Abstract
Background
Thermophilic Bacillus strains of phylogenetic Bacillus rRNA group 5 were described as a new genus Geobacillus. Their geographical distribution included oilfields, hay compost, hydrothermal vent or soils. The members from the genus Geobacillus have a growth temperatures ranging from 35 to 78°C and contained iso-branched saturated fatty acids (iso-15:0, iso-16:0 and iso-17:0) as the major fatty acids. The members of Geobacillus have similarity in their 16S rRNA gene sequences (96.5–99.2%). Thermophiles harboring intrinsically stable enzymes are suitable for industrial applications. The quest for intrinsically thermostable lipases from thermophiles is a prominent task due to the laborious processes via genetic modification.
Results
Twenty-nine putative lipase producers were screened and isolated from palm oil mill effluent in Malaysia. Of these, isolate T1T was chosen for further study as relatively higher lipase activity was detected quantitatively. The crude T1 lipase showed high optimum temperature of 70°C and was also stable up to 60°C without significant loss of crude enzyme activity. Strain T1T was a Gram-positive, rod-shaped, endospore forming bacterium. On the basic of 16S rDNA analysis, strain T1T was shown to belong to the Bacillus rRNA group 5 related to Geobacillus thermoleovorans (DSM 5366T) and Geobacillus kaustophilus (DSM 7263T). Chemotaxonomic data of cellular fatty acids supported the affiliation of strain T1T to the genus Geobacillus. The results of physiological and biochemical tests, DNA/DNA hybridization, RiboPrint analysis, the length of lipase gene and protein pattern allowed genotypic and phenotypic differentiation of strain T1T from its validly published closest phylogenetic neighbors. Strain T1T therefore represents a novel species, for which the name Geobacillus zalihae sp. nov. is proposed, with the type strain T1T (=DSM 18318T; NBRC 101842T).
Conclusion
Strain T1T was able to secrete extracellular thermostable lipase into culture medium. The strain T1T was identified as Geobacillus zalihae T1T as it differs from its type strains Geobacillus kaustophilus (DSM 7263T) and Geobacillus thermoleovorans (DSM 5366T) on some physiological studies, cellular fatty acids composition, RiboPrint analysis, length of lipase gene and protein profile.
Similar content being viewed by others
Background
The Bacillus rRNA group 5 which comprised thermophilic Bacillus strains was transferred into new genus Geobacillus which represented a phenotypically and phylogenetically coherent group of thermophilic bacilli with high levels of 16S rRNA sequence similarity (96.5–99.2%) [1]. The members of this genus are widespread in various thermophilic and mesophilic geographic areas on the earth such as oilfields, hay compost, hydrothermal vent or soils [1–5]. At present, the members of this genus included Geobacillus stearothermophilus, Geobacillus thermocatenulatus, Geobacillus thermoleovorans, Geobacillus kaustophilus, Geobacillus thermoglucosidasius, Geobacillus thermodenitrificans, Geobacillus subterraneus, Geobacillus uzenensis, Geobacillus caldoxylosilyticus, Geobacillus toebii, Geobacillus vulcani, Geobacillus lituanicus, Geobacillus tepidamans, Geobacillus gargensis, Geobacillus jurassicus, Geobacillus caldoproteolyticus, Geobacillus pallidus and Geobacillus debilis with growth temperatures ranging from 35 to 78°C [1, 4–19].
Recently, microorganisms such as Bacillus sp. RSJ-1 [20], Bacillus thermoleovorans ID-1 [21], Bacillus sp. THL027 [22], Bacillus sp. strain A30-1 [23], Bacillus sp. strain 398 [24], Bacillus thermocatenulatus [25], Bacillus spp. [26–30] were reported as thermostable lipase producers. As thermophilic bacterial strains have an optimum growth temperature of 65–70°C, lipases isolated from such strains are good candidates for lipid modifications [31].
The stability of biocatalysts is an important criterion when dealing with bioprocesses at high temperature for sustainable operation. Enzyme stability is dictated by its three-dimensional configuration, which in turn is determined by genetic and environmental factors [32]. Therefore, thermophiles are promising sources of heat-stable enzymes. In addition to higher thermostability, proteins from thermophiles often showed higher stability toward organic solvents and higher activity at elevated temperature [33]. In addition, genetic engineering in altering the stability of enzymes is a difficult task and laborious processes. Therefore, efforts have been focused on the screening of microorganisms harboring intrinsically stable biocatalysts. Putative lipase producers were screened quantitatively across various basal media to check for lipase production and the stability of lipase. The highest lipase producing strain have been identified and characterized intensively. Screening and isolation of heat-stable lipase producers are important to fulfill industrial requirements for the desired characteristics. Identification of industrially important enzyme producer is conducted to determine its phylogenetic position in systematic microbiology.
