Abstract
Background
Pyrrolizidine alkaloids (PAs) are a class of secondary metabolites that function as feeding deterrents in a range of different plant species. In perennial ryegrass (Lolium perenne L.) the only PAs that have been identified are the thesinine-rhamnoside group, which displays significant genetic variation. Homospermidine synthase (HSS) has evolved from deoxyhypusine synthase (DHS) and catalyses the first step in the PA pathway, making it a key candidate for the investigation of genes influencing observed PA trait variation.
Results
During PCR amplification and sequence analysis of DHS we identified two putative HSS genes in perennial ryegrass. One of the genes (LpHSS1) was absent in some perennial ryegrass plants. Thesinine-rhamnoside levels were measured using liquid chromatography coupled with mass spectrometry in a diverse association mapping population, consisting of 693 plants free of fungal endophytic symbionts. Association tests that accounted for population structure identified a significant association of absence of the LpHSS1 gene with lower levels of thesinine-rhamnoside PAs. HSS-like gene sequences were identified for other grass species of the Poaceae, including tall fescue, wheat, maize and sorghum.
Conclusion
HSS is situated at the crucial first step in the PA pathway making it an important candidate gene for investigation of involvement in PA phenotypic variation. In this study, PA level in perennial ryegrass was strongly associated with the presence or absence of the LpHSS1 gene. A genetic marker, developed for the presence/absence of LpHSS1, may be used for marker-assisted breeding to either lower or increase PAs in breeding populations of perennial or Italian ryegrass to investigate a potential role in the deterrence of herbivore pests. The presence of HSS-like genes in several other Poaceae species suggests that PA biosynthesis may occur in plant family members beyond perennial ryegrass and tall fescue and identifies a potential route for manipulating PA levels.
Similar content being viewed by others
Background
Pyrrolizidine alkaloids (PAs) are a diverse class of secondary metabolites produced by plants to deter herbivores [1, 2]. PAs are produced mainly in angiosperms and are predominantly found in species belonging to the Boraginaceae, Asteraceae, Fabaceae and Orchidaceae families [3, 4]. For several plant families PAs are notorious for their toxicity to livestock [5, 6] and occasionally humans, through use as herbal supplements [7] and via food contamination [8]. The lolines, produced by symbiotic endophytic fungi (Epichloe spp.), are one of the most well researched groups of PAs found in forage grasses [9]. Lolines are beneficial to the host plant by deterring a variety of insect pasture pests [10,11,12]. In contrast, little is known about plant-derived PAs in grasses and the role they could play as defence compounds. During the analysis of perennial ryegrass (Lolium perenne L.) metabolites, several abundant novel alkaloids were identified as ions using direct infusion mass spectrometry [13] and later characterised as the PAs E-thesinine-O-4′-α-rhamnoside, Z-thesinine-O-4′-α-rhamnoside and their glycoside derivatives [14]. This was the first time plant-derived PAs were identified in a grass from the Poaceae family. Whether these thesinine-rhamnosides play a role in deterring herbivores or have an alternative function is currently unknown.
Homospermidine synthase (HSS) catalyses the first major step in the PA pathway by transferring an aminobutyl group from spermidine to putrescine to form homospermidine. Homospermidine is then incorporated into the necine base which is the common moiety to all PAs. HSS genes have evolved from a duplication of deoxyhypusine synthase (DHS) [15] which is involved in the post-translational activation of eukaryotic initiation factor 5A (eIF5A) and is essential for cell proliferation [16]. This reaction is highly conserved across eukaroytes, and DHS and HSS genes have been well characterised in orchids [17] and several other angiosperms [18]. HSS differs from DHS in that it has lost its ability to bind and modify the eIF5A precursor protein [19].
Perennial ryegrass is a diploid outcrossing, self-incompatible species that is used extensively as a forage grass in temperate agricultural zones as well as being widely employed as an amenity grass. Breeding populations are highly heterogeneous, providing several challenges to breeders selecting for complex quantitative traits and to researchers applying marker-assisted selection [20]. Several large effect quantitative trait loci (QTL) have been identified in an F1 perennial ryegrass mapping population for some of the major alkaloids, including the E/Z-thesinine-rhamnoside PAs [21]. QTL mapping is often imprecise, identifying a large chromosomal region with numerous possible causative genes underlying the trait. Candidate-gene based association mapping [22] is one approach that can be subsequently used to identify the causative genes underlying trait variation, especially where strong evidence exists for the candidate gene from pathway biochemistry or mutational analysis. Rapid linkage disequilibrium (LD) decay is beneficial for high resolution association mapping to identify the gene and underlying functional nucleotide polymorphisms responsible for trait variation [23]. Linkage disequilibrium (LD) in perennial ryegrass rapidly decays for most genes, with r2 values of 0.2 within 500 bp [24, 25], where the population represents adequate diversity of the species. Consequently, several candidate gene-based association mapping studies have been performed in perennial ryegrass for a range of traits, including flowering time [26], drought tolerance [25] and fructan content [27].
In this study we analysed a diverse association mapping population and detected significant variation in thesinine-rhamnoside PAs and discovered that a homospermidine synthase gene (LpHSS1) was absent in some of the perennial ryegrass plants. To determine whether this polymorphism was responsible for variation in PA content we performed association tests that accounted for population structure. Significant associations were detected between low levels of the thesinine-rhamnoside group of PAs and the absence of the LpHSS1 gene (HSS1-). This provides an opportunity to determine the function of these plant PAs in ryegrass. Putative HSS genes identified in tall fescue, wheat, sorghum and maize may potentially enable the manipulation of PA levels in these other grass species.
