Abstract
Background
There is evidence that the knockdown resistance gene (Kdr) L1014F and acetylcholinesterase-1 gene (Ace-1R) G119S mutations involved in pyrethroid and carbamate resistance in Anopheles gambiae influence malaria transmission in sub-Saharan Africa. This is likely due to changes in the behaviour, life history and vector competence and capacity of An. gambiae. In the present study, performed as part of a two-arm cluster randomized controlled trial evaluating the impact of household screening plus a novel insecticide delivery system (In2Care Eave Tubes), we investigated the distribution of insecticide target site mutations and their association with infection status in wild An. gambiae sensu lato (s.l.) populations.
Methods
Mosquitoes were captured in 40 villages around Bouaké by human landing catch from May 2017 to April 2019. Randomly selected samples of An. gambiae s.l. that were infected or not infected with Plasmodium sp. were identified to species and then genotyped for Kdr L1014F and Ace-1R G119S mutations using quantitative polymerase chain reaction assays. The frequencies of the two alleles were compared between Anopheles coluzzii and Anopheles gambiae and then between infected and uninfected groups for each species.
Results
The presence of An. gambiae (49%) and An. coluzzii (51%) was confirmed in Bouaké. Individuals of both species infected with Plasmodium parasites were found. Over the study period, the average frequency of the Kdr L1014F and Ace-1R G119S mutations did not vary significantly between study arms. However, the frequencies of the Kdr L1014F and Ace-1R G119S resistance alleles were significantly higher in An. gambiae than in An. coluzzii [odds ratio (95% confidence interval): 59.64 (30.81–131.63) for Kdr, and 2.79 (2.17–3.60) for Ace-1R]. For both species, there were no significant differences in Kdr L1014F or Ace-1R G119S genotypic and allelic frequency distributions between infected and uninfected specimens (P > 0.05).
Conclusions
Either alone or in combination, Kdr L1014F and Ace-1R G119S showed no significant association with Plasmodium infection in wild An. gambiae and An. coluzzii, demonstrating the similar competence of these species for Plasmodium transmission in Bouaké. Additional factors including behavioural and environmental ones that influence vector competence in natural populations, and those other than allele measurements (metabolic resistance factors) that contribute to resistance, should be considered when establishing the existence of a link between insecticide resistance and vector competence.
Graphical Abstract

Similar content being viewed by others
Background
Mosquitoes of the Anopheles gambiae species complex are the main malaria vectors in sub-Saharan Africa [1]. The remarkable vector capacity of these mosquitoes [2] is largely due to their propensity to blood feed on humans and rest indoors [3]. The great ability of these mosquitoes to adapt to human behaviour has led to the development of insecticide-based vector control measures targeting indoor biting and resting. These measures primarily comprise the use of long-lasting insecticidal nets and indoor residual spraying, which are used to limit human-vector contact and reduce mosquito survival [4]. These insecticide-based vector control tools have been highly effective against malaria vectors, as shown by considerable reductions in disease burden [5]. However, the long-term effectiveness of both of these strategies is threatened by the emergence of insecticide resistance in malaria vector populations [6, 7].
There are several mechanisms responsible for insecticide resistance, of which metabolic and target site resistance are the most common [8,9,10]. Metabolic resistance leads to an increase in the activities of enzymes responsible for an insecticide’s degradation, while modification of the insecticide target site prevents the insecticide molecule from binding to the site. The molecular basis of resistance mediated by target site mutations has been characterized for several mosquito populations [11,12,13]. For example, the G119S mutation in the acetylcholinesterase-1 gene (Ace-1R) (a single amino acid substitution from glycine to serine at locus 119 at the acetylcholinesterase catalytic site) is responsible for organophosphate and carbamate resistance among malaria vectors in West Africa [14]. Likewise, the L1014F mutation of the knockdown resistance (Kdr) gene, also called the Kdr-west mutation (an amino acid substitution from leucine to phenylalanine in the voltage gated sodium channel gene, at the 1014 locus, typically causing knock down resistance) is responsible for pyrethroid and dichlorodiphenyltrichloroethane resistance in mosquito populations [12].
Despite the rise of insecticide resistance, its operational significance for vector control is controversial. In many instances, insecticide-based tools seem to continue to protect against malaria [15,16,17,18], whereas a community trial of long-lasting insecticidal nets clearly demonstrated that resistance is having an impact on their effectiveness [19]. Resistance is dynamic and therefore cannot be randomized to assess its epidemiological impact. Several studies have evaluated the association between single insecticide resistance gene mutations (of Kdr or Ace-1R) and vector competence in An. gambiae [20,21,22]. However, these involved laboratory assays utilizing mosquito colonies or wild strains infected with malaria parasites in the laboratory. The coexistence of both Kdr and Ace-1R in wild populations of An. gambiae sensu lato (s.l.) is common in west Africa, including Côte d’Ivoire [23, 24]. To our knowledge, the impact of this association on vector competence has never been studied.
We took advantage of a two-arm cluster randomized controlled trial evaluating the impact of household screening plus a novel insecticide delivery system (In2Care Eave Tubes) to capture mosquitoes in study villages around Bouaké by human landing catches, between May 2017 and April 2019. Mosquitoes were identified to species and then genotyped for Kdr L1014F and Ace-1R G119S mutations using quantitative polymerase chain reaction (qPCR) assays, and the frequencies of the two alleles were compared between Anopheles coluzzii and Anopheles gambiae and then between infected and uninfected groups for each species.
