Abstract
Genome-wide association studies of complex physiological traits and diseases consistently found that associated genetic factors, such as allelic polymorphisms or DNA mutations, only explained a minority of the expected heritable fraction. This discrepancy is known as “missing heritability”, and its underlying factors and molecular mechanisms are not established. Epigenetic programs may account for a significant fraction of the “missing heritability.” Epigenetic modifications, such as DNA methylation and chromatin assembly states, reflect the high plasticity of the genome and contribute to stably alter gene expression without modifying genomic DNA sequences. Consistent components of complex traits, such as those linked to human stature/height, fertility, and food metabolism or to hereditary defects, have been shown to respond to environmental or nutritional condition and to be epigenetically inherited. The knowledge acquired from epigenetic genome reprogramming during development, stem cell differentiation/de-differentiation, and model organisms is today shedding light on the mechanisms of (a) mitotic inheritance of epigenetic traits from cell to cell, (b) meiotic epigenetic inheritance from generation to generation, and (c) true transgenerational inheritance. Such mechanisms have been shown to include incomplete erasure of DNA methylation, parental effects, transmission of distinct RNA types (mRNA, non-coding RNA, miRNA, siRNA, piRNA), and persistence of subsets of histone marks.
Similar content being viewed by others
Introduction
Patterns of heritable traits within the human population determine body phenotypes, through a deeply intertwined interaction between genetic components and the environment. Specific genetic/DNA sequence variants are typically inherited transgenerationally as Mendelian alleles and are supposed to carry with them all the genetic information that acts as inheritable determinant [1]. Genome-wide association studies (GWAS) have recently demonstrated that multiple genomic loci are linked to complex traits, such as body development and height ([2, 3] and references therein). Several common disorders, such as type 2 diabetes, Crohn’s disease, and rheumatoid arthritis, were also shown to possess significant genetic components, as provided by multiple polymorphic loci [4–6]. These findings led to postulate models whereby numerous genetic factors provide small, independent contributions to complex phenotypes, such effects being essentially additive [5]. However, simple models of additive effects of ever-smaller components have remained as yet unproven [4–7]. On the other hand, identified genetic factors associated with complex diseases have been found to confer far less disease risk than expected from empirical estimates of heritability and typically explain only a minority of the heritable traits. As a consequence, “pure” genetic models are prone to underestimate the interactions among loci [5], globally designated as epistasis. Epistatic components need to be integrated by estimates of the contribution of non-genetic factors, globally designated as the “missing heritability” [7, 8]. Hence, the issue remained open, whereby identified genetic factors associated with complex diseases conferred far less disease risk than expected from empirical estimates of heritability. As an example, Crohn’s disease is a recessive disorder which shows about 80 % heritability. However, the genetic components identified to date only explain 20 % of this heritable fraction [9]. An additional example is that of human population stature [2, 3]. A significant fraction of height-determining genes has been identified by GWAS analysis [10–12]. Most of these genes were demonstrated to be largely overlapping in Caucasian and non-Caucasian populations [13, 14], consistent with an actual identification of the most relevant height-associated determinants. However, the identified polymorphisms were found to account only for a minor fraction of stature heritability. Although dedicated procedures for SNP-associated analyses have significantly increased their combined predictive power [15], a large amount of heritable height-associated factors remains undetectable by conventional GWAS, suggesting that such non-DNA sequence-linked information may be associated to epigenetic heredity.
Epigenetic heritability
Epigenetic modifications, such as DNA methylation, can contribute to alter gene expression in heritable manner without affecting the underlying genomic sequences. Such epigenetic contribution would be systematically missed by conventional DNA sequence-based analyses. A model of epigenetic inheritance, as additional to Mendelian heredity of polymorphic DNA sequences, would thus efficiently explain the lack of detection in conventional GWAS as “missing heritability”. It would also help explaining the cases of rapid, heritable adaptations to changing environmental conditions, such as for human stature [2, 3], and the occurrence of hereditary epistatic effects. Support for this model is provided by the evidence that phenotypic plasticity can emerge over rapid time scales, at rates that are orders of magnitude higher than the processes of natural selection [16, 17].
However, to be tenable, such a model of epigenetic inheritance poses rigorous requirements: (a) mitotic inheritance of epigenetic traits across cell generations (see discussion on DNA methylation maintenance through mitotic cycles); (b) epigenetic inheritance across successive meiotic divisions (see the paragraphs describing gamete generation and the development of primordial germ cells (PGC); and (c) true transgenerational inheritance, which requires proof of heritability beyond the first generation that has not been unexposed to the causal epigenetic modifiers (see the paragraphs describing transgenerational inheritance of DNA methylation and of chromatin states).
Evidence is now accumulating that provides insight in the mechanisms that underlie epigenetic transmission. These include the following: (1) DNA methylation, (2) histone modifications and chromatin remodeling, (3) inheritance of specific mRNAs, long non-coding RNAs (ncRNAs) and siRNAs/miRNAs, (4) feedback loops through which mRNA or protein products of a gene can stimulate its own transcription and enable “heritable states” of gene expression, and (5) the activity of chaperones such as Hsp90 that plays an important role in chromatin remodeling and can mediate epigenetic transgenerational variation. The most relevant mechanisms are described in Table 1.
DNA methylation
Approximately 60–80 % of the 28 million CpG dinucleotides in the human genome are methylated [18, 19]. DNA methyltransferases (DNMT) recognize hemimethylated sites (maintenance methylation) or specific unmethylated sequences (de novo methylation) (Fig. 1). Maintenance DNA methylation occurs during DNA replication and is predominantly dependent on DNMT1, whereas de novo DNA methylation is carried out by DNMT3A, DNMT3B [20], and DNMT3L [21, 22]. Both DNMT3A and DNMT3B localize to methylated, CpG-dense regions and preferentially bind to the bodies of transcribed genes but are excluded from regions of active promoters and enhancers [23].
