Abstract
The availability of genome sequences obtained using next-generation sequencing (NGS) has revolutionized the field of infectious diseases. Indeed, more than 38,000 bacterial and 5,000 viral genomes have been sequenced to date, including representatives of all significant human pathogens. These tremendous amounts of data have not only enabled advances in fundamental biology, helping to understand the pathogenesis of microorganisms and their genomic evolution, but have also had implications for clinical microbiology. Here, we first review the current achievements of genomics in the development of improved diagnostic tools, including those that are now available in the clinic, such as the design of PCR assays for the detection of microbial pathogens, virulence factors or antibiotic-resistance determinants, or the design of optimized culture media for ‘unculturable’ pathogens. We then review the applications of genomics to the investigation of outbreaks, either through the design of genotyping assays or the direct sequencing of the causative strains. Finally, we discuss how genomics might change clinical microbiology in the future.
Similar content being viewed by others
The impact of next-generation sequencing in infectious disease diagnostics
Infectious diseases are one of the leading causes of human mortality worldwide [1]. Therefore, accurate diagnostic methods are required to optimize the clinical management of infected patients. However, the gold standard for the diagnosis of infectious diseases has long been the culture in growth-supporting media, including the isolation, identification and antibiotic-susceptibility testing of the causative microorganism. Currently, this diagnostic scheme takes a minimum of 24 hours. The introduction of the polymerase chain reaction (PCR) [2] method in the 1980s resulted in the development of a multitude of diagnostic tools that helped improve the efficiency of diagnostics and the characterization of infectious-disease agents by detecting and identifying their DNA. However, the design of these assays remained mostly empirical, being notably based on the use of the 16S rRNA gene [3], until bacterial genome sequencing became a reality in the mid-1990s [4]. Microbial genomics, enabling a rational design of most molecular assays by selecting molecular targets according to their objective, has now had a major impact on the diagnosis and prevention of infectious diseases, with detection and identification of pathogens being directly performed within specimens without the need for culture [5].
Since 2005, the development of next-generation sequencing (NGS), together with decreasing costs for sequencers and reagents, has democratized genomics (Table 1) [6]. Currently, a bacterial genome sequence can be obtained within a few days for less than US$500 [6], and more than 38,000 genome sequences are available in public databases [7]. NGS has had many applications in medical microbiology, including the design of diagnostic and genotyping tools, the identification of virulence and antibiotic-resistance mechanisms and the development of specific culture media [8]-[12].
Here, we review the most relevant applications of genomics to the fields of molecular detection, identification and genotyping of infectious-disease agents, detection of virulence and antibiotic-resistance markers, design of culture media and investigation of outbreaks (Table 2; Figure 1), including those that are already available in clinical microbiology laboratories, and we offer our thoughts on how genomics might change clinical microbiology in the future.
Applications of bacterial genomics to the management of infectious diseases. Genome sequence analysis has enabled the development of various clinical-microbiology tools for pathogen detection, identification or genotyping by identification of sequence fragments specific at distinct taxonomic levels (genus, species, strain, clone), for the detection of genes associated with antibiotic resistance or virulence and for the identification of deficient metabolisms to aid the development of optimized culture media. However, whole-genome sequencing, by giving access to the full genetic repertoire of an isolate, has demonstrated an undisputed discriminatory power for deciphering outbreaks of infectious diseases.
Detection of pathogens in clinical specimens
Rapid detection and identification of infectious agents in clinical specimens are mandatory in order to implement appropriate therapeutic measures. Therefore, an ideal detection assay should both be sensitive, specific and rapid to maximize the chances of patient recovery and be able to minimize the occurrence of clinical complications.
Since its development in 1983, PCR remained the most widely used molecular method in clinical microbiology, notably for detection of microorganisms in clinical specimens, until 1996 when real-time PCR (RT-PCR) was developed. In contrast to long-established culture-based diagnostic methods, PCR enabled identification of microorganisms regardless of their culturability and was, therefore, especially valuable in patients who had received antibiotics before sampling or those infected by fastidious microorganisms - that is, microorganisms that do not grow in the usual culture conditions [3]. However, early PCR assays were empirically designed and often targeted a gene common to all bacteria, thus allowing the detection of any species (for example, the rRNA operon or the groEL gene). Although these broad-range PCR assays enabled the discovery of many human pathogens [13], they suffered from various drawbacks, in particular a lack of sensitivity, specificity and discriminatory power among bacterial species [14]. By contrast, RT-PCR, targeting shorter fragments and using a fluorescent probe, greatly improved the speed, sensitivity and specificity of detection, in particular when coupled to the rational selection of PCR targets in genomic sequences according to the experimental objective and the degree of specificity required (genus-, species-, subspecies-, strain- or gene-specific) [15]-[17]. As the genomes from more than 37,000 bacterial strains are currently available, including those of all major human pathogens, it is now possible for clinical microbiologists to design specific PCR assays according to their needs by using the available tools. As examples, Marshall developed ‘PerlPrimer’, a software enabling the design of target-specific PCR or RT-PCR primers [15], Pritchard and colleagues proposed an alignment-free method for designing strain-specific primers for Escherichia coli O104:H4 [18], and Hung and associates designed a stepwise computational approach mixing several publicly available softwares to identify species-specific signatures in whole-genome sequences [17]. Using Streptococcus pyogenes as a model, Hung and colleagues designed and tested the validity of 15-signature-derived primer sets, including nine that were highly species-specific in vitro[17]. In addition, RT-PCR made possible the development of syndrome-driven molecular diagnosis in which assays detecting the most common etiological agents of a given syndrome are tested concomitantly [19]. In a recent study, Sokhna and colleagues described the use of a syndrome-driven strategy for the point-of-care diagnosis of febrile illness [20]. This type of diagnostic method has the advantage of testing, in a short time and a limited number of specimens, the most common causative agents of a given syndrome and can be especially valuable, for example, in the diagnosis of meningitis, pneumonia, endocarditis, pericarditis or sexually transmitted diseases. Thus, it enables a more efficient management of patients by enabling an earlier commencement of appropriate antibiotic therapy. Furthermore, genomics has also allowed the design of multiplex PCR assays enabling simultaneous detection and discrimination of various microorganisms, as has been the case for members of the Mycobacterium tuberculosis complex and Mycobacterium canettii[8]. This is also true for microarrays, some of which can enable the detection and identification of more than 2,000 viral and 900 bacterial species at once [21]. Nsofor recently reviewed the applications of microarrays to the syndrome-based diagnosis of infectious diseases, some of which, such as the ResPlex II Panel v2.0 (Qiagen, Hilden, Germany) and the FilmArray Respiratory Panel (BioMerieux, Marcy L’Etoile, France) for respiratory infections, are commercially available [22].
In addition to the development of highly specific PCR assays, the study of genomic sequences enabled the optimization of the sensitivity of detection, either by selecting a gene or fragment of noncoding DNA present as several copies in the genome [23] or by designing nested PCR assays targeting previously unused genomic fragments [24]. Fenollar and colleagues identified a seven-copy fragment in the genome from the bacterium Tropheryma whipplei and demonstrated that a RT-PCR assay targeting this repeated fragment was significantly more sensitive than assays targeting a single-copy fragment [23]. By contrast, Drancourt and colleagues developed a strategy named 'suicide PCR' that is based on nested-PCR assays targeting genome fragments that had never been used as PCR targets previously and that will be targeted only once with single-use primers [25]. These authors also demonstrated a higher sensitivity of their method over regular PCR. Targeting multicopy fragments was demonstrated to be highly sensitive for the detection of Q fever, Whipple’s disease, brucellosis, and infections caused by Mycoplasma pneumoniae or Neisseria meningitidis, whereas ‘suicide PCR’ was successful in detecting Yersinia pestis from dental specimens of ancient plague outbreaks and Rickettsia spp. in various arthropod-borne diseases [24],[25].
To date, several genome-based PCR tests have become commercially available. These include the LightCycler SeptiFast (Roche, Mannheim, Germany) and GeneXpert (Cepheid, Sunnyvale, CA, USA) systems that offer multiplexed detection of the various pathogens potentially involved in a given infectious syndrome. The latter system also enables simultaneous discrimination of M. tuberculosis complex species and detection of rifampicin resistance. Alternative assays are based on various detection methods for PCR products, as is the case for the ResPlex II Panel (Qiagen, Hilden, Germany) and Film Array (BioMerieux), in which PCR amplicons are hybridized to a microarray for the syndrome-based detection of pathogens, the GenoType MTBDRplus assay (Hain Lifescience, Nehren, Germany) that combines PCR and hybridization to a strip to detect antibiotic resistance in M. tuberculosis, and the PLEX-ID (Abbott, Abbott Park, IL, USA), in which broad-range and clade-specific PCR products are identified through using electro-spray ionization-mass spectrometry. The latter system enables screening human specimens for bacteria, viruses, fungi, protozoa and several antibiotic-resistance-associated genes [26].
However, although PCR and, more recently, RT-PCR have revolutionized the diagnosis of infectious diseases by reducing the time to diagnosis and increasing the detection sensitivity, several challenges remain, including the spectrum of detected agents, which is limited by the specificity of the assays used. However, thanks to their decreasing cost, the development of syndrome-based multiplex PCR assays or microarrays is likely to increase in the coming years. Alternatively, NGS, already known to be used for genotyping purposes in clinical microbiology, might also be increasingly used for clinical detection of pathogens, as was recently described for the diagnosis of a case of neuroleptospirosis [27].
