Abstract
Background
Salmonella can reside in healthy animals without the manifestation of any adverse effects on the carrier. If raw products of animal origin are not handled properly during processing or cooked to a proper temperature during preparation, salmonellosis can occur. In this research, we developed bioluminescent Salmonella strains that can be used for real-time monitoring of the pathogen's growth on food products. To accomplish this, twelve Salmonella strains from the broiler production continuum were transformed with the broad host range plasmid pAKlux 1, and a chicken skin attachment model was developed.
Results
Salmonella strains carrying pAKlux 1 constitutively expressed the luxCDABE operon and were therefore detectable using bioluminescence. Strains were characterized in terms of bioluminescence properties and plasmid stability. To assess the usefulness of bioluminescent Salmonella strains in food safety studies, we developed an attachment model using chicken skin. The effect of washing on attachment of Salmonella strains to chicken skin was tested using bioluminescent strains, which revealed the attachment properties of each strain.
Conclusion
This study demonstrated that bioluminescence is a sensitive and effective tool to detect Salmonella on food products in real-time. Bioluminescence imaging is a promising technology that can be utilized to evaluate new food safety measures for reducing Salmonella contamination on food products.
Similar content being viewed by others
Background
Salmonella enterica, with over 2000 different serovars, is indigenous to the gastrointestinal tracts of many mammals, birds, and reptiles, usually in low levels. Salmonella can be pathogenic in these animals if it reaches certain numbers in vivo. However, moribund animals are usually culled before they reach slaughter. Therefore, the food safety problem relative to humans is that Salmonella can be carried into the processing plant in healthy asymptomatic animals. If cross-contamination occurs during the slaughter process, Salmonella can then be transferred to carcasses that were previously uncontaminated. A significant increase in the number of Salmonella positive broilers between exiting the scalding water immersion tank and exiting the crop extractor demonstrates how in-plant cross-contamination can occur [1, 2]. When carcasses were sampled at 6 points in processing, an increase occurred in the presence of Salmonella from 19% pre-scald to 36.9 % post-chill (after exiting the immersion chill tank).
Regardless of whether the source of contamination was pre-harvest or during processing, Salmonella is difficult to remove from carcasses due to its ability to adhere to chicken skin and endure the stages of processing [3]. Laboratory research, as well as in-plant trials, has demonstrated this relationship [4–7]. Therefore, persistence of Salmonella within the processing plant may be partially explained by interactions between chicken skin and Salmonella. Chemical treatments have been developed and shown to be effective under controlled conditions in the reduction of Salmonella levels on broiler carcasses or skin [8–11]. In spite of these efforts, Salmonella is yet to be eliminated from the process.
When testing the efficacy of different antimicrobial compounds, conventional cultural techniques are typically used. The product is sampled, microbiological culturing protocols are followed, and 48 to 72 hours later the results are known. Bioluminescence imaging (BLI) is a technique that can be used for real-time quantification and tracking of live bacteria in hosts [12–15]. To enable BLI, bacteria are tagged with bacterial luciferase, which catalyzes the oxidation of a long-chain aldehyde and FMNH2 to cause emission of visible light [16–19]. Salmonella strains constitutively expressing bacterial luciferase have no significant alterations in phenotype, including growth kinetics [20, 21] or biochemical, serological, or structural phenotypes [20, 22]. Advantages of the bacterial luciferase reporter system include negligible background [23], no toxic or phenotypic effects from accumulation of signal [24], real-time detection, and no need for addition of an exogenous substrate. In addition, bioluminescence from bacterial luciferase correlates well with the amount of luciferase protein and lux mRNA [25, 26], bacterial plate counts [20, 27], and intracellular numbers of bacteria in cell culture [28].
In this research, our aim was to develop twelve bioluminescent Salmonella enterica strains that can be used for real-time monitoring of the pathogen's growth on food products. Our study is unique in that it includes multiple Salmonella field strains isolated from the broiler production continuum, including post hatchery, prior to harvest, arrival at the plant, pre-chill tank, and post-chill tank.