DNA methylation/demethylation mechanics. a Methyl groups (green circles) are transferred to C in order to generate 5-mC. DNA methyltransferases (DNMT) catalyze this process. In the “active DNA demethylation” the TET DNA demethylase converts 5-mC to 5-hmC, which is further processed to 5-fC and 5-caC. These residues are targets for the DNA repair pathway, whose most critical component is the hTDG, which is responsible also for the repair of U:G and T:G mismatches. DNA demethylation can also occur through spontaneous cytosine deamination, which is catalyzed by AID/APOBEC enzymes. This gives rise to 5-hmU and T bases. Transient U:G and T:G mismatches can be repaired by the TDG/BER pathway. b De novo and maintenance methylation occur using unmethylated DNA and hemimethylated/post-replication DNA as templates for DNMT enzymes. In the absence of maintenance methylation, progressive dilution of 5-mC or its oxidized derivatives at DNA replication can determine the appearance of unmethylated DNA. This process is known as “passive DNA demethylation”
DNA methylation at CpG islands modulates gene transcription and is involved in alternative promoter usage and regulation of short and long ncRNA processing and of enhancer activity [24–26]. DNA methylation also affects determinants of higher-order DNA structure, e.g. in X chromosome inactivation, imprinting control regions (ICR) [27], heterochromatin folding and maintenance of genomic stability [28, 29].
DNA demethylation
The dynamic on/off switching of gene expression requires a balanced action of DNA methylation versus DNA demethylation. Both active and passive DNA demethylation have indeed been shown to occur.
Active DNA demethylation, i.e. replication-independent enzymatic removal of 5-methyl cytosine (5-mC), occurs through the processing of 5-mC to 5-hydroxymethyl cytosine (5-hmC) catalyzed by the ten-eleven translocation (TET) dioxygenases [30–32]. 5-hmC is then converted to 5-formylcytosine (5-fC) and 5-carboxylcytosine (5-caC) [30]. The thymine DNA glycosylase (TDG) efficiently excises both 5-fC and 5-caC [33, 34]. This leaves an abasic site that is subsequently processed through the base excision repair (BER) pathway [33] (Fig. 1).
Passive DNA demethylation can also proceed through the processing of 5-mC to 5-hmC [30–32] (Fig. 1). Neither maintenance methylase DNMT1 nor de novo methylases DNMT3A and DNMT3B recognize 5-hmC, and no mechanisms for 5-hmC maintenance have been as yet identified [32]. Hence, 5-hmC is inevitably lost at each replication cycle [35]. This appears to have a functional impact on PGC, which progressively lose 5-mC between embryonic day (E) E9.5 and E10.5, until 5-mC becomes undetectable by E11.5 [31]. This progressive loss of 5-mC occurs concurrently with an enrichment of 5-hmC, suggesting a genome-scale conversion of 5-mC to 5-hmC for the epigenomic reprogramming of these cells [31].
An additional path of passive DNA demethylation proceeds through destabilization of DNMT1, which is then followed by “loss by dilution” of 5-mC through successive replication cycles. Overexpression of SET7 leads to decreased DNMT1 levels via induction of proteasome-mediated degradation [36]. SET7 directly interacts with DNMT1 and specifically monomethylates Lys-142 of DNMT1 [36]. On the opposite side, AKT1 phosphorylates DNMT1 at Ser143 and stabilizes the protein [37]. Phosphorylation of DNMT1 at Ser143 interferes with monomethylation of the nearby Lys142 [37], making these two modifications mutually exclusive. Rb and ATM also affect the stability of DNMT1 [38]. The inactivation of pRB promotes a Tip60 (acetyltransferase)-dependent ATM activation. This allows activated ATM to physically bind to DNMT1, forming a complex with Tip60 and the E3 ligase ubiquitin-like containing PHD and RING finger domain protein 1 (Uhrf1) and accelerates the DNMT1 ubiquitination driven by Tip60-dependent acetylation [38, 39]. In contrast, histone deacetylase 1 (HDAC1) and the deubiquitinase HAUSP (herpes virus-associated ubiquitin-specific protease) stabilize DNMT1 [39].
Of note, 5-mC frequently undergoes deamination (Fig. 1). Hence, a DNA methylation-dependent modification can end up in a permanently fixed DNA sequence change. Physiological enzymes are involved, raising the intriguing issue of “guided” mutagenesis of Mendelian traits. The AID and APOBEC enzyme families catalyze the cytosine processing which leads to cytosine deamination. This occurs predominantly on 5-hmC and 5-mC residues, giving rise to formation of 5-hydromethyluracil (5-hmU) and thymine (T) bases, respectively [40] (Fig. 1). Consequently, transient U:G and T:G mismatches can be generated, though most of these mutations can be efficiently repaired by the TDG/BER pathway [40]. Notably, dysregulated APOBEC3B-catalyzed deamination can provide a chronic source of DNA damage, with consequent TP53 inactivation; this was shown to lead to development of breast cancer [41].
Reversible changes of epigenetic patterns
Epigenetic reprogramming through the mechanisms described above has been demonstrated in mammals over distinct, key developmental stages:
Erasure of DNA methylation patterns in the zygote
Erasure of DNA methylation patterns of the gametes (oocyte and sperm [42]) in the zygote was shown to occur immediately after fertilization. This process has been traditionally considered as a mechanism for resetting epigenetic marks between generations, to ensure the totipotency of the zygote after fertilization. Recent evidence from genome-scale DNA methylation analysis of human development confirmed a transient, highly dynamic state of global hypomethylation that affects most CpGs [43]. However, neither histone codes nor DNA methylation patterns are completely erased and are carried over through zygote divisions and generation of PGCs, thus, providing some of the means for transgenerational inheritance.
Erasure and reconstitution of DNA methylation patterns in PGCs
Epigenetic marks have been shown to undergo reprogramming across meiotic divisions of PGCs during gametogenesis. However, genome-wide DNA methylation profiling in PGCs revealed that, although the bulk of the genome becomes demethylated [44, 45], several loci escape this epigenetic erasure [46]. This leads to preserving the methylation status of more than 40 % of all 5-mC [16]. Substantial numbers of genes have been found to retain parental DNA methylation patterns in sperm and oocytes as a result of epigenetic transmission from PGCs [47]. It was recently reported that 5-hmC and 5-fC do exist in both maternal and paternal genomes and that 5-mC or its oxidized derivatives can be converted to unmodified cytosines through active demethylation rather than by passive dilution during embryonic development [48].
Erasure and reconstitution of epigenetic signatures during early body development
Reprogramming/re-establishment of epigenetic signatures was also shown to be necessary for proper development of a mature organism [49]. It has been shown that during early mitotic divisions of a mammalian embryo, daughter cells derived from the zygote have a globally hypomethylated genome [50] and a transcriptionally active chromatin due to histone H4 acetylation [51]. Therefore, they are an epigenetically homogeneous cell population [52, 53]. At the blastocyst stage, peripheral cells (that will become the extraembryonic tissue) have low levels of DNA methylation and are epigenetically different from cells of the inner cell mass (ICM, which will form the embryo) that have already undergone re-establishment of some methylation patterns [52, 53]. A major epigenetic switch then occurs during implantation at the transition from the blastocyst to the post-implantation epiblast [47].