Methods
Study area
The trial was conducted from May 2017 to April 2019 in central Côte d’Ivoire. The methodology used in this study has been well described by Sternberg et al. [25]. Briefly, 40 villages within a 60-km radius in the district of Bouaké were identified for inclusion in the study. All the households in the 40 study villages received insecticide-treated nets, while those of half of the study villages (20 villages) also had household screening (S) and In2Care Eave Tubes (ET) installed (SET).
Mosquito collection and processing
The mosquito-collection process has been previously described by Sternberg et al. [25]. Each month during the trial, mosquitoes were sampled by human landing catches (HLC) both indoors and outdoors at four randomly selected houses in each of the 40 study villages. HLC were undertaken from 6 p.m. to 8 a.m. the following day for two consecutive nights during the first 5 months of the trial and then on one night per month until the end of the trial. The collected mosquitoes were sorted and morphologically identified to species using the key described by Gillies and Meillon [26] and counted. All malaria vectors were stored for further analysis, but for the interaction study, only An. gambiae s.l., the main malaria vector in Côte d’Ivoire, was considered.
DNA extraction
PCR assays were used to assess sporozoite prevalence in a monthly random sub-sample of up to 30 female mosquitoes per village. Mosquitoes were identified to sibling species and Kdr L1014F and Ace-1R G119S mutations detected. Genomic DNA was extracted from the head and thorax of individual females using cetyltrimethylammonium bromide, as described by Yahouedo et al. [27].
Detection of Plasmodium infection
Plasmodium spp. (Plasmodium malariae, Plasmodium falciparum, Plasmodium ovale and Plasmodium vivax) infections were detected by real-time PCR in accordance with Mangold et al. [28]. The primers were synthesized and supplied by Eurofins Genomics (Ebersberg, Germany) and were as follows: forward PL1473F18 (5′-TAA CGA AGA ACG TCT TAA-3′) and reverse PL1679R18 (5′-GTT CCT CTA AGA AGC TTT-3′). The reactions were prepared in a total reaction volume of 10 μl, which contained 2 μl of 5× HOT FIREPol EvaGreen qPCR Mix Plus (Solis Biodyne, Tartu, Estonia), 0.3 μl of each primer, 6.4 μl of sterile water, and 1 μl of DNA template. The real-time PCR mixtures were pre-incubated at 95 °C for 12 min followed by amplification for 50 cycles of 10 s at 95 °C, 5 s at 50 °C and 20 s at 72 °C, with fluorescence acquisition at the end of each cycle. Characterisation of the PCR product was performed with melting curve analysis of the amplicons (95 °C for 60 s, 60 °C for 60 s, then 60–90 °C for 1 s), with fluorescence acquisition at each temperature transition. Plasmodium species were identified by melting curve generated at different temperatures (i.e., for P. malariae, 73.5–75.5 °C; for P. falciparum, 75.5–77.5 °C; for P. ovale, 77.5–79.5 °C; and for P. vivax:, 79.5–81.5 °C).
Species identification
A subsample of 1392 An. gambiae s.l. (686 infected with Plasmodium sp. and 706 uninfected, which were randomly selected) was analysed for molecular identification of sibling species. The molecular identification was performed using a classic PCR assay in accordance with Favia et al. [29]. The following primers were used: R3 (5’-GCC AAT CCG AGC TGA TAG CGC-3’), R5 (5’-CGA ATT CTA GGG AGC TCC AG-3’), Mopint (5’-GCC CCT TCC TCG ATG GCA T-3’) and B/Sint (5’-ACC AAG ATG GTT CGT TGC-3’). The reaction mixture consisted of 14 μl of sterile water, 0.75 μl of each primer R3 and R5, 1.5 μl of each primer Mopint and B/Sint, and 5 µl of Master Mix. A 23.5-µl volume of the reaction mixture was inserted into each 0.5-ml PCR tube along with 1 µl of each DNA sample. Amplification was performed on a MJ Research PTC-100 Thermal Cycler PCR machine (Marshall Scientific, Watertown, MA) with cycling conditions of 95 °C for 3 min, followed by 30 cycles at 95 °C for 30 s, 72 °C for 45 s and 72 °C for 60 s. Amplified fragments were analysed on 2% agarose gel with 4 μl of SYBR Green. The results were analysed as described in Favia et al. [29] to determine An. coluzzii [1300-bp band (R3/R5) plus 727-bp band (Mop-int)] or An. gambiae [1300-bp band (R3/R5) plus 475-bp band (B/S-int)].
Detection of Kdr L1014F mutation in An. gambiae s.l.
Detection of the Kdr L1014F mutation was performed using the TaqMan real-time PCR assay, as described by Bass et al. [30]. The reactions were carried out in a total reaction volume of 10 μl, which contained 2 μl of 5× HOT FIREPol Probe Universal qPCR Mix (Solis Biodyne), 0.125 µl primer/probe mix, 6.875 μl of sterile water, and 1 μl of DNA template.