Results and discussion
Bioluminescent Salmonella strains
pAKlux 1 was transferred to twelve Salmonella strains that we isolated in a study on the poultry production and processing continuum. The strains represented the twelve most commonly isolated serovars from our study. This result compared favorably with a previous report where transformation of a lux plasmid into Salmonella isolates from a poultry processing plant was only successful for one isolate out of seven attempted [20]. Expression of the luxCDABE operon, which encodes bacterial luciferase, was driven by the lacZ promoter on pAKlux 1. Because Salmonella does not have lacIq in its chromosome, it constitutively expresses the lacZ promoter on pAKlux 1 and hence produces continuous light while it is alive and metabolically active.
We showed that bacteria numbers and bioluminescence correlated well (R2 = 0.99) in all strains used (Figure 1). The minimum detectable numbers for all twelve strains was less than 1500 CFU/ml, and it was less than 300 CFU/ml for a majority of strains (S. Alachua, 334 CFU/ml; S. Braenderup, 217 CFU/ml; S. Enteritidis, 175 CFU/ml; S. Heidelberg, 169 CFU/ml; S. Kentucky, 229 CFU/ml; S. Mbandaka, 248 CFU/ml; S. Montevideo, 209 CFU/ml; S. Newport, 125 CFU/ml; S. Schwarzengrund, 1470 CFU/ml; S. Seftenberg, 1386 CFU/ml; S. Thompson, 1044 CFU/ml; and S. Typhimurium, 202 CFU/ml). This result was comparable to previous studies [13, 15, 29, 30].
Correlation between bioluminescence and bacterial numbers. The correlation between luminescent signals and bacteria numbers for representative serovars S. Montevideo and S. Newport. A) 96-well plate containing S. Montevideo (rows A, B, C, and D) and S. Newport (rows E, F, G, and H). 25 μl of bacteria suspension from columns 11 and 12 were spread on LB agar plates with ampicillin (100 μg ml-1) to determine the concentration of colony forming units. B) plot of bioluminescence against bacterial numbers for S. Montevideo. C) plot of bioluminescence against bacterial numbers for S. Newport.
The average theoretical light intensity per CFU was calculated for each strain and revealed a greater than 10-fold difference between some serovars (Figure 2). This difference was not due to lack of viability; bacterial plate counts showed that all the strains were viable at this stage and that viable bacterial densities for all the serovars were within a 3 fold range. The difference in luminescence could reflect a difference between serovars in efficiency of luxCDABE expression from the lacZ promoter, or it could reflect a difference between serovars in the activity of bacterial luciferase within the bacteria. Alternatively, the difference could reflect differences in metabolic activity between the serovars at this stage of growth (16 hours). Bioluminescence is known to correlate well with bacterial metabolism [31, 32].
Expression of bioluminescence in Salmonella serovars. The theoretical amount of bioluminescence produced per CFU of each Salmonella serovar following 16 hrs of growth was calculated. Bioluminescence per cfu was calculated by dividing the background subtracted bioluminescence value (in p/s/cm2/sr) in each well by the number of bacteria (determined by serial dilution and plate counts). The mean and standard error from four replicates were then determined for each strain. Light emission was almost 12 fold higher in S. Newport as compared to S. Schwarzengrund.
Plasmid stability
The stability of pAKlux 1 was determined by subculturing bioluminescent Salmonella broth cultures under selective (with ampicillin) and non-selective (no ampicillin) conditions for 15 days. Bioluminescence of Salmonella strains cultured under non-selective conditions was declining by day 2 and continued declining linearly (R2 = 0.95) until the conclusion of the experiment (Figure 3). Based on the data, the average half-life of pAKlux 1 in Salmonella was approximately 7 days under the described culture conditions. However, the stability varied between Salmonella serovars. Among the Salmonella strains, plasmid stability was lower in S. Kentucky and S. Typhimurium as compared to the others. The half-life of pAKlux 1 was about 4 days in S. Kentucky and about 5 days in S. Typhimurium (Figure 3). S. Newport, S. Slachua, and S. Enteritidis maintained pAKlux 1 longer than others, with a half life of about 9 days.
