Abstract
Background
The microbiota has been shown to play an important role in the biology of insects. In recent decades, significant efforts have been made to better understand the diversity of symbiotic bacteria associated with mosquitoes and assess their influence on pathogen transmission. Here, we report the bacterial composition found in field-caught Aedes albopictus populations by using culture-dependent methods.
Results
A total of 104 mosquito imagos (56 males and 48 females) were caught from four contrasting biotopes of Madagascar and their bacterial contents were screened by plating whole body homogenates on three different culture media. From 281 bacterial colony types obtained, amplified ribosomal DNA restriction analysis (ARDRA) showed they had 40 distinct ribotypes. Sequencing and BLAST analysis of the 16S rDNA genes responsible for each representative profile made it possible to identify 27 genera distributed in three major phyla. In female mosquitoes, bacterial isolates were mostly Proteobacteria (51.3%) followed by Firmicutes (30.3%) and Actinobacteria (18.4%). Conversely, Actinobacteria was the most abundant phylum in male mosquitoes (48%) followed by Proteobacteria (30.6%) and Firmicutes (20.4%). The relative abundance and composition of isolates also varied between sampling sites, ranging from 3 distinct families in Ankazobe to 8 in Tsimbazaza Park, and Toamasina and Ambohidratrimo. Pantoea was the most common genus in both females and males from all sampling sites, except for Ambohidratrimo. No differences in genome size were found between Pantoea isolates from mosquitoes and reference strains in pulse field gel electrophoresis. However, according to the numbers and sizes of plasmids, mosquito isolates clustered into three different groups with other strains isolated from insects but distinct from isolates from the environment.
Conclusions
The recent upsurge in research into the functional role of the insect microbiota prompts the interest to better explore the role some bacteria detected here may have in the mosquito biology. Future studies of culturable bacteria might decipher whether they have a biological role in the invasiveness of Ae. albopictus. As a possible candidate for paratransgenesis, the predominant genus Pantoea will be characterized to better understand its genetic contents and any possible influence it may have on vector competence of Ae. albopictus.
Similar content being viewed by others
Background
Insects can be considered as holobiont units in which the insect host and its microbiota are involved in complex reciprocal multipartite interactions [1]. Numerous studies have shown the beneficial impact of microbiota on their insect hosts, especially in phytophagous insects. For instance, bacterial endosymbionts contribute to different biological functions like supplying essential nutrients, inducing resistance to pathogens and parasitoids, and conferring tolerance of temperature stress [2–6]. Surprisingly, the nature and function of naturally occurring microorganisms harboured by hematophagous arthropods have been largely overlooked in research even though these aspects might be relevant in the study of pathogen transmission. There are nevertheless a few examples of the molecular characterization of bacterial species in the microbiota of mosquito vectors based on culture-dependent or independent methods or both [7–12]. Recent years have seen a growing interest in metagenomic-based studies of bacterial communities possibly displacing traditional culture-based analysis [13]. For instance, next generation sequencing technology was successfully used in Anopheles gambiae to provide a ‘deeper’ description of the bacterial community than can be achieved with conventional molecular techniques [14]. However, even though such an approach can reveal the number and richness of bacterial species, it is still important to search for culturable bacteria residing in insects for several reasons. Culturing bacteria still offers the best way of observing the diverse characteristics of the isolated organism. The physiological characteristics of bacterial isolates need to be determined to investigate properties such as antibiotic resistance, interspecies growth inhibition or population dynamics within mosquito cohorts. The availability of key representative isolates therefore allows detailed analyses of biochemical, metabolic and functional processes. For example, isolation of Actinobacteria showed that they are involved in cellulose and hemicellulose degradation pathways in termites [15, 16]. Culturable Proteobacteria associated with insects were shown to play a role in carbohydrate degradation and nutrient provision [17, 18]. In addition to phenotypic characterization of bacterial isolates, culturing also facilitates bacterial genome sequencing, a further link towards revealing functionality [19]. There have also been a number of recent studies of the use of engineered bacteria in the development of more efficient insect control strategies. Insect bacterial symbionts were genetically modified and the recombinants reintroduced into their native host. This paratransgenesis strategy involves different steps but requires the initial isolation of a suitable bacterial species [20]. In mosquitoes, paratransgenesis studies have mainly focused on anopheline mosquitoes, vectors of the malaria parasite [11]. As an efficient colonizer of Anopheles stephensi, the bacterium Asaia sp. was originally proposed as a candidate for malaria control [21], but recently it has been suggested that Pantoea agglomerans, another bacterial symbiont of Anopheles, could also be engineered to express and secrete anti-Plasmodium effector proteins [22].