Results and discussion
Screening and isolation of thermophilic lipolytic bacteria
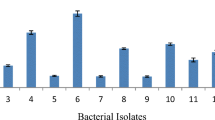
Rubbish dump sites and palm oil mill effluent are potential sites containing thermophilic lipolytic bacteria because these sites served as sewage discharge areas for household waste and palm oil factory. The oily environment may provide a good environment for lipolytic microorganisms to flourish. Samples were collected from the rubbish dump site and palm oil mill effluent at different treatment areas. Samples were enriched with enrichment medium (EM-1) containing olive oil as the sole carbon source at 60°C for 2 days under shaking condition to promote the growth of thermophilic lipolytic bacteria. Twenty nine putative lipase producers gave positive results on triolein agar plate by forming an intense blue color around the colonies (Table 1). Of these, 23 and 6 isolates were isolated from rubbish dump site and palm oil mill, respectively. Further confirmation test was performed quantitatively on the 29 putative lipase producers that gave positive results on triolein agar plates. Various basal media M1, M3, TYEM and BM-1 were tested for lipase production during the selection. Low lipase activities ranging from 0 ~ 0.044 U/ml were detected colorimetrically in culture supernatant, except for isolate T1T, which showed high lipase activity (0.150 U/ml) after 24 h incubation (Table 1).
On the basis of relatively higher lipase activity detected for isolate T1T, the effect of temperature on the activity and stability of crude T1 lipase was further investigated. The effect of temperature on the lipase activity and stability was examined from 40 to 80°C. As shown in Fig. 1, the crude enzyme from isolate T1T manifested its maximal activity at 70°C with olive oil as substrate. Crude T1 lipase was fairly stable up to 60°C for 30 min and gradually decreased upon prolonged temperature treatment. This is due to T1 lipase tended to lose its native conformation as a result of breaking of the intrinsic interaction above its stable range. This is a discrepancy between temperature activity (70°C) and stability (60°C), as the enzyme is tend to be protected by heat denaturation in the presence of olive oil. In addition, the enzyme thermostability is greatly influenced by the presence of water, because denaturation is linked to its conformational mobility in aqueous mixture [34]. The crude enzyme of isolate T1T was fairly active at higher temperature as compared to other thermostable lipases from Bacillus spp. [26, 30, 33] which exhibited maximal activity at 60°C. High temperature activity and stability of enzyme offer great potentials in industrial applications, and hence attempts have been made to identify isolate T1T.
Lipase activity (■) and stability (▲) of crude T1 lipase at various temperature. Crude T1 lipase was assayed at various temperatures ranging from 50 to 80°C with olive oil emulsion (1:1, v/v) as substrate (pH 7.0). For the lipase stability test, crude T1 lipase was assayed after heat treatment at various temperatures for 30 min.
Identification of isolate T1T
During the characterization of organism isolated from palm oil mill effluent, strain T1T was recovered on nutrient broth at 60°C. The growth condition for strain T1T was 50–70°C and between pH 5 and 9 with the optimum growth temperature and pH of 65°C and pH 6.5, respectively, in nutrient broth. These met the criteria of thermophilic bacteria, which grew at temperatures above 50°C [35]. To verify the systematic position of this bacterium, a study of morphological and physiological characteristics, 16S rRNA analysis, cellular fatty acids analysis, DNA composition, DNA/DNA hybridization, RiboPrint analysis, lipase gene analysis and protein profile were undertaken.
The cellular morphology of isolate T1T is rod-shaped, 0.8–1.0 μm width and 2.5–6.0 μm length, gram positive bacteria. The terminal spore is oval/cylindrical in shape and swollen the sporangium. The DNA base composition of strain T1T is around 52.6% mol G + C. The partial sequencing of the 16S rDNA shows 99.5% similarity to validly described Geobacillus kaustophilus (DSM 7263T) and Geobacillus thermoleovorans (DSM 5366T). The 16S rRNA sequence of strain T1T is a continuous stretch of 1519 bp (AY166603). Construction of phylogenetic trees using the neighbour-joining method in determining the evolutionary relationship among a group of validly described closely related species is indicated in Fig. 2. A comparison of biochemical, morphological and physiological properties of strain T1T with its closest phylogenetic neighbors is presented in Table 2. Strain T1T can be distinguished from Geobacillus thermoleovorans (DSM 5366T) phenotypically by oxidase test, arabinose, mannitol, inositol, lactose and casein hydrolysis. However, strain T1T differs from Geobacillus kaustophilus (DSM 7263T) by lysozyme test, arabinose, mannitol, ribose, adonitol, lactose, gelatin and casein tests. Since the sequencing result and physiological data did not allow strain T1T to be identified with one of the above mentioned species, further analysis need to be carried out to verify its phylogenetic position.
Phylogenetic position of Geobacillus zalihae T1T with other validly described species of the genus Geobacillus. The members of genus Geobacillus used include G. thermoleovorans (DSM 5366T); G. kaustophilus (DSM 7263T); G. vulcani (DSM 13174T); G. lituanicus (DSM 15325T); G. thermocatenulatus (DSM 730T); G. gargensis (DSM 15378T); G. stearothermophilus (NCDO 1768T); G. uzenensis (DSM 13551T); G. jurassicus (DSM 15726T); G. subterraneus (DSM 13553T); G. thermodenitrificans (DSM 465T); G. caldoxylosilyticus (ATCC 700356T); G. toebi (DSM 14590T); G. thermoglucosidasius (ATCC 43742T) G. tepidamans (DSM 16325T); G. caldoproteolyticus (DSM15730T); G. pallidus (DSM3670T); G. debilis (DSM16016T). Escherichia coli were used as an out-group. Phylogenetic tree was inferred by using the neighbour-joining methods. The software package MEGA 3.1 was used for analysis.