Stability of pAK lux 1 in Salmonella. Salmonella strains carrying pAKlux 1 were subcultured under ampicillin selective and non-selective conditions for 15 days. At each passage, bioluminescence was measured and normalized for cell density (OD600). The ratio between the normalized values under non-selective (nLB) and ampicillin selected conditions (nLBamp) revealed the plasmid stability in different Salmonella serovars. The black line represents the mean of eleven strains, and the red line is the linear trend line.
pAKlux 1 is derived from the broad host range plasmid pBBR1, which is relatively stable in gram-negative bacteria in the absence of antibiotic selection [33–35]. pAKlux 1 was stable in the gram-negative species Edwardsiella ictaluri for at least 10 days without antibiotic selection, and the plasmid caused no alterations in growth kinetics, native plasmids, and pathogenicity as compared to the parent strain [15]. Data from the current study indicates that the pBBR1 replicon is not as stable in Salmonella as it is in other gram-negative bacteria. However, bioluminescent Salmonella strains labeled with pAKlux 1 should be suitable for short term experiments in which antibiotic selection cannot be applied.
Characterization of skin attachment properties of Salmonella strains
We developed an in vitro skin attachment model for characterization of attachment properties of different Salmonella strains using BLI. Bioluminescence was successfully detected on chicken skin after being exposed to Salmonella strains expressing pAKlux 1. Using this model, we were able to show that Salmonella strains from different serovars vary in their ability to attach to chicken skin (Figure 4). Bacteria numbers in S. Seftenberg, S. Thompson, and S. Schwarzengrund were significantly different from each other and from other strains (P ≤ 0.05). The S. Heidelberg strain had significantly less binding to chicken skin than all the other strains and was 79 fold lower than S. Seftenberg, which had the highest amount of binding (Figure 4). This result suggests that Salmonella strains vary in their ability to bind to chicken skin; this ability may be a discriminating factor determining whether Salmonella strains persist through processing or whether they are removed.
Attachment of Salmonella strains to chicken skin. The amount of Salmonella attachment to chicken skin was determined by measuring the bioluminescence following 1 h incubation of bacterial suspension with quadruplicate skin samples. Bacteria numbers of each strain were estimated from the bioluminescence values using each strain's linear correlation formula. Unattached Salmonella were removed by gentle washing prior to bioluminescence measurements. The bacteria number for each strain is the geometric mean of four replicates from three separate experiments. Letters on the right of the graph indicate statistical groupings as determined by Tukey's test. Values with the same letter are not significantly different (P > 0.05).
The effect of washing on Salmonella removal from chicken skin
Bioluminescence was an effective tool for measuring the effects of washing for removal of Salmonella from chicken skin using our model (Figure 5). Bioluminescent Salmonella strains have been previously utilized to monitor progress of infection, effect of heat and pH treatments, growth in food samples, and toxicity [13, 20, 22, 36]. One study used a single bioluminescent Salmonella Hadar isolate to investigate the effectiveness of washing for removal of Salmonella from turkey skin [20]. The current study demonstrates, for the first time, use of BLI for real-time monitoring of twelve Salmonella strains using a chicken skin model.
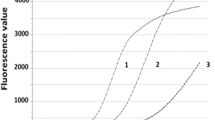
The effect of washing on removal of Salmonella from chicken skin. Bioluminescence was used to measure the ability of agitated water bath to remove Salmonella strains from chicken skin. Salmonella was allowed to attach to chicken skin for 1 h, and then the amount of Salmonella present on skin that received four 30 min washes was compared to the amount of Salmonella present on non-washed skin. Bioluminescence was measured after each 30 min wash, and the mean of four replicates from three separate experiments was determined. A) Representative plate containing 3 Salmonella strains at five time points. Half of the plate included washed skin samples (w) and the other half included unwashed skin controls (nw). Column 1 is S. Seftenberg, column 2 is S. Thompson, and column 3 is S. Typhimurium. B) Amount of bioluminescence for the three representative strains at each time point.
As a general trend observed from all strains, washing suppressed the reproduction of Salmonella on chicken skin, probably due to physical removal of bacteria (Figure 6). In non-washed skin samples, Salmonella numbers increased steadily over the two hour incubation period (Figure 5B), with final numbers showing an increase ranging from 143% (S. Schwarzengrund) to 459% (S. Newport) compared to initial measurement at time zero. In washed skin samples, the increase in Salmonella numbers was lower, demonstrating the effectiveness of simple agitation in water for suppressing Salmonella growth on chicken skin.