Primers Kdr-forward (5'-CATTTTTCTTGGCCACTGTAGTGAT-3') and Kdr-reverse (5'-CGATCTTGGTCCATGTTAATTTGCA-3') were standard oligonucleotides with no modification. The probes were labelled with two distinct fluorophores: VIC to detect the susceptible allele, and FAM to detect the resistant allele. Amplifications were performed on a LightCycler 96 Systems real-time qPCR machine (Roche LifeScience, Meylan, France) with cycling conditions of 95 °C for 10 min, followed by 45 cycles at 95 °C for 10 s, 60 °C for 45 s and 72 °C for 1 s. FAM and VIC fluorescence was captured at the end of each cycle and genotypes were called from endpoint fluorescence using LightCycler 96 software (Roche LifeScience) for the analysis of the results.
Detection of Ace-1 R G119S mutation in An. gambiae s.l.
Allelic and genotypic frequencies for insensitive acetylcholinesterase phenotypes characterized by the G119S mutation were determined for An. gambiae s.l. by using the TaqMan assay, in accordance with Bass et al. [31]. The reactions were carried out in a total reaction volume of 10 μl, which contained 2 μl of the 5× HOT FIREPol Probe Universal qPCR Mix (Solis Biodyne), 0.125 µl primer/probe mix, 6.875 μl of sterile water, and 1 μl of DNA template. Primers Ace-1-Forward (5’-GGC CGT CAT GCT GTG GAT-3’), and Ace-1-Reverse (5’-GCG GTG CCG GAG TAG A-3’) were standard oligonucleotides with no modification. The probes were labelled with two distinct fluorophores: VIC to detect the susceptible allele and FAM to detect the resistant allele. Amplifications were performed on a LightCycler 96 Systems real-time qPCR machine (Roche LifeScience) with cycling conditions of 95 °C for 10 min, followed by 55 cycles at 92 °C for 15 s, 60 °C for 60 s and 72 °C for 1 s. FAM and VIC fluorescence was captured at the end of each cycle and genotypes were called from endpoint fluorescence using LightCycler 96 software (Roche LifeScience) for the analysis of the results.
Statistical analysis
To analyse the distribution of Kdr L1014F and Ace-1R G119S genotypic and allelic frequencies, data collected for the same study arm between May 2017 and April 2019 were compared between the species. The association between genotypic and allelic frequencies for these mutations and infection status were determined using Pearson’s chi-square test in R (version 4.0.3). Kdr L1014F and Ace-1R G119S combined genotypic distribution frequencies within infection status for each species were also included. Fisher’s exact test was used when the number of individual samples available for a test was less than 30. The significance threshold was set at 5%. Odds ratio (OR) was computed to assess the strength of difference or association between resistance alleles and infection status. The allelic frequencies were tested to Hardy–Weinberg equilibrium (HWE) conformity using the exact HW test, and were calculated as follows:
where RR indicates the resistant homozygous genotype, RS the heterozygous genotype, and SS the susceptible homozygous genotype. Nota bene: Kdr L1014F and Ace-1R G119S mutations each comprise three genotypes expressing different allelic variants on the targeted loci. The resistant (R) and susceptible (S) alleles are possible versions of these genes.
Ethical clearance
Ethical approval was obtained from the ethics committee of the Côte d’Ivoire Ministry of Health (reference 039/MSLS/CNER-dkn), the Pennsylvania State University Human Research Protection Program under the Office for Research Protections (references STUDY00003899 and STUDY00004815), and the ethical review board of the London School of Hygiene and Tropical Medicine (no. 11223). Verbal and written informed consent, using the language spoken locally, was obtained from all the participants (mosquito collectors and head of each household) prior to their enrolment in the study. Mosquito collectors were vaccinated against yellow fever, and the project provided treatment of confirmed malaria cases free of charge for any study participant, in accordance with national policies.
Results
Genotypic and allelic frequency distributions of Kdr L1014F and Ace-1 R G119S mutations in An. gambiae s.l.
Out of the 1392 mosquitoes analysed by PCR, 1255 were successfully identified to species (< 10% failure rate). Both An. gambiae (n = 624; 49.7%) and An. coluzzii (n = 631; 50.3%) were found. For each species, the proportions of infected vs uninfected individuals were similar (Fig. 1). There were no significant differences in the allelic frequency of Kdr or Ace-1R between the control and Eave Tube areas for each species (P ˃ 0.05) (Table 1). Genotypic and allelic frequencies of Kdr L1014F and Ace-1R G119S mutations for An. coluzzii and An. gambiae are shown in Table 2. Kdr allelic frequency was significantly greater in An. gambiae than in An. coluzzii [OR (95% confidence interval; 95% CI): 59.64 (30.81–131.63)] (Table 2). By contrast, the frequency of heterozygous individuals was significantly higher for An. coluzzii (42.95%) than for An. gambiae (1.12%), indicating deviation from HWE in the An. gambiae populations with an excess of resistant homozygous genotypes (Table 2) (P < 0.001).
The allelic frequency of the Ace-1R G119S mutation was low in both An. coluzzii and An. gambiae, although it was significantly more prevalent in An. gambiae than in An. coluzzii [OR (95% CI): 2.79 (2.17–3.60)]. Deviation from HWE for Ace-1R G119S was observed for both An. gambiae and An. coluzzii populations.
Insecticide-resistance genes and infection status
The genotypic and allelic frequencies of Kdr L1014F and Ace-1R G119S gene mutations among infected and uninfected mosquitoes are shown in Table 3. Regardless of the species, there were no significant differences in genotypic or allelic frequencies between infected and uninfected individuals (P ˃ 0.05) (Table 3).