Methods
Plant material
The association mapping population included 693 diverse individuals derived from a range of elite breeding populations, cultivars and wild ecotypes of perennial ryegrass (Additional file 1). All population entries were heat and fungicide-treated, prior to the trial, to remove fungal endophytic symbionts (Epichloë festucae var. lolii). This was necessary to eliminate variation in endophyte strain or endophyte status (infected vs. not infected) as potentially confounding factors in downstream analyses. Successful removal of endophyte was confirmed in each individual using tiller immuno-detection blots [28]. Sixteen plants from various Italian ryegrass cultivars (Lolium multiflorum Lam.) (Additional file 2) were also used to extract DNA for marker testing. Plants from the association mapping population were grown outdoors in 1.5 L pots filled with a commercial sand-peat potting mixture, at AgResearch Grasslands, Palmerston North (latitude 40.3523° S, longitude 175.6082° E), New Zealand. Each genotype was clonally replicated five times in a repeated (n = 5 replicate blocks) randomised row-column design which was constrained to avoid clonal replicates being present more than once in the same row or column. Plants were cut back every 3 weeks to simulate rotational grazing and plant material (predominantly consisting of the leaf blades) was harvested to a level of 40 mm above ground in May 2012 (Southern hemisphere late autumn).
LC-MS analysis of thesinine-rhamnoside content
Plant material was immediately placed into liquid nitrogen, freeze dried and ground to a fine powder in a laboratory sample mill. Fifty mg of ground plant material was weighed into a 2.0 mL micro-centrifuge tube along with one 3 mm zirconium bead to enhance extraction. One mL of extraction solvent (50% acetonitrile:50% water, 0.1% formic acid, containing 10 μg/mL dichloroflourescein and d2-tyrosine as internal standards) was added and the tube was capped, shaken for 1 min (Qiagen Tissue Lyser II) and then centrifuged in a Minifuge at 14,000 g for 3 min. Approximately 500 μL of the supernatant was placed in a 2 mL glass autosampler vial, capped and stored at − 20 °C until analysis. Samples were analysed by reverse phase (RP) LC-MS using an Agilent RRHD SB-C18 column and Thermo Exactive mass spectrometer with ESI in positive mode. The resulting data were subjected to peak detection process with PhenoAnalyzer (SpectralWorks Ltd., Manchester, UK), using the key data collection and processing parameter settings described previously [29]. The resulting peak data matrices underwent a series of quality control filtering before statistical analysis was conducted to probe into biological effects. These filtering steps, including missing value imputation, peak merge and deisotoping, run-order correction and batch effect corrections were accomplished using in-house scripts (available upon request). The peak intensity of four targeted metabolites were selected for this investigation. These are E-thesinine-rhamnoside (m/z 434.2159, RT 4.51), Z-thesinine-rhamnoside (m/z 434.2159, RT 4.73), E-thesinine-rhamnoside-glycoside (m/z 596.2681, RT 4.33), and Z-thesinine-rhamnoside-glycoside (m/z 596.2681, RT 4.55).
Statistical analysis of thesinine-rhamnoside traits
Thesinine-rhamnoside signal intensity data were analysed using the variance component analysis procedure, residual maximum likelihood (REML) option, in GenStat [30]. A completely random linear model was used in the analysis using the REML algorithm. The final, adjusted genotypic means were based on Best Linear Unbiased Predictors (BLUP’s). Estimation of the significance of the genotypic variance component for each trait was based on the log-likelihood ratio test. The variance components generated from the REML analysis were used to estimate clonal mean repeatability (Rc) [31], for the four metabolites, using the model Rc = σ2g/(σ2g + σ2ε/nr), where; σ2g is the genotypic component of variance, σ2ε the residual variance and nr is the number of replications. Pearson correlation coefficients amongst traits were obtained using GenStat [30]. BLUP’s were square root and log-transformed to assess the effect on association tests.
PCR marker development and linkage mapping
Two perennial ryegrass expressed sequence tags (ESTs) previously submitted to GenBank (GR522183 and GR514484) were identified as LpDHS and LpHSS1 and plasmid clones fully sequenced in the forward and reverse directions using Sanger sequencing. A third gene (LpHSS2) was identified during PCR amplification of the initial genes. All three perennial ryegrass genes are annotated according to function which has been validated by in vitro protein assays (Dietrich Ober, pers. comm). Genomic sequences were sourced from inhouse gene-enriched Genethresher sequences [32] and several gaps filled by the cloning and sequencing of PCR products (Additional file 3). PCR products were cloned into p-GEM-T (Promega) according to manufacturer’s directions. Colony PCR’s were performed on at least eight clones, purified using ExoSAP-IT (Applied Biosystems) according to manufacturer’s directions and Sanger sequenced. Sequences were assembled and consensus derived using the software Sequencher v4.10.1 (GeneCodes Corporation). Genomic DNA sequence and inferred protein sequences were deposited in GenBank for LpDHS (MF375394), LpHSS1 (MF375393) and LpHSS2 (MF375392). PCR primers specific to the LpHSS1 gene (Additional file 3) were used to test for the presence of the gene. Genomic DNA extractions for the entire association population and 16 Italian ryegrass plants were performed using the DNAeasy plant maxi kit (Qiagen) following the manufacturer’s recommended protocol. PCR amplification was performed in a total volume of 20 μl containing 20 ng of genomic DNA, 1.5 mmol/L MgCl2, 0.5 mmol/L dNTPs, 1 U Platinum Taq polymerase (Invitrogen), 1× Invitrogen PCR buffer, and 0.5 μmol/L primer. Primers F5, F9 and R8 were selected for evaluating the entire association mapping population for the presence or absence of LpHSS1 in a multiplex PCR reaction. In the multiplex reaction F9 and R8 primers were at 0.5 μmol/L and F5 at 0.125 μmol/L. To ensure a negative result for LpHSS1 was not a consequence of a failed PCR, assays were also multiplexed with an internal control based on the cab gene [33] using primers (cab-f 5’GTCTCGACTACCTCGGCAAC3’ cab-r 5’ACCGAACATGGAGAACATGG3’) at 0.075 μmol/L each. This internal positive control amplifies an expected product of 254 bp. Thermocycling was performed in a 9700 Gene Amp (Perkin Elmer) PCR machine with an initial cycle of 5 min at 94 °C; 35 cycles at 94 °C for 10 s, 57 °C for 10 s and 72 °C for 1 min; final extension at 72 °C for 5 min. In all other PCR reactions thermocycling was as above except annealing temperature varied as indicated in Additional file 3.