Comparison of the amount of Salmonella before and after 2 h incubation. The change in bioluminescence on chicken skin following 2 h incubation (relative to pre-wash bioluminescence) for washed and non-washed treatments is shown. Letters at the top of the graph indicate statistical groupings for the mean differences between washed and non-washed treatments within each strain as determined by Tukey's test. Values with the same letter are not significantly different (P > 0.05).
For all 12 strains, the washed treatments had a significantly lower change in bioluminescence than the corresponding non-washed treatments after the 2 h wash period. However, there was strain variation in the effectiveness of washing (Figure 6). For three strains (S. Kentucky, S. Mbandaka and S. Montevideo), washing reduced the number of Salmonella present on chicken skin at 120 min compared to 0 min (pre-wash). For the other strains, although washing did not decrease the number of Salmonella on chicken skin, it was effective in significantly reducing the growth relative to non-washed samples.
When the differences in bioluminescence between washed and non-washed treatments at 120 min were compared across strains, the decrease in bioluminescence for S. Newport caused by washing was significantly greater than the decrease in bioluminescence for S. Alachua, S. Braenderup, S. Enteritidis, and S. Schwarzengrund (Figure 6). Thus, based on the difference in bioluminescence between washed and non-washed treatments, washing was apparently most effective in removing S. Newport and least effective for S. Alachua, S. Braenderup, S. Enteritidis, and S. Schwarzengrund. However, it is interesting to note that the two strains that attached to chicken skin most effectively (S. Seftenberg and S. Thompson; Figure 4) were also able to increase their numbers in the wash treatment most effectively (3.7 and 2.7 fold increases, respectively, relative to pre-wash) over the 2 hr incubation.
Conclusion
Our results demonstrate that pAKlux 1 is effective for labeling Salmonella enterica strains with bioluminescence. Using this tool, we were able to develop twelve bioluminescent Salmonella strains that were isolated from the poultry production/processing continuum. However, stability results indicated that pAKlux 1 is not as stable in Salmonella as it is in other gram-negative species. Therefore, this labeling system may not be useful for long-term experiments, and work is already in progress in our group to develop a labeling system for Salmonella that is more stable.
Our work also shows that bioluminescence is an effective and sensitive method to monitor Salmonella on food products. In particular, our study demonstrates that Salmonella strains vary in their ability to attach to chicken skin. Future research aimed at determining whether the ability to attach to chicken skin correlates with the ability to persist in poultry production/processing environments is warranted. Finally, our results indicate that simple washing in an agitated water bath can remove contaminating Salmonella from chicken skin, but this method alone cannot eliminate the Salmonella completely due to strong bacterial attachment to skin. Therefore, additional methodologies should be used in poultry processing to eliminate Salmonella from the chicken skin to prevent disease outbreaks. The skin model developed in this study should be very useful for testing alternative strategies in addition to washing procedures.
Methods
Bacterial strains and growth media
As part of a previous study, we have collected, isolated, and serotyped thousands of Salmonella specimens derived and cataloged at five different sites along the broiler production continuum: post hatchery, prior to harvest, arrival at the plant, pre-chill tank, and post-chill tank. Salmonella isolates were collected from 66 different flocks during the years 2003–2006. The cataloged information ascribed to each isolate includes location within the production continuum, flock environmental and production parameters, as well as processing plant information. Strains from twelve Salmonella enterica serovars from the poultry production continuum were selected for this study (S. Alachua, S. Braenderup, S. Enteritidis, S. Heidelberg, S. Kentucky, S. Mbandaka, S. Montevideo, S. Newport, S. Schwarzengrund, S. Seftenberg, S. Thompson, and S. Typhimurium). Salmonella strains were grown using Luria-Bertani broth and agar plates at 37°C, and bioluminescent Salmonella strains were grown in the same medium containing ampicillin (100 μg ml-1) for plasmid maintenance.