Genotyping
In situations when understanding the source and spread of microorganisms is crucial, as is the case for outbreaks caused by multidrug-resistant or hypervirulent bacteria and nosocomial or pandemic infections, a higher discriminatory power is needed to be able to trace pathogens at the strain level. Identifying bacteria at the strain level - or bacterial strain typing - is particularly important for epidemiological surveillance of infections. Strain typing also has applications in studying bacterial population dynamics. Over the past three decades, molecular typing (or molecular fingerprinting) methods have largely superseded phenotypic methods, including the morphology of colonies on various culture media, biochemical tests, serology, killer toxin susceptibility and pathogenicity, which exhibit insufficient discriminatory power, inability to quantify genetic relationships between isolates, limited reagent availability, poor intra- and inter-laboratory reproducibility and difficulties in comparing results obtained in different laboratories. In a similar fashion as described for PCR assay design, genomic sequences can be a source of genotyping targets. Molecular typing methods can be classified as non-sequence-based and sequence-based genotyping methods, depending on their design (Figure 2). Non-sequence-based genotyping methods include pulsed-field gel electrophoresis (PFGE), PCR-restriction fragment length polymorphism (PCR-RFLP), multiple-locus variable-number tandem-repeat analysis (MLVA), single-nucleotide polymorphisms (SNPs) and microarrays. Sequence-based genotyping methods include multilocus sequence typing (MLST), multispacer sequence typing (MST) and whole-genome sequence typing. The choice of genotyping method should be made according to the population structure of the investigated microorganism. This is particularly crucial for clonal bacteria, such as M. tuberculosis or Bacillus anthracis, for which structural genes are poorly polymorphic and PCR-RFLP or MLST are inadequate, whereas MLVA is able to discriminate among strains [28].
Principles of genome-based genotyping methods. By genomic comparison, investigators can identify specific sequence signatures that can be used in non-sequence-based methods (DNA banding-pattern-, PCR- or hybridization-based methods) or sequence-based methods (partial or complete genome sequencing). MLST, multi-locus sequence typing; MLVA, multiple locus variable number tandem repeat analysis; MST, muti-spacer sequence typing; PCR-RFLP, PCR-restriction fragment length polymorphism; PFGE, pulsed-field gel electrophoresis; RFLP, restriction fragment length polymorphism; SNP, single nucleotide polymorphism.
Non-sequence-based genotyping methods
PFGE and PCR-RFLP have long been considered as 'gold standard' genotyping methods. These methods are DNA-banding-pattern-based methods that compare the electrophoretic profiles of restriction-enzyme-cut genomes or PCR-amplified genes from various strains. Initially, these methods relied on uncharacterized genomic differences or empirically selected target genes. By contrast, genome sequences, as was the case for M. tuberculosis or Y. pestis[9], can be used to rationally improve the sensitivity and specificity of PFGE or PCR-RFLP by enabling the ‘in silico’ prediction of the most appropriate restriction profiles of rare-cutter enzymes for a given bacterium.
In an alternative approach, Yang and colleagues have used genomics to design the ‘Pan-PCR’ software, dedicated to the identification of strain-specific PCR targets in genome sequences in a ‘presence/absence’ mode, that is, the amplification of a series of unrelated genes that were differentially present in the genomes from the studied strains [29]. As an example, in Acinetobacter baumannii, the presence or absence of six genetic loci, as determined by six locus-specific PCR assays, discriminated 29 tested strains [29]. Such a method is rapid, easy to perform and only requires a real-time thermal cycler, but it might not be adapted to species with highly conserved genomes such as B. anthracis in which the gene content does not vary among strains.
Another non-sequence-based genotyping method that benefited from the availability of genome sequences is MLVA. This method is based on the determination of the number and length of variable number of tandem repeats (VNTRs) present in a genome and is applicable to a variety of pathogens [30],[31]. Currently, MLVA is a reference genotyping method for many bacteria, such as M. tuberculosis[28],[32], and has also been used to investigate outbreaks of infections, as was demonstrated by Paranthaman and colleagues, who accurately identified the source of a multidrug-resistant Salmonella enterica serovar Typhimurium outbreak that occurred in England in 2011 [31]. MLVA is a rapid, easy-to-perform, affordable and reproducible genotyping method with high discriminatory power, but it has been demonstrated to be non-adaptable for some species, such as Mycoplasma hyopneumoniae, which lacks tandem repeats [33], and in long-term epidemiology for Mycobacterium leprae in which variations in the VNTR pattern were observed not only between isolates but also between specimens from the same patient [16].
The detection of single nucleotide polymorphisms (SNPs), another widely used typing method for bacteria, has also been improved through using genome sequences. This method, based on point-nucleotide changes between strains of a given species, has enabled the genotyping of several bacterial pathogens [9],[34]-[39], including Coxiella burnetii[40]. Using SNP genotyping, Huijsmans and colleagues identified five genotypes of C. burnetii that were involved in the large outbreak of Q fever that occurred in the Netherlands between 2007 and 2012 [40]. By comparison with other genotyping methods, SNP-based methods are rapid, sensitive, easy to perform and unambiguous in result interpretation. However, it should be noted that interpreting SNP genotyping data is highly dependent on the algorithm, the reference sequence and the sequencing platform used, which highlights a need for standardization of the methods used.
Genome-based DNA microarrays, an intermediate between non-sequence-based and sequence-based methods, contain probes specific for some or all genes present in a genome [41]. This method enables simultaneous strain comparisons at a whole-genome level. It can be automated and is a fast, sensitive and high-throughput genotyping tool [16],[42]. Genome-based DNA microarrays were developed to genotype a number of human pathogens, including Escherichia coli[43], for which Geue and colleagues were able to discriminate 446 Shiga-toxin-producing E. coli[44]. DNA microarrays can also be used to detect and identify microorganisms in complex floras [30],[45]. However, although highly discriminatory, microarray-based methods suffer from the major drawback that they cannot identify genetic fragments for which no probe is used.
Sequence-based genotyping methods
By comparison with non-sequence-based methods, sequence-based genotyping has the major advantage of being highly reproducible because the sequence fragments on which it is based are stored in public databases. Sequence-based genotyping methods can rely on the selection of one or several genomic targets or on the whole genome sequence. Single-locus sequence-typing methods require the in silico identification of a highly variable gene, such as the coagulase- and protein-A-encoding genes that are the genomic targets of coa or spa typing, respectively, two broadly used tools for Staphylococcus aureus[46],[47].
MLST, developed in 1998, is one of the most frequently used sequence-based genotyping methods. It is based on the combination of genotypes obtained from several individual genes, usually housekeeping genes, for characterizing bacterial strains [48]. Genome-sequence-designed MLST assays have been useful for typing pathogens that have highly variable genomes among strains, such as E. coli, N. meningitidis or S. aureus[30],[49],[50], but they demonstrated limited discriminatory power among those bacteria with highly conserved genomes such as B. anthracis[30]. In 2012, rMLST, based on a combination of 53 ribosomal protein subunits, was demonstrated to discriminate strains within the genus Neisseria[51]. However, whole-genome MLST, incorporating more than 500 loci, was able to identify bacteria at the clone level [52]. This method is especially valuable when implemented with the BIGSdb platform that enables standardization of data [53]. In a similar fashion, multi-spacer typing (MST), based on the assumption that intergenic spacers are more variable than genes owing to a lower selection pressure, combines sequences from the most variable intergenic spacers between aligned genomes of bacterial strains instead of genes [54]. First developed for Y. pestis[54], MST has also been efficient at typing strains from various other bacteria, including C. burnetii[30],[55]-[57]. Glazunova and colleagues, by using a combination of 10 intergenic spacer sequences, were able to classify 159 C. burnetii isolates within 30 distinct genotypes [55]. MST was demonstrated to be more discriminatory than MLST for R. conorii strains [56].
However, bacterial whole-genome sequencing (WGS) using NGS, by giving access to the whole genetic content of a strain, is the ultimate discriminatory sequence-based genotyping method and has already demonstrated its usefulness for epidemiological investigations, showing the rapid global transmission of infectious diseases [38],[58],[59] (Table 3). WGS was used to compare 86 human M. tuberculosis isolates from a German outbreak and has demonstrated its superiority over other genotyping methods for tracing and investigating micro-epidemics [60],[61]. In 2010, WGS was used to study 63 strains of methicillin-resistant Staphylococcus aureus (MRSA) from various countries and enabled reconstruction of intercontinental transmissions over four decades as well as the potential transmission within a hospital environment [38]. WGS was also used to investigate the cholera outbreak in Haiti that occurred in 2010 [58],[59], revealing that Haitian strains were closely related to strains from Nepal. These pioneering studies demonstrated the potential of WGS for retrospective genotyping. The major challenge is to make WGS a genotyping tool during the course of outbreaks, and for this it will be necessary to facilitate access to sequencing platforms.
Detection of virulence factors
In addition to identifying bacteria at various taxonomic levels, WGS offers the opportunity to detect various genetic markers, such as virulence factors or antibiotic resistance-associated genes. Identifying and characterizing the virulence factors of pathogens are crucial for understanding the pathogenesis of the diseases that they cause and for developing dedicated molecular tools to detect specific virulence markers. However, among the currently known virulence markers, only toxins are important for optimizing the management of patients, as these agents are able to cause hospital outbreaks of severe infections with high mortality rates, such as the hypervirulent ribotype O27 Clostridium difficile[62], or because the administration of antibiotics can have a significant impact on the outcome. This is notably the case for S. aureus, in which the secretion of the Panton-Valentine leukocidin is induced by oxacillin or depressed by clindamycin [63],[64], for the Shiga-toxin production in E. coli that is stimulated by β-lactams, sulfonamides and fluoroquinolones [65], and for Rickettsia conorii, in which fluoroquinolones upregulate a toxin-antitoxin module [66]. Therefore, determining the toxinic repertoire of strains of selected bacterial species can be crucial for effective clinical management.
