTTCCAGGGAACTCGGAAGGCACCTCGCCAAAAGCATCCTCTACAAATTACAACTCGGACCCGAGGGCCAGATTTCAAATTTGAGCTGTTGCCGCTTCACTCGCCGTTACTGGGGCAATCCCTGTTGGTTTCTTTTCCTCCGCTTATTGATATGCTTAAT) was found by antisense primer TW13.
With the help of aligner software, FS1 and RS2 were compared in order to find out the consensus sequence, and therefore, a consensus sequence of 600 bp (CS) (5′-ATTAAGCATATCAATAAGCGGAGAAAAGAAACCAACAGGGATTGCCCCAGTAACGGCGAGTGAAGCGGCAACAGCTCAAATTTGAAATCTGGCCCTCGGGTCCGAGTTGTAATTTGTAGAGGATGCTTTTGGCGAGGTGCCTTCCGAGTTCCCTGGAACGGGACGCCATAGAGGGTGAGAGCCCCGTCTGGTCGGACACCGAGCCTCTGTAAAGCTCCTTCAACGAGTCGAGTAGTTTGGGAATGCTGCTCAAAATGGGAGGTATATGTCTTCTAAAGCTAAATATTGGCCAGAGACCGATAGCGCACAAGTAGAGTGATCGAAAGATGAAAAGCACTTTGAAAAGAGGGTTAAAAAGTACGTGAAATTGTTGAAAGGGAAGCGCCTATGACCAGACTTGCGCCCGGTGAATCATCCAGCGTTCTCGCTGGTGCACTTTGCCGGGCACAGGCCAGCATCAGTTTGGCGCGGGGGAAAAAGGCTTCGGGAATGTGGCTCCCTCGGGAGTGTTATAGCCCGCTGCGTAATACCCTGCGCCGGACTGAGGTACGCGCATCGCAAGGATGCTGGCGTAATGGTCATCAGCGACCCGTCTTGAA) was found and considered as the consensus sequence of D2 region of 28S rDNA.
With a view to identify the isolated B. bassiana sp., a BLAST search was carried out between CS and NCBI database and 15 sequences, with their respective host organisms, were selected based on maximum identity score (Table 1).
Based on the BLAST search, presented in Table 1, the CS sequence of isolated B. bassiana showed maximum identity (100%) with fungal strain CCFC225554 (accession no: AY283555.1). The consensus DNA sequence, derived from D2 region in 28S rDNA of B. bassiana isolated from Bogidhola Tea Estate, was deposited in NCBI database under the accession no KT359346.
Phylogenetic analysis
A distance matrix, based on the nucleotide substitution per site at D2 region in 28 s rDNA of fungi, listed in Table 1, is attempted and represented in Table 2.
Whether the values represent the rate of nucleotide base substitution, values on top upper right of the matrix represent rates of transversion (A or G  C or T) and the values on the lower left represent rates of transition (A
C or T) and the values on the lower left represent rates of transition (A  G;C
G;C  T). From the matrix, it could be observed that B. bassiana isolate had a nucleotide substitution score of 0.000 with AY283555.1 and had a maximum nucleotide substitution value of 0.019 with KC510278.1 and HQ591387.1. Therefore, based on the score achieved from distance matrix, it could be concluded that there was no variation in nucleotide alignment spanning D2 region in 28 s rDNA of isolated B. bassiana and its strain AY283555.1 (Park et al., 2002; Holmes, 2003 and Soltis and Soltis, 2003).
T). From the matrix, it could be observed that B. bassiana isolate had a nucleotide substitution score of 0.000 with AY283555.1 and had a maximum nucleotide substitution value of 0.019 with KC510278.1 and HQ591387.1. Therefore, based on the score achieved from distance matrix, it could be concluded that there was no variation in nucleotide alignment spanning D2 region in 28 s rDNA of isolated B. bassiana and its strain AY283555.1 (Park et al., 2002; Holmes, 2003 and Soltis and Soltis, 2003).
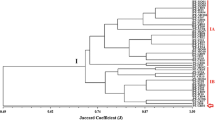
Based on the homology of sequence across D2 region in 28S r DNA of isolated Beauveria strain with 15 other closely related strains cited from gene bank, a phylogenetic relationship is assessed and presented in Fig. 4.
In the figure, horizontal lines, considered as branches, represent evolutionary lineages changing over time because of nucleotide substitution. The amounts of changes were directly proportional to the length of the branches in horizontal dimension. A phylogenetic tree was generated from original dataset with the aid of neighbourhood-joining method. Additionally, 1000 bootstrap pseudoreplicate datasets of original dataset were carried out, followed by generation of phylogenetic tree against each of the bootstrapped data sets. Afterwards, the topology of the trees, generated from bootstrap data sets, was compared to the topology of the original phylogenetic tree, with an aim to infer the bootstrap consensus phylogenetic tree and taken to represent the evolutionary history of the isolated B. bassiana strain. During the topology comparison, based on the support of the each node, bootstrap values were assigned against each interior branch node of original tree. As bootstrap value was used to verify the solidity of each node, bootstrap values of 95% or greater was considered statistically significant whether nodes can be rejected if they occur in less than 5% of the bootstrap estimates. Although two nodes in consensus phylogenetic tree somewhat achieved comparatively low bootstrap replication values (31 and 33%), but other nodes were supported by bootstrap values between 93 and 100%, which were statistically significant and indicate ‘support’ for a clade (Park et al., 2002; Peterson, 2008 and Ergashev et al., 2009).