Result of chemotaxonomic analyses is given in the species description. The fatty acid profile of strain T1T is typical for the Bacillus rRNA-group 5 (thermophilic Bacillus strains). The major content of cellular fatty acids of strain T1T is iso-fatty acids. Among them, iso-branched pentadecanoic acid (iso-C15), hexadecanoic acid (iso-C16) and heptadecanoic acid (iso-C17) making up 78.33% of the total fatty acids for strain T1T. Strain T1T can be differentiated from Geobacillus thermoleovorans DSM 5366T based on percent composition of iso-fatty acids (iso-C15, iso-C16 and iso-C17 making up 62.1%). There was only 6.14% of iso-C16 for strain T1T but 21% for Geobacillus thermoleovorans DSM 5366T (Table 3).
Although FAME is not always a reliable tool for preliminary identification of bacilli, it could be used in combination with other identification methods [36]. DNA/DNA hybridization experiments were performed in DSMZ (Germany) with strain T1T and its type strains of closest phylogenetical neighbors. The genomic DNA/DNA relatedness between strain T1T and its type strains Geobacillus kaustophilus (DSM7263T) Geobacillus thermoleovorans (DSM 5366T) were 73.6% and 68.2%, respectively. Whereas, the type strains Geobacillus kaustophilus (DSM 7263T) and Geobacillus thermoleovorans (DSM 5366T) showed a DNA/DNA similarity of 71.8%. The DNA/DNA reassociation values were below the threshold value of 70% DNA/DNA similarity for definition of species [37] between strain T1T and Geobacillus thermoleovorans DSM 5366T but above the threshold between strain T1T and Geobacillus kaustophilus DSM 7263T. Neither the species Geobacillus kaustophilus DSM 7263T and Geobacillus thermoleovorans DSM 5366T can be differentiated from one another nor strain T1T can be differentiated at the species level from its closest phylogenetic neighbors by DNA/DNA hybridization. However, DNA/DNA hybridization tests between Geobacillus kaustophilus DSM 7263T and Geobacillus thermoleovorans DSM 5366T were 84% and 54% as reported by Sunna et al. [3] and Nazina et al. [13]. This disagreement may be due to the absence of adequate hybridization controls in the experiments. Therefore, further tests need to be carried out to accurately place the strain T1T phylogenetically.
The RiboPrint analysis was carried out for the decision on the affiliation of strain T1T. However, the RiboPrint pattern of strain T1T was not identified by the Dupont identification library to give rise to the identification at the species level (>0.85). Its RiboPrint pattern showed the highest similarity to Geobacillus kaustophilus DSM 7263T (0.69). The similarity to the pattern of Geobacillus thermoleovorans DSM 5366T was somewhat lower (0.57). The patterns between type strains Geobacillus kaustophilus DSM 7263T and Geobacillus thermoleovorans DSM 5366T show a binary similarity of 0.64.
Further analysis was also carried out by amplifying full-length thermostable lipase gene using primers as described in materials and methods [38]. Fig. 3 showed amplified full-length lipase genes of strain T1T and its type strains. The amplified lipase gene of strain T1T was around 2 kb but 1.8 kb for its type strains Geobacillus kaustophilus DSM 7263T and Geobacillus thermoleovorans DSM 5366T. A stretch of about 220 bp insertion could be seen at downstream of the open reading frame of thermostable lipase gene (Fig. 4). In addition, the intracellular protein profiles were determined by SDS-PAGE. Samples (30 μg) were separated on 12% SDS-polyacrylamide gel and stained using Coomassie blue. Strain T1T showed obvious different protein profile as compared to its type strains at region between 34 to 47 kDa (Fig. 5).
Downstream sequence alignment of lipase genes derived from Geobacillus zalihae T1T and its phylogenetic neighbors. The alignment was generated using Lip7263 (G. Kaustophilus DSM 7263T), Lip5366 (G. thermoleovorans DSM 5366T), and LipT1 (G. zalihae T1T). The C-terminal of the open reading frame coding sequences is bracket in red, and the stop codon was labeled as STOP.
