Abstract
Background
The survival of coronaviruses are influenced by weather conditions and seasonal coronaviruses are more common in winter months. We examine the seasonality of respiratory infections in England and Wales and the associations between weather parameters and seasonal coronavirus cases.
Methods
Respiratory virus disease data for England and Wales between 1989 and 2019 was extracted from the Second-Generation Surveillance System (SGSS) database used for routine surveillance. Seasonal coronaviruses from 2012 to 2019 were compared to daily average weather parameters for the period before the patient’s specimen date with a range of lag periods.
Results
The seasonal distribution of 985,524 viral infections in England and Wales (1989–2019) showed coronavirus infections had a similar seasonal distribution to influenza A and bocavirus, with a winter peak between weeks 2 to 8. Ninety percent of infections occurred where the daily mean ambient temperatures were below 10 °C; where daily average global radiation exceeded 500 kJ/m2/h; where sunshine was less than 5 h per day; or where relative humidity was above 80%. Coronavirus infections were significantly more common where daily average global radiation was under 300 kJ/m2/h (OR 4.3; CI 3.9–4.6; p < 0.001); where average relative humidity was over 84% (OR 1.9; CI 3.9–4.6; p < 0.001); where average air temperature was below 10 °C (OR 6.7; CI 6.1–7.3; p < 0.001) or where sunshine was below 4 h (OR 2.4; CI 2.2–2.6; p < 0.001) when compared to the distribution of weather values for the same time period. Seasonal coronavirus infections in children under 3 years old were more frequent at the start of an annual epidemic than at the end, suggesting that the size of the susceptible child population may be important in the annual cycle.
Conclusions
The dynamics of seasonal coronaviruses reflect immunological, weather, social and travel drivers of infection. Evidence from studies on different coronaviruses suggest that low temperature and low radiation/sunlight favour survival. This implies a seasonal increase in SARS-CoV-2 may occur in the UK and countries with a similar climate as a result of an increase in the R0 associated with reduced temperatures and solar radiation. Increased measures to reduce transmission will need to be introduced in winter months for COVID-19.
Similar content being viewed by others
Background
Coronaviruses (CoVs) are enveloped, single-strand, positive-sense RNA viruses that occur in a variety of animals, including birds, reptiles and amphibians [1], and humans, causing respiratory and gastrointestinal infections. They appear to have originated over three hundred million years ago [2]. The seven coronaviruses that are known to infect humans are currently restricted to the Alphacoronaviruses and Betacoronaviruses, all deriving from animal sources. There are four seasonal coronaviruses species (HCoV-NL63, HCoV-HKU1, HCoV-OC43, HCoV-229E). Bats are hosts of more than 30 sequenced CoVs and only Alphacoronavirus and Betacoronavirus have been detected in bats [3]. Both genera have been found in bats in Asia, Europe, Africa, North and South America and Australasia; but Alphacoronaviruses are more widespread than Betacoronaviruses and their detection rate is also higher.
The outbreak of coronavirus disease 2019 (COVID-19) caused by Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) in Wuhan, China [4, 5], with subsequent person-to-person spread to many Chinese cities and then across six continents initiated outbreaks of infection around the world that became a pandemic [6, 7]. The clinical features of COVID-19 are varied, ranging from an asymptomatic state to acute respiratory distress syndrome and multiple organ dysfunction [8]. The start of both the Severe Acute Respiratory Syndrome (SARS) in 2002/2003 and the COVID-19 epidemic in 2019/2020 occurred in cold dry winter seasons that coincided with major national public holidays [9].
With the rapid increase in cases of COVID-19 worldwide [10,11,12], the main strategies to limit the damage caused by SARS-CoV-2 have included a range of non-pharmaceutical interventions including closing air-routes and borders, case and contact tracing, isolation/quarantine, social distancing, increased use of personal protective equipment (PPE) and cleaning and disinfection procedure, restriction of non-essential activity in most countries, restriction of normal business activity to online/remote only, that is designed to delay or reduce the viral spread and lower the peak of the outbreak [13, 14]. It has been suggested that the weather in European temperate northern-hemisphere summer conditions has contributed to keeping the virus at bay as a result of the same drivers that contribute to the occurrence of the seasonal coronaviruses. Winter pressure result from closed wards as a result of norovirus outbreaks, high bed occupancy, high rates of respiratory disease, increased vulnerability from cold weather [15] and increased illness due to seasonal respiratory viruses such as influenza [16]. There is a known seasonal element to the spread of many respiratory infections, including seasonal coronaviruses, and a better understanding of this may aid predictive models for COVID-19. A previous study showed influenza A cases increase in Europe and Asia from December to March and in southern continents from June to October, while the seasonal peak of respiratory syncytial virus (RSV) occurs slightly earlier than with influenza A [17]. For viruses that are transmitted through person-to-person routes and that have an R0 of more than 1 the modelling suggests that the size of the population that are naïve to the virus (the non-immune population) will control the epidemic progression. The weather parameters involved will contribute to the seasonal focus of the epidemic. The areas of the world that have more warm, humid and tropical climates have one or two peaks, but with more cases throughout the year [18, 19].
Respiratory viruses, including coronaviruses, are more frequent in autumn/winter (November to March) in the Northern Hemisphere and April to August in the Southern Hemisphere [20,21,22,23,24,25,26,27,28,29]. In more tropical countries, however, there is less evidence of any seasonality [30], or an increase in wet or dry seasons [31]. In Brazil respiratory infections were increased at higher temperatures, although cases occurred year-round [32]. In urban Italy, childhood infections with RSV were negatively correlated with temperature while positively correlated with humidity [33]. Human metapneumovirus in Korea peaked between weeks 15 and 20 [34,35,36,37] which is later than for the UK [38]. In China, human metapneumovirus infections were associated with sunshine duration, wind speed and diurnal temperature variation [38], and human bocavirus showed both summer and winter increases in different years [39]. Infections with influenza are strongly seasonal, and the weather variables associated with these have been examined [40]. Low temperature and UV indexes were the most predictive for influenza A virus seasonal epidemics in northern Europe [41]. Diurnal temperature range has been found to correlate with influenza A in South Korea [42], although the mechanism for this is unclear. Temperature across the globe has been matched to influenza A and B [43].
The proposed mechanisms for increased respiratory disease in the winter include the effects of weather on virus survival, behavioural changes in winter months, and changes to people’s disease susceptibility. Behavioural changes that may influence cases of infection include increased winter crowding as a result of spending more time indoors in close proximity with other people [44], more time being spent outdoors in summer, changes in schooling and university attendance, travel and other behavioural, cultural and religious influences [44]. The physiological elements that may change include day length, which has been suggested as a parameter contributing to influenza morbidity and mortality, host chilling which may increase susceptibility [44], transitions from cold exterior to hot interior environments that may cause alterations in nasal mucosal physiology, and seasonal effects from changing vitamin D levels that are a product of UV exposure. Day length is a standard seasonal parameter that is independent of the measured sunlight levels. Mortality in the 1918–1920 influenza pandemic was examined in a multiple regression model [45], showing enhanced survival in populations that experienced a short day-length (≤ 10 h light/day). It was suggested that exposure to short day lengths, typical of winter periods in non-tropical areas, yields “robust and enduring reductions in the magnitude of cytokine, febrile, and behavioural responses to infection” and suggested that a proportion of the global variance in morbidity and mortality may be explained by effects of day length. School attendance has also been linked to winter infections (e.g., increases in influenza) and day length [46]. Day length and global radiation in the UK are likely to show a degree of correlation.