Using MAPMAKER/EXP 3.0 [34] the LpDHS1, LpHSS1 and LpHSS2 genes were mapped by genetic linkage analysis in an F1 population comprised of 94 perennial ryegrass individuals, using gene markers that reveal polymorphisms at the three loci (Additional file 4). Framework SSR markers are reported in a previously published linkage map [32].
Diversity analysis of DHS and HSS gene sequences
Festuca arundinacea (tall fescue) EST sequences were retrieved from Genbank by BLAST using LpDHS, LpHSS1 and LpHSS2. After assembly of ESTs into nucleotide consensus sequences protein translations for two tall fescue gene paralogs (FaDHS and FaHSS1) (Additional file 5) were obtained and used for sequence homology and phylogenetic analysis.
Sequences covering a range of Poaceae species were chosen from GenBank for sequence alignment and phylogenetic analysis and are as follows: NP_001051218 (Oryza sativa japonica DHS), XP_006650561 (Oryza brachyantha DHS), XP_003559916 (Brachypodium distachyon DHS), EMT19875 (Aegilops tauschii DHS), ACP28133 (Triticum aestivum DHS), AER42619 (Hordeum brevisubulatum DHS), XP_004981833 (Setaria italica DHS), XP_002444933 (Sorghum bicolor 07 g001640 DHS), XP_002466487 (Sorghum bicolor 01 g008630 DHS), XP_0024446268 (Sorghum bicolor 06 g011780 HSS), NP_001149084 (Zea mays DHS), NP_001130806 (Zea mays HSS), XP_020196793 (Aegilops tauschii HSS). The Triticum aestivum HSS was retrieved from the UniProt database (accession A0A1DSYB60). The Hordeum vulgare DHS amino acid sequence was obtained from a protein translation of the full length EST sequence AK248438. Two functionally characterised protein sequences were used from the monocot orchid Phalaenopsis amabilis CAR31338.1 (HSS), CAL69908.1 (DHS).
Protein and DNA alignments were performed using the Geneious software v7.1.5 (http://www.geneious.com [35]) with the ClustalW alignment option with a BLOSUM cost matrix. Gblocks v0.91b (web-based- http://molevol.cmima.csic.es/castresana/Gblocks_server.html) was used to eliminate poorly aligned positions and gaps using default parameters providing 238 amino acids for final analysis. An unrooted phylogenetic tree of DHS and HSS genes from the Poaceae family was constructed in Geneious using the PhyML (v3.2) plug in using the WAG amino acid substitution model with 1000 replicates to obtain bootstrap proportions.
Population structure and association testing
A core set of 308 SNP markers from 308 genes (Additional file 6), with minor allele frequency > 0.1 and which were dispersed relatively evenly across the genome (based on synteny with rice), were used for evaluating population structure (Q) and kinship (K). Population structure was assessed using the software STRUCTURE v2.3.4 [36]. The number of subpopulations (k) was determined by running k from 1 to 10 using the admixture model with a burn in time of 50,000 and 100,000 iterations for each run. Each k was run ten times and the software STRUCTURE HARVESTER [37] used to analyse results. The method of Evanno et al. [38] was used to identify the best estimate of k and was subsequently used to assign each individual to a Q-matrix. Association tests utilising BLUP’s for the PAs were performed in the software TASSEL version 5.2.9 [39] using GLM (Q) and MLM (Q + K) with the kinship matrix (K) calculated using the centred IBS option (Additional file 7) and the population structure matrix Q described above.
Results
Statistical analysis of thesinine-rhamnoside traits
The abundance in perennial ryegrass of the four targeted metabolites, E-thesinine-rhamnoside, Z-thesinine-rhamnoside, E-thesinine-rhamnoside-glycoside and Z-thesinine-rhamnoside-glycoside, were estimated based on peak intensities. Frequency distributions for each trait appeared right-skewed or bimodal (Additional file 8A). Both non-transformed BLUP’s and square root and log-transformed BLUP’s were tested in association analyses but all led to similar conclusions. Therefore, only results based on non-transformed BLUP’s are described from here on. All four metabolites demonstrated highly significant genotypic variation in the association population (Table 1). Rc, the upper limit of the degree of genetic determination [31], was also high for all four metabolites (> 0.948, Table 1), which further confirms a strong genetic basis to the observed phenotypic variation. Thesinine-rhamnoside traits were all significantly correlated to each other (P < 0.001) as determined by Pearson’s correlation coefficient analysis (Additional file 8B).