Description of Geobacillus zalihae sp. nov
Geobacillus zalihae (za.li'ha.e. N.L. gen. n. zalihae of Zaliha). The novel species is isolated from palm oil mill effluent in Selangor, Malaysia, with the type strain T1T (DSM 18318T; NBRC 101842T). Cells are rod-shaped, 0.8–1.0 width and 2.5–6.0 length, gram positive bacteria. The terminal spores are oval/cylindrical and swollen the sporangium. Growth occurs at 50–70°C with an optimum temperature of 65°C. Growth at 65°C occurs between pH 5 and 9 with maximal growth at pH 6.5. The DNA base composition of strain T1T was around 52.6% mol G + C. The iso-fatty acids were in major amount according to cellular fatty acid profile in which iso-C15 (32.42%), and iso-C17 (39.77%) were in abundant (77.19%). Growth is aerobic and tolerant up to 2% NaCl. It can not perform anaerobic growth. It shows positive in catalase test but not oxidase test. It is able to hydrolyze starch but not gelatin and casein. Acids are produced from L-arabinose, D-lactose but not D-mannitol.
Conclusion
As a consequence, the strain T1T merits recognition as a member of a novel species through morphological and physiological studies, cellular fatty acids composition, DNA composition, DNA/DNA hybridization, RiboPrint analysis. Sizes of full-length lipase genes and protein profiles were additional evidences. The Geobacillus zalihae strain T1T was deposited in DSMZ (DSM 18318T) and NITE (NBRC 101842T).
Methods
Bacterial isolation
Numerous enrichment cultures that were derived from samples of rubbish dump site and palm oil mill effluent were screened for their ability to degrade olive oil. The strain described here was isolated from a sample from palm oil mill effluent in Semenyih, Malaysia. Samples (5%) were inoculated into enrichment medium (EM1) with olive oil as the sole carbon source and incubated at 60°C under shaking condition (150 rpm) for 2 days at an initial pH 7.0. The composition of EM1 is as follows: olive oil 2%; NaCl 0.2%; MgSO4.7H2O 0.04%; MgCl2.6H2O 0.07%; CaCl2.2H2O 0.05%; KH2PO4 0.03%; K2HPO4 0.03%; (NH4)2SO4 0.05% [21], to which 0.01% of trace elements solution containing 0.026% B, 0.05% Cu, 0.05% Mn, 0.006% Mo and 0.07% Zn were added [27]. The enriched cultures were further screened by using triolein agar plate. Triolein agar comprising of triolein (0.25%), bacteriological agar (1%), nutrient broth (0.8%) and Victoria Blue (0.01%) was adjusted to pH 7.0. The medium was homogenized for 5 minutes before sterilization. The sterilized triolein agar was poured into petri dishes. Isolates that showed positive results on the triolein agar were then tested for their lipase production in basal media.
Selection of thermostable lipase producer
Lipase production was determined aerobically at 60°C in 500 ml blue cap bottle containing 100 ml basal medium. The composition of basal mineral media used in this study was (g/L): BM1 (NaNO3: 7; K2HPO4: 2; KH2PO4: 1; KCl: 0.1; MgSO4.7H2O: 0.5; CaCl2: 0.01; FeSO4.7H2O: 0.012; yeast extract: 1 in which 0.01% trace elements and 2% olive oil were supplemented) [39]; M1 (peptone: 3; yeast extract: 1; NaCl: 0.5 in which 1% olive oil was supplemented) [30]; M3 (nutrient broth: 0.325; gum Arabic: 1; CaCl2.2H2O: 0.05; Tween.80: 1 in which 1% olive oil was supplemented) [40] and TYEM (tryptone: 6; yeast extract: 2; CaCl2.2H2O: 0.2; MgSO4.7H2O: 0.1; FeCl3.6H2O: 0.4 in which 1.5% olive oil was supplemented) [21]. The pH was adjusted to 7.0 and the medium was sterilized for 15 min at 121°C. Bacterial inoculum (3 ml) was then inoculated into 100 ml basal medium and incubated by shaking at 150 rpm, 60°C. The cell free supernatant was obtained by centrifugation at 10, 000 rpm, 4°C for 10 min prior to lipase assay.
Lipase assay
The lipase activity was assayed by colorimetry [41]. Culture filtrate (1 ml) was shaken with 2.5 ml of olive oil (70% oleate residues) emulsion (1:1, v/v) and 20 μl of 0.02 M CaCl2 in a water bath shaker at an agitation rate of 200 rpm. The emulsion was prepared by mixing together an equal volume of olive oil (Bertoli, Italy) and 50 mM phosphate buffer (pH 7.0) with a magnetic stirrer for 10 min. The reaction mixture was shaken for 30 min at 50°C. The enzyme reaction in the emulsion system was stopped by adding 6 M HCl (1 ml) and isooctane (5 ml), followed by mixing using a vortex mixer for 30 s. The upper isooctane layer (4 ml) containing the fatty acid was transferred to a test tube for analysis. Copper reagent (1 ml) was added and again mixed with a vortex mixer for 30 s. The reagent was prepared by adjusting the solution of 5% (w/v) copper (II) acetate-1-hydrate to pH 6.1 with pyridine. The absorbance of the upper layer was read at 715 nm. Lipase activity was measured by measuring the amount of free fatty acids released based on the standard curve of free fatty acid. One unit of lipase activity was defined as the amount of enzyme releasing 1 μmole of fatty acid per minute.