Possible mechanisms contributing to the seasonality of respiratory virus survival and increased viral transmission include sunlight and UV, which may reduce virus survival in aerosols and on surfaces [44, 47, 48]. Ambient humidity has been examined as a factor contributing to influenza A and B occurrence [43]. A study of droplet size and influenza survival suggested that virus survival at one hour is lower with increasing temperature and raised absolute humidity [49]. The humidity may cause droplets to increase in size and come out of the air more quickly. However, the authors thought that relative humidity (which is strongly linked to temperature) is a better predictor of virus survival than absolute humidity. Indoor and outdoor absolute humidity are high in summer and low in winter and show similar values. Indoor relative humidity is lower in the winter and higher in summer, while outdoor relative humidity is higher in winter and lower in summer [50]. The humidity of environments in buildings during the winter may encourage viral transmission [51].
Coronavirus seasonal characteristics
Coronaviruses are relatively robust enveloped RNA viruses. Their survival characteristics reflect this, and they are sensitive to surface-active agents. Survival of virus in nasal, salivary, respiratory or faecal secretions, or with cultured virus in a suitable medium, gives some insights into the mechanisms by which weather might contribute to seasonality. Virus survival studies suggests that environmental variables, such as humidity, temperature, sunlight and UV radiation, might contribute to the seasonal transmission of coronaviruses by influencing the inactivation of virus in air and on surfaces. Studies suggest that all coronaviruses tend to survive longer in colder and dryer conditions.
SARS, MERS and COVID-19
The SARS outbreak showed little evidence of being influenced by climate, although the risk of increased daily incidence of SARS in Hong Kong was 18.2 fold higher on days with lower temperature than days with higher temperature [52]. SARS-CoV remained viable in faeces, serum and sputum for 72 h, and survived on a variety of surfaces when tested using cytopathic effect in Vero-E6 cells [53]. The virus was relatively stable at 4 °C, and survived for 2 h at 20 °C and 37 °C, but was sensitive to exposure at 56 °C, at 67 °C and at 75 °C, for 90, 60 and 30 min respectively, and to UV radiation for 60 min. SARS-CoV can survive for 4 days in diarrheal stool samples with an alkaline pH, and it may remain infectious in respiratory specimens for > 7 days at room temperature [54]. SARS-CoV was inactivated at 65 °C for 10 min [55] and by ultraviolet light (UVC) for 10 min [55]. MERS-CoV is also not very seasonal [56], is not strongly influenced by weather and transmission is more influenced by other measures [57,58,59], including camel-related events [60]. A case-crossover analysis of the impact of weather on primary cases of MERS in Saudi Arabia found more cases when conditions were relatively cold and dry [61]. A study of the weather influences on 712 MERS cases found higher incidences of infection at times when there was higher temperatures, low relative humidity and low wind speed [62]. MERS virus is inactivated with a variety of UV based treatments [63,64,65] and this can be used for large-scale inactivation of SARS-CoV for vaccine production purposes [66].
An examination of COVID-19 in cities in China showed evidence that low temperature, mild diurnal temperature range and low humidity favoured transmission [67]. However, examining the associations between weather and disease during the early stages of the pandemic is complicated by the dynamics of the disease emergence. A Study of SARS-CoV-2 survival in simulated saliva or culture media dried onto stainless steel was reduced in count by 90% in 6.8 min and 14.3 min at room temperature when exposed to simulated sunlight [68]. Similar results have been found for aerosols [69]. The survival of SARS-CoV-2 on surfaces at room temperature is longer at low relative humidity than at high, and longer at 24 °C than at 37 °C [70]. A review has examined the influence of climatic conditions on the stability and survival of SARS-CoV-2 and persistence at lower temperatures and lower relative humidity [71].
Seasonal coronaviruses
The seasonality of seasonal coronaviruses has been examined in 21 countries across the world and showed seasonal patterns similar to influenza and RSV in temperate climates [72]. Heat maps of the four species show most cases occur within the months December to March, while southern hemisphere distributions were from July to September. It has been suggested that forecasting based on weather might help in targeting countries most at risk [73]. Coronaviruses 229E and OC43 survived for up to six days but were substantially reduced in titre on surfaces such as aluminium, latex gloves and sponges, surviving for hours rather than days [74]. HCoV-229E was also able to survive on lettuce at 4 °C for a couple of days [75]. Freeze dried coronavirus NL63 cultured in LLC-MK2 cells was viable for 14 days in liquid recovery media at ambient temperature (21 °C) but survived at higher concentrations at 4 °C [76]. Human coronavirus 229E survival in a stabilised aerosol was found to remain viable for longest at 4 °C, with little difference made by relative humidity of 30%, 50% or 80%, while at 20 °C survival was better at 30% and 50% than at 80% relative humidity [77]. Survival was lowest at room temperature with a high relative humidity [77], but although viral titres for HCoV-229E and SARS-CoV reduce at room temperature within a couple of days, lower numbers may survive for several days [78].
Studies of other coronaviruses
Other coronaviruses that have been used as potential surrogates of SARS-CoV, MERS-CoV and SARS-CoV-2 include porcine transmissible gastroenteritis virus (TGEV), bovine coronavirus (BCoV), canine coronavirus (CCV), turkey coronavirus (TCoV) and murine hepatitis virus (MHV). These viruses survived longest at low relative humidity (20%), and lowest at moderate to high humidity (> 50% RH) [79]. TGEV can survive on protective equipment for up to 24 days, but with significant log reduction of viral counts [80]. CCV does not survive for long periods above 4 °C [81], can survive at 56 °C for 30 min but is inactivated at 75 °C [82]. TCoV shows reduced viral titres at room temperature compared to 4 °C [83]. A cell culture of BCoV in growth medium survived for 14 days but for shorter periods in bovine faecal suspensions on the surface of romaine lettuce [84]. A study of MHV found this virus to be highly susceptible to UV inactivation [85]. Both SARS-CoV and MHV-A59 were reduced in numbers by > 5 logs to undetectable levels by UVC disinfection at 1.22 m distance. Review of the effects of ultraviolet light on different coronaviruses suggests much of the differences between studies reflect the experimental conditions and, overall, coronaviruses are very sensitive to ultraviolet light [86].
This paper aims to contribute to understanding COVID-19 by reviewing seasonal respiratory/enteric virus infections in England and Wales and relationships between seasonal coronavirus cases and individual weather variables that might contribute to this seasonality. The pathways linking season to transmission and human susceptibility of these viruses, including the dynamics of infections in children, may be informed by the survival characteristics of coronaviruses on surfaces, in aerosols, particles and droplets.
Methods
Respiratory virus disease data was extracted from the Second-Generation Surveillance System (SGSS) database. This is the main infectious diseases database used for routine surveillance in England and Wales [87]. The data includes some viruses that may be transmitted through water and food as a comparator for viruses predominantly transmitted through the respiratory route. Individual case episodes of infectious diseases in England and Wales between 1989 and 2019 were diagnosed at local laboratory or national reference laboratory level. This surveillance data represents laboratory diagnoses that include different detection and identification methods across the 31-year time period, and RT-PCR is the method used to detect coronavirus over the chosen 8-year study period. In order to examine the relationship between weather and season the coronavirus cases occurring in the period of the year when cases were decreasing (from day 43 or 12th February to day 224 or 12th August) were when cases are declining from the late winter seasonal peak are here called the Down period. The days of the year 225–366 and 1–42 (13th August to 11th February) are here called the Up period when cases are increasing. These cut-off values were based on the averaged maximum and minimum cases over the 8-year period used for coronaviruses. This allows the associations between cases and weather to be compared across the year.