The branches holding AY283555.1 and isolated B. bassiana strain were originated from a single node, containing a bootstrap value of 99, where both branches had the same length and described that both had been evolved from same ancestor, with exactly equal and same genetic changes (Fig. 4). Whether bootstrap value of 100 signifies the maximum support of respective nodes, the node holding the AY283555.1 and branches achieved a bootstrap value of 99 which was almost proximate to 100. Bootstrap value of 99 at the inner node means that the species AY283555.1 and B. bassiana isolate were siblings in 99% of the bootstrap replications and were grouped together in monophyletic clade in 99% of the bootstrap replications (Park et al., 2002; Peterson, 2008 and Ergashev et al., 2009). Additionally, even though AY184966.1and HQ591387.1 were also grouped in a monophyletic clade attaining bootstrap replication value of 99, but they were originated from an entirely different node as their ancestor was different from the ancestor evolved to generate AY283555.1 and B. bassiana isolate. This result is consistent with that of D2 region of LSU rDNA sequence analysis.
Presence of Beauveria bassiana in other tea garden soil samples
Although a very poor mycelial growth was found on a few petri dishes inoculated with the soil samples taken from other tea gardens, their occurrence was ignored as their mycelia growth was found very poor compared to the mycelia growth of B. bassiana, found in P1, P2, P3, P4, P5, and P6. However, the growth of B. bassiana was not observed on most of the petri dishes, following inoculation with the soil of other tea gardens. It was revealed from the nutritionists of Bogidhola Tea Estate that, in addition to the regular nutritional components, they often add shrimp powder into the soil, as a nutritional supplement. Rao et al. (2006) reported that shrimp powder was a good source of carbohydrate entitled trehalose which supported the development and growth of B. bassiana (Campbell et al., 1983 and Rao et al., 2006). With this information, it could be concluded that supply of trehalose into the soil of Bogidhola Tea Estate endorses the growth of B. bassiana in soil. On the contrary, no such additional nutritional supplements, containing trehalose, have been supplied to the soil of the other tea gardens, and for this reason, a very insignificant existence of B. bassiana was found in their soil.
Conclusion
It could be concluded from the topology of the consensus phylogenetic tree that since both of B. bassiana isolate and B. bassiana strain CCFC225554, evolved from the same ancestor through same course of genetic evolution, must be identical. It was observed that the soil sample from Bogidhola tea estate contains substantial amount of B. bassiana, while a very negligible existence of that fungus was detected in the soil of the other tea gardens. Obtained results proved that B. bassiana is a crucial component of soil that render protection of the tea plantation of Bogidhola tea estate, while lack of this fungus may make tea plantation of other areas more vulnerable to insect pest attack. Despite the fact that the nutritional components present in the soil across the Golaghat district are nearly the same, an additional component, named trehalose, present only in the soil of Bogidhola Tea estate, endorses growth of B. bassiana inside that particular tea garden.
References
Al-Samarrai TH, Schmid J (2000) A simple method for extraction of fungal genomic DNA. Lett Appl Microbiol 30(1):53–56
Campbell RK, Barnes GL, Cartwright BO, Eikenbary RD (1983) Growth and sporulation of Beauveria bassiana and Metarhizium anisopliae in a basal medium containing various carbohydrate sources. J Invertebr Pathol 41(1):117–121
Castrillo LA, Vandenberg JD, Wraight SP (2003) Strain-specific detection of introduced Beauveria bassiana in agricultural fields by use of sequence-characterized amplified region markers. J Invertebr Pathol 82(2):75–83
da Costa GL, Sarquis MIM, de Moraes AML, Bittencourt VREP (2001) Isolation of Beauveria bassiana and Metarhizium anisopliae var. anisopliae from Boophilus microplus tick (Canestrini, 1887), in Rio de Janeiro State, Brazil. Mycopathologia 154(4):207–209
Ergashev K, Guzalova AG, Leclerque A (2009) Phylogenetic characterization of entomopathogenic fungi from Uzbekistan. IOBC/wprs Bulletin IOBC/wprs Bulletin 45:311–314
Feng MG, Poprawski TJ, Khachatourians GG (1994) Production, formulation and application of the entomopathogenic fungus Beauveria bassiana for insect control: current status. Biocontrol Sci Tech 4(1):3–34