Genomics has played an important role in the identification of virulence factors in bacteria. Three main strategies are used to identify virulence-factor-encoding genes in genomes [67]: first, comparison of genomes from strains or species exhibiting diverse degrees of virulence; second, identification of laterally transferred genomic islands, assuming that virulence genes are often acquired by this mechanism [67]; and, third, running the genome against databases of known virulence markers. The first approach was used in studies between Y. pestis, the causative agent of plague, and the less-virulent but closely related species Y. pseudotuberculosis[10], between a pathogenic strain of E. coli O157:H7 and a non-pathogenic laboratory strain of E. coli K-12 [68],[69], between a highly virulent Staphylococcus epidermidis causing community-acquired endocarditis and commensal strains [70], and between Klebsiella pneumoniae strains [71]. The second strategy enabled the identification of pathogenicity islands in various species [72]-[75], such as E. coli or S. aureus. The third method enabled identification of virulence genes in a variety of species [76]-[87], notably Listeria monocytogenes and M. tuberculosis. All three strategies are complementary but cannot replace functional studies for confirmation of the real role of the identified virulence factors in pathogenesis.
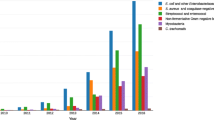
Paradoxically, genomic comparisons have also questioned the paradigm of virulence by gene acquisition. In many genera, genome reduction, rather than acquisition of additional genetic material, can be associated with increased virulence, as many of the most virulent bacterial pathogens have smaller genomes than closely related species [88]. The comparison of rickettsial genomes showed that Rickettsia prowazekii, the agent of epidemic typhus, the deadliest rickettsiosis, had the smallest genome in this genus (Figure 2) [89]. Similar findings were reported for Mycobacterium ulcerans[90]. In addition, the presence of ‘non-virulence’ genes was described as discriminating Shigella dysenteriae from E. coli or Y. pestis from Y. pseudotuberculosis[88]. In Y. pestis, for example, the loss of the rcsA and nghA genes, which encode a repressor of biofilm synthesis and an inhibitor of biofilm formation, respectively, might have contributed to a more efficient flea-borne transmission [91]. Therefore, the pathogenic repertoire of a bacterium should not only take into account the presence or absence of virulence factors but also of ‘non-virulence’ genes.
However, it should be noted that the virulence of a bacterial strain might not systematically be predicted from its genome sequence, in particular when the identified virulence markers are not expressed. Indeed, Priest and colleagues could overcome this limitation by using systems biology to predict virulence in S. aureus[92]. Briefly, these authors not only considered the presence of virulence genes but also took into account the known regulatory networks of these genes.
Detection of antibiotic resistance
Currently, antimicrobial resistance is a major public health concern worldwide, especially as some pathogenic multidrug-resistant bacteria are already resistant to all antibiotics in use in the clinic [93]. Detection of bacterial resistance determinants, and identification of new arrangements of known resistance genes, as well as new putative resistance markers can be achieved with WGS. This might help predict the resistance phenotype, set up enhanced in-hospital infection-control measures, adapt a specific therapy and enable the identification of resistance-causing genes or mutations that could be detected by PCR from clinical isolates or specimens and serve as targets for routine detection tools [94]. The strategies for identifying resistance markers are very similar to those aimed at identifying virulence genes [6]. However, as incomplete data link genotype to phenotype in terms of drug resistance, WGS genomic-based detection is particularly suited for antibiotics for which resistance-associated mutations or genes are known and notably for fastidious bacteria such as M. tuberculosis[95].
Genomic comparisons of phenotypically resistant and susceptible strains has enabled investigation of the resistome - that is, the repertoire of genetic markers associated with antibiotic resistance of multidrug-resistant strains of Enterococcus faecium[11] and S. pneumoniae[96]. Genome sequencing has also enabled identification of resistance mechanisms in fastidious bacteria that express few phenotypic characteristics, as was the case for T. whipplei, the causative agent of Whipple’s disease, that is resistant to fluoroquinolones owing to mutations in the gyrA and parC genes [97], Rickettsia felis, which expresses a β-lactamase activity that was first found in the genome [98], and M. tuberculosis, in which a putative rRNA methyltransferase might explain its resistance to macrolide antibiotic drugs [95].
Several PCR assays used in clinical practice derive from genomic sequences. The recent discovery of the mecC gene - a homolog of the mecA gene encoding methicillin resistance, responsible for false susceptibility testing results - in the genome of a methicillin-resistant S. aureus[99] elicited the design of specific PCR assays [100]. The spread of carbapenemase-producing enterobacteriaceae also prompted the sequencing of genomes from various MDR strains, including an NDM-1-producing E. coli strain [101] and a blaKPC2-producing K. pneumoniae[102], which in turn enabled the development of dedicated PCR assays [103]. Therefore, although many genome-based molecular tests facilitating the management of infections have already been developed to date, there is no doubt that WGS data will be used extensively in future assay design.
Culturing unculturable pathogens
Despite the breakthrough of molecular methods, culture remains the cornerstone of routine microbiology as it provides insight into their ecology and pathogenicity. However, a majority of microorganisms in nature are not cultivable using standard techniques. Many fastidious bacteria grow poorly on commonly used media, and others are considered uncultivable on axenic media, possibly owing to deficient or partial metabolic pathways. Thus, genome sequences might enable identification of incomplete metabolic pathways [104] and the essential nutrients that a bacterium is unable to produce [105], which could then be incorporated into a specifically designed culture medium. T. whipplei, causing Whipple’s disease, was the first ‘unculturable’ human pathogen [106],[107] to benefit from such an in silico design of a culture medium. An axenic culture medium specifically designed to contain the nine amino acids that this bacterium was unable to synthesize enabled its axenic growth [12]. A similar approach was used for Xyllela fastidiosa[108], Leptospirillum ferrodiazotrophum[109] and C. burnetii[110]. Alternatively, genomics might help improve culture media, as was the case for E. coli and M. pneumoniae[111],[112]. However, this strategy might not be efficient for just any bacterium, as was the case for M. leprae. Despite the many important metabolic activities missing in the genome [113] of this bacterium, no specifically complemented axenic medium has enabled any growth to date. However, although it is important to improve culture methods for fastidious microorganisms, the investigation of unusual infections or outbreaks needs rapid and informative methods that may help influence the management of patients and course of the outbreaks. Such progress is now made possible by NGS.
Real-time genomics for the diagnosis of infections or the investigation of outbreaks
The development of NGS bench-top sequencers such as the MiSeq (Illumina) and Ion Torrent Personal Genome Sequencer (PGM; Life Technologies) has made genome sequencing compatible with the routine clinical-microbiology workflow [6]. Such a strategy enables, within a few hours, exhaustive access to the genotype [39], virulence markers and antibiotic-resistance repertoire. Real-time genomics has notably been used to investigate several nosocomial [70],[114] or community-acquired infections [115]-[118] (Table 3). Sherry and colleagues used PGM sequencing of four MDR E. coli strains to confirm that the nosocomial outbreak that had occurred in a neonatal unit in Melbourne, Australia, had been caused by a unique clone and to characterize the resistance genes for this outbreak strain [118]. In Germany, Mellmann and colleagues compared the genomes from two E. coli O104:H4 strains from two hemolytic uremic syndrome outbreaks and concluded that the strains had diverged from a common ancestor and that NGS was suitable for the characterization of a pathogen in the early stages of an outbreak [115]. In both cases, genome sequences were obtained in a few days (five and three days, respectively). These findings demonstrated how rapid and precise genomic sequencing, although limited to a few clinical-microbiology laboratories currently, could transform patient management or improve hospital infection control in routine clinical practice.
Although only a few studies to date have described a turnaround time sufficiently short to enable WGS data to influence the course of outbreaks [119], the increasing number of teams using WGS for epidemiological purposes (Table 3) leaves little doubt as to the likelihood of its systematic use as a first-line tool to track and understand epidemics in the near future.
How will next-generation sequencing change clinical microbiology?
NGS has the potential to change clinical microbiology in several ways. First, the increasing number of genome sequences will enable the development of new and improved pathogen-specific or syndrome-based single or multiplexed RT-PCR assays and will aid the refinement of DNA targets, primers and probes used in existing tests [120]. Second, the increase in speed, decreasing costs and discriminatory power of NGS make it an ideal tool for routine use in diagnostic microbiology laboratories. NGS has the potential to replace several existing tests performed on the same isolate, notably identification of antibiotic-resistance mechanisms, virulence determinants and genotype, in particular for microorganisms that are difficult to grow. As such, it is especially well suited for infection control. In addition, NGS without the need for culture, in particular single-cell sequencing, might be relevant for the routine characterization of unculturable bacteria. Third, NGS has proven its usefulness to decipher complex microbiotas in various metagenomic studies [121]. Recent studies have demonstrated its ability not only to discriminate among microorganisms present in human specimens, and thus possibly detect co-infections, but also uncover unexpected or new pathogens [122]-[124].
However, several challenges remain, the most important being a facilitated and rapid access of clinical microbiology laboratories to sequencing platforms, and a need for standardized and fully automated sequence interpretation that would ideally be independent of both the sequencing platform and the exact species of microorganism [125]-[127]. Also needed is the ability to translate the data into relevant information enabling microbiologists, clinicians and public-health epidemiologists to implement control measures in real-time and alter the course of outbreaks. This implies a constant update and curation of public databases as well as the development of systems-biology-based softwares that will enable prediction of virulence and antibiotic resistance from genome sequences.