Bioluminescence tagging of Salmonella strains
Bioluminescent Salmonella enterica strains were established using the broad host range plasmid pAKlux 1 containing the luxCDABE operon from Photorhabdus luminescens [15]. Salmonella strains were grown to logarithmic phase (OD600 of 0.6–0.8), made electrocompetent by washing with 10% cold glycerol solution four times, and stored at -80°C. pAKlux 1 was isolated from Escherichia coli DH5α strain by QIAprep Spin Miniprep Kit (Qiagen, Valencia, CA). Salmonella strains were transformed with pAKlux 1 by electroporation using a Gene Pulser II system at 2.5 kV, 25 uF, and 400 Ω (Bio-Rad, Hercules, CA). Cells were allowed to recover for 1 h at 37°C in SOC media (Invitrogen Corp., Carlsbad, CA). Following recovery, bacteria were spread on LB plates with ampicillin and placed in an incubator at 37°C for approximately 16 h. Ampicillin resistant bioluminescent Salmonella colonies were detected using a ChemiImager 5500 imaging system with AlphaEaseFC software (Alpha Innotech, San Leandro, CA) or using an IVIS Imaging System 100 Series with Living Image Software v2.50 (Xenogen Corp., Alameda, CA).
Characterizing the bioluminescence properties of Salmonella strains
Bioluminescent Salmonella strains were grown overnight, and OD600 values of each strain were measured from quadruplicate samples in a 96-well plate using ThermoMax spectrometer (Molecular Devices, Sunnyvale, USA). Following OD600 measurements, four separate dilution series were prepared from each strain in black 96-well microtiter plates. Each series contained 2 × 10-3, 4 × 10-3, 8 × 10-3, 1.6 × 10-4, 3.2 × 10-4, 6.4 × 10-4, 1.3 × 10-5, and 2.6 × 10-5 dilutions. Bioluminescence was measured for 5 s at 37°C using an IVIS Imaging System 100 Series, and bioluminescence was quantified using Living Image software v2.50 (Xenogen Corp.). The last two dilutions were spread on LB agar with ampicillin to determine viable bacterial densities.
The linear correlation between population densities and bioluminescence was determined for each strain by plotting bioluminescence against bacteria numbers determined by duplicate plate counts. The minimum detectible number for each strain was determined using the number of bacteria present in the last dilution that had detectable luminescence above background. Colony counts were also used to calculate the theoretical amount of bioluminescence produced per CFU for each strain, which was used to compare expression of bioluminescence among serovars.
Plasmid stability in vitro
Plasmid stability of pAKlux 1 in eleven Salmonella strains was analyzed by subculturing bioluminescent Salmonella strains in LB medium with and without ampicillin 15 times. For each passage, new 0.1 ml cultures were inoculated in microplates in quadruplicate from 16 h cultures at a 40-fold dilution. Bacterial density (OD600) and bioluminescence were determined for each of the 16 h cultures prior to starting new subcultures. Bioluminescence was measured using an IVIS Imaging System for 5 s at 37°C and normalized by dividing total flux by OD600 readings. The average normalized bioluminescence for each strain and passage was determined under non-selective and ampicillin-selected conditions, and plasmid stability was determined by calculating the ratio between normalized bioluminescence under non-selective conditions versus ampicillin-selected conditions. This revealed the capability of each serovar to maintain pAKlux 1 under non-selective conditions.
Development of skin attachment model
An experimental model for investigating bacterial attachment to chicken skin was established. Chicken skin was obtained from a commercial poultry processing plant and submerged in 0.26% sodium hypochlorite solution for approximately 2 h and stored at 4°C until use. Circular sections of skin approximately 8 mm in diameter were made using a circular cutting blade and placed in black 24-well culture plates. To determine the optimal bacterial dose to prevent loss of experimental data due to image saturation, chicken skin was incubated with different numbers of bioluminescent Salmonella (1 × 108 to 2.5 × 104 CFU) in 1 ml of phosphate-buffered saline. Based on these results, we used a bacterial concentration of 1 × 106 CFU/ml for subsequent experiments because this dose does not cause saturation and because it falls in the linear detection interval (1 × 103-1 × 108 photons/s/cm2/steradean [p/s/cm2/sr]).