Characterization of crude lipase
The effect of temperature of the crude lipase was evaluated by assaying at temperatures ranging from 50 to 80°C. Crude enzyme (1 ml) was shaken with 2.5 ml of olive oil (70% oleate residues) emulsion (1:1, v/v) and 20 μl of 0.02 M CaCl2 in a water bath shaker at an agitation rate of 200 rpm. The lipase activity was measured colorimetrically. Assessment of the thermostability of crude lipase was performed by measuring the residual activity after 30 min pre-incubation at various temperatures ranging from 50 to 80°C. The treated enzyme was immediately put in ice-bath for 10 min before measuring the residual activity at 50°C for 30 min.
Identification of strain T1T
A study of morphological and 16S rRNA analysis was conducted in UPM. Its physiological characteristics, cellular fatty acids analysis, DNA composition, DNA/DNA hybridization and RiboPrint analysis was undertaken in Deutsche Sammlung Von Mikroorganismen (DSMZ), Germany.
Morphological and physiological study
For the morphological study, pure bacterial strain was streaked on nutrient agar plate and incubated for 24 h at 60°C prior to gram staining. It was then observed under a light microscope. Morphological and physiological characteristics were further determined in DSMZ (Germany). The physiological characteristics study included catalase and oxidase test, anaerobic growth, Voges-Proskauer test, growth at 30, 40 and 70°C, growth in medium at pH 5.7, 2% and 5% NaCl, lysozyme broth, fermentation of D-glucose, L-arabinose, D-xylose, D-mannitol, D-fructose, M-inosit, D-ribose, D-cellobiose, L-rhamnose, sorbitol, D-galactose, adonit, D-lactose, hydrolysis of starch, gelatin, casein and Tween 80, decomposition of tyrosine, use of citrate and propionate, nitrate reduction, indol production, phenylalanine deaminase and arginine dihydrolase test.
Cellular fatty acids analysis
Fatty acids were extracted and analysed following the instructions of Sherlock microbial identification system. The culture of Bacillus thermocatenulatus was used as control during cellular fatty acids analysis.
16S rDNA analysis
The 16S rDNA was amplified by PCR using two universal primers: 16S-F (5'-GAG TTT GAT CCT GGC TCA G-3') and 16S-R (5'-CGG CTA CCT TGT TAC GAC TT-3'). The PCR product was purified using QIAquick gel extraction kit (Qiagen, Germany). The purified PCR product was cloned into TOPO TA PCR 2.1 cloning vector (Invitrogen, USA). The recombinant plasmid was extracted with QIAprep plasmid extraction kit (Qiagen, Germany) and was then sequenced using an ABI PRISM 377 DNA sequencer (Applied Biosystems, USA) as recommended by the manufacturers. The 16S rDNA sequence of Geobacillus zalihae T1T was analyzed using software package MEGA 3.1 [42].
DNA base composition
The chromosomal DNA was isolated and purified according to the procedure of Cashion et al. [43]. The G + C contents were determined by using chromatography conditions adopted from Tamaoka and Komagata [44]. The DNA was hydrolysed and the resultant nucleotides were analysed by reverse-phase HPLC [45].
DNA/DNA hybridization
DNA/DNA hybridization was carried out as described by De Ley et al. [46], with the modifications described by Huss et al. [47] and Escara & Hutton [48], using a model 2600 spectrophotometer equipped with a model 2527-R thermoprogrammer and plotter (Gilford Instrument Laboratories). Renaturation rates were compared with the TRANSFER.BAS program by Jahnke [49].
Ribotyping analysis
Standardized automated ribotyping is performed using the QualiconTM RiboPrinter system as described by Bruce [50]. The RiboPrinter system combines molecular processing steps for ribotyping in a stand-alone, automated instrument. Steps included cell lysis, digestion of chromosomal DNA with restriction enzyme Eco R1, separation of fragments by electrophoresis, transfer of DNA fragments to a nylon membrane, hybridization to a probe generated from the rrn B operon from E. coli, chemiluminescent detection of the probe to the fragments containing rrn operon sequences, image detection and computerized analysis of RiboPrint patterns.
Amplification of full-length thermostable lipase gene
Genomic DNA was extracted by using conventional method [51]. In order to amplify the full length sequence of thermostable lipase gene, a set of primers was designed based on the thermostable lipase gene sequence of Bacillus thermoleovorans (AF134840), as follows: BTL-F: 5'-GGC GGT GAT GGA ACG CTG CCA TGA-3' and BTL-R: 5'-CCG ACG ATA GAC TGG CGG ACA AAT G-3'. Polymerase Chain Reaction (PCR) was carried out in a reaction mixture (100 μl) containing DNA template (10–100 ng), 10 mM deoxynucleotide triphosphates (dNTPs) (0.2 mM), 10 × PCR buffer (10.0 μl), 25 mM MgCl2 (2 mM), oligonucleotide primers: BTL-F (30 pmol) and BTL-R (30 pmol), and Taq DNA polymerase (2 U). The gene was amplified with a thermocycler (Gene Amp PCR system 2400, Perkin Elmer, Foster, CA) with the temperature program of predenaturation at 94°C for 4 min, 30 cycles PCR of 1 min denaturation at 94°C, 2 min annealing at 55°C and 2 min extension at 72°C. The final extension step at 72°C was 7 min and preservation was at 4°C. The amplified products were electrophoresed on 1.0% agarose gel (w/v) with 1 kb marker as the standard marker. The PCR products were purified and sequenced as described earlier. The sequences alignment was generated using CLUSTALW and TEXSHADE in Biology Workbench 3.2 [52].