Frequencies of Kdr and Ace-1 R genotypic combinations and infection status
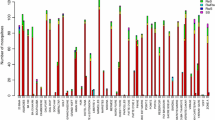
Nine possible genotypic combinations for the Kdr L1014F and Ace-1R G119S mutations were recorded in this study (Fig. 2). For all genotypic combinations, the first two alleles refer to Kdr genotypes whereas the last two alleles refer to Ace-1R genotypes: Kdr-Ace-1R (RRRR), Kdr-Ace-1R (RRRS), Kdr-Ace-1R (RRSS), Kdr-Ace-1R (RSRR), Kdr-Ace-1R (RSRS), Kdr-Ace-1R (RSSS), Kdr-Ace-1R (SSRR), Kdr-Ace-1R (SSRS), and Kdr-Ace-1R (SSSS). Figure 2 shows that the frequency of individuals bearing Kdr RR genotypes, either when present alone or together with Ace-1R genotypes, was significantly higher in wild An. gambiae than in wild An. coluzzii; this was observed in both control and SET areas. By contrast, the frequencies of mosquitoes bearing the Kdr heterozygous genotype were significantly higher for An. coluzzii than for An. gambiae, confirming that the former species is better adapted to insecticide pressure than the latter one (Fig. 2). Overall, there were no significant differences between infected and uninfected groups for each of the genotypic combinations for An. coluzzii or An. gambiae.
Frequencies of Kdr and Ace-1R genotypic combinations between infected and uninfected groups in each study arm. Error bars represent 95% CIs. For all combined genotypes, the first two alleles refer to Kdr genotypes and the last two refer to Ace-1R genotypes. RR Resistant homozygous genotype, RS heterozygous genotype, SS susceptible homozygous genotype; for other abbreviations, see Fig. 1
Discussion
This study evaluated the effects of the Kdr L1014F and Ace-1R G119S gene mutations on Plasmodium spp. infection status in natural An. gambiae s.l. populations. The presence of both An. coluzzii and An. gambiae in similar proportions in this longitudinal study was consistent with the results of previous studies carried out in the area of Bouaké [24, 32], but it contrasts with the results of another study conducted in adjacent areas within Bouaké that found An. coluzzii to be predominant [33]. The observed difference is likely due to the study sampling period covering both the rainy and dry seasons in our study compared to the rainy season only in the other study [33]. We observed no difference in infection rate between An. gambiae and An. coluzzii. This aligns with the results of previous studies conducted in Burkina Faso and Senegal [21, 34], which reported equivalent susceptibility of these species to Plasmodium. The results presented here demonstrate that these sibling species are equally competent vectors of malaria in humans in the central region of Côte d'Ivoire.
With regard to resistance genes, there were no significant differences in the allelic frequency of Kdr or Ace-1R between the control and Eave Tube areas regardless of mosquito species. This is because Kdr was already close to fixation (> 80%) in An. gambiae s.l. species prior to the intervention employing the Eave Tubes [24], leaving a tiny window for further selection. Also, the insecticide deployed in the Eave Tube trial was a pyrethroid (β-cyfluthrin) [35] which could not induce selection pressure on Ace-1R since this gene is associated with organophosphate and carbamate resistance [14, 24].
We found significantly higher Kdr L1014F and Ace1R G119S genotypic and allelic frequencies in An. gambiae than in An. coluzzii, which was in agreement with observations of Koukpo et al. [36] in Benin and Zogo et al. [37] in Côte d’Ivoire. There was a 59 times greater probability of encountering the Kdr L1014F resistance allele in An. gambiae than in An. coluzzii, whereas the frequency of individuals heterozygous for Kdr L1014F was higher for An. coluzzii (42.95%) than for An. gambiae (1.12%). These results clearly highlight a deviation from HWE within both malaria vector species for the Kdr L1014F mutation. It is possible that evolutionary factors affect mosquito population structure through the excessive use of insecticides. These factors induce the selection of rare and existing mutations in natural populations of both species which later become variably widespread [38].
Furthermore, Ace-1R G119S allelic frequency was significantly higher in An. gambiae than in An. coluzzii, although the amplitude was moderate. The low proportion (< 10%) of homozygous resistant (RR) genotypes observed in An. gambiae and An. coluzzii populations could indicate a high fitness cost associated with the Ace-1R G119S gene [39, 40]. Conversely, this potential fitness cost associated with Ace-1R may be counteracted by the duplication of this gene, which induced various heterozygous genotypes by increasing their proportions [41]. Further studies focusing on Ace-1R genotype distribution, including duplication in An. gambiae s.l., are needed. Our study showed that in areas where Kdr L1014F and Ace-1R G119S coexist in An. gambiae s.l., the frequency of individuals bearing the Kdr L1014F RR genotype appeared to be significantly higher for An. gambiae than for An. coluzzii. By contrast, the frequencies of those bearing the Kdr L1014F heterozygous genotype were significantly higher for An. coluzzii than for An. gambiae, confirming the trend seen when this genotype is present in isolation. To our knowledge, this is the first study to evaluate the distribution of An. gambiae s.l. individuals bearing both of these mutations. The results presented here call for further studies to better understand the genotypic structure of their combinations.