Genotyping and linkage mapping
A panel of 16 diverse perennial ryegrass plants from the association mapping population were used to validate the designed primer sets within the LpHSS1 gene (Fig. 1a). Two plants failed to amplify the LpHSS1 gene across all 6 primer sets tested (blue and red bar regions depicted in Fig. 1a). The plants that amplified PCR products gave positive reactions for all 6 primer sets. In a similar survey of 16 plants from nine Italian ryegrass cultivars, plants from six cultivars failed to amplify the LpHSS1 gene with all six primer sets (Additional file 2). Two primers (F5 + F9) were multiplexed with primer R8 (Fig. 1a and b) and a positive control primer set (to detect failed reactions) and used to genotype the full association mapping population for the presence (HSS1+) or absence (HSS1-) of the LpHSS1 gene. HSS1+ plants can either be homozygous or hemizygous for the LpHSS1 gene. The incidence of HSS1- across the entire association mapping population was 0.18. HSS1- was at a much higher incidence in some populations than others (Additional file 1). In a relatively restricted sample of wild ecotypes, HSS1- was observed in material sourced from regions as diverse as Greece, Italy, Tunisia and Uzbekistan. HSS1- was observed in many European cultivars (19/39) but not identified in any of the Australian cultivars and only 3 out of 18 New Zealand cultivars. Two of the New Zealand cultivars have a high incidence of HSS1- (Tolosa 8/13 plants; Grasslands Impact 2/4 plants). Genetic linkage analysis of the LpDHS, LpHSS1, and LpHSS2 mapped all genes from perennial ryegrass to the same location on the distal half of linkage group (LG) 4 (Fig. 2).
Genotyping assays for the presence (HSS1+) or absence (HSS1-) of the LpHSS1 gene. a Location of primers within the LpHSS1 gene. Black arrow bars represent exon locations of the LpHSS1 gene and the position of the start (ATG) and stop (TGA) codons are indicated. Blue bars represent PCR products from various primer combinations used to genotype plants. Red bars represent the F9 + R8 and F5 + R8 PCR products which were developed into a multiplex PCR assay for the genotyping of the entire association mapping population. Numbers given on blue and red bars are PCR product sizes. b Screening plants from the association mapping population for LpHSS1 using the F5 + F9 + R8 multiplex assay. The positive control product from the endogenous cab gene confirms a successful PCR when no LpHSS1 gene product is amplified.
Location of the DHS and HSS genes on linkage group 4 in perennial ryegrass, determined by genetic linkage analysis. Map distance in centimorgans is shown at the left and marker names to the right. Bold markers represent framework SSR markers [32]
Diversity Analysis of DHS and HSS gene sequences in the Poaceae
Orthologs of the LpDHS and LpHSS1 genes in tall fescue shared an amino acid identity of 97.7 (FaDHS) and 96.3% (FaHSS1) respectively (Fig. 3a and b). No tall fescue sequences for LpHSS2 were identified in GenBank. Sequences from Poaceae species were putatively assigned function as DHS or HSS based on combined lines of evidence. Genes were assigned as HSS’s if the sequence was divergent from the main DHS phylogenetic clade, they contained conserved amino acid changes known to favour HSS activity [40], and if a second gene that resided in the DHS phylogenetic clade was present. Species with a single gene present from a sequenced genome also supported assignment as DHS. From the phylogenetic tree (Fig. 4) most Poaceae amino acid sequences cluster with DHS in respective subfamilies. However, both wheat, Aegilops tauschii (ancestor of wheat), maize and sorghum possessed sequences divergent from the main DHS clade, as observed for Lolium/Festuca HSS’s, suggesting they may function as HSS’s. In one particular key region of these two proteins, several amino acid differences (255, 277 and 312) are conserved with HSS-like sequences compared to DHS sequences in Poaceae (Fig. 5). The valine 277 is also conserved in a homospermidine synthase functionally characterised in orchids (Fig. 5) [17].
Population structure
The software STRUCTURE HARVESTER was used to analyse output from STRUCTURE (testing k = 1–10) to determine that the optimal number of population subgroups within the association mapping population was three (Additional file 9). The three subpopulations reflected geographical origins to some extent. Group 1 was comprised of cultivars developed in Europe, natural populations sourced from Europe and the majority of cultivars from New Zealand and Australia. Group 2 mainly identified North African–sourced populations and New Zealand breeding populations with a known North African contribution (Dr Alan Stewart, pers. comm). The third group principally included New Zealand cultivars and breeding populations with a recognised basis in germplasm sourced from Northwest Spain (e.g. Tolosa and Impact cultivars) [41] and derived breeding populations (Dr Alan Stewart pers. comm).
Association of HSS1- with thesinine-rhamnoside levels
Statistical tests were performed for the association of thesinine-rhamnoside PA levels with the presence or absence of the LpHSS1 gene. Significant associations were detected for all four thesinine-rhamnoside PAs using either the MLM (Q + K) or GLM method (Q) implemented in the software TASSEL (Table 2). The strongest association using MLM analysis, which corrects for population structure and kinship, was for E-Thesinine-rhamnoside (p = 1 × 10− 17) with a marker R2 value of 0.110. The E/Z-thesinine glycosides had higher p values (8.5 × 10− 13 and 2.9 × 10− 10) and lower R2 values (0.076 and 0.059) but were still highly significant associations. The HSS1- genotype has a large effect on reducing thesinine-rhamnoside content when compared to plants that possess the HSS1+ allele in the association population (Fig. 6a). For example, a 5-fold difference in E-thesinine rhamnoside levels occurred between HSS1- and HSS1+ plants (Fig. 6a). Decreased thesinine-rhamnoside levels in HSS1- plants is also evident when analysed by subpopulation (Fig. 6b).
a Effect of HSS1- on the content of different thesinine-rhamnosides in the association mapping population of 693 plants. b Effect of HSS1- on E-thesinine-rhamnoside content in different population subgroups as identified by the programme STRUCTURE. ** and *** indicate significance at P<0.01 and P<0.001 respectively. Error bars represent standard error of the mean
Discussion
Plant-derived PAs have been identified and studied mainly in angiosperm species [3, 4] and less so in the commercially important grasses of the Poaceae family where they have only been identified in a few Lolium and Festuca species [14]. In this study, the identification of a homospermidine synthase gene (LpHSS1) that is present/absent in a perennial ryegrass association mapping population has allowed us to determine that the LpHSS1 gene is actively involved in the biosynthesis of PAs and is partially responsible for phenotypic variation of PA levels.