Erasure and reconstitution of DNA methylation patterns in adult stem cells
Somatic cell nuclear transfer has been utilized as one of the procedures to obtain reprogramming of somatic cells toward a totipotent state, with the generation of induced pluripotent stem cells [54, 55]. This, and the possibility to isolate embryonic stem (ES) cells from the blastocyst ICM, provided unprecedented opportunities to investigate the mechanics of erasure and re-establishment of epigenetic patterns and to define the molecular components involved in these processes. Declining levels of TETs during differentiation were shown to be associated with decreased hydroxymethylation levels at the promoters of ES cell-specific genes [32]. Thus, the balance between hydroxymethylation and methylation in the genome appears to be linked with the balance between pluripotency and lineage commitment [32]. Moreover, reprogramming-resistant regions strongly enriched for H3K9me3 were demonstrated to be critical barriers for efficient reprogramming [56]. Hence, modulation of epigenetic inheritance appears to play a key role in stem cell differentiation/de-differentiation.
Gamete-carried epigenetic traits
Nucleosomes are largely replaced by protamines in mature human sperm, thus erasing most chromatin patterns. However, not all histones in sperm are replaced by protamines, and epigenetic marks such as H3K4me2 and H3K27me3 have been detected in sperm [57, 58]. These retained nucleosomes are significantly enriched at loci of developmental importance, including imprinted gene clusters, microRNA clusters, HOX gene clusters, and the promoters of stand-alone developmental transcription and signalling factors [57]. H3K4me2 was found to be enriched at promoters of genes coding for developmental transcription factors, whereas H3K4me3 was found predominantly localized at promoters of genes important for spermatogenesis and rearrangement of nuclear architecture and presumably active during the gametogenesis [57].
Our findings indicated that sperm methylation pattern of the CD5/Leu1 and CD8/Leu2 genes is incompletely erased. This had a heavy impact on gene function and tightly prevented the expression of the CD5 gene, though not of CD8 [59]. Genes encoding olfactory receptors, in cases where mice associated specific odors with fearful experience, were also found differentially methylated in sperm, and this methylation pattern was transmitted to F1 and F2 generations [60].
RNA molecules packaged in sperm represent an additional contributor to transgenerational transmission of epigenetic traits and have been shown to profoundly affect offspring phenotypes [61, 62]. Injecting sperm RNA from traumatized males into fertilized wild type oocytes reproduced the altered behavior in the offspring. Moreover, miRNA-mediated signals can change DNA methylation patterns in the F2 sperm, and this signature can be maintained and replicated through subsequent mitotic and meiotic cycles [63].
Notably, RNA expressed in somatic cells can be transferred to gametes via extracellular vesicles [64]. Subcutaneous injection of human melanoma cells stably expressing enhanced green fluorescent protein (EGFP) led to the transfer of EGFP mRNA in murine sperm heads, likely through exosomes-mediated transport [65]. Furthermore, in C. elegans, it has recently been shown that neurons can transmit double-stranded RNA (dsRNA) to the germ cells to initiate transgenerational silencing of their target genes [16, 66]. Thus, extracellular vesicles have been revealed as an important route for transgenerational inheritance of epigenetic signatures.
Transgenerational inheritance of DNA methylation patterns
Transgenerational inheritance of epigenetic patterns [16, 66] is key to a model of “epigenetic missing heredity”. In this regard, it should be noted that purely parental effects, such as the impact of direct in utero exposure to particular nutritional, hormonal, or stress/toxin environments, do not represent true transgenerational heredity [67]. In addition, F1 gametes are potentially exposed in utero to maternal experiences, and this may subsequently affect F2 offsprings. Hence, proof of transgenerational transmission of ancestral memory requires demonstrating the passing of the epigenetic trait through unexposed F2 gametes to F3 offsprings [16].
Physiological traits, such as body stature, were shown to rapidly and progressively adaptate to changing environments, e.g., nutritional status [2]. The epigenetic inactivation of height-associated genes (e.g., BMP2, BMP6, CABLES1, DLEU7, GNAS, GNASAS, HHIP, MOS, PLAGL1) was shown to be functionally equivalent to Mendelian physical loss of the corresponding alleles. Moreover, distinct epigenetic defects, such as in Beckwith–Wiedemann, Prader–Willi, Angelman’s, Rett, and Silver–Russell syndromes ([2] and references therein), were correspondingly shown to cause hereditary growth anomalies, indicating that body stature is under a heritable epigenetic control.
Correspondingly, the appearance of several pathological conditions with heritable components, such as diabetes and Crohn’s disease, is affected by interaction with dynamic environmental factors, such as host pathogens or nutritional status [68]. Exposure of parents to distinct diet regimens, stress, drugs, or endocrine/metabolic dysfunctions, was shown to additionally affect the transgenerational transmission of altered DNA methylation patterns [69–75]. As an example, F2 generation offspring (i.e., the grandchildren) of alcohol-abusing women have a higher tendency to show fetal alcohol syndrome than the F2 progeny of control women [76, 77].
Epigenetic modifications were also shown to be caused by exposure to environmental toxins, including metals (cadmium, arsenic, nickel, chromium, and methylmercury), solvents (trichloroethylene), air pollutants (black carbon, benzene), food-chain contaminants (dioxins), and tobacco smoke (nicotine, benzo(a)pyrene) [78, 79]. In utero and neonatal exposure to low doses of Bisphenol A and/or phthalates causes epigenetic alterations [71], such as differential methylation at CpG islands, histone modifications, and altered expression of ncRNAs and miRNAs [80, 81]. DNA methylation was shown to be affected by periconceptional maternal plasma concentrations of micronutrients involved in one-carbon metabolism, such as folate, B2 vitamin, methionine, and betaine [70]. The first evidence in mammals for a true transgenerational transmission of exposure-determined epigenetic traits was obtained in rats exposed to endocrine disruptors vinclozolin or methoxychlor during gestation. This induced in the F1 generation an adult phenotype of decreased spermatogenic capacity and of increased incidence of male infertility. Remarkably, transgenerational transmission of these effects through the male germ line was then observed in F1 to F4 generations [82]. DNMT3L has been reported to be necessary for maternal methylation imprinting [22]. DNMT3L is enzymatically inactive but acts as a stimulatory factor for de novo methylation by DNMT3A [21]. DNA methylation signatures driving altered behavioral/metabolic phenotypes (from exposure to maternal stress) were subsequently shown to be transgenerationally transmitted to the offspring [83], through miRNA delivery by sperm cells to the oocyte [63, 83].