Protein pattern analysis
SDS-PAGE was done on 12% running gels by using the method of Laemmli [53]. A broad range of protein standard (MBI Fermentas, Germany) was used as a molecular mass marker. Extracellular protein was concentrated with 5,000 MWCO cut-off vivaspin 15R (Vivascience, Germany). Protein samples (30 μg) were loaded for analysis.
References
Nazina TN, Tourova TP, Poltaraus AB: Taxonomic study of aerobic thermophilic bacilli: descriptions of Geobacillus subterraneus gen. nov., sp. nov. and Geobacillus uzenensis sp. nov. from petroleum reservoirs and transfer of Bacillus stearothermophilus, Bacillus thermocatenulatus, Bacillus thermoleovorans, Bacillus kaustophilus, Bacillus thermoglucosidasius and Bacillus thermodenitrificans to Geobacillus as the new combinations Geobacillus stearothermophilus, Geobacillus thermocatenulatus, Geobacillus thermoleovorans, Geobacillus kaustophilus, Geobacillus thermoglucosidasius and Geobacillus thermodenitrificans. Int J Syst Evol Microbiol. 2001, 51: 433-446.
Sharp RJ, Riley PW, White D: Heterotrophic thermophilic bacilli. Thermophilic Bacteria. Edited by: Kristjansson JK. 1992, Boca Raton: CRC Press, 19-50.
Sunna A, Tokajian S, Burghardt J, Rainey F, Antranikian G, Hashwa F: Identification of Bacillus kaustophilus, Bacillus thermocatenulatus and Bacillus strain HSR as members of Bacillus thermoleovorans. Syst Appl Microbiol. 1997, 20: 232-237.
Coccamo D, Gugliandolo C, Stackebrandt E, Maugeri TL: Bacillus vulcani sp. nov., a novel thermophilic species isolated from a shallow marine hydrothermal vent. Int J Syst Evol Microbiol. 2000, 50: 2009-2012.
Sung MH, Kim H, Bae JW, 9 other authors: Geobacillus toebii sp. nov., a novel thermophilic bacterium isolated from hay compost. Int J Syst Evol Microbiol. 2002, 52: 2251-2255. 10.1099/ijs.0.02181-0.
Golovacheva RS, Loginova LG, Salokhov TA, Kolesnikov AA, Zaitseva GN: A new thermophilic species, Bacillus thermocatenulatus nov. sp. Microbiology (English translation of Mikrobiologiya). 1975, 44: 230-233.
Suzuki Y, Kishigami T, Inoue K, 4 other authors: Bacillus thermoglucosidasius sp. nov., a new species of obligately thermophilic bacilli. Syst Appl Microbiol. 1983, 4: 487-495.
Priest FG, Goodfellow M, Todd C: A numerical classification of the genus Bacillus. J Gen Microbiol. 1988, 134: 1847-1882.
White D, Sharp RJ, Priest FG: A polyphasic taxonomic study of thermophilic bacilli from a wide geographical area. Antonie van Leeuwenhoek. 1993, 64: 357-386. 10.1007/BF00873093.
Zarilla K, Perry JJ: Bacillus thermoleovorans sp. nov., a species of obligately thermophilic hydrocarbon utilizing endospore-forming bacteria. Syst Appl Microbiol. 1987, 9: 258-264.
Manachini PL, Mora D, Nicastro G, Parini C, Stackebrandt E, Pukall R, Fortina MG: Bacillus thermodenitrificans sp. nov., nom. rev. Int J Syst Bacteriol. 2000, 50 (Pt 3): 1331-1337.
Fortina MG, Mora D, Schumann P, Parini C, Manachini PL, Stackebrandt E: Reclassification of Saccharococcus caldoxylosilyticus as Geobacillus caldoxylosilyticus (Ahmad et al., 2000) comb. nov. Int J Syst Evol Microbiol. 2001, 51: 2063-2071.
Nazina TN, Lebedeva EV, Poltaraus AB, Tourova TP, Grigoryan AA, Sokolova DS, Lysenko AM, Osipov GA: Geobacillus gargensis sp. nov., a novel thermophile from a hot spring, and the reclassification of Bacillus vulcani as Geobacillus vulcani comb. nov. Int J Syst Evol Microbiol 5. 2004, 4: 2019-2024. 10.1099/ijs.0.02932-0.
Kuisiene N, Raugalas J, Chitavichius D: Geobacillus lituanicus sp. nov. Int J Syst Evol Microbiol. 2004, 54: 1991-1995. 10.1099/ijs.0.02976-0.