The study area is a temperate oceanic climate according to the Köppen-Geiger climate classification. We used daily averages of climate observations obtained from the Met Office from within a box that covers England and Wales, specified by boundaries at 50 degrees N and 56 degrees N latitude and 6 degrees W and 2 degrees E longitude. The weather stations included 1623 sites measuring air temperature, 206 measuring global radiation, 888 measuring sunshine duration, 1617 measuring relative humidity, 1571 measuring dewpoint temperature and 564 measuring precipitation rate. Weather parameters included daily averages for air temperature (°C), dewpoint temperature (°C), relative humidity (%), hourly precipitation (mm/h), hourly global radiation (average of total radiation including sun, UV, cosmic, measured hourly) (kJ/m2/h), and sunshine (h/day). A range of lag periods were used to estimate an optimum across the six weather parameters (Additional file 1: S1), making assumptions that in a person-to-person transmitted outbreak it is likely that short-term weather influences (1–2 weeks) will be more important than longer-term ones (1–3 months). We used 14-day averages for direct associations and 28-day averages for more seasonally smoothed associations. Some of the datasets were split into quantiles (e.g. deciles by temperature). In scatter plots for cases in the whole coronavirus dataset the data points were jittered to reduce overplotting (small random additions to data to prevent multiple values being expressed in a single dot; Fig. 3s–x only).
Analysis used Stata, Microsoft Excel and R. Tests for significance used Chi square to test coronavirus weather distributions with the distribution of weather in the days of the study period. Literature was reviewed using Endnote extractions from PubMed.
Results
The number of weekly laboratory-confirmed cases of the common respiratory viral diseases in England and Wales between 1989 and 2019 is presented in Fig. 1 along with some viruses that are transmitted by food, water or other routes. This gives some sense of the changes to surveillance over longer time periods and the more summer focus of enteric viruses compared to the winter focus of respiratory ones. There were 985,524 patients with a viral infection over this period. The data was used to select the coronavirus time series. Coronavirus infections (11,741 cases) were from 41 laboratories in England and Wales over the period 2012–2019, and included all age groups, although 29% of cases were in children under 3 years of age. Coronavirus infections had a similar seasonal distribution to influenza A and bocavirus, with a winter peak between weeks 2–8 (January and February). The distribution of cases through the year shows the different seasonal distribution of the annual peak for each virus. Seasonal coronavirus infections have a distribution that is similar to that of influenza A and bocavirus with a winter peak in weeks 2–8. Influenza B and human metapneumovirus were increased a few weeks earlier at around the New Year but the period of infection extends into the spring. Parainfluenza 1 and 2 both have a peak earlier in the late autumn/winter period between weeks 40 and 50 while parainfluenza 3 has a spring peak (weeks 10–25) that differs from all of the other viruses. Parainfluenza 1 and 2 viruses peak in the autumn/winter period between weeks 40 and 50, parainfluenza 3 has a spring peak (weeks 10–25), human metapneumovirus and influenza B virus increase at the end of the year, with infection extending into the spring. Adenovirus and rhinoviruses are relatively unseasonal. The more enteric viruses (coxsackievirus A and B; echovirus; enterovirus) have a summer peak (weeks 25–33), that show marked seasonality even with relatively sporadic cases. Most of the viral diseases exhibit an annual cycle of infection, with some of the diseases exhibiting a biennial cycle (i.e. every 2 years; e.g. parainfluenza 1 and 2; influenza B; Fig. 2).
Weekly seasonality of respiratory viruses 1989–2019. Cases per week of laboratory diagnosed viral infections reported in England and Wales. There are changes in laboratory surveillance over 31 years and includes changes resulting from improved diagnostic tests, improved sampling and testing, reductions in disease, introductions in vaccine, and improved surveillance as a result of the H1N1 outbreak in 2009. Many pathogens have an annual cycle, some have a biannual cycle (parainfluenza 1; parainfluenza 2; parainfluenza 4; influenza B), some are relatively unseasonal (adenovirus; poliovirus; rhinovirus), and some have a more sporadic nature (influenza A H3N2; echovirus; coxsackie A; coxsackie B
Cases per week for diagnosed respiratory viruses in England and Wales 1989–2019 and the percentage of cases per week that are in the 0–2 years old age group. a Coronavirus; b influenza A; c respiratory syncytial virus (RSV); d influenza B; e human metapneumovirus; f human bocavirus; g adenovirus; h rhinovirus. Weekly coronavirus cases and the cases as a percentage of all cases for i coronavirus cases 2009–2019; j influenza A cases 2008–2019 (covering the H1N1 outbreak in 2009); k Influenza cases 1989–2007
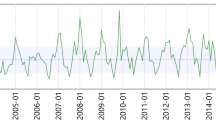
The increase in seasonal coronavirus cases in England and Wales in the autumn/winter period was associated with a period in which the percentage of the cases in patients under 3 years increased from 20 to 50% (Fig. 2a). Such large increases were not seen in influenza A (Fig. 2b), respiratory syncytial virus (Fig. 2c), influenza B (Fig. 2d), human metapneumovirus (Fig. 2e), or human bocavirus (Fig. 2f), adenovirus (Fig. 2g), rhinovirus (Fig. 2h). Weekly reports of coronavirus between 2012 and 2019 show an annual increase in the percentage of cases in children in the weeks 43–50 (October to December) as the annual rise in cases begins (Fig. 2i). This suggests that the start of the seasonal outbreak of coronavirus is frequently associated with an increase in infections in young children (0–2 years old) that may drive, at least in part, the seasonal increase. Some of the seasonality of respiratory viruses may be driven by the child population and results from a continuing increase in the susceptibility of this population cohort as a result of new births through the year. Data for influenza A before the H1N1 pandemic shows an increase in older children at the start of the pandemic (Fig. 2j), but an increase in the percentage of younger children can be seen in influenza cases before this period. (Fig. 2k).
The climate of the study area (mean year air temperature, dew point temperature, relative humidity, precipitation, global radiation, sunshine hours, etc.) are shown in Figs. 3s–x. Global radiation, sunshine hours and temperature were increased in summer months and relative humidity higher in winter, with rainfall having a distribution that varied from week to week with a seasonal distribution that was higher in autumn and winter.