Transgenerational inheritance of chromatin states
Distinct types of histone post-translational modifications (PTMs) play a critical role in the nucleosome-dependent regulation of gene transcription. The largest body of knowledge has been gathered on histone methylation and acetylation [84], which cooperates in re-shaping chromatin organization. This histone code interacts closely with DNA methylation: unmethylated Lys-4 on H3 histone (H3K4) acts as a docking site for DNMT3A, which is recruited on nucleosomes and methylates associated target nucleotides [21, 22]. Recent findings showed that in the absence of histone H3, DNMT3A exists in an autoinhibitory form, in which the ATRX–DNMT3–DNMT3L (ADD) domain binds to the catalytic domain and hinders its DNA-binding capacity [85]. Once the DNMT3A–DNMT3L complex is recruited to the nucleosome, unmethylated H3K4 binds to the ADD domain and stimulates DNMT3A to undergo a significant conformational change from an autoinhibitory form to an active form that can bind DNA and exert DNA methylation activity [85]. Moreover, interaction of Dppa3 with histone H3K9me2 blocks the activity of TET3, favoring the maintenance of DNA methylation [86]. Less common histone PTMs have been recently identified [87, 88]. Among them, histone H1 arginine 54 citrullination (H1R54ci) determines histone displacement from chromatin and chromatin decondensation. Histone PTMs were further shown to be a critical mechanism for maintaining stem cell pluripotency [89].
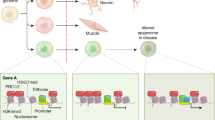
Histone PTMs and corresponding DNA methylation patterns can affect imprinting in mammalian cells [90, 91] through the selective recruitment of effector proteins, known as “readers” (e.g. the bromodomain motif that docks onto acetylated lysines [92]), which drive chromatin packaging around nucleosomes [88, 91] (Fig. 2).
Histone modifications and DNA cooperate in re-shaping chromatin organization and regulating gene expression. a (left) De novo DNA methylation occurs on unmethylated DNA. It is catalyzed by the DNMT3, whose subunits can be positioned in proximity of their target sites through physical interaction with unmethylated H3K4. a (right) Maintenance DNA methylation occurs during the DNA replication and is catalyzed by the DNMT1. Uhrf1 and proliferating cell nuclear antigen (PCNA) associates to DNMT1 and recruit it to the replication fork, concentrating its activity on hemimethylated DNA. The Uhrf1 TTD domain interacts with H3K9me. This binding allows a faithful propagation of DNA methylation patterns throughout mitosis. b (top) HDAC and the transcription factor complex (TFC) can be recruited on sensitive promoters, leading to histone deacetylation. HMT-driven methylation of the histone tails causes tight wrapping of DNA around nucleosome cores and inhibition of gene expression. b (bottom) The accumulation of HAT-driven histone acetylation determines DNA relaxation around the nucleosomes surrounding HAT-sensitive promoters; this leads to increased transcription and gene expression. c Methylation of H3K9 plays a central role in non-DNA-dependent mechanisms of regulation of gene activity (top). Hsp90 has a strong effect on the histone code via stabilization of KDM4B, which demethylases H3K9 (middle). Non-coding RNAs alter the histone code through siRNA-dependent mechanisms that lead to direct competition between BORDERLINE ncRNAs and H3K9me for binding to the HP1 proteins, such as Swi6. This occurs at heterochromatin/euchromatin boundary sites and counteracts the spreading of heterochromatin into neighboring euchromatin (bottom). A acetyl groups, M methyl groups, HAT histone acetyltranferase, HDAC histone deacetylase, HMT histone methyltransferase
Histone PTM-driven heritable silencing of gene expression is also affected by various chromatin remodeling factors. Among them, the polycomb-group (PcG) proteins recognize specific histone modifications such as the H3K27me3 and participate in maintenance of repressed chromatin domains [93]. PcG proteins take part to the regulation of X-chromosome inactivation and maintenance of stem cell identity [94]. Correspondingly, the molecular chaperone Hsp90 was shown to alter the histone code via interaction and stabilization of KDM4B, which demethylates H3K9. Pharmacological inhibition of Hsp90 results in ubiquitin-dependent proteasomal degradation of KDM4B, which is accompanied by increased methylation of H3K9 [95] (Fig. 2).
Although the mechanisms through which DNA methylation states can propagate across cell divisions have been studied in depth, it is still unclear how the histone code (e.g., the histone PTM levels) is restored through multiple rounds of DNA replication. During DNA replication, nucleosomes are disrupted ahead of the replication machinery and reassembled on the two newly synthesized DNA strands. Histones PTMs are transmitted with high efficiency at replication forks, indicating a specific recycling of old histones. However, incorporation of new, largely unmodified histones also occurs [96–98]. Reincorporated parental, modified histones may serve as a blueprint to modify neighboring new histones, limiting the possible “dilution” of the corresponding code. Recent findings suggested that most PTM levels are maintained according to the simple paradigm that new histones acquire modifications to become identical to the old ones [99]. These and other findings also demonstrated that H3K9me3 and H3K27me3 are central marks for cellular memory [97, 100], and are propagated by continuous modification of both new and old histones through the generations. Epigenetic heritability of H3K9 methylation was recently investigated in fission yeast and demonstrated to involve the activity of a single H3K9 methyltransferase, Clr4, that directs all H3K9 methylation and heterochromatin through a “read-write” mechanism [101, 102]. Hence, histones act as carriers of epigenetic information, and the kinetics of PTM restoration appears to play a critical role in epigenetic inheritance [90, 98].
Epigenetic inheritance in model organisms
Non-DNA methylation-mediated epigenetic heritability has been demonstrated also in non-mammalian species, such as C. elegans [1], D. melanogaster [103], and S. pombe [104].
C. elegans lacks DNA methylation and has evolved histone methylation/demethylation pathways to regulate the transmission of epigenetic information through multiple generations [1, 105]. In C. elegans, transgenerational inheritance has been shown to be mediated by transmission of piRNAs. piRNAs induce a highly-stable, long-term gene silencing, which persists at least through 20 generations. The inheritance of the phenotype then becomes independent of the original piRNA, as is taken over by siRNAs [106]. These siRNAs act by modulating transcriptional gene silencing of histone methyltransferases, with consequent rearrangement of the chromatin structure [106]. Such heterochromatin-like configuration is required for stable silencing [16].