The vector competence in association with resistance genes was investigated. We found no evidence of an association between Plasmodium infection status and Kdr L1014F or Ace-1R G119S gene mutations. These results are similar to those found in a study undertaken in Guinea where these target site mutations (Kdr L1014F or Ace-1R G119S) were not associated with Plasmodium infection in wild An. gambiae [42], but phenotypic resistance was rather associated with infection. By contrast, a study in Tanzania found a link between Kdr-east and Plasmodium infection in wild An. gambiae [43].
The lack of an association between Plasmodium infection status and resistance genes under natural conditions contrasts with the findings of several other studies, which reported that resistance-associated genes affect vector competence for the transmission of Plasmodium parasites [20, 21, 44]. There are three possible reasons for these differences. First, these contrasting results could derive from studies that used colonies maintained in the laboratory over years, which can decrease resistance, including a loss of genetic diversity [45, 46]. Second, some genetic susceptibility studies do not take into account additional factors that influence competence in natural vector populations, e.g. mosquito blood-feeding rate, age at infection, longevity, and exposure to an insecticide and to other pathogens that could influence mosquito immune status [47,48,49,50,51]. A natural infection study also implies that the effects of ecology and behaviour on vector competence have been assessed [52, 53]. Third, resistance involves mutations and metabolic components with different functions; therefore studying one in isolation from the other may not be representative of phenotypic resistance. The absence of an association between a combination of genotypes (Kdr L1014F-Ace-1R G119S) and infection status in An. coluzzii or An. gambiae needs to be considered further in the context of control programmes, given that this is now a common observation in many parts of west Africa [13, 24].
Conclusions
We found no significant association between the Kdr L1014F and Ace-1R G119S mutations, when present alone or together, and infection status in wild An. gambiae and An. coluzzii, which demonstrates the similar competence of these species for Plasmodium transmission within areas of Bouaké. However, the frequencies of the Kdr and Ace-1R genotypes and alleles were significantly higher in An. gambiae than in An. coluzzii. Additional factors that influence vector competence in natural vector populations and measurements of factors besides alleles or genotypes that contribute to resistance should be considered when investigating the existence of a link between insecticide resistance and vector competence.
Availability of data and materials
The data supporting the conclusions of this manuscript are included within the manuscript and are available from the corresponding author on reasonable request.
Abbreviations
- Ace-1 R :
-
Acetylcholinesterase-1 gene
- CI:
-
Confidence interval
- HWE:
-
Hardy–Weinberg equilibrium
- Kdr :
-
Knockdown resistance gene
- OR:
-
Odds ratio
- qPCR:
-
Quantitative polymerase chain reaction
- SET:
-
Screening plus In2Care Eave Tubes
- SNP:
-
Single nucleotide polymorphism
References
Sinka ME, Bangs MJ, Manguin S, Rubio-Palis Y, Chareonviriyaphap T, Coetzee M, et al. A global map of dominant malaria vectors. Parasit Vectors. 2012;5:69. https://doi.org/10.1186/1756-3305-5-69.
Garrett-Jones C, Shidrawi GR. Malaria vectorial capacity of a population of Anopheles gambiae. Bull WHO. 1969;40:531–545. https://apps.who.int/iris/handle/10665/267721.
Coluzzi M, Sabatini A, Petrarca V, Di Deco MA. Chromosomal differentiation and adaptation to human environments in the Anopheles gambiae complex. Trans R Soc Trop Med Hyg. 1979;73:483–97. https://doi.org/10.1016/0035-9203(79)90036-1.
Zaim M, Aitio A, Nakashima N. Safety of pyrethroid-treated mosquito nets. Med Vet Entomol. 2000;14:1–5. https://doi.org/10.1046/j.1365-2915.2000.00211.x.
Bhatt S, Weiss DJ, Cameron E, Bisanzio D, Mappin B, Dalrymple U, et al. The effect of malaria control on Plasmodium falciparum in Africa between 2000 and 2015. Nature. 2015;526:207–11. https://doi.org/10.1038/nature15535.
Ranson H, N’Guessan R, Lines J, Moiroux N, Nkuni Z, Corbel V. Pyrethroid resistance in African anopheline mosquitoes: what are the implications for malaria control? Trends Parasitol. 2011;27:91–8. https://doi.org/10.1016/j.pt.2010.08.004.
Strode C, Donegan S, Garner P, Enayati AA, Hemingway J. The impact of pyrethroid resistance on the efficacy of insecticide-treated bed nets against African anopheline mosquitoes: systematic review and meta-analysis. PLoS Med. 2014;11: e1001619. https://doi.org/10.1371/journal.pmed.1001619.
Mitchell SN, Rigden DJ, Dowd AJ, Lu F, Wilding CS, Weetman D, et al. Metabolic and target-site mechanisms combine to confer strong DDT resistance in Anopheles gambiae. PLoS ONE. 2014;9: e92662. https://doi.org/10.1371/journal.pone.0092662.
Chouaïbou M, Zivanovic GB, Knox TB, Jamet HP, Bonfoh B. Synergist bioassays: a simple method for initial metabolic resistance investigation of field Anopheles gambiae s.l. populations. Acta Trop. 2014;130:108–11. https://doi.org/10.1016/j.actatropica.2013.10.020.