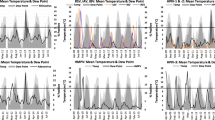
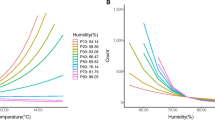
Seasonal coronavirus infections based on the week of infection and daily measures of the average weather for England and Wales between 2012 and 2019. a–f Coronavirus cases per week as a percentage of all cases and mean of weather parameters over the week. Each is lagged by a different number of weeks before the specimen date; a global radiation (kJ/m2/h), b relative humidity (%), c air temperature (°C), d sunshine (hours per day), e dewpoint temperature (°C), f precipitation (mm/hour); g–l Coronavirus cases as a percentage of cases per year and weekly mean of weather parameters separated into periods when the cases were declining (Down—days of year 43–224 and Up—days of year 225–366 and 1–42). g Global radiation (kJ/m2/h), h relative humidity (%), i air temperature (°C), j sunshine (hours per day), k dewpoint temperature (°C), l precipitation (mm/h); m–r Coronavirus case numbers split by quantiles of the weather parameters with a 2-week lag. m Global radiation (kJ/m2/h), n relative humidity (%), o air temperature (°C), p sunshine (hours per day), q dewpoint temperature (°C), r precipitation (mm/h); s–x Coronavirus cases by day of year and weather parameters with a 2-week lag. s Global radiation (kJ/m2/h), t relative humidity (%), u air temperature (°C), v sunshine (hours per day), w dewpoint temperature (°C), x precipitation (mm/h). y–ad Average weather parameters in the previous 28 days were split into ten quantiles based on the weather values. y Global radiation (kJ/m2/h), z relative humidity (%), aa air temperature (°C), ab sunshine (hours per day), ac dewpoint temperature (°C), ad precipitation (mm/h)
There are strong correlations between coronavirus cases and average global radiation (the total solar radiation), with little recorded infection where this exceeds 300 kJ/m2/hour (Fig. 3a, m, s) using a 2-week lag before the specimen date. The cases occurring in the period of the year from day 43 (12 February) to day 224 (12 August) when cases are declining after the late winter peak (here called the Down period) had a closer relationship with global radiation and few cases where this was over 300 kJ/m2/h, compared to the Up period (days 225–366 and 1–42) where there were more cases over 300 kJ/m2/h (Fig. 2g). The difference between Up and Down periods was less at longer lags, with the closest at 5 weeks but increased thereafter (Additional file 1: Fig. S2 a–h). Coronavirus infections were significantly more common where daily average global radiation was under 300 kJ/m2/hour compared to that expected by the global radiation based on days over the same time period (OR 4.3; CI 3.9–4.6; p < 0.001); (Fig. 3m). The distribution of cases in relation to average global radiation over the previous 4 weeks was a better fit than all other weather parameters (r2 = 0.91). Air temperature was strongly inversely correlated with cases (Fig. 2c, o, u) and the relationship was relatively similar between Down and Up periods (Fig. 2i), with less than 10% of cases where the air temperature was over 10 °C (Fig. 3aa). For air temperatures of 10 °C or more 2 weeks before, there were significantly fewer cases than that expected by the air temperature distribution based on days over the same time period (OR 0.15; CI 0.13–0.164; p < 0.001) and for air temperatures below 10 °C there were significantly larger case numbers (OR 6.72; CI 6.135–7.366; p < 0.001) (Fig. 3o). Relative humidity also correlated with coronavirus cases (Fig. 2b, n, t) and the Up and Down periods showed different relationships (Fig. 2h). Infection was uncommon where the relative humidity was below 75%. For relative humidity of 84% or more in the 2 weeks before, there were significantly more cases than that expected by the relative humidity distribution based on days over the same time period (OR 1.906; CI 1.755–2.07; p < 0.001) and for relative humidity below 84% there were significantly smaller case numbers (OR 0.524; CI 0.483–0.569; p < 0.001) (Fig. 3n). Sunshine hours showed a similar relationship to global radiation, but with a less close correlation (Figs. 2d, 2p, 2v). The decline with sunshine hours was more gradual than with global radiation (Fig. 2p) and the difference between the Down and Up periods was similar to that seen with global radiation (Fig. 2j). Infections were rare where daily sunshine exceeded 7 h. For daily sunshine of 4 h or more 2 weeks before, there were significantly fewer cases than that expected by the sunshine hours distribution based on days over the same time period (OR 0.406; CI 0.373–0.442; p < 0.001) and for sunshine below 4 h there were significantly larger case numbers (OR 2.457; CI 2.257–2.674; p < 0.001) (Fig. 3p). Dewpoint temperature was similar to air temperature (Figs. 2e, q, w), showing no difference between Down and Up periods (Fig. 2k) and less than 10% of infections occurred where the dewpoint temperature was over 7 °C (Fig. 2ac). There was no evidence that rainfall had a significant impact on the occurrence of infections with coronavirus (Figs. 2f, l, r, x, 2ad). The coronavirus cases were linked to averaged weather parameters in the previous 4 weeks, were sorted, separated into ten quantiles and expressed as the weather upper and lower standard deviations above the average, with maximum and minimum range and compared to all weather values over the days between 2012 and 2019 (Fig. 3y–ad). Coronavirus infections had lower global radiation than expected (Fig. 3y), relative humidity of cases was higher than expected (Fig. 3z), air temperature of cases was lower than expected (Fig. 3aa), sunshine hours of cases was lower than expected (Fig. 3ab), dewpoint temperature was lower than expected (Fig. 3ac), and precipitation was the same as expected (Fig. 3ad).
When weather parameters were examined in combination for the Down and Up periods, there was consistent evidence that the Up period was much more closely correlated with weather variables than the Down period (Fig. 3g–l), and with the mass of infections in the early weeks of the year and a gradual decline through to summer months (Fig. 3s–x). The association between global radiation and coronavirus infections was not constant and can be seen to cycle through the year (Additional file 1: Fig. S2h). Combinations of air temperature and global radiation, air temperature and relative humidity, and air temperature and sunlight showed a closer association in Up than Down periods (Additional file 1: Fig. S3a–S3f) and distribution of weather relationships across the days of the year (Additional file 1: Fig. S3g–S3j). The density of cases by combinations of weather parameters was also examined (Additional file 1: Fig. S4a–S4f).
Discussion
The development of the pandemic is dynamic and dominated by the immunological naivety of the world population to SARS-CoV-2. This study has found that seasonal coronavirus infections in England and Wales have a broadly similar distribution to influenza A and human bocavirus infections, but with a characteristic peak in days 23 to 54 (weeks 3 to 8; January to February) and reduced disease in days 131 to 303 (weeks 19 to 43; May to October). Parainfluenza 1, 2, 3, 4 and influenza B have a biannual cycle, differing from other respiratory infections occurring in England and Wales and confirming previously reported seasonality trends [87]. Respiratory infections have repeating cycles every 1 to 2 years, suggesting that their seasonal distribution is driven by the degree of susceptibility of the population to each infection and the environmental determinants that change over the year, affecting virus survival and transmission and population behaviour. Some of the changed susceptibility may reflect gradual loss of immunity to circulating viruses, and some to new babies being born. The biennial viruses (e.g. Parainfluenza 1 and 2) are presumed to have too small a susceptible population in the year after a winter increase to allow an epidemic the following year (Fig. 1). A study from the Flu Watch cohort study showed eight infections with three different seasonal coronaviruses occurred in the following season, but all were of a different type [88], suggesting reinfection with the same species is uncommon over short timescales. There is evidence in Japan and Norway the individual Coronavirus types are bi-annual while fitting into an overall annual cycle [89, 90]. There is no evidence for the distribution if these types influencing each other.
In this study, the seasonal reductions in coronavirus showed disease associated with average outside air temperatures above 10 °C, global radiation over 300 kJ/m2/h, average sunshine hours over 5 h per day and no association with precipitation. The association with dewpoint temperature was similar to air temperature but at lower temperatures. The even relationship to average air temperature between spring (Down) and autumn (Up) periods suggests temperature is the principal environmental factor associated with disease occurrence. However, the different relationships with weather in periods after the annual epidemic (Down) and the period at the start of the new epidemic (Up) suggest that temperature and global radiation may inhibit the start of the epidemic more effectively than it reduces the end of one, perhaps reflecting the strength of respiratory transmission due to the susceptibility of the population.
While many of the features of COVID-19 are different to influenza, some of the aspects of the survival and transmission of seasonal coronaviruses maybe similar to SARS-CoV-2. Of over 140 studies examining COVID-19 and weather published to date, a majority examined the short period at the start of the pandemic. The dynamics of the COVID-19 pandemic suggest that at the start the high percentage of susceptible people means that the effects of weather were relatively small during the initial phase, but may increase in subsequent years as the virus becomes endemic and the percentage of the population that is susceptible declines [91]. From April through to October seasonal coronavirus cases are relatively uncommon in England and Wales, suggesting that their ability to transmit within the community is slightly reduced compared to winter months. We infer that there is a reduced R0 value in this period linked to temperatures above 10 °C and global radiation above 300 kJ/m2/h, although the reductions are not sufficient to prevent transmission in summer months. For COVID-19, the seasonal changes in weather likely contribute to a reduction in R0 during summer months but the size of the susceptible population means that these reductions are small, likely to break down with changes in interventions and increase the risks of increased transmission during the winter. The demonstration of increased rates of transmission in young children (0 to 2 years old) at the start of the winter increase in seasonal coronaviruses suggests that additional measures to control transmission in young children should be considered.