Characterization of skin attachment properties of Salmonella strains
After establishing the skin attachment model, differential attachment properties of twelve Salmonella strains were determined. For each strain, overnight bioluminescent Salmonella cultures were diluted to approximately 1 × 106 CFU/ml in distilled water, and 1 ml was added to skin sections in quadruplicate. Plates were incubated at room temperature for 1 h to allow Salmonella to attach to skin. Then bacterial suspension was removed, and each well was gently washed twice with distilled water to remove unattached bacteria. Immediately after washing, plates were warmed to 37°C for 5 min and bioluminescence was measured for 15 s using the IVIS Imaging System. The experiment was repeated three times.
Total flux was calculated from the pseudo color images. To normalize for bacterial density differences between strains, the total flux from each well was divided by the OD600 of the bacterial suspension used to infect that well. Because the amount of bioluminescence from different strains varied, theoretical bacterial numbers were calculated from bioluminescence using the linear correlation formulas calculated for each strain (Figure 1). The data were transformed by taking the base 10 logarithm of the calculated bacterial numbers to improve normality. To compare bacterial numbers of the 12 Salmonella strains following skin attachment (after 1 h incubation), a two-way analysis of variance (ANOVA) of the transformed data was conducted using PROC GLM SAS 9.1 (SAS Institute Inc., Carey NC). Variables for replicate, strain, and their interaction were included in the model. Pairwise comparison of the means was done using Tukey procedure. A significance level of P ≤ 0.05 was used. Data were then retransformed to percent mortality for interpretation.
The effect of washing on Salmonella removal from chicken skin
The effect of washing chicken skin in an agitated water bath for removal of Salmonella was determined. The same 12 bioluminescent Salmonella strains were allowed to attach to chicken skin sections using the exact method described for the attachment experiment. In this experiment, two treatments were included for each strain: a washed treatment and a control (non-washed) treatment. Each treatment was set up in quadruplicate, and the experiment was repeated three times. Each bioluminescence measurement was conducted after warming plates to 37°C for 5 min. Following the 1 h incubation for attachment and washes to remove unattached bacteria, bioluminescence was measured. One ml of distilled water was added to the washed treatments, and control treatments received no water. Plates were incubated at room temperature for 30 min with agitation at 200 rpm. Water was removed from the washed treatments, and bioluminescence was measured. One ml of water was added to the washed skin treatments, and the wash procedure was repeated three more times (2 h total). After each wash, bioluminescence was measured. Total flux was measured from the pseudo color images for each time point, and samples were normalized as described for the attachment experiment.
To determine the effect of washing within each strain, the mean difference in bioluminescence between 0 minutes (prior to washing) and 120 minutes was calculated for both washed and non-washed treatments for each strain. The mean differences of washed and non-washed treatments within each strain were then compared by conducting a two-way analysis of variance using PROC GLM SAS 9.1 (SAS Institute Inc., Carey NC) for each strain. Variables for replicate, treatment, and their interaction were included in each model. To compare the effect of washing across strains, the mean difference in bioluminescence between washed and non-washed treatments after 120 minutes was calculated for each strain. A two-way analysis of variance was conducted on the mean differences using PROC GLM SAS 9.1 (SAS Institute Inc., Carey NC). Variables for replicate, strain, and their interaction were included in the model. Pairwise comparison of the means was done using Tukey procedure. A significance level of 0.05 was used in all analyses.
References
Sarlin LL, Barnhart ET, Caldwell DJ, Moore RW, Byrd JA, Caldwell DY, Corrier DE, Deloach JR, Hargis BM: Evaluation of alternative sampling methods for Salmonella critical control point determination at broiler processing. Poult Sci. 1998, 77: 1253-1257.
Lillard HS: Incidence and recovery of Salmonellae and other bacteria from commercially processed poultry carcasses at selected pre- and post-evisceration steps. J Food Prot. 1989, 52: 88-91.
Lillard HS: The impact with commercial processing procedures on the bacterial contamination and cross-contamination of broiler carcasses. J Food Prot. 1990, 53: 202-204.
Lillard HS: Factors affecting the persistence of Salmonella during the processing of poultry. J Food Prot. 1989, 52: 829-832.