The DHS and HSS genes of perennial ryegrass were mapped using genetic linkage analysis to the distal half of linkage group LG4. All 3 genes are co-located (Fig. 2) indicating that they most likely evolved through tandem duplication. Their map location is not in the same region as large effect QTLs identified previously for thesinine-rhamnoside PAs [21], which were proximally located in LG4. Other genes residing further down the PA pathway or transcriptional regulator genes not yet identified may explain the earlier identified QTLs.
The majority of New Zealand perennial ryegrass cultivars originate from a narrow base of two ecotypes (Hawkes Bay and Mangere) [41] which is the likely reason for the low incidence of HSS1- in this germplasm source. The New Zealand cultivars Tolosa and Grasslands Impact, have a notably high incidence of HSS1- attributed to introgression of Northwest Spain germplasm [41]. The further introduction of NW Spain and other European germplasm into New Zealand breeding populations [41] may disperse HSS1- more widely. The relatively wide-spread dispersal of HSS1- within perennial ryegrass and Italian ryegrass indicates that this locus is not essential and probably incurs no major penalty to fitness.
The three subpopulations (k) identified in this study were similar to the subpopulations determined in an earlier perennial ryegrass association mapping study [42]. In both these and other studies [25, 43] the main subpopulation group identified is from Europe, New Zealand and Australia and represents the majority of commercial cultivars and elite breeding populations. The association mapping population in our study was biased towards this major subpopulation group so as to identify markers in commercially relevant breeding populations. Wild ecotypes from Africa and Europe are less adapted to the New Zealand environment, with a high mortality rate. As a consequence our association population did not cover the full diversity of the species and could explain why several minor subpopulation groups reported in another study [25] were not detected here. Typically candidate gene-based association mapping approaches rely on testing multiple SNPs that tag gene haplotypes in the hope of identifying an underlying functional nucleotide polymorphism (FNP) or one in LD to the FNP responsible for trait variation [22]. In this study the presence/absence of a gene among individuals from a population at a relatively high incidence allowed us to quickly evaluate whether the gene was responsible for trait variation avoiding an initial laborious SNP discovery and haplotype determination phase. For the presence/absence of the LpHSS1 gene the effect on level of thesinine-rhamnosides is relatively large (Fig. 6), as would be expected for a candidate gene at the crucial first step of the pathway. This strong functional association is substantiated by the relatively large-sized population (693 plants) coupled with the very small p-values obtained (< 6 × 10− 10) for marker-PA associations. Marker p-values using GLM (Q) analysis were probably inflated compared to MLM (Q + K) which additionally accounts for kinship within the population and results in improved control of type I and type II error rates [44, 45]. Reduced levels of PAs in HSS1- plants were found for all 3 sub-population groups identified (Fig. 6b) providing additional assurance that associations are not falsely attributed to population structure effects.
In most species a single functional DHS gene exists [15]. Of interest is the identification of two sequences from sorghum that are relatively similar to each other and grouped with other DHS genes. The sorghum protein from locus 07 g001640 appears truncated and a result of an incorrect protein prediction as exon1 and exon2 are present in the genome sequence. If this locus encodes a functional gene it would be of interest to determine whether both sorghum genes can function as DHS as has been found in the species Crotalaria juncea [46]. In species that produce PAs, the DHS gene has been duplicated and recruited as a HSS [17, 46]. When analysing sequences of the Poaceae family wheat, maize and sorghum were found to possess sequences that had diverged significantly from DHS. These divergent sequences, as identified in perennial ryegrass and tall fescue, are more likely to function as HSS but require functional validation to be certain. Further evidence for their role as HSSs includes several amino acid changes that are different when compared to DHS gene sequences in Poaceae and extend beyond this family to a number of angiosperm plant families [40]. One of the key amino acid differences when comparing HSS sequences across species is amino acid number 277 (valine) (Fig. 5), which has been shown to favour the HSS reaction substrate putrescine rather than the DHS substrate eIF5A [40]. No PAs have been identified or reported from maize or sorghum but now the identification of putative HSS genes from these two species suggests that a range of undiscovered PAs could be present. Given the absence of the LpHSS1 gene regularly observed in perennial ryegrass and pseudogenes detected in other plant species [17, 40, 46] naturally occurring mutations in HSS genes could explain PA variation in plants more broadly, given their key location in the PA pathway.
After a gene duplication, as in the case of HSS from DHS, pseudogenization can occur as two genes of identical function will not be stabilised in the genome [47] unless an additional product of the gene is advantageous. In the case of HSS in many plant species, the gene has undergone adaptive evolution to be recruited into a new role of herbivore defence through PA production [18, 19]. Symbiotic Epichloё fungal endophytes of perennial ryegrass [11] produce PAs and a range of other alkaloids, which can be strain specific [48], and which deter various insect pests. More effective host-endophyte relationships might contribute to functional redundancy of HSS genes and thereby hasten the pseudogenization process, resulting in eventual elimination from the perennial ryegrass genome.