Limitations
The weather data was based on national average values, rather than linked to patient postcode and therefore represents the national weather rather than the local weather. The weather variables used are representative of conditions across a broad area, local variations in conditions (e.g. local microclimates) are not captured, and many of the weather factors auto-correlate to some extent because of their natural relationships in the environment (for example relative humidity is influenced by temperature). In England people spend the majority of their time indoors, and while there is a relationship between indoor and outdoor temperature, particularly for warm conditions [92], sufficient data on indoor conditions were not available for this study. In reality the effects of weather are likely to be combinations of parameters that have direct and indirect influences on disease occurrence. The respiratory infections reported are likely to derive from the more severe infections such as pneumonia or infections in children or more vulnerable patients. The data may not capture a majority of the milder infections, such as the common cold that are better examined using syndromic surveillance sources. The respiratory cases are influenced by current practices over 31 years. We included the respiratory virus data over 31 years to 1. Show how the time-period for coronavirus data was selected; 2. highlight changes in surveillance over 30 years; 3. Illustrate the variations in seasonality of viral respiratory infections over a long time period; 4. To demonstrate the bi-annual epidemics with some viral infections whilst most are annual. For the coronaviruses we used data from 8 years from 2012 to 2019 only. While it can be assumed that survival properties of SARS-CoV-2 might be somewhat similar to SARS-CoV, MERS-CoV and other coronaviruses, examining the degree of similarity was beyond the scope of our study. A discussion of the similarities and differences between influenza, other respiratory viruses, seasonal coronaviruses and the SARS like coronaviruses would help to understand COVID-19 infections and a more generalised world model of the factors contributing to respiratory virus seasonality is needed. It has been suggested that there is no independent relationship with weather and no teasing out of the relative contributions of the weather parameters to the seasonality. The results we show demonstrate individual associations between weather parameters and disease but do not attempt to attribute how much of the seasonality is due to each, or whether the effects of the weather are direct or indirect. While the study shows associations between individual weather parameters and disease occurrence it is recognised that the drivers of disease involve combined effects of physical, physiological, social and behavioural elements. Figure 4 shows the combined contributions of some of the main infection drivers for seasonal epidemics.
Some of the drivers influencing the seasonality of UK respiratory infections. Seasonal coronavirus infections are thought to be influenced by the size of the susceptible child population, which drives an annual epidemic in children that, in turn, infects susceptible adults. The timing of the epidemic is influenced by changing transmission dynamics through the year. Low temperature, low humidity, short daylength and low UV all probably contribute to better survival of the virus in winter months than summer, pushing the immunity driven epidemic to occur in the winter months. Travel abroad introduces new viruses that differ from the currently circulating strains. Travel variation will differ by country and holidays/festivals (including school/university holidays) (Additional file 1: S5). Many social and public interactions that contribute to infection are relatively constant through the year. In addition to these drivers there will be the gradual increase in susceptibility as a result of declining antibody levels and genetic drift within the viruses
Conclusions
Identifying and understanding the seasonal distribution of viral diseases and what factors drive their circulation and the severity of symptoms can help in preparing health care systems and populations to better protect those most at risk from serious complications from becoming infected, and support timing of interventions to help achieve this. Travel abroad, which also has a seasonal pattern, is likely to introduce new viruses that differ from the currently circulating strains. The timing of the annual coronavirus epidemic is influenced by changing transmission dynamics through the year. Low temperature, low relative humidity indoors, short daylength and low UV contribute to better survival of the virus in winter months than in the summer, pushing the immunity driven epidemic to occur in the winter months.
Availability of data and materials
The datasets generated and/or analysed during the current study are not publicly available. The datasets of anonymous routine surveillance data are confidential, and care needs to be taken to avoid deductive disclosure. The data are available from the corresponding author on reasonable request.
Abbreviations
- COVID-19:
-
The disease caused by SARS-CoV-2
- CoVs:
-
Coronaviruses
- Down period:
-
The period between day of the year 43 (12 February) to day 224 (12 August) are when cases are declining from the late winter seasonal peak
- MERS:
-
Middle eastern respiratory syndrome
- MRC:
-
UK Medical Research Council
- NERC:
-
UK Natural Environment Research Council
- NIHR HPRU:
-
National Institute for Health Research Health Protection Research Unit
- PHE:
-
Public Health England
- RSV:
-
Respiratory syncytial virus
- SARS:
-
Severe acute respiratory syndrome
- SARS-CoV-2:
-
Severe Acute Respiratory Syndrome 2 virus
- SGSS:
-
Second Generation Surveillance System
- Up period:
-
The period between day of year 225–366 and 1–42 (13th August to 11th February) when cases are increasing
- UV:
-
Ultraviolet light
References
Shi M, Lin XD, Chen X, Tian JH, Chen LJ, Li K, Wang W, Eden JS, Shen JJ, Liu L, et al. The evolutionary history of vertebrate RNA viruses. Nature. 2018;556(7700):197–202.
Wertheim JO, Chu DK, Peiris JS, Kosakovsky Pond SL, Poon LL. A case for the ancient origin of coronaviruses. J Virol. 2013;87(12):7039–45.
Wong ACP, Li X, Lau SKP, Woo PCY. Global epidemiology of bat coronaviruses. Viruses. 2019;11(2):174.
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395(10224):565–74.
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, Si HR, Zhu Y, Li B, Huang CL, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270–3.
Harypursat V, Chen YK. Six weeks into the 2019 coronavirus disease (COVID-19) outbreak- it is time to consider strategies to impede the emergence of new zoonotic infections. Chin Med J (Engl). 2020;133:1118.
Jalava K. First respiratory transmitted food borne outbreak? Int J Hyg Environ Health. 2020;226:113490.
Singhal T. A review of coronavirus disease-2019 (COVID-19). Indian J Pediatr. 2020;87:281–6.
Sun Z, Thilakavathy K, Kumar SS, He G, Liu SV. Potential factors influencing repeated SARS outbreaks in China. Int J Environ Res Public Health. 2020;17(5):1633.
WHO Situation Reports https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports/
Situation Update Worldwide https://www.ecdc.europa.eu/en/geographical-distribution-2019-ncov-cases
Coronavirus (COVID-19): Advice https://www.gov.uk/government/collections/coronavirus-covid-19-list-of-guidance
Flaxman S, Mishra S, Gandy A, Unwin HJT, Mellan TA, Coupland H, Whittaker C, Zhu H, Berah T, Eaton JW, et al. Estimating the effects of non-pharmaceutical interventions on COVID-19 in Europe. Nature. 2020;584(7820):257–61.
Hsiang S, Allen D, Annan-Phan S, Bell K, Bolliger I, Chong T, Druckenmiller H, Huang LY, Hultgren A, Krasovich E, et al. The effect of large-scale anti-contagion policies on the COVID-19 pandemic. Nature. 2020;584(7820):262–7.
White C. Office for National Statistics. Excess winter mortality in England and Wales: 2018 to 2019 (provisional) and 2017 to 2018 (final). Office Natl Stat. 2018;2019:1–22.
Surveillance of influenza and other respiratory viruses in the UK Winter 2018 to 2019 https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/839350/Surveillance_of_influenza_and_other_respiratory_viruses_in_the_UK_2018_to_2019-FINAL.pdf
Li Y, Reeves RM, Wang X, Bassat Q, Brooks WA, Cohen C, Moore DP, Nunes M, Rath B, Campbell H, et al. Global patterns in monthly activity of influenza virus, respiratory syncytial virus, parainfluenza virus, and metapneumovirus: a systematic analysis. Lancet Glob Health. 2019;7(8):e1031–45.
Hirve S, Newman LP, Paget J, Azziz-Baumgartner E, Fitzner J, Bhat N, Vandemaele K, Zhang W. Influenza seasonality in the tropics and subtropics—when to vaccinate? PLoS ONE. 2016;11(4):e0153003.