D. melanogaster shows “paramutations,” a form of epigenetic inheritance whereby one allele at a gene locus is capable of inducing a structural modification in the paired allele, in the absence of DNA sequence changes, which is then inherited through meiotic divisions [103]. The paramutated allele itself becomes paramutagenic and is capable to epigenetically convert a new paramutable allele. The paramutation has been shown to occur without any chromosome pairing between the paramutagenic and the paramutated loci and is mediated by maternal inheritance of piRNAs.
In the yeast S. pombe, transgenerational inheritance has been demonstrated to be mediated by multiple long ncRNAs termed BORDERLINEs, which act in a sequence-independent but locus-dependent manner [104] (Fig. 2). BORDERLINE ncRNAs are processed by Dicer into short RNAs referred to as brdrRNAs [104]. brdrRNAs then compete with H3K9me for binding to the HP1 protein Swi6. This prevents spreading of the HP1 protein Swi6 and histone H3K9 methylation beyond pericentromeric repeat regions and leads to Swi6 removal from chromatin, which counteracts the spreading of heterochromatin into neighboring euchromatin by preventing the spreading of H3K9me [104].
Phenotypic impact of epigenetic heredity
Epigenetic alterations can have strong impact on hereditary disease phenotypes. Altered balance of epigenetic networks has been reported to cause major pathologies, including complex phenotype syndromes and cancer (Tables 2 and 3).
Most hereditary diseases linked to defect of epigenetic control present with multi-organ abnormalities and overall developmental defects. Several of these syndromes are characterized by mental retardation and other central nervous system defects. Bone and cartilage growth abnormalities are also frequent, consistent with a key role of epigenetic regulation in body development [2, 3].
Notably, several epigenetic, hereditary syndromes are also characterized by gross chromosomal anomalies. This is consistent with the role that epigenetic mechanisms that play in the regulation of chromosome architecture and maintenance of genomic stability [28, 29], as well as in modulation of regulatory networks that involve p53.
Epigenetic changes can also have a major role in the development of cancer [107]. Most studied examples include patients with sporadic colorectal cancer with a microsatellite instability phenotype that shows methylation and silencing of the gene encoding MLH1 (Table 3), indicating that epigenetic silencing can result directly in genomic instability in transformed cells [108, 109]. However, epigenetic regulation of key oncogenes/tumor suppressor genes appears much more widespread than commonly appreciated, as major targets include cyclins, cyclin inhibitors, APC, BRCA1, retinoic acid receptors, protease modulators, IGF2, and transcription factors associated with epithelial-mesenchymal transition, e.g. Twist.
Conclusions
The genomes of eukaryotic organisms are adaptable to non-genetic/environmental-driven changes through epigenetic modulation of gene expression across generation cycles (Fig. 3). Epigenetic modifications include DNA methylation and histone modifications. These have the ability to alter gene expression patterns without affecting the nucleotide sequence of the underlying genome, the only exception being deamination of 5-mC to thymine. Tight regulation of the activity of DNA methyltransferases as well as of demethylases plays a mechanistic role in the establishment, maintenance and transient erasure of DNA methylation patterns. Correspondingly, post-translational modifications of histones and of regulatory proteins were shown to play a role in hereditary transmission of chromatin composition and configuration states, both in mammals and in model organisms. Thus, epigenetic programs contribute to the transgenerational inheritance of complex traits, which may help accounting for the “missing heritability” in current GWAS studies.
Epigenetic factors influencing human development and growth. The human life cycle is represented in the scheme. Major factors influencing the epigenetic programs and the maintenance of epigenetic patterns at both DNA and chromatin (histone code) levels are the maternal lifestyle during pregnancy and the personal exposure to harmful environments during post-natal growth and adult life
Abbreviations
- 5-caC:
-
5-carboxylcytosine
- 5-fC:
-
5-formylcytosine
- 5-hmC:
-
5-hydroxymethyl cytosine
- 5-hmU:
-
5-hydromethyluracil
- 5-mC:
-
5-methyl cytosine
- BER:
-
base excision repair
- DNMT:
-
DNA methyltransferase
- dsRNA:
-
double-stranded RNA
- EGFP:
-
enhanced green fluorescent protein
- ES:
-
embryonic stem
- GWAS:
-
genome-wide association studies
- HDAC:
-
histone deacetylase
- ICM:
-
inner cell mass
- ICR:
-
imprinting control regions
- ncRNA:
-
non-coding RNA
- PcG:
-
polycomb-group
- PGC:
-
primordial germ cells
- PTMs:
-
post-translational modifications
- T:
-
thymine
- TDG:
-
thymine DNA glycosylase
- TET:
-
ten-eleven translocation
References
Lim JP, Brunet A. Bridging the transgenerational gap with epigenetic memory. Trends Genet. 2013;29:176–86.
Tripaldi R, Stuppia L, Alberti S. Human height genes and cancer. BBA Reviews Cancer. 1836;2013:27–41.
Simeone P, Alberti S. Epigenetic heredity of human height. Physiological reports. 2014;2:e12047.
Marian AJ. Elements of “missing heritability”. Curr Opin Cardiol. 2012;27:197–201.
Zuk O, Hechter E, Sunyaev SR, Lander ES. The mystery of missing heritability: Genetic interactions create phantom heritability. Proc Natl Acad Sci U S A. 2012;109:1193–8.
Zuk O, Schaffner SF, Samocha K, Do R, Hechter E, Kathiresan S, et al. Searching for missing heritability: designing rare variant association studies. Proc Natl Acad Sci U S A. 2014;111:E455–464.
Koch L. Epigenetics: an epigenetic twist on the missing heritability of complex traits. Nat Rev Genet. 2014;15:218.
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, et al. Finding the missing heritability of complex diseases. Nature. 2009;461:747–53.
Park JH, Wacholder S, Gail MH, Peters U, Jacobs KB, Chanock SJ, et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nat Genet. 2010;42:570–5.
Gudbjartsson DF, Walters GB, Thorleifsson G, Stefansson H, Halldorsson BV, Zusmanovich P, et al. Many sequence variants affecting diversity of adult human height. Nat Genet. 2008;40:609–15.
Lettre G, Jackson AU, Gieger C, Schumacher FR, Berndt SI, Sanna S, et al. Identification of ten loci associated with height highlights new biological pathways in human growth. Nat Genet. 2008;40:584–91.