Lillard HS: Bacterial cell characteristics and conditions influencing Salmonella adhesion to poultry skin. J Food Prot. 1985, 48: 803-807.
Lillard HS: Distribution of "attached" Salmonella typhimurium cells between poultry skin and a surface film following water immersion. J Food Prot. 1986, 49: 449-454.
McMeekin TA, Thomas CJ: Retention of bacteria on chicken skin after immersion in bacterial suspensions. J Food Prot. 1978, 48: 939-943.
Li Y, Slavik MF, Walker JT, Xiong H: Pre-chill spray of chicken carcasses to reduce Salmonella Typhimurium. J Food Sci. 1997, 62: 605-607. 10.1111/j.1365-2621.1997.tb04441.x.
Mullerat J, Sheldon BW, Klapes NA: Inactivation of Salmonella species and other food-borne pathogens with Salmide, a sodium chlorite-based oxyhalogen disinfectant. J Food Prot. 1995, 58: 535-540.
Rathgeber BM, Waldroup AL: Antibacterial activity of a sodium acid pyrophosphate product in chiller water against bacteria on broiler carcasses. J Food Prot. 1995, 58: 530-534.
Basti AA, Razavilar V: Growth response and modeling of the effects of selected factors on the time-to-detection and probability of growth initiation of Salmonella Typhimurium. Food Microbiology. 2004, 21: 431-438. 10.1016/j.fm.2003.10.006.
Contag PR, Olomu IN, Stevenson DK, Contag CH: Bioluminescent indicators in living mammals. Nat Med. 1998, 4: 245-247. 10.1038/nm0298-245.
Contag CH, Contag PR, Mullins JI, Spilman SD, Stevenson DK, Benaron DA: Photonic detection of bacterial pathogens in living hosts. Mol Microbiol. 1995, 18: 593-603. 10.1111/j.1365-2958.1995.mmi_18040593.x.
Contag CH, Bachmann MH: Advances in in vivo bioluminescence imaging of gene expression. Annu Rev Biomed Eng. 2002, 4: 235-260. 10.1146/annurev.bioeng.4.111901.093336.
Karsi A, Menanteau-Ledouble S, Lawrence ML: Development of bioluminescent Edwardsiella ictaluri for noninvasive disease monitoring. FEMS Microbiol Lett. 2006, 260: 216-223. 10.1111/j.1574-6968.2006.00310.x.
Meighen EA: Molecular biology of bacterial bioluminescence. Microbiol Rev. 1991, 55: 123-142.
Hastings JW: Chemistries and colors of bioluminescent reactions: a review. Gene. 1996, 173: 5-11. 10.1016/0378-1119(95)00676-1.
Frackman S, Anhalt M, Nealson KH: Cloning, organization, and expression of the bioluminescence genes of Xenorhabdus luminescens. J Bacteriol. 1990, 172: 5767-5773.
Wilson T, Hastings JW: Bioluminescence. Annu Rev Cell Dev Biol. 1998, 14: 197-230. 10.1146/annurev.cellbio.14.1.197.
Bautista DA, Chen J, Barbut S, Griffiths MW: Use of an autobioluminescent Salmonella Hadar to monitor the effects of acid and temperature treatments on cell survival and viability on lactic acid-treated poultry carcasses. J Food Prot. 1998, 61: 1439-1445.
Beyer W, Bohm R: Labeling Salmonella live vaccine strains with the lux operon from Vibrio fischeri improves their detection and discrimination from wild type. Microbiol Res. 1996, 151: 407-419.
Chen J, Griffiths MW: Luminescent Salmonella strains as real time reporters of growth and recovery from sublethal injury in food. Int J Food Microbiol. 1996, 31: 27-43. 10.1016/0168-1605(96)00941-5.
Troy T, Jekic-McMullen D, Sambucetti L, Rice B: Quantitative comparison of the sensitivity of detection of fluorescent and bioluminescent reporters in animal models. Mol Imaging. 2004, 3: 9-23. 10.1162/153535004773861688.
Billard P, DuBow MS: Bioluminescence-based assays for detection and characterization of bacteria and chemicals in clinical laboratories. Clin Biochem. 1998, 31: 1-14. 10.1016/S0009-9120(97)00136-7.