Stevenson BJ, Bibby J, Pignatelli P, Muangnoicharoen S, O’Neill PM, Lian L-Y, et al. Cytochrome P450 6M2 from the malaria vector Anopheles gambiae metabolizes pyrethroids: sequential metabolism of deltamethrin revealed. Insect Biochem Mol Biol. 2011;41:492–502. https://doi.org/10.1016/j.ibmb.2011.02.003.
Chouaïbou M, Kouadio FB, Tia E, Djogbenou L. First report of the East African kdr mutation in an Anopheles gambiae mosquito in Côte d’Ivoire. Wellcome Open Res. 2017; 2:8. https://doi.org/10.12688/wellcomeopenres.10662.1.
Martinez-Torres D, Chandre F, Williamson MS, Darriet F, Berge JB, Devonshire AL, et al. Molecular characterization of pyrethroid knockdown resistance (Kdr) in the major malaria vector Anopheles gambiae s.s. Insect Mol Biol. 1998;7:179–84. https://doi.org/10.1046/j.1365-2583.1998.72062.x.
Dabiré RK, Namountougou M, Diabaté A, Soma DD, Bado J, Toé HK, et al. Correction: distribution and frequency of kdr mutations within Anopheles gambiae s.l. populations and first report of the Ace-1 G119S mutation in Anopheles arabiensis from Burkina Faso (West Africa). PLoS ONE. 2015;10:e0141645. https://doi.org/10.1371/journal.pone.0101484.
Essandoh J, Yawson AE, Weetman D. Acetylcholinesterase (Ace-1) target site mutation 119S is strongly diagnostic of carbamate and organophosphate resistance in Anopheles gambiae s.s. and Anopheles coluzzii across southern Ghana. Malaria J. 2013;12:404. https://doi.org/10.1186/1475-2875-12-404.
Cook J, Hergott D, Phiri W, Rivas MR, Bradley J, Segura L, et al. Trends in parasite prevalence following 13 years of malaria interventions on Bioko island, Equatorial Guinea: 2004–2016. Malaria J. 2018;17:62. https://doi.org/10.1186/s12936-018-2213-9.
Dossou-Yovo J, Guillet P, Rogier C, Chandre F, Carnevale P, Assi S-B, et al. Protective efficacy of lambda-cyhalothrin treated nets in Anopheles gambiae pyrethroid resistance areas of Côte d’Ivoire. Am J Trop Med Hyg. 2005;73:859–64. https://doi.org/10.4269/ajtmh.2005.73.859.
Kleinschmidt I, Bradley J, Knox TB, Mnzava AP, Kafy HT, Mbogo C, et al. Implications of insecticide resistance for malaria vector control with long-lasting insecticidal nets: a WHO-coordinated, prospective, international, observational cohort study. Lancet Infect Dis. 2018;18:640–9. https://doi.org/10.1016/S1473-3099(18)30172-5.
Tokponnon FT, Sissinto Y, Ogouyémi AH, Adéothy AA, Adechoubou A, Houansou T, et al. Implications of insecticide resistance for malaria vector control with long-lasting insecticidal nets: evidence from health facility data from Benin. Malaria J. 2019;11:550. https://doi.org/10.1186/s13071-018-3101-4.
Protopopoff N, Mosha JF, Lukole E, Charlwood JD, Wright A, Mwalimu CD, et al. Effectiveness of a long-lasting piperonyl butoxide-treated insecticidal net and indoor residual spray interventions, separately and together, against malaria transmitted by pyrethroid-resistant mosquitoes: a cluster, randomised controlled, two-by-two factorial design trial. Lancet. 2018;391:1577–88. https://doi.org/10.1016/S0140-6736(18)30427-6.
Alout H, Ndam NT, Sandeu MM, Djégbe I, Chandre F, Dabiré RK, et al. Insecticide resistance alleles affect vector competence of Anopheles gambiae s.s. for Plasmodium falciparum field isolates. PLoS ONE. 2013;8:e63849. https://doi.org/10.1371/journal.pone.0063849.
Ndiath M, Cailleau A, Diedhiou S, Gaye A, Boudin C, Richard V, et al. Effects of the kdr resistance mutation on the susceptibility of wild Anopheles gambiae populations to Plasmodium falciparum: a hindrance for vector control. Malaria J. 2014;13:340. https://doi.org/10.1186/1475-2875-13-340.
Mitri C, Markianos K, Guelbeogo WM, Bischoff E, Gneme A, Eiglmeier K, et al. The kdr-bearing haplotype and susceptibility to Plasmodium falciparum in Anopheles gambiae: genetic correlation and functional testing. Malaria J. 2015;14:391. https://doi.org/10.1186/s12936-015-0924-8.
Dabiré RK, Namountougou M, Diabaté A, Soma DD, Bado J, Toé HK, et al. Distribution and frequency of kdr mutations within Anopheles gambiae s.l. populations and first report of the Ace1G119S mutation in Anopheles arabiensis from Burkina Faso (West Africa). PLoS ONE. 2014;9:e101484. https://doi.org/10.1371/journal.pone.0101484.
Camara S, Koffi AA, Ahoua Alou LP, Koffi K, Kabran JPK, Koné A, et al. Mapping insecticide resistance in Anopheles gambiae (s.l.) from Côte d’Ivoire. Parasit Vectors. 2018;11:19. https://doi.org/10.1186/s13071-017-2546-1.