Weedon MN, Lango H, Lindgren CM, Wallace C, Evans DM, Mangino M, et al. Genome-wide association analysis identifies 20 loci that influence adult height. Nat Genet. 2008;40:575–83.
Cho YS, Go MJ, Kim YJ, Heo JY, Oh JH, Ban HJ, et al. A large-scale genome-wide association study of Asian populations uncovers genetic factors influencing eight quantitative traits. Nat Genet. 2009;41:527–34.
Okada Y, Kamatani Y, Takahashi A, Matsuda K, Hosono N, Ohmiya H, et al. A genome-wide association study in 19 633 Japanese subjects identified LHX3-QSOX2 and IGF1 as adult height loci. Hum Mol Genet. 2010;19:2303–12.
Yang J, Benyamin B, McEvoy BP, Gordon S, Henders AK, Nyholt DR, et al. Common SNPs explain a large proportion of the heritability for human height. Nat Genet. 2010;42:565–9.
Szyf M. Nongenetic inheritance and transgenerational epigenetics. Trends Mol Med. 2015;21:134–44.
Laforsch C, Tollrian R. Embryological aspects of inducible morphological defenses in Daphnia. Journal of Morphology. 2004;262:701–7.
Ziller MJ, Gu H, Muller F, Donaghey J, Tsai LT, Kohlbacher O, et al. Charting a dynamic DNA methylation landscape of the human genome. Nature. 2013;500:477–81.
Rivera CM, Ren B. Mapping human epigenomes. Cell. 2013;155:39–55.
Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99:247–57.
Chen ZX, Mann JR, Hsieh CL, Riggs AD, Chedin F. Physical and functional interactions between the human DNMT3L protein and members of the de novo methyltransferase family. J Cell Biochem. 2005;95:902–17.
Suetake I, Shinozaki F, Miyagawa J, Takeshima H, Tajima S. DNMT3L stimulates the DNA methylation activity of Dnmt3a and Dnmt3b through a direct interaction. J Biol Chem. 2004;279:27816–23.
Baubec T, Colombo DF, Wirbelauer C, Schmidt J, Burger L, Krebs AR, et al. Genomic profiling of DNA methyltransferases reveals a role for DNMT3B in genic methylation. Nature. 2015;520(7546):243–7.
Kulis M, Queiros AC, Beekman R, Martin-Subero JI. Intragenic DNA methylation in transcriptional regulation, normal differentiation and cancer. Biochim Biophys Acta. 1829;2013:1161–74.
Lujambio A, Portela A, Liz J, Melo SA, Rossi S, Spizzo R, et al. CpG island hypermethylation-associated silencing of non-coding RNAs transcribed from ultraconserved regions in human cancer. Oncogene. 2010;29(48):6390–401.
Ehrlich M, Lacey M. DNA methylation and differentiation: silencing, upregulation and modulation of gene expression. Epigenomics. 2013;5:553–68.
Henckel A, Nakabayashi K, Sanz LA, Feil R, Hata K, Arnaud P. Histone methylation is mechanistically linked to DNA methylation at imprinting control regions in mammals. Hum Mol Genet. 2009;18(18):3375–83.
Alberti S, Nutini M, Herzenberg LA. DNA methylation prevents the amplification of TROP1, a tumor associated cell surface antigen gene. Proc Natl Acad Sci USA. 1994;91:5833–7.
Nasr AF, Nutini M, Palombo B, Guerra E, Alberti S. Mutations of TP53 induce loss of DNA methylation and amplification of the TROP1 gene. Oncogene. 2003;22:1668–77.
Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009;324:930–5.
Hackett JA, Sengupta R, Zylicz JJ, Murakami K, Lee C, Down TA, et al. Germline DNA demethylation dynamics and imprint erasure through 5-hydroxymethylcytosine. Science. 2013;339:448–52.
Ficz G, Branco MR, Seisenberger S, Santos F, Krueger F, Hore TA, et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature. 2011;473:398–402.
He YF, Li BZ, Li Z, Liu P, Wang Y, Tang Q, et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science. 2011;333:1303–7.
Song CX, Szulwach KE, Dai Q, Fu Y, Mao SQ, Lin L, et al. Genome-wide profiling of 5-formylcytosine reveals its roles in epigenetic priming. Cell. 2013;153:678–91.
Sarkar DK. Male germline transmits fetal alcohol epigenetic marks for multiple generations: a review. Addiction Biology. 2015. doi:10.1111/adb.12186.
Esteve PO, Chin HG, Benner J, Feehery GR, Samaranayake M, Horwitz GA, et al. Regulation of DNMT1 stability through SET7-mediated lysine methylation in mammalian cells. Proc Natl Acad Sci U S A. 2009;106:5076–81.
Esteve PO, Chang Y, Samaranayake M, Upadhyay AK, Horton JR, Feehery GR, et al. A methylation and phosphorylation switch between an adjacent lysine and serine determines human DNMT1 stability. Nat Struct Mol Biol. 2011;18:42–8.
Shamma A, Suzuki M, Hayashi N, Kobayashi M, Sasaki N, Nishiuchi T, et al. ATM mediates pRB function to control DNMT1 protein stability and DNA methylation. Mol Cell Biol. 2013;33:3113–24.
Du Z, Song J, Wang Y, Zhao Y, Guda K, Yang S, et al. DNMT1 stability is regulated by proteins coordinating deubiquitination and acetylation-driven ubiquitination. Sci Signal. 2010;3:ra80.
Pastor WA, Aravind L, Rao A. TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nat Rev Mol Cell Biol. 2013;14:341–56.
Burns MB, Lackey L, Carpenter MA, Rathore A, Land AM, Leonard B, et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature. 2013;494:366–70.
Gannon JR, Emery BR, Jenkins TG, Carrell DT. The sperm epigenome: implications for the embryo. Adv Exp Med Biol. 2014;791:53–66.
Smith ZD, Chan MM, Humm KC, Karnik R, Mekhoubad S, Regev A, et al. DNA methylation dynamics of the human preimplantation embryo. Nature. 2014;511:611–5.
Guo F, Yan L, Guo H, Li L, Hu B, Zhao Y, et al. The transcriptome and DNA methylome landscapes of human primordial germ cells. Cell. 2015;161:1437–52.
Tang WW, Dietmann S, Irie N, Leitch HG, Floros VI, Bradshaw CR, et al. A unique gene regulatory network resets the human germline epigenome for development. Cell. 2015;161:1453–67.