Conclusions and perspectives
The expansion of genomics, giving access to the genomes of virtually all human pathogens, has greatly changed our approach regarding management of infectious diseases by shedding light on their genetic diversity, pathogenesis, evolution, detection and treatment. With access to the full genetic content of microorganisms, rational selection of DNA fragments has enabled creation of a wide array of detection and typing methods as well as specialized tools for the identification of genes encoding factors affecting virulence or antibiotic resistance. In addition, NGS methods have reached a point, both in terms of cost and speed, where they might enter the routine microbiology laboratory and be used routinely for the rapid sequencing of microorganisms that exhibit unusual pathogenicity, are antibiotic-resistant or cause outbreaks. However, the major challenge in order to include genome sequencing in the routine workflow of the clinical-microbiology laboratory, in addition to a need for a multiplication of sequencing platforms, is a clear need for improved sequence analysis, both in terms of numbers and data handling of bioinformatic facilities, and storage capacity, as well as homogenized gene-function assignment.
It is likely that NGS, by permitting genome sequencing from single cells or single colonies, will also constitute a major step forward in the comprehension of bacterial genome dynamics [128]. This strategy has the advantage over other sequencing methods in that it is applicable to microorganisms that are unculturable and/or part of complex floras [129],[130]. However, single-cell genomics also currently suffers from several limitations, which include genome amplification biases, chimeric DNA rearrangements and a need for the improved de novo assembly of DNA sequences of previously non-sequenced microorganisms.
Abbreviations
- MLST:
-
multi-locus sequence typing
- MLVA:
-
multiple variable number tandem repeat analysis
- MRSA:
-
methicillin-resistant Staphylococcus aureus
- MST:
-
multi-spacer typing
- NGS:
-
next-generation sequencing
- PCR-RFLP:
-
PCR-restriction fragment length polymorphism
- PFGE:
-
pulsed-field gel electrophoresis
- RFLP:
-
restriction fragment length polymorphism
- RT-PCR:
-
real-time polymerase chain reaction
- SNP:
-
single nucleotide polymorphism
- VNTRs:
-
variable number of tandem repeats
- WGS:
-
whole-genome sequencing
References
Lozano R, Naghavi M, Foreman K, Lim S, Shibuya K, Aboyans V, Abraham J, Adair T, Aggarwal R, Ahn SY, Alvarado M, Anderson HR, Anderson LM, Andrews KG, Atkinson C, Baddour LM, Barker-Collo S, Bartels DH, Bell ML, Benjamin EJ, Bennett D, Bhalla K, Bikbov B, Bin Abdulhak A, Birbeck G, Blyth F, Bolliger I, Boufous S, Bucello C, Burch M, et al: Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet. 2012, 380: 2095-2128.
Mullis KB, Faloona FA: Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction. Methods Enzymol. 1987, 155: 335-350.
Relman DA, Falkow S: Identification of uncultured microorganisms: expanding the spectrum of characterized microbial pathogens. Infect Agents Dis. 1992, 1: 245-253.
Fleischmann RD, Adams MD, White O, Clayton RA, Kirkness EF, Kerlavage AR, Bult CJ, Tomb JF, Dougherty BA, Merrick JM, McKenney K, Sutton G, FitzHugh W, Fields C, Gocyne JD, Scott J, Shirley R, Liu L-I, Glodek A, Kelley JM, Weidman JF, Phillips CA, Spriggs T, Hedblom E, Cotton MD, Utterback TR, Hanna MC, Nguyen DT, Saudek DM, Brandon RC, et al: Whole-genome random sequençing and assembly of Hamophilus influenzae Rd. Science. 1995, 269: 496-512.
Logares R, Haverkamp TH, Kumar S, Lanzen A, Nederbragt AJ, Quince C, Kauserud H: Environmental microbiology through the lens of high-throughput DNA sequencing: synopsis of current platforms and bioinformatics approaches. J Microbiol Methods. 2012, 91: 106-113.
Didelot X, Bowden R, Wilson DJ, Peto TE, Crook DW: Transforming clinical microbiology with bacterial genome sequencing. Nat Rev Genet. 2012, 13: 601-612.
The Genomes Online Database. [], [http://www.genomesonline.org/]
Reddington K, O'Grady J, Dorai-Raj S, Niemann S, van SD, Barry T: A novel multiplex real-time PCR for the identification of mycobacteria associated with zoonotic tuberculosis. PLoS One. 2011, 6: e23481-
Achtman M, Morelli G, Zhu P, Wirth T, Diehl I, Kusecek B, Vogler AJ, Wagner DM, Allender CJ, Easterday WR, Chenal-Francisque V, Worsham P, Thomson NR, Parkhill J, Lindler LE, Carniel E, Keim P: Microevolution and history of the plague bacillus, Yersinia pestis . Proc Natl Acad Sci U S A. 2004, 101: 17837-17842.
Chain PS, Carniel E, Larimer FW, Lamerdin J, Stoutland PO, Regala WM, Georgescu AM, Vergez LM, Land ML, Motin VL, Brubaker RR, Fowler J, Hinnebusch J, Marceau M, Medigue C, Simonet M, Chenal-Francisque V, Souza B, Dacheux D, Elliott JM, Derbise A, Hauser LJ, Garcia E: Insights into the evolution of Yersinia pestis through whole-genome comparison with Yersinia pseudotuberculosis . Proc Natl Acad Sci U S A. 2004, 101: 13826-13831.
van Schaik W, Top J, Riley DR, Boekhorst J, Vrijenhoek JE, Schapendonk CM, Hendrickx AP, Nijman IJ, Bonten MJ, Tettelin H, Willems RJ: Pyrosequencing-based comparative genome analysis of the nosocomial pathogen Enterococcus faecium and identification of a large transferable pathogenicity island. BMC Genomics. 2010, 11: 239-
Renesto P, Crapoulet N, Ogata H, La Scola B, Vestris G, Claverie JM, Raoult D: Genome-based design of a cell-free culture medium for Tropheryma whipplei . Lancet. 2003, 362: 447-449.
Goh SH, Potter S, Wood JO, Hemmingsen SM, Reynolds RP, Chow AW: HSP60 gene sequences as universal targets for microbial species identification: studies with coagulase-negative staphylococci. J Clin Microbiol. 1996, 34: 818-823.
Janda JM, Abbott SL: 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. J Clin Microbiol. 2007, 45: 2761-2764.
Marshall O: Graphical design of primers with PerlPrimer. Methods Mol Biol. 2007, 402: 403-414.
Li W, Raoult D, Fournier PE: Bacterial strain typing in the genomic era. FEMS Microbiol Rev. 2009, 33: 892-916.
Hung GC, Nagamine K, Li B, Lo SC: Identification of DNA signatures suitable for use in development of real-time PCR assays by whole-genome sequence approaches: use of Streptococcus pyogenes in a pilot study. J Clin Microbiol. 2012, 50: 2770-2773.
Pritchard L, Holden NJ, Bielaszewska M, Karch H, Toth IK: Alignment-free design of highly discriminatory diagnostic primer sets for Escherichia coli O104:H4 outbreak strains. PLoS One. 2012, 7: e34498-
Bouricha M, Samad MA, Levy PY, Raoult D, Drancourt M: Point-of-care syndrome-based, rapid diagnosis of infections on commercial ships. J Travel Med. 2014, 21: 12-16.
Sokhna C, Mediannikov O, Fenollar F, Bassene H, Diatta G, Tall A, Trape JF, Drancourt M, Raoult D: Point-of-care diagnosis of emerging pathogens in rural Senegal. PLoS Negl Trop Dis. 2013, 7: e1999-
Gardner SN, Jaing CJ, McLoughlin KS, Slezak TR: A microbial detection array (MDA) for viral and bacterial detection. BMC Genomics. 2010, 11: 668-
Nsofor CA: DNA microarrays and their applications in medical microbiology. Biotechnol Mol Biol Rev. 2014, 9: 1-11.
Fenollar F, Fournier PE, Robert C, Raoult D: Use of genome-selected repeated sequences increases the sensitivity of PCR detection of Tropheryma whipplei . J Clin Microbiol. 2004, 42: 401-403.
Fournier PE, Raoult D: Suicide PCR on skin biopsy specimens for diagnosis of rickettsioses. J Clin Microbiol. 2004, 42: 3428-3434.
Raoult D, Aboudharam G, Crubezy E, Larrouy G, Ludes B, Drancourt M: Molecular identification by "suicide PCR" of Yersinia pestis as the agent of Medieval Black Death. Proc Natl Acad Sci U S A. 2000, 97: 12800-12803.
Wolk DM, Kaleta EJ, Wysocki VH: PCR-electrospray ionization mass spectrometry: the potential to change infectious disease diagnostics in clinical and public health laboratories. J Mol Diagn. 2012, 14: 295-304.
Wilson MR, Naccache SN, Samayoa E, Biagtan M, Bashir H, Yu G, Salamat SM, Somasekar S, Federman S, Miller S, Sokolic R, Garabedian E, Candotti F, Buckley RH, Reed KD, Meyer TL, Seroogy CM, Galloway R, Henderson SL, Gern JE, DeRisi JL, Chiu CY: Actionable diagnosis of neuroleptospirosis by next-generation sequencing. N Engl J Med. 2014, 370: 2408-2417.
Supply P, Warren RM, Bañuls AL, Lesjean S, Van Der Spuy GD, Lewis LA, Tibayrenc M, Van Helden PD, Locht C: Linkage disequilibrium between minisatellite loci supports clonal evolution of Mycobacterium tuberculosis in a high tuberculosis incidence area. Mol Microbiol. 2003, 47: 529-538.