Specialist plant insect herbivores have adapted to PAs through various methods of detoxification [2]. PAs introduced into grass plants from symbiotic endophytic fungi on the other hand have proven to be successful as feeding deterrents to invertebrates in several pasture grasses [10, 12]. It is currently unknown whether the thesinine-rhamnoside PAs and others as yet unidentified in perennial ryegrass play a role in insect deterrence. It would be of interest to select families of plants that possess the full range of concentrations of these particular metabolites to investigate feeding response by a range of insects. PAs from some plant species are also renowned for their toxicity to mammalian livestock, where livers can be damaged by pyrrols that are released as breakdown products [6, 49]. Livestock that forage on widely-used pasture grasses, such as perennial ryegrass, are likely to have adapted to the specific assortment of plant defensive PAs ingested, which are presumably negligible in toxicity, with detoxification by rumen bacteria as one potential mechanism of adaption [50]. If plants are bred for extremely high levels of thesinine-PAs, livestock feeding trials to evaluate the potential for any adverse effect should be conducted. The marker developed for the LpHSS1 gene will facilitate the selection of high and low PA plants to investigate whether it is beneficial to increase levels to deter insect pests or reduce levels if any adverse effect occurs in livestock.
In this study we investigated the thesinine-rhamnoside PAs as these are the only perennial ryegrass PAs identified to date. As the metabolome of perennial ryegrass is further characterized, other PAs of interest will inevitably be identified. Given that the LpHSS1 gene is at a crucial step in the beginning of the PA pathway it is likely that HSS1- genotypes will similarly reduce the levels of any other PAs. The conclusive results of the LpHSS1 gene being implicated in PA variation suggests future investigation into the LpHSS1, LpHSS2 and LpDHS genes for FNPs associated with PA levels is a worthwhile pursuit.
Conclusions
In this study we have identified a HSS gene that is absent in a diverse range of perennial and Italian ryegrass populations. The absence of the gene is strongly associated with reduced levels of the thesinine-rhamnoside group of PAs in perennial ryegrass. The presence of HSS-like genes in several other Poaceae species provides evidence that PA biosynthesis may not be limited to perennial ryegrass and tall fescue within this plant family and identifies a potential route for manipulating PA levels.
Abbreviations
- DHS:
-
Deoxyhypusine synthase
- ESTs:
-
Expressed sequence tags
- FNP:
-
Functional nucleotide polymorphism
- GLM:
-
General linear model
- HSS:
-
Homospermidine synthase
- LC-MS:
-
Liquid chromatography-mass spectrometry
- LD:
-
Linkage disequilibrium
- LG:
-
Linkage group
- MLM:
-
Mixed linear model
- PA:
-
Pyrrolizidine alkaloid
- QTL:
-
Quantitative trait loci
- SSR:
-
Simple sequence repeat
References
Macel M. Attract and deter: A dual role for pyrrolizidine alkaloids in plant-insect interactions. Phytochem Rev. 2011;10:75–82.
Wei X, Vrieling K, Mulder PPJ, Klinkhamer PGL. Testing the generalist-specialist dilemma: The role of pyrrolizidine alkaloids in resistance to invertebrate herbivores in Jacobaea species. J Chem Ecol. 2015;41:159–67.
Roeder E. Medicinal plants in China containing pyrrolizidine alkaloids. Pharmazie. 2000;55:711–26.
Stegelmeier BL, Edgar JA, Colegate SM, Gardner DR, Schoch TK, Coulombe RA, Molyneux RJ. Pyrrolizidine alkaloid plants, metabolism and toxicity. J Nat Toxins. 1999;8:95–116.
Fletcher MT, McKenzie RA, Blaney BJ, Reichmann KG. Pyrrolizidine alkaloids in Crotalaria taxa from northern Australia: Risk to grazing livestock. J Agric Food Chem. 2009;57:311–9.
Shimshoni JA, Mulder PPJ, Bouznach A, Edery N, Pasval I, Barel S, Abd-El Khaliq M, Perl S. Heliotropium europaeum poisoning in cattle and analysis of its pyrrolizidine alkaloid profile. J Agric Food Chem. 2015;63:1664–72.
Teschke R, Eickhoff A. Herbal hepatotoxicity in traditional and modern medicine: Actual key issues and new encouraging steps. Front Pharmacol. 2015;6:72.
Edgar JA, Roeder E, Molyneux RJ. Honey from plants containing pyrrolizidine alkaloids: A potential threat to health. J Agric Food Chem. 2002;50:2719–30.
Schardl CL, Grossman RB, Nagabhyru P, Faulkner JR, Mallik UP. Loline alkaloids: Currencies of mutualism. Phytochemistry. 2007;68:980–96.
Bryant RH, Cameron NE, Edwards GR. Response of black beetle and red-headed pasture cockchafer larvae to loline alkaloids in meadow fescue roots. N Z Plant Prot. 2010;63:219–23.
Popay AJ, Bonos SA. Biotic responses in endophytic grasses. In: Roberts CA, West CP, Spiers DE, editors. Neotyphodium in cool-season grasses. Oxford, UK: Blackwell Publishing Ltd; 2005. p. 163–85.
Riedell WE, Kieckhefer RE, Petroski RJ, Powell RG. Naturally-occurring and synthetic loline alkaloid derivatives: Insect feeding behaviour modification and toxicity. J Entomol Sci. 1991;26:122–9.
Koulman A, Tapper BA, Fraser K, Cao M, Lane GA, Rasmussen S. High-throughput direct-infusion ion trap mass spectrometry: A new method for metabolomics. Rapid Commun Mass Spectrom. 2007;21:421–8.