Gkountela S, Zhang KX, Shafiq TA, Liao WW, Hargan-Calvopina J, Chen PY, et al. DNA demethylation dynamics in the human prenatal germline. Cell. 2015;161:1425–36.
Borgel J, Guibert S, Li Y, Chiba H, Schubeler D, Sasaki H, et al. Targets and dynamics of promoter DNA methylation during early mouse development. Nat Genet. 2010;42:1093–100.
Wang L, Zhang J, Duan J, Gao X, Zhu W, Lu X, et al. Programming and inheritance of parental DNA methylomes in mammals. Cell. 2014;157:979–91.
Seisenberger S, Peat JR, Hore TA, Santos F, Dean W, Reik W. Reprogramming DNA methylation in the mammalian life cycle: building and breaking epigenetic barriers. Philos Trans R Soc Lond B Biol Sci. 2013;368:20110330.
Meissner A, Mikkelsen TS, Gu H, Wernig M, Hanna J, Sivachenko A, et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature. 2008;454:766–70.
Reik W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature. 2007;447:425–32.
Santos F, Hendrich B, Reik W, Dean W. Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev Biol. 2002;241:172–82.
Santos J, Pereira CF, Di-Gregorio A, Spruce T, Alder O, Rodriguez T, et al. Differences in the epigenetic and reprogramming properties of pluripotent and extra-embryonic stem cells implicate chromatin remodelling as an important early event in the developing mouse embryo. Epigenetics Chromatin. 2010;3:1.
Krishnakumar R, Blelloch RH. Epigenetics of cellular reprogramming. Curr Opin Genet Dev. 2013;23:548–55.
Papp B, Plath K. Epigenetics of reprogramming to induced pluripotency. Cell. 2013;152:1324–43.
Matoba S, Liu Y, Lu F, Iwabuchi KA, Shen L, Inoue A, et al. Embryonic development following somatic cell nuclear transfer impeded by persisting histone methylation. Cell. 2014;159:884–95.
Hammoud SS, Nix DA, Zhang H, Purwar J, Carrell DT, Cairns BR. Distinctive chromatin in human sperm packages genes for embryo development. Nature. 2009;460(7254):473–8.
Brykczynska U, Hisano M, Erkek S, Ramos L, Oakeley EJ, Roloff TC, et al. Repressive and active histone methylation mark distinct promoters in human and mouse spermatozoa. Nat Struct Mol Biol. 2010;17:679–87.
Alberti S, Herzenberg LA. DNA methylation prevents transfection of genes for specific surface antigens. Proc Natl Acad Sci USA. 1988;85:8391–4.
Dias BG, Ressler KJ. Parental olfactory experience influences behavior and neural structure in subsequent generations. Nat Neurosci. 2014;17:89–96.
Carone BR, Fauquier L, Habib N, Shea JM, Hart CE, Li R, et al. Paternally induced transgenerational environmental reprogramming of metabolic gene expression in mammals. Cell. 2010;143:1084–96.
Rassoulzadegan M, Grandjean V, Gounon P, Vincent S, Gillot I, Cuzin F. RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature. 2006;441:469–74.
Liebers R, Rassoulzadegan M, Lyko F. Epigenetic regulation by heritable RNA. PLoS Genet. 2014;10:e1004296.
Johnson GD, Mackie P, Jodar M, Moskovtsev S, Krawetz SA. Chromatin and extracellular vesicle associated sperm RNAs. Nucleic Acids Res. 2015 [Epub ahead of print].
Cossetti C, Lugini L, Astrologo L, Saggio I, Fais S, Spadafora C. Soma-to-germline transmission of RNA in mice xenografted with human tumour cells: possible transport by exosomes. PLoS One. 2014;9:e101629.
Sharma A. Transgenerational epigenetic inheritance requires a much deeper analysis. Trends Mol Med. 2015;21:269–70.
Heard E, Martienssen RA. Transgenerational epigenetic inheritance: myths and mechanisms. Cell. 2014;157:95–109.
Silventoinen K, Kaprio J, Lahelma E, Koskenvuo M. Relative effect of genetic and environmental factors on body height: differences across birth cohorts among Finnish men and women. Am J Public Health. 2000;90:627–30.
Waterland RA, Kellermayer R, Laritsky E, Rayco-Solon P, Harris RA, Travisano M, et al. Season of conception in rural gambia affects DNA methylation at putative human metastable epialleles. PLoS Genet. 2010;6:e1001252.
Dominguez-Salas P, Moore SE, Baker MS, Bergen AW, Cox SE, Dyer RA, et al. Maternal nutrition at conception modulates DNA methylation of human metastable epialleles. Nat Commun. 2014;5:3746.
Dolinoy DC, Huang D, Jirtle RL. Maternal nutrient supplementation counteracts bisphenol A-induced DNA hypomethylation in early development. Proc Natl Acad Sci U S A. 2007;104:13056–61.
Davis AP, Murphy CG, Saraceni-Richards CA, Rosenstein MC, Wiegers TC, Mattingly CJ. Comparative Toxicogenomics Database: a knowledgebase and discovery tool for chemical-gene-disease networks. Nucleic Acids Res. 2009;37:D786–792.
Singh S, Li SS. Phthalates: toxicogenomics and inferred human diseases. Genomics. 2011;97:148–57.
Singh S, Li SS. Bisphenol A and phthalates exhibit similar toxicogenomics and health effects. Gene. 2012;494:85–91.
Berdasco M, Esteller M. Aberrant epigenetic landscape in cancer: how cellular identity goes awry. Dev Cell. 2010;19:698–711.
Kvigne VL, Leonardson GR, Borzelleca J, Brock E, Neff-Smith M, Welty TK. Alcohol use, injuries, and prenatal visits during three successive pregnancies among American Indian women on the Northern Plains who have children with fetal alcohol syndrome or incomplete fetal alcohol syndrome. Maternal and Child Health Journal. 2008;12 Suppl 1:37–45.
Kvigne VL, Leonardson GR, Borzelleca J, Welty TK. Characteristics of grandmothers who have grandchildren with fetal alcohol syndrome or incomplete fetal alcohol syndrome. Maternal and Child Health Journal. 2008;12:760–5.
Baccarelli A, Bollati V. Epigenetics and environmental chemicals. Current Opinion in Pediatrics. 2009;21:243–51.