Schäffer C, Franck WL, Scheberl A, Kosma P, McDermott TR, Messner P: Classification of isolates from locations in Austria and Yellowstone National Park as Geobacillus tepidamans sp. nov. Int J Syst Evol Microbiol. 2004, 54: 2361-2368. 10.1099/ijs.0.63227-0.
Nazina TN, Sokolova DS, Grigoryan AA, 6 other authors: Geobacillus jurassicus sp. nov., a new thermophilic bacterium isolated from a high-temperature petroleum reservoir, and the validation of the Geobacillus species. Syst Appl Microbiol. 2005, 28: 43-53. 10.1016/j.syapm.2004.09.001.
Chen XG, Stabnikova O, Tay JH, Wang JY, Tay TL: Thermoactive extracellular proteases of Geobacillus caldoproteolyticus, sp. nov., from sewage sludge. Extremophiles. 2004, 8: 489-498. 10.1007/s00792-004-0412-5.
Scholz T, Demharter W, Hensel R, Kandler O: Bacillus pallidus sp. nov., a new thermophilic species from sewage. Syst Appl Microbiol. 1987, 9: 91-96.
Banat IM, Marchant R, Rahman TJ: Geobacillus debilis sp. nov., a novel obligately thermophilic bacterium isolated from a cool soil environment, and reassignment of Bacillus pallidus to Geobacillus pallidus comb. nov. Int J Syst Evol Microbiol. 2004, 54: 2197-2201. 10.1099/ijs.0.63231-0.
Sharma R, Soni SK, Vohra RM, Gupta LK, Gupta JK: Purification and characterization of a thermostable alkaline lipase from a new thermophilic Bacillus sp. RSJ-1. Process Biochem. 2001, 37: 1075-1084. 10.1016/S0032-9592(01)00316-8.
Lee DW, Koh YS, Kim KJ, Kim BC, Choi HJ, Kim DS, Suhartono MT, Pyun YR: Isolation and characterization of a thermophilic lipase from Bacillus thermoleovorans ID-1. FEMS Microbiol Lett. 1999, 179: 393-400. 10.1111/j.1574-6968.1999.tb08754.x.
Dharmsthiti S, Luchai S: Production, purification and characterization of thermophilic lipase from Bacilus sp. THL027. FEMS Microbiol Lett. 1999, 179: 241-246. 10.1111/j.1574-6968.1999.tb08734.x.
Wang Y, Srivastava KC, Shen GJ, Wang HY: Thermostable alkaline lipase from a newly isolated thermophilic Bacillus, strain A30-1 (ATCC 53841). J Ferment Bioeng. 1995, 79 (5): 433-438. 10.1016/0922-338X(95)91257-6.
Kim HK, Sung MH, Kim HK, Oh TK: Occurrence of thermostable lipase in thermophilic Bacilus sp. strain 398. Biosci Biotechnol Biochem. 1994, 58 (5): 961-962.
Schmidt-Dannert C, Sztajer H, Stöcklein W, Menge U, Schmid RD: Screening, purification and properties of a thermophilic lipase from Bacillus thermocatenulatus. Biochim Biophys Acta. 1994, 1214 (1): 43-53.
Nawani N, Dosanjh NS, Kaur J: A novel thermostable lipase from a thermophilic Bacillus sp.: characterization and esterification studies. Biotechnol Lett. 1998, 20 (10): 997-1000. 10.1023/A:1005430215849.
Llarch A, Logan NA, Castellví J, Prieto MJ, Guinea J: Isolation and characterization of thermophilic Bacillus spp. from geothermal environments on deception island, south Shetlanh Archipelago. Microbiological Ecology. 1997, 34: 58-65. 10.1007/s002489900034.
Becker P, Reesh IA, Markossian S, Antranikian G, Märkl H: Determination of the kinetic parameters during continuous cultivation of the lipase-producing thermophile Bacillus sp. IHI-91 on olive oil. Appl Microbiol Biotechnol. 1997, 48: 184-190. 10.1007/s002530051036.
Handelsman T, Shoham Y: Production and characterization of an extracellular thermostable lipase from a thermophilic Bacillus sp. J Gen Appl Microbiol. 1994, 40: 435-443.
Sugihara A, Tani T, Tominaga Y: Purification and characterization of a novel thermostable lipase from Bacillus sp. J Biochem. 1991, 109 (2): 211-215.
Sigurgísladóttir S, Konráðsdóttir M, Jónsson A, Kristjánsson JK, Matthiasson E: Lipase activity of thermophilic bacteria from icelandic hot springs. Biotechnol Lett. 1993, 15 (4): 361-366. 10.1007/BF00128277.
Lllanes A: Stability of biocatalysis. Electron J Biotechnol. 1999, 2 (1): 1-9.
Schmidt-Dannert C, Rúa ML, Atomi H, Schmid RD: Thermoalkalophilic lipase of Bacillus thermocatenulatus. I. Molecular cloning, nucleotide sequence, purification and some properties. Biochim Biophys Acta. 1996, 1301: 105-114.