Yang JY, Brooks S, Meyer JA, Blakesley RR, Zelazny AM, Segre JA, Snitkin ES: Pan-PCR, a computational method for designing bacterium-typing assays based on whole-genome sequence data. J Clin Microbiol. 2013, 51: 752-758.
Fournier PE, Drancourt M, Raoult D: Bacterial genome sequencing and its use in infectious diseases. Lancet Infect Dis. 2007, 7: 711-723.
Paranthaman K, Haroon S, Latif S, Vinnyey N, de Souza V, Welfare W, Tahir M, Cooke E, Stone K, Lane C, Peters T, Puleston R: Emergence of a multidrug-resistant (ASSuTTm) strain of Salmonella enterica serovar Typhimurium DT120 in England in 2011 and the use of multiple-locus variable-number tandem-repeat analysis in supporting outbreak investigations. Foodborne Pathog Dis. 2013, 10: 850-855.
Hill V, Zozio T, Sadikalay S, Viegas S, Streit E, Kallenius G, Rastogi N: MLVA based classification of Mycobacterium tuberculosis complex lineages for a robust phylogeographic snapshot of its worldwide molecular diversity. PLoS One. 2012, 7: e41991-
Minion FC, Lefkowitz EJ, Madsen ML, Cleary BJ, Swartzell SM, Mahairas GG: The genome sequence of Mycoplasma hyopneumoniae strain 232, the agent of swine mycoplasmosis. J Bacteriol. 2004, 186: 7123-7133.
Pallen MJ, Loman NJ, Penn CW: High-throughput sequencing and clinical microbiology: progress, opportunities and challenges. Curr Opin Microbiol. 2010, 13: 625-631.
Read TD, Salzberg SL, Pop M, Shumway M, Umayam L, Jiang L, Holtzapple E, Busch JD, Smith KL, Schupp JM, Solomon D, Keim P, Fraser CM: Comparative genome sequencing for discovery of novel polymorphisms in Bacillus anthracis . Science. 2002, 296: 2028-2033.
Zhang W, Qi W, Albert TJ, Motiwala AS, Alland D, Hyytia-Trees EK, Ribot EM, Fields PI, Whittam TS, Swaminathan B: Probing genomic diversity and evolution of Escherichia coli O157 by single nucleotide polymorphisms. Genome Res. 2006, 16: 757-767.
Roumagnac P, Weill FX, Dolecek C, Baker S, Brisse S, Chinh NT, Le TA, Acosta CJ, Farrar J, Dougan G, Achtman M: Evolutionary history of Salmonella typhi . Science. 2006, 314: 1301-1304.
Harris SR, Feil EJ, Holden MT, Quail MA, Nickerson EK, Chantratita N, Gardete S, Tavares A, Day N, Lindsay JA, Edgeworth JD, de Lencastre H, Parkhill J, Peacock SJ, Bentley SD: Evolution of MRSA during hospital transmission and intercontinental spread. Science. 2010, 327: 469-474.
Beres SB, Carroll RK, Shea PR, Sitkiewicz I, Martinez-Gutierrez JC, Low DE, McGeer A, Willey BM, Green K, Tyrrell GJ, Goldman TD, Feldgarden M, Birren BW, Fofanov Y, Boos J, Wheaton WD, Honisch C, Musser JM: Molecular complexity of successive bacterial epidemics deconvoluted by comparative pathogenomics. Proc Natl Acad Sci U S A. 2010, 107: 4371-4376.
Huijsmans CJ, Schellekens JJ, Wever PC, Toman R, Savelkoul PH, Janse I, Hermans MH: Single-nucleotide-polymorphism genotyping of Coxiella burnetii during a Q fever outbreak in The Netherlands. Appl Environ Microbiol. 2011, 77: 2051-2057.
Bryant PA, Venter D, Robins-Browne R, Curtis N: Chips with everything: DNA microarrays in infectious diseases. Lancet Infect Dis. 2004, 4: 100-111.
Sabat AJ, Budimir A, Nashev D, Sá-Leão R, van Dijl Jm, Laurent F, Grundmann H, Friedrich AW: Overview of molecular typing methods for outbreak detection and epidemiological surveillance. Euro Surveill. 2013, 18: 20380-
Miller MB, Tang YW: Basic concepts of microarrays and potential applications in clinical microbiology. Clin Microbiol Rev. 2009, 22: 611-633.
Geue L, Stieber B, Monecke S, Engelmann I, Gunzer F, Slickers P, Braun SD, Ehricht R: Development of a rapid microarray-based DNA subtyping assay for the alleles of Shiga toxins 1 and 2 of Escherichia coli . J Clin Microbiol. 2014, 52: 2898-2904.
Wu L, Thompson DK, Liu X, Fields MW, Bagwell CE, Tiedje JM, Zhou J: Development and evaluation of microarray-based whole-genome hybridization for detection of microorganisms within the context of environmental applications. Environ Sci Technol. 2004, 38: 6775-6782.
Koreen L, Ramaswamy SV, Graviss EA, Naidich S, Musser JM, Kreiswirth BN: spa typing method for discriminating among Staphylococcus aureus isolates: implications for use of a single marker to detect genetic micro- and macrovariation. J Clin Microbiol. 2004, 42: 792-799.
Shopsin B, Gomez M, Waddington M, Riehman M, Kreiswirth BN: Use of coagulase gene (coa) repeat region nucleotide sequences for typing of methicillin-resistant Staphylococcus aureus strains. J Clin Microbiol. 2000, 38: 3453-3456.
Maiden MC, Bygraves JA, Feil E, Morelli G, Russell JE, Urwin R, Zhang Q, Zhou J, Zurth K, Caugant DA, Feavers IM, Achtman M, Spratt BG: Multilocus sequence typing: A portable approach to the identification of clones within populations of pathogenic microorganisms. Proc Natl Acad Sci U S A. 1998, 95: 3140-3145.
Kang CI, Cha MK, Kim SH, Ko KS, Wi YM, Chung DR, Peck KR, Lee NY, Song JH: Clinical and molecular epidemiology of community-onset bacteremia caused by extended-spectrum beta-lactamase-producing Escherichia coli over a 6-year period. J Korean Med Sci. 2013, 28: 998-1004.
Zhang YJ, Chen YS, Wang ZW, Li YQ, Wang DX, Shang Y, Fu RR, Hu YH, Geng R, Wei LP, Yang JP, Li JS, Yu Q, DU J, Gao ZC: Serological and molecular capsular typing, antibiotic susceptibility and multilocus sequence typing of Streptococcus pneumoniae isolates from invasive and non-invasive infections. Chin Med J (Engl). 2013, 126: 2296-2303.
Jolley KA, Bliss CM, Bennett JS, Bratcher HB, Brehony C, Colles FM, Wimalarathna H, Harrison OB, Sheppard SK, Cody AJ, Maiden MC: Ribosomal multilocus sequence typing: universal characterization of bacteria from domain to strain. Microbiology. 2012, 158: 1005-1015.
Maiden MC, Jansen van Rensburg MJ, Bray JE, Earle SG, Ford SA, Jolley KA, McCarthy ND: MLST revisited: the gene-by-gene approach to bacterial genomics. Nat Rev Microbiol. 2013, 11: 728-736.
Jolley KA, Maiden MC: BIGSdb: Scalable analysis of bacterial genome variation at the population level. BMC Bioinformatics. 2010, 11: 595-
Drancourt M, Roux V, Dang LV, Tran-Hung L, Castex D, Chenal-Francisque V, Ogata H, Fournier PE, Crubézy E, Raoult D: Genotyping, Orientalis-like Yersinia pestis, and plague pandemics. Emerg Infect Dis. 2004, 10: 1585-1592.
Glazunova O, Roux V, Freylikman O, Sekeyova Z, Fournous G, Tyczka J, Tokarevich N, Kovacava E, Marrie TJ, Raoult D: Coxiella burnetii genotyping. Emerg Infect Dis. 2005, 11: 1211-1217.
Fournier PE, Zhu Y, Ogata H, Raoult D: Use of highly variable intergenic spacer sequences for multispacer typing of Rickettsia conorii strains. J Clin Microbiol. 2004, 42: 5757-5766.
Zhu Y, Fournier PE, Ogata H, Raoult D: Multispacer typing of Rickettsia prowazekii enabling epidemiological studies of epidemic typhus. J Clin Microbiol. 2005, 43: 4708-4712.
Hendriksen RS, Price LB, Schupp JM, Gillece JD, Kaas RS, Engelthaler DM, Bortolaia V, Pearson T, Waters AE, Upadhyay BP, Shrestha SD, Adhikari S, Shakya G, Keim PS, Aarestrup FM: Population genetics of Vibrio cholerae from Nepal in 2010: evidence on the origin of the Haitian outbreak. MBio. 2011, 2: e00157-11.
Chin CS, Sorenson J, Harris JB, Robins WP, Charles RC, Jean-Charles RR, Bullard J, Webster DR, Kasarskis A, Peluso P, Paxinos EE, Yamaichi Y, Calderwood SB, Mekalanos JJ, Schadt EE, Waldor MK: The origin of the Haitian cholera outbreak strain. N Engl J Med. 2011, 364: 33-42.
Roetzer A, Diel R, Kohl TA, Rückert C, Nübel U, Blom J, Wirth T, Jaenicke S, Schuback S, Rüsch-Gerdes S, Supply P, Kalinowski J, Niemann S: Whole genome sequencing versus traditional genotyping for investigation of a Mycobacterium tuberculosis outbreak: a longitudinal molecular epidemiological study. PLoS Med. 2013, 10: e1001387-
Köser CU, Holden MT, Ellington MJ, Cartwright EJ, Brown NM, Ogilvy-Stuart AL, Hsu LY, Chewapreecha C, Croucher NJ, Harris SR, Sanders M, Enright MC, Dougan G, Bentley SD, Parkhill J, Fraser LJ, Betley JR, Schulz-Trieglaff OB, Smith GP, Peacock SJ: Rapid whole-genome sequencing for investigation of a neonatal MRSA outbreak. N Engl J Med. 2012, 366: 2267-2275.