Xu ZW, Li ZJ, Hu WB. Global dynamic spatiotemporal pattern of seasonal influenza since 2009 influenza pandemic. Infect Dis Poverty. 2020;9(1):2.
Mair C, Nickbakhsh S, Reeve R, McMenamin J, Reynolds A, Gunson RN, Murcia PR, Matthews L. Estimation of temporal covariances in pathogen dynamics using Bayesian multivariate autoregressive models. PLoS Comput Biol. 2019;15(12):e1007492.
Galanti M, Birger R, Ud-Dean M, Filip I, Morita H, Comito D, Anthony S, Freyer GA, Ibrahim S, Lane B, et al. Longitudinal active sampling for respiratory viral infections across age groups. Influenza Other Respir Viruses. 2019;13(3):226–32.
Brini Khalifa I, Hannachi N, Guerrero A, Orth-Holler D, Bhiri S, Bougila J, Boughamoura L, Merchaoui SN, Sboui H, Mahdhaoui N, et al. Demographic and seasonal characteristics of respiratory pathogens in neonates and infants aged 0 to 12 months in the Central-East region of Tunisia. J Med Virol. 2019;91(4):570–81.
Friedman N, Alter H, Hindiyeh M, Mendelson E, Shemer Avni Y, Mandelboim M. Human coronavirus infections in Israel: epidemiology, clinical symptoms and summer seasonality of HCoV-HKU1. Viruses. 2018;10(10):515.
van Asten L, Bijkerk P, Fanoy E, van Ginkel A, Suijkerbuijk A, van der Hoek W, Meijer A, Vennema H. Early occurrence of influenza A epidemics coincided with changes in occurrence of other respiratory virus infections. Influenza Other Respir Viruses. 2016;10(1):14–26.
Litwin CM, Bosley JG. Seasonality and prevalence of respiratory pathogens detected by multiplex PCR at a tertiary care medical center. Arch Virol. 2014;159(1):65–72.
Ambrosioni J, Bridevaux PO, Wagner G, Mamin A, Kaiser L. Epidemiology of viral respiratory infections in a tertiary care centre in the era of molecular diagnosis, Geneva, Switzerland, 2011–2012. Clin Microbiol Infect. 2014;20(9):O578-584.
Varghese L, Zachariah P, Vargas C, LaRussa P, Demmer RT, Furuya YE, Whittier S, Reed C, Stockwell MS, Saiman L. Epidemiology and clinical features of human coronaviruses in the pediatric population. J Pediatric Infect Dis Soc. 2018;7(2):151–8.
Killerby ME, Biggs HM, Haynes A, Dahl RM, Mustaquim D, Gerber SI, Watson JT. Human coronavirus circulation in the United States 2014–2017. J Clin Virol. 2018;101:52–6.
Price OH, Sullivan SG, Sutterby C, Druce J, Carville KS. Using routine testing data to understand circulation patterns of influenza A, respiratory syncytial virus and other respiratory viruses in Victoria, Australia. Epidemiol Infect. 2019;147:e221.
Kenmoe S, Tchendjou P, Vernet MA, Moyo-Tetang S, Mossus T, Njankouo-Ripa M, Kenne A, Penlap Beng V, Vabret A, Njouom R. Viral etiology of severe acute respiratory infections in hospitalized children in Cameroon, 2011–2013. Influenza Other Respir Viruses. 2016;10(5):386–93.
Owusu M, Annan A, Corman VM, Larbi R, Anti P, Drexler JF, Agbenyega O, Adu-Sarkodie Y, Drosten C. Human coronaviruses associated with upper respiratory tract infections in three rural areas of Ghana. PLoS ONE. 2014;9(7):e99782.
Gurgel RQ, Bezerra PG, Duarte Mdo C, Moura AA, Souza EL, Silva LS, Suzuki CE, Peixoto RB. Relative frequency, possible risk factors, viral codetection rates, and seasonality of respiratory syncytial virus among children with lower respiratory tract infection in Northeastern Brazil. Medicine (Baltimore). 2016;95(15):e3090.
Nenna R, Evangelisti M, Frassanito A, Scagnolari C, Pierangeli A, Antonelli G, Nicolai A, Arima S, Moretti C, Papoff P, et al. Respiratory syncytial virus bronchiolitis, weather conditions and air pollution in an Italian urban area: an observational study. Environ Res. 2017;158:188–93.
Zhu R, Guo C, Zhao L, Deng J, Wang F, Sun Y, Qian Y. Epidemiological and genetic characteristics of human metapneumovirus in pediatric patients across six consecutive seasons in Beijing, China. Int J Infect Dis. 2020;91:137–42.
Schreiner D, Groendahl B, Puppe W, Off HNT, Poplawska K, Knuf M, Meyer CU, Reischl AT, Gehring S. High antibiotic prescription rates in hospitalized children with human metapneumovirus infection in comparison to RSV infection emphasize the value of point-of-care diagnostics. Infection. 2019;47(2):201–7.
Price RHM, Graham C, Ramalingam S. Association between viral seasonality and meteorological factors. Sci Rep. 2019;9(1):929.
Liu WK, Chen DH, Tan WP, Qiu SY, Xu D, Zhang L, Gu SJ, Zhou R, Liu Q. Paramyxoviruses respiratory syncytial virus, parainfluenza virus, and human metapneumovirus infection in pediatric hospitalized patients and climate correlation in a subtropical region of southern China: a 7-year survey. Eur J Clin Microbiol Infect Dis. 2019;38(12):2355–64.
Lim YK, Kweon OJ, Kim HR, Kim TH, Lee MK. Clinical features, epidemiology, and climatic impact of genotype-specific human metapneumovirus infections: long-term surveillance of hospitalized patients in South Korea. Clin Infect Dis. 2019;70:3683.
Liu WK, Liu Q, Chen DH, Tan WP, Cai Y, Qiu SY, Xu D, Li C, Li X, Lin ZS, et al. Epidemiology of HBoV1 infection and relationship with meteorological conditions in hospitalized pediatric patients with acute respiratory illness: a 7-year study in a subtropical region. BMC Infect Dis. 2018;18(1):329.
Roussel M, Pontier D, Cohen JM, Lina B, Fouchet D. Quantifying the role of weather on seasonal influenza. BMC Public Health. 2016;16:441.
Ianevski A, Zusinaite E, Shtaida N, Kallio-Kokko H, Valkonen M, Kantele A, Telling K, Lutsar I, Letjuka P, Metelitsa N, et al. Low temperature and low UV Indexes correlated with peaks of influenza virus activity in Northern Europe during 2010(–)2018. Viruses. 2019;11(3):207.
Park JE, Son WS, Ryu Y, Choi SB, Kwon O, Ahn I. Effects of temperature, humidity, and diurnal temperature range on influenza incidence in a temperate region. Influenza Other Respir Viruses. 2020;14(1):11–8.
Chong KC, Lee TC, Bialasiewicz S, Chen J, Smith DW, Choy WSC, Krajden M, Jalal H, Jennings L, Alexander B, et al. Association between meteorological variations and activities of influenza A and B across different climate zones: a multi-region modelling analysis across the globe. J Infect. 2020;80(1):84–98.
Shaw Stewart PD. Seasonality and selective trends in viral acute respiratory tract infections. Med Hypotheses. 2016;86:104–19.
Prendergast BJ. Can photoperiod predict mortality in the 1918–1920 influenza pandemic? J Biol Rhythms. 2011;26(4):345–52.
Zerbini G, van der Vinne V, Otto LKM, Monecke S, Kantermann T, Merrow M. tardiness increases in winter: evidence for annual rhythms in humans. J Biol Rhythms. 2019;34(6):672–9.