Fernandes EKK, Keyser CA, Rangel DEN, Foster RN, Roberts DW (2010) CTC medium: a novel dodine-free selective medium for isolating entomopathogenic fungi, especially Metarhizium acridum, from soil. Biol Control 54(3):197–205
Hazarika LK, Bhuyan M, Hazarika BN (2009) Insect pests of tea and their management. Annu Rev Entomol 54:267–284
Hillis DM, Dixon MT (1991) Ribosomal DNA: molecular evolution and phylogenetic inference. Q Rev Biol 66(4):411–453
Holmes S (2003) Bootstrapping phylogenetic trees: theory and methods. Statist Sci 18(2):241–255
Kurtzman CP, Robnett CJ (1995) Molecular relationships among hyphal ascomycetous yeasts and yeast like taxa. Can J Bot 73(S1):824–830
Kwiatkowski NP, Babiker WM, Merz WG, Carroll KC, Zhang SX (2012) Evaluation of nucleic acid sequencing of the D1/D2 region of the large subunit of the 28S rDNA and the internal transcribed spacer region using SmartGene IDNS software for identification of filamentous fungi in a clinical laboratory. J Mol Diagn 14(4):393–401
Mondal S, Mukherjee A, Datta S, Bhattacharya P (2015) Studies on isolation, optimization and bioprocess engineering behaviour of entomopathogenic fungi, Beauveria bassiana. Indian Chemical Engineer 59(1):41–56
Ninet B, Jan I, Bontems O, Chenne BL, Jousson O, Panizzon R, Lew D, Monod M (2003) Identification of dermatophyte species by 28S ribosomal DNA sequencing with a commercial kit. J Clin Microbiol 41(2):826–830
Park HS, Kim GY, Nam B, Lee S, Lee J (2002) The determination of the partial 28s ribosomal DNA sequences and rapid detection of Phellinus linteus and related species. Mycobiology 30(2):82–87
Park J, Kim G, Park H, Nam B, An W, Cha J, Lee TH, Lee J (2001) Phylogenetic analysis of caterpillar fungi by comparing ITS 1-5.8S-ITS 2 ribosomal DNA sequences. Mycobiology 29(3):121–131
Peterson SW (2008) Phylogenetic analysis of Aspergillus species using DNA sequences from four loci. Mycologia 100(2):205–226
Porter TM, Golding GB (2012) Factors that affect large subunit ribosomal DNA amplicon sequencing studies of fungal communities: classification method, primer choice, and error. PLoS One 7(4):e35749
Rao YK, Lu S, Liu B, Tzeng Y (2006) Enhanced production of an extracellular protease from Beauveria bassiana by optimization of cultivation processes. Biochem Eng J 28(1):57–66
Richard GF, Kerrest A, Dujon B (2008) Comparative genomics and molecular dynamics of DNA repeats in eukaryotes. Microbiol Mol Biol Rev 72(4):686–727
Rio D, Stewart AJ, Mullen W, Burns J, Lean MEJ, Brighenti F, Crozier A (2004) HPLC-MSn analysis of phenolic compounds and purine alkaloids in green and black tea. J Agric Food Chem 52(10):2807–2815
Safavi SA (2010) Isolation, identification and pathogenicity assessment of a new isolate of entomopathogenic fungus, Beauveria bassiana in Iran. J Plant Prot Res 50(2):158–163
Soltis PS, Soltis DE (2003) Applying the bootstrap in phylogeny reconstruction. Stat Sci 18(2):256–267
Suzuki S, Okuda T, Komatsubara S (1999) Selective isolation and distribution of Sporichthya strains in soil. Appl Environ Microbiol 65(5):1930–1935.
Tedersoo L, Anslan S, Bahram M, Põlme S, Riit T, Liiv I, Kõljalg U, Kisand V, Nilsson RH, Hildebrand F, Bork P, Abarenkov K (2015) Shotgun metagenomes and multiple primer pair-barcode combinations of amplicons reveal biases in metabarcoding analyses of fungi. MycoKeys 10:1–43
Ui WH, Won K (1999) General properties and phylogenetic utilities of nuclear ribosomal DNA and mitochondrial DNA commonly used in molecular systematic. Korean J Parasitol 37(4):215–228
Urtz BE, Rice WC (1997) RAPD-PCR characterization of Beauveria bassiana isolates from the rice water weevil Lissorhoptrus oryzophilus. Lett Appl Microbiol 25(6):405–409
Acknowledgements
The authors thank the University Grant Commission (Govt. of India) for supporting this work and Hungarian scholarship board to carry on further work.
Disclosure
All the experiments undertaken in this study comply with the current laws of the country where they were performed.
Author information
Authors and Affiliations
Contributions
The authors, SM and SB equally took part in all the experiments carried out for the present study at the Department of Chemical Engineering, Jadavpur University, Kolkata, India. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Mondal, S., Baksi, S. Strain-specific identification of Beauveria bassiana isolated from a novel habitat, using rDNA-based sequence analogy. Egypt J Biol Pest Control 28, 29 (2018). https://doi.org/10.1186/s41938-018-0033-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s41938-018-0033-4