Loo VG, Poirier L, Miller MA, Oughton M, Libman MD, Michaud S, Bourgault AM, Nguyen T, Frenette C, Kelly M, Vibien A, Brassard P, Fenn S, Dewar K, Hudson TJ, Horn R, René P, Monczak Y, Dascal A: A predominantly clonal multi-institutional outbreak of Clostridium difficile-associated diarrhea with high morbidity and mortality. N Engl J Med. 2005, 353: 2442-2449.
Dumitrescu O, Boisset S, Badiou C, Bes M, Benito Y, Reverdy ME, Vandenesch F, Etienne J, Lina G: Effect of antibiotics on Staphylococcus aureus producing Panton-Valentine leukocidin. Antimicrob Agents Chemother. 2007, 51: 1515-1519.
Casalta JP, Zaratzian C, Hubert S, Thuny F, Gouriet F, Habib G, Grisoli D, Deharo JC, Raoult D: Treatment of Staphylococcus aureus endocarditis with high doses of trimethoprim/sulfamethoxazole and clindamycin-Preliminary report. Int J Antimicrob Agents. 2013, 42: 190-191.
Wong CS, Jelacic S, Habeeb RL, Watkins SL, Tarr PI: The risk of the hemolytic-uremic syndrome after antibiotic treatment of Escherichia coli O157:H7 infections. N Engl J Med. 2000, 342: 1930-1936.
Botelho-Nevers E, Edouard S, Leroy Q, Raoult D: Deleterious effect of ciprofloxacin on Rickettsia conorii-infected cells is linked to toxin-antitoxin module up-regulation. J Antimicrob Chemother. 2012, 67: 1677-1682.
Karlin S: Detecting anomalous gene clusters and pathogenicity islands in diverse bacterial genomes. Trends Microbiol. 2001, 9: 335-343.
Dobrindt U, Hochhut B, Hentschel U, Hacker J: Genomic islands in pathogenic and environmental microorganisms. Nat Rev Microbiol. 2004, 2: 414-424.
Perna NT, Plunkett G, Burland V, Mau B, Glasner JD, Rose DJ, Mayhew GF, Evans PS, Gregor J, Kirkpatrick HA, Pósfai G, Hackett J, Klink S, Boutin A, Shao Y, Miller L, Grotbeck EJ, Davis NW, Lim A, Dimalanta ET, Potamousis KD, Apodaca J, Anantharaman TS, Lin J, Yen G, Schwartz DC, Welch RA, Blattner FR: Genome sequence of enterohaemorrhagic Escherichia coli O157:H7. Nature. 2001, 409: 529-533.
Fournier PE, Gouriet F, Gimenez G, Robert C, Raoult D: Deciphering genomic virulence traits of a Staphylococcus epidermidis strain causing native-valve endocarditis. J Clin Microbiol. 2013, 51: 1617-1621.
Lery LM, Frangeul L, Tomas A, Passet V, Almeida AS, Bialek-Davenet S, Barbe V, Bengoechea JA, Sansonetti P, Brisse S, Tournebize R: Comparative analysis of Klebsiella pneumoniae genomes identifies a phospholipase D family protein as a novel virulence factor. BMC Biol. 2014, 12: 41-
Hayashi T, Makino K, Ohnishi M, Kurokawa K, Ishii K, Yokoyama K, Han CG, Ohtsubo E, Nakayama K, Murata T, Tanaka M, Tobe T, Iida T, Takami H, Honda T, Sasakawa C, Ogasawara N, Yasunaga T, Kuhara S, Shiba T, Hattori M, Shinagawa H: Complete genome sequence of enterohemorrhagic Escherichia coli O157:H7 and genomic comparison with a laboratory strain K-12. DNA Res. 2001, 8: 11-22.
Parkhill J, Dougan G, James KD, Thomson NR, Pickard D, Wain J, Churcher C, Mungall KL, Bentley SD, Holden MT, Sebaihia M, Baker S, Basham D, Brooks K, Chillingworth T, Connerton P, Cronin A, Davis P, Davies RM, Dowd L, White N, Farrar J, Feltwell T, Hamlin N, Haque A, Hien TT, Holroyd S, Jagels K, Krogh A, Larsen TS, et al: Complete genome sequence of a multiple drug resistant Salmonella enterica serovar Typhi CT18. Nature. 2001, 413: 848-852.
Baba T, Takeuchi F, Kuroda M, Yuzawa H, Aoki K, Oguchi A, Nagai Y, Iwama N, Asano K, Naimi T, Kuroda H, Cui L, Yamamoto K, Hiramatsu K: Genome and virulence determinants of high virulence community-acquired MRSA. Lancet. 2002, 359: 1819-1827.
Fitzgerald JR, Sturdevant DE, Mackie SM, Gill SR, Musser JM: Evolutionary genomics of Staphylococcus aureus: insights into the origin of methicillin-resistant strains and the toxic shock syndrome epidemic. Proc Natl Acad Sci U S A. 2001, 98: 8821-8826.
Hoffmaster AR, Ravel J, Rasko DA, Chapman GD, Chute MD, Marston CK, De BK, Sacchi CT, Fitzgerald C, Mayer LW, Maiden MC, Priest FG, Barker M, Jiang L, Cer RZ, Rilstone J, Peterson SN, Weyant RS, Galloway DR, Read TD, Popovic T, Fraser CM: Identification of anthrax toxin genes in a Bacillus cereus associated with an illness resembling inhalation anthrax. Proc Natl Acad Sci U S A. 2004, 101: 8449-8454.
Fouts DE, Mongodin EF, Mandrell RE, Miller WG, Rasko DA, Ravel J, Brinkac LM, DeBoy RT, Parker CT, Daugherty SC, Dodson RJ, Durkin AS, Madupu R, Sullivan SA, Shetty JU, Ayodeji MA, Shvartsbeyn A, Schatz MC, Badger JH, Fraser CM, Nelson KE: Major structural differences and novel potential virulence mechanisms from the genomes of multiple campylobacter species. PLoS Biol. 2005, 3: e15-
Carlson JH, Porcella SF, McClarty G, Caldwell HD: Comparative genomic analysis of Chlamydia trachomatis oculotropic and genitotropic strains. Infect Immun. 2005, 73: 6407-6418.
Broekhuijsen M, Larsson P, Johansson A, Byström M, Eriksson U, Larsson E, Prior RG, Sjöstedt A, Titball RW, Forsman M: Genome-wide DNA microarray analysis of Francisella tularensis strains demonstrates extensive genetic conservation within the species but identifies regions that are unique to the highly virulent F. tularensis subsp. tularensis . J Clin Microbiol. 2003, 41: 2924-2931.
Carroll IM, Khan AA, Ahmed N: Revisiting the pestilence of Helicobacter pylori: insights into geographical genomics and pathogen evolution. Infect Genet Evol. 2004, 4: 81-90.
Buchrieser C, Rusniok C, Kunst F, Cossart P, Glaser P: Comparison of the genome sequences of Listeria monocytogenes and Listeria innocua: clues for evolution and pathogenicity. FEMS Immunol Med Microbiol. 2003, 35: 207-213.
Behr MA, Sherman DR: Mycobacterial virulence and specialized secretion: same story, different ending. Nat Med. 2007, 13: 286-287.
Smoot JC, Barbian KD, Van Gompel JJ, Smoot LM, Chaussee MS, Sylva GL, Sturdevant DE, Ricklefs SM, Porcella SF, Parkins LD, Beres SB, Campbell DS, Smith TM, Zhang Q, Kapur V, Daly JA, Veasy LG, Musser JM: Genome sequence and comparative microarray analysis of serotype M18 group A Streptococcus strains associated with acute rheumatic fever outbreaks. Proc Natl Acad Sci U S A. 2002, 99: 4668-4673.
Beres SB, Richter EW, Nagiec MJ, Sumby P, Porcella SF, DeLeo FR, Musser JM: Molecular genetic anatomy of inter- and intraserotype variation in the human bacterial pathogen group A Streptococcus . Proc Natl Acad Sci U S A. 2006, 103: 7059-7064.
Brochet M, Couvé E, Zouine M, Vallaeys T, Rusniok C, Lamy MC, Buchrieser C, Trieu-Cuot P, Kunst F, Poyart C, Glaser P: Genomic diversity and evolution within the species Streptococcus agalactiae . Microbes Infect. 2006, 8: 1227-1243.
Fraser CM, Norris SJ, Weinstock GM, White O, Sutton GG, Dodson R, Gwinn M, Hickey EK, Clayton R, Ketchum KA, Sodergren E, Hardham JM, McLeod MP, Salzberg S, Peterson J, Khalak H, Richardson D, Howell JK, Chidambaram M, Utterback T, McDonald L, Artiach P, Bowman C, Cotton MD, Fujii C, Garland S, Hatch B, Horst K, Roberts K, Sandusky M, et al: Complete genome sequence of Treponema pallidum, the syphilis spirochete. Science. 1998, 281: 375-388.
Chain PS, Hu P, Malfatti SA, Radnedge L, Larimer F, Vergez LM, Worsham P, Chu MC, Andersen GL: Complete genome sequence of Yersinia pestis strains Antiqua and Nepal516: evidence of gene reduction in an emerging pathogen. J Bacteriol. 2006, 188: 4453-4463.