Nawani N, Kaur J: Purification, characterization and thermostability of lipase from a thermophilic Bacillus sp. J33. Mol Cell Biochem. 2000, 206: 91-96. 10.1023/A:1007047328301.
Perry JJ, Staley JT: Taxonomy of eubacteria and archaea. Microbiology: Dynamics and Diversity. Edited by: Perry JJ, Staley JT. 1997, Fort Worth: Saunders College Publishing, 388-413.
Suihko ML, Stackebrandt E: Identification of aerobic mesophilic bacilli isolated from board and paper products containing recycled fibers. J Appl Microbiol. 2003, 94: 25-34. 10.1046/j.1365-2672.2003.01803.x.
Wayne LG, Brenner DJ, Colwell RR, 9 other authors: Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol. 1987, 37: 463-464.
Leow TC, Rahman RNZRA, Basri M, Salleh AB: High level expression of thermostable lipase from Geobacillus sp. strain T1. Biosci Biotechnol Biochem. 2004, 68 (1): 96-103. 10.1271/bbb.68.96.
Haba E, Bresco O, Ferrer C, Marqués A, Busquets M, Manresa A: Isolation of lipase-secreting bacteria by deploying used frying oil as selective substrate. Enzyme Microb Technol. 2000, 26: 40-44. 10.1016/S0141-0229(99)00125-8.
Hun CJ, Rahman RNZRA, Salleh AB, Basri M: A newly isolated organic solvent tolerant Bacillus sphaericus 205y producing organic solvent-stable lipase. Biochem Eng J. 2003, 15 (2): 147-151. 10.1016/S1369-703X(02)00185-7.
Kwon DK, Rhee JS: A simple and rapid colorimetric method for determination of free fatty acids for lipase assay. J Am Oil Chem Soc. 1986, 63 (1): 89-92. 10.1007/BF02676129.
Kumar S, Tamura K, Nei M: MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Briefings in Bioinformatics. 2004, 5 (2): 150-163. 10.1093/bib/5.2.150.
Cashion P, Holder-Franklin MA, McCully J, Franklin M: A rapid method for the base ratio determination of bacterial DNA. Anal Biochem. 1977, 81: 461-466. 10.1016/0003-2697(77)90720-5.
Tamaoka J, Komagata K: Determination of DNA base composition by reversed-phase high-performance liquid chromatography. FEMS Microbiol Lett. 1984, 25: 125-128. 10.1111/j.1574-6968.1984.tb01388.x.
Mesbah M, Premachandran U, Whitman W: Precise measurement of the G+C content of deoxyribonucleic acid by high performance liquid chromatography. Int J Sys Bact. 1989, 39: 159-167.
De Ley J, Cattoir H, Reynaerts A: The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem. 1970, 12: 133-142. 10.1111/j.1432-1033.1970.tb00830.x.
Huss VAR, Festl H, Schleifer KH: Studies on the spectrophotometric determination of DNA hybridization from renaturation rates. Syst Appl Microbiol. 1983, 4: 184-192.
Escara JF, Hutton JR: Thermal stability and renaturation of DNA in dimethylsulphoxide solutions: Acceleration of renaturation rate. Biopolymers. 1980, 19: 1315-1327. 10.1002/bip.1980.360190708.
Jahnke KD: Basic computer program for evaluation of spectroscopic DNA renaturation data from GILFORD System 2600 spectrometer on a PC/XT/AT type personal computer. J Microbiol Methods. 1992, 15: 61-73. 10.1016/0167-7012(92)90069-G.
Bruce JL: Automated system rapidly identifies and characterizes microorganisma in food. Food Technol. 1996, 50 (1): 77-81.
Sambrook J, Fritsch EF, Maniatis T: Molecular cloning: a laboratory manual. 1989, New York: Cold Spring Harbor Laboratory Press
The Biology Workbench 3.2. [http://workbench.sdsc.edu/]
Laemmli UK: Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970, 227 (5259): 680-5. 10.1038/227680a0.
Markossian S, Becker P, Märkl H, Antranikian G: Isolation and characterization of lipid-degrading Bacillus thermoleovorans IHI-91 from an Icelandic hot spring. Extremophiles. 2000, 4: 365-371. 10.1007/s007920070006.
Acknowledgements
This research was supported financially by Ministry of Science, Technology and Innovation, Malaysia (09-02-04-0336-EA001).
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
TCL performed most of the experiments described in this paper, contributed to their design and analysis, and helped to draft the manuscript. RNZRAR, ABS and MB conceived of the study and experimental design, and prepared the manuscript. All authors read and approved the final manuscript.
Raja Noor Zaliha Raja Abd Rahman, Thean Chor Leow, Abu Bakar Salleh and Mahiran Basri contributed equally to this work.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Rahman, R.N.Z.R.A., Leow, T.C., Salleh, A.B. et al. Geobacillus zalihae sp. nov., a thermophilic lipolytic bacterium isolated from palm oil mill effluent in Malaysia. BMC Microbiol 7, 77 (2007). https://doi.org/10.1186/1471-2180-7-77
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2180-7-77