Sternberg ED, Cook J, Ahoua Alou LP, Aoura CJ, Assi SB, Doudou DT, et al. Evaluating the impact of screening plus eave tubes on malaria transmission compared to current best practice in central Côte d’Ivoire: a two armed cluster randomized controlled trial. BMC Public Health. 2018;18:894. https://doi.org/10.1186/s12889-018-5746-5.
Gillies MT, Coetzee M. A supplement to the Anophelinae of Africa south of the Sahara (Afrotropical Region). Johannesburg. 1987;143:15.
Yahouédo GA, Chandre F, Rossignol M, Ginibre C, Balabanidou V, Mendez NGA, et al. Contributions of cuticle permeability and enzyme detoxification to pyrethroid resistance in the major malaria vector Anopheles gambiae. Sci Rep. 2017;7:11091. https://doi.org/10.1038/s41598-017-11357-z.
Mangold KA, Manson RU, Koay ESC, Stephens L, Regner M, Thomson RB, et al. Real-Time PCR for detection and identification of Plasmodium spp. J Clin Microbiol. 2005;43:2435–40. https://doi.org/10.1128/JCM.43.5.2435-2440.2005.
Favia G, Lanfrancotti A, Spanos L, Sidén-Kiamos I, Louis C. Molecular characterization of ribosomal DNA polymorphisms discriminating among chromosomal forms of Anopheles gambiae s.s.: An. gambiae s.s. rDNA polymorphisms. Insect Mol Biol. 2001;10:19–23. https://doi.org/10.1046/j.1365-2583.2001.00236.x.
Bass C, Nikou D, Donnelly MJ, Williamson MS, Ranson H, Ball A, et al. Detection of knockdown resistance (kdr) mutations in Anopheles gambiae: a comparison of two new high-throughput assays with existing methods. Malaria J. 2007;6:111. https://doi.org/10.1186/1475-2875-6-111.
Bass C, Nikou D, Vontas J, Williamson MS, Field LM. Development of high-throughput real-time PCR assays for the identification of insensitive acetylcholinesterase (ace-1R) in Anopheles gambiae. Pestic Biochem Physio. 2010;96:80–5. https://doi.org/10.1016/j.pestbp.2009.09.004.
Koffi AA, Ahoua-Alou LP, Djenontin A, Kabran JPK, Dosso Y, Kone A, et al. Efficacy of Olyset ® Duo, a permethrin and pyriproxyfen mixture net against wild pyrethroid-resistant Anopheles gambiae s.s. from Côte d’Ivoire: an experimental hut trial. Parasite. 2015;22:28. https://doi.org/10.1051/parasite/2015028.
Zoh DD, Ahoua Alou LP, Toure M, Pennetier C, Camara S, Traore DF, et al. The current insecticide resistance status of Anopheles gambiae (s.l.) (Culicidae) in rural and urban areas of Bouaké, Côte d’Ivoire. Parasit Vectors. 2018;11:118. https://doi.org/10.1186/s13071-018-2702-2.
Gnémé A, Guelbéogo WM, Riehle MM, Sanou A, Traoré A, Zongo S, et al. Equivalent susceptibility of Anopheles gambiae M and S molecular forms and Anopheles arabiensis to Plasmodium falciparum infection in Burkina Faso. Malar J. 2013;12:204. https://doi.org/10.1186/1475-2875-12-204.
Sternberg ED, Cook J, Alou LPA, Assi SB, Koffi AA, Doudou DT, et al. Impact and cost-effectiveness of a lethal house lure against malaria transmission in central Côte d’Ivoire: a two-arm, cluster-randomised controlled trial. Lancet. 2021;397:805–15. https://doi.org/10.1016/S0140-6736(21)00250-6.
Koukpo CZ, Fassinou AJYH, Ossè RA, Agossa FR, Sovi A, Sewadé WT, et al. The current distribution and characterization of the L1014F resistance allele of the kdr gene in three malaria vectors (Anopheles gambiae, Anopheles coluzzii, Anopheles arabiensis) in Benin (West Africa). Malaria J. 2019;18:175. https://doi.org/10.1186/s12936-019-2808-9.
Zogo B, Soma DD, Tchiekoi BN, Somé A, Ahoua Alou LP, Koffi AA, et al. Anopheles bionomics, insecticide resistance mechanisms, and malaria transmission in the Korhogo area, northern Côte d’Ivoire: a pre-intervention study. Parasite. 2019;26:40. https://doi.org/10.1051/parasite/2019040.
Nkya TE, Poupardin R, Laporte F, Akhouayri I, Mosha F, Magesa S, et al. Impact of agriculture on the selection of insecticide resistance in the malaria vector Anopheles gambiae: a multigenerational study in controlled conditions. Parasit Vectors. 2014;7:480. https://doi.org/10.1186/s13071-014-0480-z.
Alout H, Dabiré RK, Djogbénou LS, Abate L, Corbel V, Chandre F, et al. Interactive cost of Plasmodium infection and insecticide resistance in the malaria vector Anopheles gambiae. Sci Rep. 2016;6:29755. https://doi.org/10.1038/srep29755.
Djogbénou L, Noel V, Agnew P. Costs of insensitive acetylcholinesterase insecticide resistance for the malaria vector Anopheles gambiae homozygous for the G119S mutation. Malar J. 2010;9:12. https://doi.org/10.1186/1475-2875-9-12.