Screening culturable bacteria using traditional microbiological techniques is an important method in mosquito-associated microbiota investigation. One of the key mosquito species for pathogen transmission is Aedes albopictus, which is a vector of several arboviruses pathogenic to humans, some having a devastating impact worldwide [23]. This species has been identified as the primary vector responsible for recent outbreaks of Dengue and Chikungunya which emerged in Madagascar and other neighbouring islands [24, 25]. Until now, no bacterial species has been reported as being essential for mosquito biology, while only Wolbachia has been proposed as a gene driver system in Aedes mosquitoes. Here we present an in-depth investigation of culturable bacteria in natural populations of Ae. albopictus. Our main objective was to assess the abundance and phylogenetic diversity of culturable bacteria in a set of adult male and female mosquitoes from different regions of Madagascar. This deeper screening of the bacterial isolates retrieved significantly extends our previous work on the prevalence of Acinetobacter and Asaia associated with Madagascarian populations of Ae. albopictus[26].
Methods
Sampling areas and mosquito collection
The sampling areas and capture procedure were approved by Madagascar National Parks. Aedes albopictus specimens were sampled in December 2010 at four sites in two regions of Madagascar, Analamanga and Antsinanana. The main characteristics of the sampling sites are summarized in Table 1. Briefly, the two regions have a similar tropical climate, but different biotopes according to the vegetation or the presence of human or animal hosts susceptible to mosquito bites. Butterfly netting was used to collect both female and male mosquitoes flying near the grass or ground, as previously described [27]. The live mosquitoes collected were identified using morphological characteristics keys [28] and transported to the local laboratory.
Enrichment and isolation of bacteria from mosquitoes
Only non-blooded mosquitoes were used for the analysis. Specimens of Ae. albopictus were anaesthetised with ether and surface-disinfected as previously described [12], then crushed individually in 150 μl of sterile 0.8% NaCl with sterile piston pellets. After a brief vortexing, the homogenate was used in different isolation procedures using various media, from generalist to selective. All solid media were supplemented with 2.5 μg ml-1 amphotericin B to prevent the growth of fungi. An aliquot of the homogenate (10 μl) was streaked onto a modified rich solid Luria-Bertani medium (LBm, LB with 5 mg ml-1 NaCl) and incubated at 28°C for 24 to 48 h. Another aliquot (20 μl) was inoculated into 1 ml of selective enrichment medium I (0.2% KNO3, 0.02% MgSO4.7H2O, 0.2% sodium acetate, 0.04 M KH2PO4, pH 6), a medium which is suitable for the isolation of Acinetobacter species [29]. Cultures were incubated at 30°C for 24 to 48 h with shaking. When microbial growth occurred, an aliquot (10 μl) of the culture was streaked onto Herellea agar plates (Biolife, Italy), a medium suitable for the isolation of Gram-negative bacteria especially members of the Acinetobacter genus and the Enterobacteriaceae family [30]. These cultures were further incubated at 37°C for 24 to 48 h. In parallel, 1 ml of pre-enrichment liquid medium (pH 3.5), which is suitable for the isolation of acetic acid bacteria [31], was inoculated with an aliquot of homogenate (20 μl). These cultures were incubated with shaking at 30°C for 3 days. When microbial growth occurred, an aliquot (10 μl) was streaked onto CaCO3 agar plates (pH 6.8), a medium suitable for the isolation of members of the genus Asaia, and the plate was incubated at 30°C for 3 days as previously described [32]. Colonies were selected according to various characteristics including colour, shape, or size. Individual colonies were then re-inoculated onto fresh agar plates of the appropriate isolation medium. Newly formed colonies were streaked again to check for purity and stored in 25% glycerol at -20°C for two weeks before they were transported to the laboratory in Lyon, France. Isolates were re-streaked and new glycerol stocks were made and stored at -80°C. Brief morphological descriptions of colony size, shape and colour were recorded for each isolate.
PCR and amplified ribosomal DNA restriction analysis (ARDRA)
For PCR, a sterile toothpick was used to transfer bacteria from a single colony freshly grown on appropriate medium into 20 μl sterile water in a 0.5 ml Eppendorf tube. The homogenate was placed on a heating block at 95°C for 2 min followed by 2 min on ice. This step was repeated and the tube was centrifuged at 16,000 g for 5 min. The supernatant (2 μl) was used as template in a 50-μl PCR reaction. Bacterial 16S rrs genes were amplified with eubacterial-specific primers pA (5′-AGAGTTTGATCCTGGCTCAG-3′) and pH (5′-AAGGAGGTGATCCAGCCGCA-3′) in reaction mixture containing 1 × polymerase reaction buffer, 0.2 μM MgCl2, 200 μM of each deoxynucleoside triphosphate, 10 pmol of each primer and 1 U of Taq polymerase (Invitrogen). PCR amplifications consisted of 3 min at 95°C, 35 cycles of 30 sec at 94°C, 40 sec at 55°C and 1 min 30 sec at 72°C, and finally 10 min at 72°C. Amplified DNA fragments were purified using the QIAquick PCR Purification Kit (Qiagen). ARDRA was performed to screen the rrs genes of bacterial isolates in 20 μl reactions containing 200 ng of DNA template, 1 × Buffer Tango™ and 10 U each of endonucleases RsaI and HhaI (Fermentas, France), as previously described [12]. DNA fragments were separated on 2% agarose gels stained with ethidium bromide with a 50-bp DNA ladder marker (Fermentas). Isolates showing the same restriction pattern with the two endonucleases were considered to be similar.