Grant WB, Lahore H, McDonnell SL, Baggerly CA, French CB, Aliano JL, Bhattoa HP. Evidence that vitamin D supplementation could reduce risk of influenza and COVID-19 infections and deaths. Nutrients. 2020;12(4):988.
Schuit M, Gardner S, Wood S, Bower K, Williams G, Freeburger D, Dabisch P. The influence of simulated sunlight on the inactivation of influenza virus in aerosols. J Infect Dis. 2020;221(3):372–8.
Marr LC, Tang JW, Van Mullekom J, Lakdawala SS. Mechanistic insights into the effect of humidity on airborne influenza virus survival, transmission and incidence. J R Soc Interface. 2019;16(150):20180298.
Lai S, Ruktanonchai NW, Zhou L, Prosper O, Luo W, Floyd JR, Wesolowski A, Santillana M, Zhang C, Du X, et al. Effect of non-pharmaceutical interventions to contain COVID-19 in China. Nature. 2020;12:988.
Netz RR. Mechanisms of airborne infection via evaporating and sedimenting droplets produced by speaking. J Phys Chem B. 2020;124(33):7093–101.
Lin K, Yee-Tak Fong D, Zhu B, Karlberg J. Environmental factors on the SARS epidemic: air temperature, passage of time and multiplicative effect of hospital infection. Epidemiol Infect. 2006;134(2):223–30.
Duan SM, Zhao XS, Wen RF, Huang JJ, Pi GH, Zhang SX, Han J, Bi SL, Ruan L, Dong XP, et al. Stability of SARS coronavirus in human specimens and environment and its sensitivity to heating and UV irradiation. Biomed Environ Sci. 2003;16(3):246–55.
Lai MY, Cheng PK, Lim WW. Survival of severe acute respiratory syndrome coronavirus. Clin Infect Dis. 2005;41(7):e67-71.
Darnell ME, Subbarao K, Feinstone SM, Taylor DR. Inactivation of the coronavirus that induces severe acute respiratory syndrome, SARS-CoV. J Virol Methods. 2004;121(1):85–91.
Al-Tawfiq JA, Memish ZA. Lack of seasonal variation of middle east respiratory syndrome coronavirus (MERS-CoV). Travel Med Infect Dis. 2019;27:125–6.
Da’ar OB, Ahmed AE. Underlying trend, seasonality, prediction, forecasting and the contribution of risk factors: an analysis of globally reported cases of Middle East Respiratory Syndrome Coronavirus. Epidemiol Infect. 2018;146(11):1343–9.
Al-Ahmadi K, Alahmadi S, Al-Zahrani A. Spatiotemporal clustering of middle east respiratory syndrome coronavirus (MERS-CoV) incidence in Saudi Arabia, 2012–2019. Int J Environ Res Public Health. 2019;16(14):2520.
Darling ND, Poss DE, Schoelen MP, Metcalf-Kelly M, Hill SE, Harris S. Retrospective, epidemiological cluster analysis of the Middle East respiratory syndrome coronavirus (MERS-CoV) epidemic using open source data. Epidemiol Infect. 2017;145(15):3106–14.
Alkhamis MA, Fernandez-Fontelo A, VanderWaal K, Abuhadida S, Puig P, Alba-Casals A. Temporal dynamics of Middle East respiratory syndrome coronavirus in the Arabian Peninsula, 2012–2017. Epidemiol Infect. 2018;147:1–10.
Gardner EG, Kelton D, Poljak Z, Van Kerkhove M, von Dobschuetz S, Greer AL. A case-crossover analysis of the impact of weather on primary cases of Middle East respiratory syndrome. BMC Infect Dis. 2019;19(1):113.
Altamimi A, Ahmed AE. Climate factors and incidence of Middle East respiratory syndrome coronavirus. J Infect Public Health. 2019;13:704–8.
Hindawi SI, Hashem AM, Damanhouri GA, El-Kafrawy SA, Tolah AM, Hassan AM, Azhar EI. Inactivation of Middle East respiratory syndrome-coronavirus in human plasma using amotosalen and ultraviolet A light. Transfusion. 2018;58(1):52–9.
Eickmann M, Gravemann U, Handke W, Tolksdorf F, Reichenberg S, Muller TH, Seltsam A. Inactivation of Ebola virus and Middle East respiratory syndrome coronavirus in platelet concentrates and plasma by ultraviolet C light and methylene blue plus visible light, respectively. Transfusion. 2018;58(9):2202–7.
Keil SD, Bowen R, Marschner S. Inactivation of Middle East respiratory syndrome coronavirus (MERS-CoV) in plasma products using a riboflavin-based and ultraviolet light-based photochemical treatment. Transfusion. 2016;56(12):2948–52.
Tsunetsugu-Yokota Y. Large-scale preparation of UV-inactivated SARS coronavirus virions for vaccine antigen. Methods Mol Biol. 2008;454:119–26.
Liu J, Zhou J, Yao J, Zhang X, Li L, Xu X, He X, Wang B, Fu S, Niu T, et al. Impact of meteorological factors on the COVID-19 transmission: a multi-city study in China. Sci Total Environ. 2020;726:138513.
Ratnesar-Shumate S, Williams G, Green B, Krause M, Holland B, Wood S, Bohannon J, Boydston J, Freeburger D, Hooper I, et al. Simulated Sunlight Rapidly Inactivates SARS-CoV-2 on Surfaces. J Infect Dis. 2020;222(2):214–22.
Schuit M, Ratnesar-Shumate S, Yolitz J, Williams G, Weaver W, Green B, Miller D, Krause M, Beck K, Wood S, et al. Airborne SARS-CoV-2 is rapidly inactivated by simulated sunlight. J Infect Dis. 2020;222:564–71.
Biryukov J, Boydston JA, Dunning RA, Yeager JJ, Wood S, Reese AL, Ferris A, Miller D, Weaver W, Zeitouni NE, et al. Increasing temperature and relative humidity accelerates inactivation of SARS-CoV-2 on surfaces. Msphere. 2020;5(4):e00441.
Aboubakr HA, Sharafeldin TA, Goyal SM. Stability of SARS-CoV-2 and other coronaviruses in the environment and on common touch surfaces and the influence of climatic conditions: a review. Transbound Emerg Dis. 2020;68:296–312.
Li Y, Wang X, Nair H. Global seasonality of human seasonal coronaviruses: a clue for post-pandemic circulating season of SARS-CoV-2 virus? J Infect Dis. 2020;222:1090.
Scafetta N. Distribution of the SARS-CoV-2 pandemic and its monthly forecast based on seasonal climate patterns. Int J Environ Res Public Health. 2020;17(10):3493.
Sizun J, Yu MW, Talbot PJ. Survival of human coronaviruses 229E and OC43 in suspension and after drying onsurfaces: a possible source ofhospital-acquired infections. J Hosp Infect. 2000;46(1):55–60.
Yepiz-Gomez MS, Gerba CP, Bright KR. Survival of respiratory viruses on fresh produce. Food Environ Virol. 2013;5:150–6.
Florek D, Burmistrz M, Potempa J, Pyrc K. Stability of infectious human coronavirus NL63. J Virol Methods. 2014;205:87–90.
Ijaz MK, Brunner AH, Sattar SA, Nair RC, Johnson-Lussenburg CM. Survival characteristics of airborne human coronavirus 229E. J Gen Virol. 1985;66(Pt 12):2743–8.
Rabenau HF, Cinatl J, Morgenstern B, Bauer G, Preiser W, Doerr HW. Stability and inactivation of SARS coronavirus. Med Microbiol Immunol. 2005;194(1–2):1–6.