Koulman A, Seeliger C, Edwards PJB, Fraser K, Simpson W, Johnson L, Cao M, Rasmussen S, Lane GA. E/Z-Thesinine-O-4′-α-rhamnoside, pyrrolizidine conjugates produced by grasses (Poaceae). Phytochemistry. 2008;69:1927–32.
Ober D, Hartmann T. Homospermidine synthase, the first pathway-specific enzyme of pyrrolizidine alkaloid biosynthesis, evolved from deoxyhypusine synthase. Proc Natl Acad Sci U S A. 1999;96:14777–82.
Nishimura K, Murozumi K, Shirahata A, Park MH, Kashiwagi K, Igarashi K. Independent roles of eIF5A and polyamines in cell proliferation. Biochem J. 2005;385:779–85.
Nurhayati N, Gondé D, Ober D. Evolution of pyrrolizidine alkaloids in Phalaenopsis orchids and other monocotyledons: Identification of deoxyhypusine synthase, homospermidine synthase and related pseudogenes. Phytochemistry. 2009;70:508–16.
Reimann A, Nurhayati N, Backenköhler A, Ober D. Repeated evolution of the pyrrolizidine alkaloid-mediated defense system in separate angiosperm lineages. Plant Cell. 2004;16:2772–84.
Ober D, Harms R, Witte L, Hartmann T. Molecular evolution by change of function: Alkaloid-specific homospermidine synthase retained all properties of deoxyhypusine synthase except binding the eIF5a precursor protein. J Biol Chem. 2003;278:12805–12.
Conaghan P, Casler MD. A theoretical and practical analysis of the optimum breeding system for perennial ryegrass. Ir J Agric Food Res. 2011;50:47–63.
Koulman A, Cao M, Faville M, Lane G, Mace W, Rasmussen S. Semi-quantitative and structural metabolic phenotyping by direct infusion ion trap mass spectrometry and its application in genetical metabolomics. Rapid Commun Mass Spectrom. 2009;23:2253–63.
Zhu CS, Gore M, Buckler ES, Yu JM. Status and prospects of association mapping in plants. Plant Genome. 2008;1:5–20.
Carlton VEH, Ireland JS, Useche F, Faham M. Functional single nucleotide polymorphism-based association studies. Hum Genomics. 2006;2:391–402.
Xing Y, Frei U, Schejbel B, Asp T, Lübberstedt T. Nucleotide diversity and linkage disequilibrium in 11 expressed resistance candidate genes in Lolium perenne. BMC Plant Biol. 2007;7:43.
Yu X, Bai G, Liu S, Luo N, Wang Y, Richmond DS, Pijut PM, Jackson SA, Yu J, Jiang Y. Association of candidate genes with drought tolerance traits in diverse perennial ryegrass accessions. J Exp Bot. 2013;64:1537–51.
Skøt L, Sanderson R, Thomas A, Skøt K, Thorogood D, Latypova G, Asp T, Armstead I. Allelic variation in the perennial ryegrass FLOWERING LOCUS T gene is associated with changes in flowering time across a range of populations. Plant Physiol. 2011;155:1013–22.
Skøt L, Humphreys J, Humphreys MO, Thorogood D, Gallagher J, Sanderson R, Armstead IP, Thomas ID. Association of candidate genes with flowering time and water-soluble carbohydrate content in Lolium perenne (L.). Genetics. 2007;177:535–47.
Hahn H, Huth W, Schöberlein W, Diepenbrock W. Detection of endophytic fungi in Festuca spp. by means of tissue print immunoassay. Plant Breed. 2003;122:217–22.
Fraser K, Lane GA, Otter DE, Harrison SJ, Quek SY, Hemar Y, Rasmussen S. Non-targeted analysis by LC-MS of major metabolite changes during the oolong tea manufacturing in New Zealand. Food Chem. 2014;151:394–403.
GenStat:GenStat for Windows 17th Edition. https://www.vsni.co.uk/downloads/genstat/17th-edition/.
Falconer DS. Introduction to quantitative genetics 3rd ed. Harlow: Longman Scientific & Technical; 1989.
Gill GP, Wilcox PL, Whittaker DJ, Winz RA, Bickerstaff P, Echt CE, Kent J, Humphreys MO, Elborough KM, Gardner RC. A framework linkage map of perennial ryegrass based on SSR markers. Genome. 2006;49:354–64.
Patel M, Milla-Lewis S, Zhang W, Templeton K, Reynolds WC, Richardson K, Biswas M, Zuleta MC, Dewey RE, Qu R, Sathish P. Overexpression of ubiquitin-like LpHUB1 gene confers drought tolerance in perennial ryegrass. Plant Biotechnol J. 2015;13:689–99.
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L. MAPMAKER: An interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics. 1987;1:174–81.
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, et al. Geneious basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28:1647–9.
Pritchard JK, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics. 2000;155:945–59.
Earl DA, von Holdt BM. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Gen Res. 2012;4:359–61.
Evanno G, Regnaut S, Goudet J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol Ecol. 2005;14:2611–20.
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES. TASSEL: Software for association mapping of complex traits in diverse samples. Bioinformatics. 2007;23:2633–5.
Kaltenegger E, Eich E, Ober D. Evolution of homospermidine synthase in the Convolvulaceae: A story of gene duplication, gene loss, and periods of various selection pressures. Plant Cell. 2013;25:1213–27.
Stewart AV. Genetic origins of perennial ryegrass (Lolium perenne) for New Zealand pastures. In: Mercer CF, editor. Advances in pasture plant breeding grassland research and practice series no 12. Dunedin: New Zealand Grassland Association; 2006. p. 55–62.