Maccani MA, Avissar-Whiting M, Banister CE, McGonnigal B, Padbury JF, Marsit CJ. Maternal cigarette smoking during pregnancy is associated with downregulation of miR-16, miR-21, and miR-146a in the placenta. Epigenetics. 2010;5:583–9.
Avissar-Whiting M, Veiga KR, Uhl KM, Maccani MA, Gagne LA, Moen EL, et al. Bisphenol A exposure leads to specific microRNA alterations in placental cells. Reprod Toxicol. 2010;29:401–6.
Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358:1148–59.
Anway MD, Cupp AS, Uzumcu M, Skinner MK. Epigenetic transgenerational actions of endocrine disruptors and male fertility. Science. 2005;308:1466–9.
Gapp K, Jawaid A, Sarkies P, Bohacek J, Pelczar P, Prados J, et al. Implication of sperm RNAs in transgenerational inheritance of the effects of early trauma in mice. Nat Neurosci. 2014;17:667–9.
Rose NR, Klose RJ. Understanding the relationship between DNA methylation and histone lysine methylation. Biochim Biophys Acta. 2014;1839(12):1362–72.
Guo X, Wang L, Li J, Ding Z, Xiao J, Yin X, et al. Structural insight into autoinhibition and histone H3-induced activation of DNMT3A. Nature. 2014;517(7536):640–4.
Nakamura T, Liu YJ, Nakashima H, Umehara H, Inoue K, Matoba S, et al. PGC7 binds histone H3K9me2 to protect against conversion of 5mC to 5hmC in early embryos. Nature. 2012;486:415–9.
Arnaudo AM, Garcia BA. Proteomic characterization of novel histone post-translational modifications. Epigenetics Chromatin. 2013;6:24.
Rothbart SB, Strahl BD. Interpreting the language of histone and DNA modifications. Biochim Biophys Acta. 1839;2014:627–43.
Christophorou MA, Castelo-Branco G, Halley-Stott RP, Oliveira CS, Loos R, Radzisheuskaya A, et al. Citrullination regulates pluripotency and histone H1 binding to chromatin. Nature. 2014;507:104–8.
Campos EI, Stafford JM, Reinberg D. Epigenetic inheritance: histone bookmarks across generations. Trends Cell Biol. 2014;24(11):664–74.
Strahl BD, Allis CD. The language of covalent histone modifications. Nature. 2000;403:41–5.
Dhalluin C, Carlson JE, Zeng L, He C, Aggarwal AK, Zhou MM. Structure and ligand of a histone acetyltransferase bromodomain. Nature. 1999;399:491–6.
Fouse SD, Shen Y, Pellegrini M, Cole S, Meissner A, Van Neste L, et al. Promoter CpG methylation contributes to ES cell gene regulation in parallel with Oct4/Nanog, PcG complex, and histone H3 K4/K27 trimethylation. Cell Stem Cell. 2008;2:160–9.
Yu Q. Cancer gene silencing without DNA hypermethylation. Epigenetics. 2008;3:315–7.
Ipenberg I, Guttmann-Raviv N, Khoury HP, Kupershmit I, Ayoub N. Heat shock protein 90 (Hsp90) selectively regulates the stability of KDM4B/JMJD2B histone demethylase. J Biol Chem. 2013;288:14681–7.
Annunziato AT. Split decision: what happens to nucleosomes during DNA replication? J Biol Chem. 2005;280:12065–8.
Probst AV, Dunleavy E, Almouzni G. Epigenetic inheritance during the cell cycle. Nat Rev Mol Cell Biol. 2009;10:192–206.
Alabert C, Groth A. Chromatin replication and epigenome maintenance. Nat Rev Mol Cell Biol. 2012;13:153–67.
Alabert C, Barth TK, Reveron-Gomez N, Sidoli S, Schmidt A, Jensen ON, et al. Two distinct modes for propagation of histone PTMs across the cell cycle. Genes Dev. 2015;29:585–90.
Apostolou E, Hochedlinger K. Chromatin dynamics during cellular reprogramming. Nature. 2013;502:462–71.
Ragunathan K, Jih G, Moazed D. Epigenetics. Epigenetic inheritance uncoupled from sequence-specific recruitment Science. 2015;348:1258699.
Audergon PN, Catania S, Kagansky A, Tong P, Shukla M, Pidoux AL, et al. Epigenetics. Restricted epigenetic inheritance of H3K9 methylation. Science. 2015;348:132–5.
de Vanssay A, Bouge AL, Boivin A, Hermant C, Teysset L, Delmarre V, et al. Paramutation in Drosophila linked to emergence of a piRNA-producing locus. Nature. 2012;490:112–5.
Keller C, Kulasegaran-Shylini R, Shimada Y, Hotz HR, Buhler M. Noncoding RNAs prevent spreading of a repressive histone mark. Nat Struct Mol Biol. 2013;20:994–1000.
Greer EL, Beese-Sims SE, Brookes E, Spadafora R, Zhu Y, Rothbart SB, et al. A histone methylation network regulates transgenerational epigenetic memory in C. elegans. Cell Rep. 2014;7:113–26.
Castel SE, Martienssen RA. RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat Rev Genet. 2013;14:100–12.
Esteller M, Corn PG, Baylin SB, Herman JG. A gene hypermethylation profile of human cancer. Cancer Res. 2001;61:3225–9.
Arnold CN, Goel A, Compton C, Marcus V, Niedzwiecki D, Dowell JM, et al. Evaluation of Microsatellite Instability, hMLH1 Expression and hMLH1 Promoter Hypermethylation in Defining the MSI Phenotype of Colorectal Cancer. Cancer Biol Ther. 2004;3.
Hawkins NJ, Ward RL. Sporadic colorectal cancers with microsatellite instability and their possible origin in hyperplastic polyps and serrated adenomas. J Natl Cancer Inst. 2001;93:1307–13.
Acknowledgements
This study was supported by grants by the Fondazione of the Cassa di Risparmio della Provincia di Chieti, the Oncoxx Biotech, the Italian Ministry of Health (RicOncol RF-EMR-2006-361866), the Compagnia di San Paolo (Grant 2489IT), and the Ministry of Development—Made in Italy (contract MI0100424).
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
MT, PS, and SA wrote the manuscript. MT and VR generated the figure panels. All authors contributed to discussing the content of the article. All authors read and approved the final manuscript.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Trerotola, M., Relli, V., Simeone, P. et al. Epigenetic inheritance and the missing heritability. Hum Genomics 9, 17 (2015). https://doi.org/10.1186/s40246-015-0041-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s40246-015-0041-3