Liu Y, Golden SS, Kondo T, Ishiura M, Johnson CH: Bacterial luciferase as a reporter of circadian gene expression in cyanobacteria. J Bacteriol. 1995, 177: 2080-2086.
Gu MB, Min J, LaRossa RA: Bacterial bioluminescent emission from recombinant Escherichia coli harboring a recA::luxCDABE fusion. J Biochem Biophys Methods. 2000, 45: 45-56. 10.1016/S0165-022X(00)00100-7.
Siragusa GR, Nawotka K, Spilman SD, Contag PR, Contag CH: Real-time monitoring of Escherichia coli O157:H7 adherence to beef carcass surface tissues with a bioluminescent reporter. Appl Environ Microbiol. 1999, 65: 1738-1745.
Maurer JJ, Doggett TA, Burns-Keliher L, Curtiss R: Expression of the rfa, LPS biosynthesis promoter in Salmonella typhimurium during invasion of intestinal epithelial cells. Curr Microbiol. 2000, 41: 172-176. 10.1007/s002840010113.
Wiles S, Clare S, Harker J, Huett A, Young D, Dougan G, Frankel G: Organ specificity, colonization and clearance dynamics in vivo following oral challenges with the murine pathogen Citrobacter rodentium. Cell Microbiol. 2004, 6: 963-972. 10.1111/j.1462-5822.2004.00414.x.
Francis KP, Joh D, Bellinger-Kawahara C, Hawkinson MJ, Purchio TF, Contag PR: Monitoring bioluminescent Staphylococcus aureus infections in living mice using a novel luxABCDE construct. Infect Immun. 2000, 68: 3594-3600. 10.1128/IAI.68.6.3594-3600.2000.
Marincs F, White DWR: Immobilization of Escherichia coli expressing the lux genes of Xenorhabdus luminescens. Applied and Environmental Microbiology. 1994, 60: 3862-3863.
Unge A, Tombolini R, Molbak L, Jansson JK: Simultaneous monitoring of cell number and metabolic activity of specific bacterial populations with a dual gfp-luxAB marker system. Appl Environ Microbiol. 1999, 65: 813-821.
Elzer PH, Kovach ME, Phillips RW, Robertson GT, Peterson KM, Roop RM: In vivo and in vitro stability of the broad-host-range cloning vector pBBR1MCS in six Brucella species. Plasmid. 1995, 33: 51-57. 10.1006/plas.1995.1006.
Kovach ME, Elzer PH, Hill DS, Robertson GT, Farris MA, Roop RM, Peterson KM: Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene. 1995, 166: 175-176. 10.1016/0378-1119(95)00584-1.
Antoine R, Locht C: Isolation and molecular characterization of a novel broad-host-range plasmid from Bordetella bronchiseptica with sequence similarities to plasmids from gram-positive organisms. Mol Microbiol. 1992, 6: 1785-1799. 10.1111/j.1365-2958.1992.tb01351.x.
Rabbow E, Stojicic N, Walrafen D, Baumstark-Khan C, Rettberg P, Schulze-Varnholt D, Franz M, Reitz G: The SOS-LUX-TOXICITY-Test on the International Space Station. Res Microbiol. 2006, 157: 30-36. 10.1016/j.resmic.2005.08.005.
Acknowledgements
This study was funded by the U.S. Department of Agriculture, Agricultural Research Service (agreement no. 58-6202-5-083). We gratefully acknowledge Dr. Scott Willard and Dr. Peter Ryan for use of the IVIS Living Image System in the MSU Laboratory for Organismal and Cellular Imaging. We thank Michelle Banes and Terry Doler for technical assistance during this project.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Authors' contributions
HB and RW isolated the Salmonella strains. ML, HB, and AK designed the bacteriological and genetic studies. AK, KH, and TK performed the experiments and analyses. RW and AK designed and conducted statistical analyses. AK, KH, ML, and HB drafted the manuscript. All authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Karsi, A., Howe, K., Kirkpatrick, T.B. et al. Development of bioluminescent Salmonella strains for use in food safety. BMC Microbiol 8, 10 (2008). https://doi.org/10.1186/1471-2180-8-10
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2180-8-10