Merhej V, Georgiades K, Raoult D: Postgenomic analysis of bacterial pathogens repertoire reveals genome reduction rather than virulence factors. Brief Funct Genomics. 2013, 12: 291-304.
Fournier PE, El Karkouri K, Leroy Q, Robert C, Giumelli B, Renesto P, Socolovschi C, Parola P, Audic S, Raoult D: Analysis of the Rickettsia africae genome reveals that virulence acquisition in Rickettsia species may be explained by genome reduction. BMC Genomics. 2009, 10: 166-
Kaser M, Pluschke G: Differential gene repertoire in Mycobacterium ulcerans identifies candidate genes for patho-adaptation. PLoS Negl Trop Dis. 2008, 2: e353-
Sun YC, Hinnebusch BJ, Darby C: Experimental evidence for negative selection in the evolution of a Yersinia pestis pseudogene. Proc Natl Acad Sci U S A. 2008, 105: 8097-8101.
Priest NK, Rudkin JK, Feil EJ, van den Elsen JM, Cheung A, Peacock SJ, Laabei M, Lucks DA, Recker M, Massey RC: From genotype to phenotype: can systems biology be used to predict Staphylococcus aureus virulence?. Nat Rev Microbiol. 2012, 10: 791-797.
Roca I, Espinal P, Vila-Farres X, Vila J: The Acinetobacter baumannii oxymoron: commensal hospital dweller turned pan-drug-resistant menace. Front Microbiol. 2012, 3: 148-
Sundsfjord A, Simonsen GS, Haldorsen BC, Haaheim H, Hjelmevoll SO, Littauer P, Dahl KH: Genetic methods for detection of antimicrobial resistance. APMIS. 2004, 112: 815-837.
Buriánková K, Doucet-Populaire F, Dorson O, Gondran A, Ghnassia JC, Weiser J, Pernodet JL: Molecular basis of intrinsic macrolide resistance in the Mycobacterium tuberculosis complex. Antimicrob Agents Chemother. 2004, 48: 143-150.
Pillai DR, Shahinas D, Buzina A, Pollock RA, Lau R, Khairnar K, Wong A, Farrell DJ, Green K, McGeer A, Low DE: Genome-wide dissection of globally emergent multi-drug resistant serotype 19A Streptococcus pneumoniae . BMC Genomics. 2009, 10: 642-
Masselot F, Boulos A, Maurin M, Rolain JM, Raoult D: Molecular evaluation of antibiotic susceptibility: Tropheryma whipplei paradigm. Antimicrob Agents Chemother. 2003, 47: 1658-1664.
Ogata H, Renesto P, Audic S, Robert C, Blanc G, Fournier PE, Parinello H, Claverie JM, Raoult D: The genome sequence of Rickettsia felis identifies the first putative conjugative plasmid in an obligate intracellular parasite. PLoS Biol. 2005, 3: e248-
García-Álvarez L, Holden MT, Lindsay H, Webb CR, Brown DF, Curran MD, Walpole E, Brooks K, Pickard DJ, Teale C, Parkhill J, Bentley SD, Edwards GF, Girvan EK, Kearns AM, Pichon B, Hill RL, Larsen AR, Skov RL, Peacock SJ, Maskell DJ, Holmes MA: Meticillin-resistant Staphylococcus aureus with a novel mecA homologue in human and bovine populations in the UK and Denmark: a descriptive study. Lancet Infect Dis. 2011, 11: 595-603.
Paterson GK, Harrison EM, Holmes MA: The emergence of mecC methicillin-resistant Staphylococcus aureus . Trends Microbiol. 2014, 22: 42-47.
Poirel L, Bonnin RA, Nordmann P: Analysis of the resistome of a multidrug-resistant NDM-1-producing Escherichia coli strain by high-throughput genome sequencing. Antimicrob Agents Chemother. 2011, 55: 4224-4229.
Liu P, Li P, Jiang X, Bi D, Xie Y, Tai C, Deng Z, Rajakumar K, Ou HY: Complete genome sequence of Klebsiella pneumoniae subsp. pneumoniae HS11286, a multidrug-resistant strain isolated from human sputum. J Bacteriol. 2012, 194: 1841-1842.
Diene SM, Rolain JM: Investigation of antibiotic resistance in the genomic era of multidrug-resistant Gram-negative bacilli, especially Enterobacteriaceae, Pseudomonas and Acinetobacter . Expert Rev Anti Infect Ther. 2013, 11: 277-296.
Papin JA, Price ND, Wiback SJ, Fell DA, Palsson BO: Metabolic pathways in the post-genome era. Trends Biochem Sci. 2003, 28: 250-258.
Tyson GW, Banfield JF: Cultivating the uncultivated: a community genomics perspective. Trends Microbiol. 2005, 13: 411-415.
Bentley SD, Maiwald M, Murphy LD, Pallen MJ, Yeats CA, Dover LG, Norbertczak HT, Besra GS, Quail MA, Harris DE, von Herbay A, Goble A, Rutter S, Squares R, Squares S, Barrell BG, Parkhill J, Relman DA: Sequencing and analysis of the genome of the Whipple's disease bacterium Tropheryma whipplei . Lancet. 2003, 361: 637-644.
Raoult D, Ogata H, Audic S, Robert C, Suhre K, Drancourt M, Claverie JM: Tropheryma whipplei twist: a human pathogenic Actinobacteria with a reduced genome. Genome Res. 2003, 13: 1800-1809.
Lemos EG, Alves LM, Campanharo JC: Genomics-based design of defined growth media for the plant pathogen Xylella fastidiosa . FEMS Microbiol Lett. 2003, 219: 39-45.
Tyson GW, Lo I, Baker BJ, Allen EE, Hugenholtz P, Banfield JF: Genome-directed isolation of the key nitrogen fixer Leptospirillum ferrodiazotrophum sp. nov. from an acidophilic microbial community. Appl Environ Microbiol. 2005, 71: 6319-6324.
Omsland A, Cockrell DC, Howe D, Fischer ER, Virtaneva K, Sturdevant DE, Porcella SF, Heinzen RA: Host cell-free growth of the Q fever bacterium Coxiella burnetii . Proc Natl Acad Sci U S A. 2009, 106: 4430-4434.
Edwards JS, Ibarra RU, Palsson BO: In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nat Biotechnol. 2001, 19: 125-130.
Ibarra RU, Edwards JS, Palsson BO: Escherichia coli K-12 undergoes adaptive evolution to achieve in silico predicted optimal growth. Nature. 2002, 420: 186-189.
Cole ST, Eiglmeier K, Parkhill J, James KD, Thomson NR, Wheeler PR, Honoré N, Garnier T, Churcher C, Harris D, Mungall K, Basham D, Brown D, Chillingworth T, Connor R, Davies RM, Devlin K, Duthoy S, Feltwell T, Fraser A, Hamlin N, Holroyd S, Hornsby T, Jagels K, Lacroix C, Maclean J, Moule S, Murphy L, Oliver K, Quail MA, et al: Massive gene decay in the leprosy bacillus. Nature. 2001, 409: 1007-1011.
Lewis T, Loman NJ, Bingle L, Jumaa P, Weinstock GM, Mortiboy D, Pallen MJ: High-throughput whole-genome sequencing to dissect the epidemiology of Acinetobacter baumannii isolates from a hospital outbreak. J Hosp Infect. 2010, 75: 37-41.
Mellmann A, Harmsen D, Cummings CA, Zentz EB, Leopold SR, Rico A, Prior K, Szczepanowski R, Ji Y, Zhang W, McLaughlin SF, Henkhaus JK, Leopold B, Bielaszewska M, Prager R, Brzoska PM, Moore RL, Guenther S, Rothberg JM, Karch H: Prospective genomic characterization of the German enterohemorrhagic Escherichia coli O104:H4 outbreak by rapid next generation sequencing technology. PLoS One. 2011, 6: e22751-
Lavezzo E, Toppo S, Franchin E, Di Camillo B, Finotello F, Falda M, Manganelli R, Palù G, Barzon L: Genomic comparative analysis and gene function prediction in infectious diseases: application to the investigation of a meningitis outbreak. BMC Infect Dis. 2013, 13: 554-
Fournier PE, Levy PY, Million M, Croce O, Blanc-Tailleur C, Brouqui P, Raoult D: Genome of a chronic osteitis-causing Clostridium tetani . N Microbes N Infect. 2014, 2: 25-26.
Sherry NL, Porter JL, Seemann T, Watkins A, Stinear TP, Howden BP: Outbreak investigation using high-throughput genome sequencing within a diagnostic microbiology laboratory. J Clin Microbiol. 2013, 51: 1396-1401.
Török ME, Reuter S, Bryant J, Köser CU, Stinchcombe SV, Nazareth B, Ellington MJ, Bentley SD, Smith GP, Parkhill J, Peacock SJ: Rapid whole-genome sequencing for investigation of a suspected tuberculosis outbreak. J Clin Microbiol. 2013, 51: 611-614.
Steensels D, Vankeerberghen A, De BH: Towards multitarget testing in molecular microbiology. Int J Microbiol. 2013, 2013: 121057-
Cho I, Blaser MJ: The human microbiome: at the interface of health and disease. Nat Rev Genet. 2012, 13: 260-270.
Lim YW, Evangelista JS, Schmieder R, Bailey B, Haynes M, Furlan M, Maughan H, Edwards R, Rohwer F, Conrad D: Clinical insights from metagenomic analysis of sputum samples from patients with cystic fibrosis. J Clin Microbiol. 2014, 52: 425-437.