Tang J, Yu X, Luo N, Xiao F, Camberato JJ, Jiang Y. Natural variation of salinity response, population structure and candidate genes associated with salinity tolerance in perennial ryegrass accessions. Plant Cell Environ. 2013;36:2021–33.
Yu X, Pijut PM, Byrne S, Asp T, Bai G, Jiang Y. Candidate gene association mapping for winter survival and spring regrowth in perennial ryegrass. Plant Sci. 2015;235:37–45.
Yu J, Pressoir G, Briggs WH, Bi IV, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, et al. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet. 2006;38:203–8.
Zhao K, Aranzana MJ, Kim S, Lister C, Shindo C, Tang C, Toomajian C, Zheng H, Dean C, Marjoram P, Nordborg M. An Arabidopsis example of association mapping in structured samples. PLoS Genet. 2007;3:e4.
Nurhayati N, Ober D. Recruitment of alkaloid-specific homospermidine synthase (HSS) from ubiquitous deoxyhypusine synthase: Does Crotalaria possess a functional HSS that still has DHS activity? Phytochemistry. 2005;66:1346–57.
Nowak MA, Boerlijst MC, Cooke J, Smith JM. Evolution of genetic redundancy. Nature. 1997;388:167–70.
Takach JE, Mittal S, Swoboda GA, Bright SK, Trammell MA, Hopkins AA, Young CA. Genotypic and chemotypic diversity of Neotyphodium endophytes in tall fescue from Greece. Appl Environ Microbiol. 2012;78:5501–10.
Seawright AA. Directly toxic effects of plant chemicals which may occur in human and animal foods. Nat Toxins. 1995;3:227–32.
Wachenheim DE, Blythe LL, Craig AM. Characterization of rumen bacterial pyrrolizidine alkaloid biotransformation in ruminants of various species. Vet Hum Toxicol. 1992;34:513–7.
Acknowledgements
Authors would like to thank Alan Stewart (PGG Wrightson Seeds) for provision of elite germplasm for the association mapping population. Casey Flay, Michelle Ebett and Brian Maw from AgResearch who co-ordinated harvest activities and maintained the trial. Catherine Carter, David Whittaker, Sathish Puthigae, Zac Hanley and Kerry Templeton from ViaLactia Biosciences for assistance with DNA extractions. Leo Liu, Daniel Hughes, Arvind Subbaraj and Tom Featonby for assistance with metabolomics extractions and analyses. The authors would like to thank Dr. Tony Conner and Dr. Jeanne Jacobs for critical review of the manuscript.
Funding
The research was funded by Pastoral Genomics, a joint venture co-funded by DairyNZ, Beef+Lamb New Zealand, Dairy Australia, AgResearch Ltd., New Zealand Agriseeds Ltd., Grasslands Innovation Ltd., DEEResearch, and the Ministry of Business, Innovation and Employment (New Zealand).
Availability of data and materials
The data sets supporting the results of this article are provided as additional files.
Author information
Authors and Affiliations
Author notes
Jan Huege is deceased. This paper is dedicated to his memory.
- Jan Huege
Contributions
GG contributed to the conception of the study, experimental design, genetic linkage mapping, population structure analysis, association testing, sequence analysis and wrote the manuscript. CB performed sequence analysis, primer design, genotyping and DNA extractions. MF contributed by performing genotyping and DNA extractions. JH performed the bulk of the metabolite data extraction and data analysis, KF contributed to analytical measurements of the PAs, assisted with data extraction and data analysis and wrote the PA analysis methods section of the manuscript. CJ contributed to the conception of the study, experimental design of the trial and writing of the manuscript. MC assisted with metabolite data extraction and data analysis. MJF contributed to experimental design, statistical analysis of traits, development of association population, structure analysis and writing of the manuscript. All authors read, edited, and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional files
Additional file 1:
Perennial ryegrass germplasm identity and association mapping data. Includes plant number, cultivar/breeding population name, country origin, population type, Margot Forde germplasm accession number, thesinine-rhamnoside BLUPs, presence/absence of HSS1 polymorphism and population structure Q values. (XLSX 79 kb)
Additional file 3:
Primer sequences used for PCR and sequencing of the DHS and HSS genes. (XLSX 13 kb)
Additional file 4:
Genetic markers used for linkage mapping of the LpDHS, LpHSS1 and LpHSS2 genes in a F1 population of perennial ryegrass. (PDF 428 kb)
Additional file 5:
Tall Fescue FaDHS and FaHSS1 translated protein sequences derived from ESTs retrieved from GenBank. (FASTA 1 kb)
Additional file 6:
Information for 308 SNPs used for population structure analysis of the perennial ryegrass association mapping population. (XLSX 28 kb)
Additional file 7:
Kinship matrix of the association population calculated using 308 SNP markers and the TASSEL kinship centred IBS option. (XLSX 5357 kb)
Additional file 8:
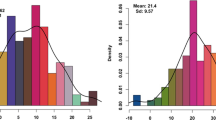
Statistical analysis of thesinine-rhamnoside traits in perennial ryegrass. A) Frequency distribution of untransformed thesinine-rhamnoside BLUP’s. B) Pearson correlation coefficients among levels of thesinine-rhamnoside PAs. (PDF 537 kb)
Additional file 9:
STRUCTURE HARVESTER results used to determine number of subpopulations (k) in the perennial ryegrass association mapping population. (PDF 494 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Gill, G.P., Bryant, C.J., Fokin, M. et al. Low pyrrolizidine alkaloid levels in perennial ryegrass is associated with the absence of a homospermidine synthase gene. BMC Plant Biol 18, 56 (2018). https://doi.org/10.1186/s12870-018-1269-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12870-018-1269-6