Djogbénou LS, Assogba B, Essandoh J, Constant EAV, Makoutodé M, Akogbéto M, et al. Estimation of allele-specific Ace-1 duplication in insecticide-resistant Anopheles mosquitoes from West Africa. Malar J. 2015;14:507. https://doi.org/10.1186/s12936-015-1026-3.
Collins E, Vaselli NM, Sylla M, Beavogui AH, Orsborne J, Lawrence G, et al. The relationship between insecticide resistance, mosquito age and malaria prevalence in Anopheles gambiae s.l. from Guinea. Sci Rep. 2019;9:8846. https://doi.org/10.1038/s41598-019-45261-5.
Kabula B, Tungu P, Rippon EJ, Steen K, Kisinza W, Magesa S, et al. A significant association between deltamethrin resistance, Plasmodium falciparum infection and the Vgsc-1014S resistance mutation in Anopheles gambiae highlights the epidemiological importance of resistance markers. Malaria J. 2016;15:289. https://doi.org/10.1186/s12936-016-1331-5.
Ndiath MO, Cohuet A, Gaye A, Konate L, Mazenot C, Faye O, et al. Comparative susceptibility to Plasmodium falciparum of the molecular forms M and S of Anopheles gambiae and Anopheles arabiensis. Malar J. 2011;10:269. https://doi.org/10.1186/1475-2875-10-269.
Fournier DA, Skaug HJ, Ancheta J, Ianelli J, Magnusson A, Maunder MN, et al. AD Model Builder: using automatic differentiation for statistical inference of highly parameterized complex nonlinear models. Optim Methods Softw. 2012;27:233–49. https://doi.org/10.1080/10556788.2011.597854.
Bolker BM, Brooks ME, Clark CJ, Geange SW, Poulsen JR, Stevens MHH, et al. Generalized linear mixed models: a practical guide for ecology and evolution. Trends Ecol Evol. 2009;24:127–35. https://doi.org/10.1016/j.tree.2008.10.008.
Cohuet A, Harris C, Robert V, Fontenille D. Evolutionary forces on Anopheles: what makes a malaria vector? Trends Parasitol. 2010;26:130–6. https://doi.org/10.1016/j.pt.2009.12.001.
Glunt KD, Thomas MB, Read AF. The effects of age, exposure history and malaria infection on the susceptibility of Anopheles mosquitoes to low concentrations of pyrethroid. PLoS ONE. 2011;6: e24968. https://doi.org/10.1371/journal.pone.0024968.
Manguin S. Biodiversity of malaria in the world. English version completely updated. Paris: John Libbey Eurotext; 2008; 133:427. http://hdl.handle.net/10390/2213.
Churcher TS, Lissenden N, Griffin JT, Worrall E, Ranson H. The impact of pyrethroid resistance on the efficacy and effectiveness of bednets for malaria control in Africa. eLife. 2016;2(5):e16090. https://doi.org/10.7554/eLife.16090.
Mbepera S, Nkwengulila G, Peter R, Mausa EA, Mahande AM, Coetzee M, et al. The influence of age on insecticide susceptibility of Anopheles arabiensis during dry and rainy seasons in rice irrigation schemes of northern Tanzania. Malaria J. 2017;16:364. https://doi.org/10.1186/s12936-017-2022-6.
Simard F, Ayala D, Kamdem G, Pombi M, Etouna J, Ose K, et al. Ecological niche partitioning between Anopheles gambiae molecular forms in Cameroon: the ecological side of speciation. BMC Ecol. 2009;9:17. https://doi.org/10.1186/1472-6785-9-17.
Gimonneau G, Bouyer J, Morand S, Besansky NJ, Diabate A, Simard F. A behavioral mechanism underlying ecological divergence in the malaria mosquito Anopheles gambiae. Behav Ecol. 2010;21:1087–92. https://doi.org/10.1093/beheco/arq114.
Acknowledgements
We would like to thank the technical staff at the Institut Pierre Richet, Bouaké, Côte d’Ivoire for their valuable support during the mosquito collection surveys and laboratory analysis. The authors are very grateful to colleagues working in various disciplines in Côte d’Ivoire, especially at the Unité de Recherche et de Pédagogie de Génétique, UFR Biosciences, Université Félix Houphouët-Boigny, Abidjan, for their useful contributions. We also thank the volunteer mosquito collectors in the villages for their participation in the study.
Funding
This study is supported by a grant to the Pennsylvania State University from the Bill & Melinda Gates Foundation (OPP1131603) for the evaluation of the impact of an intervention comprising household screening plus a novel insecticide delivery system called In2Care Eave Tubes.
Author information
Authors and Affiliations
Contributions
RZW, AAK and RN designed the study. RZW, FHAY, AAPL, EDS, IZT, WAO and SC conducted the field, laboratory and data management work. RZW, AD, and MHK analysed the data. RZW wrote the manuscript. KAA, ONA, EDS, AAPL, SPAN, MBT and NR supervised the study and revised the manuscript. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Ethical clearance and consent information are included within the manuscript.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Wolie, R.Z., Koffi, A.A., Ahoua Alou, L.P. et al. Evaluation of the interaction between insecticide resistance-associated genes and malaria transmission in Anopheles gambiae sensu lato in central Côte d’Ivoire. Parasites Vectors 14, 581 (2021). https://doi.org/10.1186/s13071-021-05079-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13071-021-05079-5