Loman NJ, Constantinidou C, Christner M, Rohde H, Chan JZ, Quick J, Weir JC, Quince C, Smith GP, Betley JR, Aepfelbacher M, Pallen MJ: A culture-independent sequence-based metagenomics approach to the investigation of an outbreak of Shiga-toxigenic Escherichia coli O104:H4. JAMA. 2013, 309: 1502-1510.
Hasman H, Saputra D, Sicheritz-Ponten T, Lund O, Svendsen CA, Frimodt-Møller N, Aarestrup FM: Rapid whole-genome sequencing for detection and characterization of microorganisms directly from clinical samples. J Clin Microbiol. 2014, 52: 139-146.
Kaas RS, Leekitcharoenphon P, Aarestrup FM, Lund O: Solving the problem of comparing whole bacterial genomes across different sequencing platforms. PLoS One. 2014, 9: e104984-
Dugan VG, Emrich SJ, Giraldo-Calderón GI, Harb OS, Newman RM, Pickett BE, Schriml LM, Stockwell TB, Stoeckert CJ, Sullivan DE, Singh I, Ward DV, Yao A, Zheng J, Barrett T, Birren B, Brinkac L, Bruno VM, Caler E, Chapman S, Collins FH, Cuomo CA, Di Francesco V, Durkin S, Eppinger M, Feldgarden M, Fraser C, Fricke WF, Giovanni M, Henn MR, et al: Standardized metadata for human pathogen/vector genomic sequences. PLoS One. 2014, 9: e99979-
Naccache SN, Federman S, Veeraraghavan N, Zaharia M, Lee D, Samayoa E, Bouquet J, Greninger AL, Luk KC, Enge B, Wadford DA, Messenger SL, Genrich GL, Pellegrino K, Grard G, Leroy E, Schneider BS, Fair JN, Martínez MA, Isa P, Crump JA, DeRisi JL, Sittler T, Hackett J, Miller S, Chiu CY: A cloud-compatible bioinformatics pipeline for ultrarapid pathogen identification from next-generation sequencing of clinical samples. Genome Res. 2014, 24: 1180-1192.
Köser CU, Fraser LJ, Ioannou A, Becq J, Ellington MJ, Holden MT, Reuter S, Török ME, Bentley SD, Parkhill J, Gormley NA, Smith GP, Peacock SJ: Rapid single-colony whole-genome sequencing of bacterial pathogens. J Antimicrob Chemother. 2014, 69: 1275-1281.
Lasken RS, McLean JS: Recent advances in genomic DNA sequencing of microbial species from single cells. Nat Rev Genet. 2014, 15: 577-584.
McLean JS: Advancements toward a systems level understanding of the human oral microbiome. Front Cell Infect Microbiol. 2014, 4: 98-
Theofiles AG, Cunningham SA, Chia N, Jeraldo PR, Quest DJ, Mandrekar JN, Patel R: Pertussis outbreak, southeastern Minnesota, 2012. Mayo Clin Proc. 2014, 89: 1378-1388.
Eyre DW, Cule ML, Wilson DJ, Griffiths D, Vaughan A, O'Connor L, Ip CL, Golubchik T, Batty EM, Finney JM, Wyllie DH, Didelot X, Piazza P, Bowden R, Dingle KE, Harding RM, Crook DW, Wilcox MH, Peto TE, Walker AS: Diverse sources of C. difficile infection identified on whole-genome sequencing. N Engl J Med. 2013, 369: 1195-1205.
Reuter S, Ellington MJ, Cartwright EJ, Köser CU, Török ME, Gouliouris T, Harris SR, Brown NM, Holden MT, Quail M, Parkhill J, Smith GP, Bentley SD, Peacock SJ: Rapid bacterial whole-genome sequencing to enhance diagnostic and public health microbiology. JAMA Intern Med. 2013, 173: 1397-1404.
Rohde H, Qin J, Cui Y, Li D, Loman NJ, Hentschke M, Chen W, Pu F, Peng Y, Li J, Xi F, Li S, Li Y, Zhang Z, Yang X, Zhao M, Wang P, Guan Y, Cen Z, Zhao X, Christner M, Kobbe R, Loos S, Oh J, Yang L, Danchin A, Gao GF, Song Y, Li Y, Yang H, et al: Open-source genomic analysis of Shiga-toxin-producing E. coli O104:H4. N Engl J Med. 2011, 365: 718-724.
Rasko DA, Webster DR, Sahl JW, Bashir A, Boisen N, Scheutz F, Paxinos EE, Sebra R, Chin CS, Iliopoulos D, Klammer A, Peluso P, Lee L, Kislyuk AO, Bullard J, Kasarskis A, Wang S, Eid J, Rank D, Redman JC, Steyert SR, Frimodt-Møller J, Struve C, Petersen AM, Krogfelt KA, Nataro JP, Schadt EE, Waldor MK: Origins of the E. coli strain causing an outbreak of hemolytic-uremic syndrome in Germany. N Engl J Med. 2011, 365: 709-717.
Johansson A, Lärkeryd A, Widerström M, Mörtberg S, Myrtännäs K, Ohrman C, Birdsell D, Keim P, Wagner DM, Forsman M, Larsson P: An outbreak of respiratory tularemia caused by diverse clones of Francisella tularensis . Clin Infect Dis. 2014, 59: 1546-1553.
Stoesser N, Giess A, Batty EM, Sheppard AE, Walker AS, Wilson DJ, Didelot X, Bashir A, Sebra R, Kasarskis A, Sthapit B, Shakya M, Kelly D, Pollard AJ, Peto TE, Crook DW, Donnelly P, Thorson S, Amatya P, Joshi S: Genome sequencing of an extended series of NDM-Klebsiella pneumoniae neonatal infections in a nepali hospital characterizes the extent of community versus hospital-associated transmission in an endemic setting. Antimicrob Agents Chemother. 2014, 58: 7347-7357.
Lévesque S, Plante PL, Mendis N, Cantin P, Marchand G, Charest H, Raymond F, Huot C, Goupil-Sormany I, Desbiens F, Faucher SP, Corbeil J, Tremblay C: Genomic characterization of a large outbreak of Legionella pneumophila serogroup 1 strains in Quebec City, 2012. PLoS One. 2014, 9: e103852-
Gilmour MW, Graham M, Van Domselaar G, Tyler S, Kent H, Trout-Yakel KM, Larios O, Allen V, Lee B, Nadon C: High-throughput genome sequencing of two Listeria monocytogenes clinical isolates during a large foodborne outbreak. BMC Genomics. 2010, 11: 120-
Bryant JM, Grogono DM, Greaves D, Foweraker J, Roddick I, Inns T, Reacher M, Haworth CS, Curran MD, Harris SR, Peacock SJ, Parkhill J, Floto RA: Whole-genome sequencing to identify transmission of Mycobacterium abscessus between patients with cystic fibrosis: a retrospective cohort study. Lancet. 2013, 381: 1551-1560.
Gardy JL, Johnston JC, Ho Sui SJ, Cook VJ, Shah L, Brodkin E, Rempel S, Moore R, Zhao Y, Holt R, Varhol R, Birol I, Lem M, Sharma MK, Elwood K, Jones SJ, Brinkman FS, Brunham RC, Tang P: Whole-genome sequencing and social-network analysis of a tuberculosis outbreak. N Engl J Med. 2011, 364: 730-739.
Jolley KA, Hill DM, Bratcher HB, Harrison OB, Feavers IM, Parkhill J, Maiden MC: Resolution of a meningococcal disease outbreak from whole-genome sequence data with rapid Web-based analysis methods. J Clin Microbiol. 2012, 50: 3046-3053.
Byrne L, Fisher I, Peters T, Mather A, Thomson N, Rosner B, Bernard H, McKeown P, Cormican M, Cowden J, Aiyedun V, Lane C: A multi-country outbreak of Salmonella Newport gastroenteritis in Europe associated with watermelon from Brazil, confirmed by whole genome sequencing: October 2011 to January 2012. Euro Surveill. 2014, 19: 6-13.
den Bakker HC, Allard MW, Bopp D, Brown EW, Fontana J, Iqbal Z, Kinney A, Limberger R, Musser KA, Shudt M, Strain E, Wiedmann M, Wolfgang WJ: Rapid whole-genome sequencing for surveillance of Salmonella enterica serovar enteritidis. Emerg Infect Dis. 2014, 20: 1306-1314.
Harris SR, Cartwright EJ, Török ME, Holden MT, Brown NM, Ogilvy-Stuart AL, Ellington MJ, Quail MA, Bentley SD, Parkhill J, Peacock SJ: Whole-genome sequencing for analysis of an outbreak of meticillin-resistant Staphylococcus aureus: a descriptive study. Lancet Infect Dis. 2013, 13: 130-136.
Long SW, Beres SB, Olsen RJ, Musser JM: Absence of patient-to-patient intrahospital transmission of Staphylococcus aureus as determined by whole-genome sequencing. MBio. 2014, 5: e01692-14.
Rebaudet S, Mengel MA, Koivogui L, Moore S, Mutreja A, Kande Y, Yattara O, Sarr Keita V, Njanpop-Lafourcade BM, Fournier PE, Garnotel E, Keita S, Piarroux R: Deciphering the origin of the 2012 cholera epidemic in Guinea by integrating epidemiological and molecular analyses. PLoS Negl Trop Dis. 2014, 8: e2898-
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Fournier, PE., Dubourg, G. & Raoult, D. Clinical detection and characterization of bacterial pathogens in the genomics era. Genome Med 6, 114 (2014). https://doi.org/10.1186/s13073-014-0114-2
Published:
DOI: https://doi.org/10.1186/s13073-014-0114-2