Casanova LM, Jeon S, Rutala WA, Weber DJ, Sobsey MD. Effects of air temperature and relative humidity on coronavirus survival on surfaces. Appl Environ Microbiol. 2010;76(9):2712–7.
Casanova L, Rutala WA, Weber DJ, Sobsey MD. Coronavirus survival on healthcare personal protective equipment. Infect Control Hosp Epidemiol. 2010;31(5):560–1.
Tennant BJ, Gaskell RM, Gaskell CJ. Studies on the survival of canine coronavirus under different environmental conditions. Vet Microbiol. 1994;42(2–3):255–9.
Pratelli A. Canine coronavirus inactivation with physical and chemical agents. Vet J. 2008;177(1):71–9.
Guionie O, Courtillon C, Allee C, Maurel S, Queguiner M, Eterradossi N. An experimental study of the survival of turkey coronavirus at room temperature and + 4 °C. Avian Pathol. 2013;42(3):248–52.
Mullis L, Saif LJ, Zhang Y, Zhang X, Azevedo MS. Stability of bovine coronavirus on lettuce surfaces under household refrigeration conditions. Food Microbiol. 2012;30(1):180–6.
Walker CM, Ko G. Effect of ultraviolet germicidal irradiation on viral aerosols. Environ Sci Technol. 2007;41(15):5460–5.
Hessling M, Hones K, Vatter P, Lingenfelder C. Ultraviolet irradiation doses for coronavirus inactivation - review and analysis of coronavirus photoinactivation studies. GMS Hyg Infect Control. 2020;15:Doc08.
Cherrie MPCN, LoIacono G, Sarran C, Hajat S, Fleming L. Pathogen seasonality and links with weather in England and Wales: a big data time series analysis. BMC Public Health. 2018;18:1067.
Aldridge RWL, Beale S, Johnson AM, Zambon M, Hayward AC, Fragaszy EB. Flu Watch Group: Seasonality and immunity to laboratory-confirmed seasonal coronaviruses HCoV-NL63, HCoV-OC43, and HCoV-229E: results from the Flu Watch cohort study version 1; peer review: 2 approved with reservations. Wellcome Open Res. 2020;5:1.
Komabayashi K, Seto J, Matoba Y, Aoki Y, Tanaka S, Ikeda T, Matsuzaki Y, Itagaki T, Mizuta K. Seasonality of human coronavirus OC43, NL63, HKU1, and 229E infection in Yamagata, Japan, 2010–2019. Jpn J Infect Dis. 2020;73(5):394–7.
Heimdal I, Moe N, Krokstad S, Christensen A, Skanke LH, Nordbo SA, Dollner H. Human coronavirus in hospitalized children with respiratory tract infections: a 9-year population-based study from Norway. J Infect Dis. 2019;219(8):1198–206.
Baker RE, Yang W, Vecchi GA, Metcalf CJE, Grenfell BT. Susceptible supply limits the role of climate in the early SARS-CoV-2 pandemic. Science. 2020;369(6501):315–9.
Nguyen JL, Schwartz J, Dockery DW. The relationship between indoor and outdoor temperature, apparent temperature, relative humidity, and absolute humidity. Indoor Air. 2014;24(1):103–12.
Acknowledgements
We thank Aarti Khanna from PHE Colindale for providing us with the SGSS data. The views expressed are those of the authors not necessarily those of the NHS, the NIHR, the Department of Health and Social Care, Public Health England or UK Health Security Agency. We would like to thank the scientists and physicians involved in making these diagnoses.
Funding
The research was funded in part by the National Institute for Health Research Health Protection Research Unit (NIHR HPRU) in Environmental Change and Health at the London School of Hygiene and Tropical Medicine in partnership with Public Health England (PHE), and in collaboration with the University of Exeter, University College London, and the Met Office; the UK Medical Research Council (MRC) and UK Natural Environment Research Council (NERC) for the MEDMI Project (https://www.data-mashup.org.uk). The NIHR HPRU-2 in Environmental Change and Health at the London School of Hygiene and Tropical Medicine in partnership with Public Health England (PHE) and University College London is also supporting this work. NIHR had no role in the design of the study and collection, analysis, and interpretation of data or in writing the manuscript.
Author information
Authors and Affiliations
Contributions
Paper writing GN; EG; HM; SV; RP; SH; CS, data extraction and analysis AK; GN; CS; EG; HM, experimental design GN; SH, weather analysis EG; HM, GN, table construction GN, DA, figures GN; EG, literature review GN, SV; EG, discussion GN; EG; HM; SV, manuscript revision GN; EG; HM; SV; RP; CS; HS; RP.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
No ethics approval, consent to participate or consent for publication was required as this was anonymised routine surveillance data. Approval for the use of data for this study was through PHE Colindale.
Consent for publication
Not applicable.
Competing interests
All authors have indicated that they have no competing interests other than the funding from the National Institute for Health Research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: FiguresS1a-S1f.
Examination of the effect of weather measured on the date of the specimen andin the previous eight weeks, based on cases as a proportion of all thecoronavirus. S1a. Coronavirus and mean air temperature with 0 to8 weeks lag. S1b. Coronavirus and Mean dewpoint temperature with0 to 8 weeks lag. S1c. Coronavirus and mean sunshine hours with 0 to 8weeks lag. S1d. Coronavirus and mean daily precipitation with 0to 8 weeks lag. Figure S1e. Coronavirus and mean relative humiditywith 0 to 8 weeks lag. Figure S1f Coronavirus and mean global radiationwith 0 to 8 weeks lag. FigureS2a. to h. Seasonal coronavirus infections and daily average of globalradiation (kJ/m2/day) for England and Wales between 2012 and 2019 based on theweek of infection with different lag periods. S2a. no lag; S2b. 2-week lag; S2c.3-week lag; S2d. 4-week lag; S2e. 5-week lag; S2f. 6-week lag; S2g. 8-week lag;S2h. Global radiation by day of year. FigureS3.Coronavirus cases in England and Wales 2012-2019 (n=12,374) as a percentage ofall cases and weather parameters. S3a., S3c., S3e. coronavirus cases in the‘Down’ period and S3b., S3d., S3f. (Down – day of year 43-224 and Up – day ofyear 225-366 and 1-42) against two weather parameters. S3a, S3b. average airtemperature and global radiation over the previous 4 weeks, S3c., S3d., averageair temperature and relative humidity over the previous 4 weeks, S3e., S3f. Average air temperature and average sunshine hours over the previous 4 weeks; S3g.,S3h., S3i., S3j. Distribution of coronavirus cases by day of year and weatherparameters with a two week lag, S3g. average air temperature (oC), S3h. globalradiation (kJ/m2), S3i. relative humidity, S3j. sunshine hours; S3k. Weeklycoronavirus cases as a proportion of annual cases and global radiation. FigureS4. a.-f.Individual seasonal coronavirus infections based on cases and dailymeasures of two weather parameters for England and Wales between 2012 and 2019,averaged over the previous 28 days. S4a.sunshine (hours per day)/air temperature (oC), S4b. sunshine(hours per day)/relative humidity (%), S4c. sunshine (hours per day)/globalradiation (kJ/m2/day), S4d. air temperature (oC)/relativehumidity (%), S4e. relative humidity (%)/global radiation (kJ/m2/day),S4f. air temperature (oC)/global radiation (kJ/m2/day). FigureS5. Monthlytraffic for the global data series over a seven-year period. Regions separatedinto Asia-Pacific, Europe and North America. From https://blog.aci.aero/airport-markets-and-seasonal-variations/
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Nichols, G.L., Gillingham, E.L., Macintyre, H.L. et al. Coronavirus seasonality, respiratory infections and weather. BMC Infect Dis 21, 1101 (2021). https://doi.org/10.1186/s12879-021-06785-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12879-021-06785-2