Sequencing of rrsrRNA genes and phylogenetic analyses
Both strands of 16S rDNA amplified from isolates representative of each ARDRA profile were sequenced at Biofidal-DTAMB (FR Bio-Environment and Health, Lyon, France). Sequences were manually curated and assembled from forward and reverse primer-generated sequences. Curated sequences were then compared to available bacterial sequences in GenBank using the BLASTn program in the National Center for Biotechnology Information (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The Ribosomal Database Project II Chimera Check was used (http://wdcm.nig.ac.jp/RDP/html/analyses.html) to discard any chimeric sequences.
Phylogenetic analyses were performed on a set of Pantoea sequences. Sequences of 16S rRNA genes from Pantoea isolates from mosquitoes were compared to all available sequences of Pantoea retrieved from GenBank that originated from other insect species and environments. Sequences were aligned using ClustalW then corrected manually using Bioedit software [33]. The resulting alignment was used to construct a maximum-likelihood tree using Seaview v.4.2.12. (http://pbil.univ-lyon1.fr/software/seaview.html). The tree topology was tested by bootstrap analysis with 1,000 resamplings.
Pulse field gel electrophoresis (PFGE) of bacterial genomes
Undigested genomes of Pantoea isolates were analysed by PFGE according to published protocols with some modifications [26, 34]. Briefly, isolates were grown in 10 ml of LBm liquid medium for 18 h at 30°C. Cell cultures were centrifuged at 5,000 g for 20 min at 4°C. The pellet was resuspended in 1 ml of 1 × Tris-EDTA buffer to obtain an optical density between 1.8 and 2.0. Cell suspensions (0.5 ml) were mixed volume to volume with 1.6% low melting point agarose (Biorad) and the mixture was distributed per 0.1 ml in the plug molds (Biorad) and cooled at 4°C. Cells were lysed in lysis solution (2 × Tris NaCl EDTA, 10% sodium lauroyl sarcosinate, 1.4 mg ml-1 lysozyme) at 37°C for 24 h and proteins were digested with proteinase K (Euromedex) in 0.5 M EDTA pH.8 containing 1% N-lauryl-sarcosine at 37°C for 48 h. The agarose plugs were incubated in 0.4 mg ml-1 phenylmethylsulfonyl fluoride (Sigma-Aldrich) at 50°C for 1 h, washed in 0.5 M EDTA pH.8 and electrophoresed in 0.8% chromosomal-grade agarose in 1 × TAE buffer using a CHEF Mapper XA (Biorad, France) at 14°C, a constant pulse of 500 ms and a field angle of 106° for 48 h at 3 V cm-1.
Plasmid content
The procedure of Eckhardt [35] was used to identify high molecular weight plasmids in Pantoea as already described [36]. Briefly, 300 μl of bacterial culture (OD600 nm equal to 0.5) was placed on 0.3% sodium lauroyl sarcosinate in 1 × Tris-borate-EDTA (TBE) buffer. After centrifugation at 2,300 g for 5 min at 4°C, the pellet was resuspended in 25 μl of lysis solution (9% saccharose, 1.9 mg ml-1 Lysozyme and 0.38 mg ml-1 RNase) and homogenates were loaded into 0.75% agarose gels in TBE containing 1% SDS. Electrophoresis was carried out at 10 V for 20 min then 85 V for 210 min. To identify lower-molecular-weight plasmids, a second method was used as described previously [37]. Plasmid sizes were estimated by comparing their relative mobility in agarose gels with those of plasmids from sequenced Azospirillum genomes [38, 39], standard supercoiled plasmids (Life Technologies, Inc., USA) and two reference strains of Pantoea (Pantoea stewartii CFBP 3614 and Pantoea agglomerans CFBP 4740) retrieved from the French collection of phytopathogenic bacteria (http://www-intranet.angers.inra.fr/cfbp/).
Statistical analysis
Differences between mosquito genders were tested by a chi-square test using R software [40].
Results
Bacterial diversity in Ae. albopictusfrom Madagascar
Culturable bacteria from 104 field-caught Ae. albopictus adults (56 males and 48 females) were analysed by plating homogenates of whole mosquito bodies onto different culture media. The bacterial isolates obtained from each mosquito were first screened on the basis of colony characteristics including colony size, shape, colour, margin, opacity, and elevation consistency. Only one colony per type was selected per plate, with the result that 62 colonies were selected from Herellea medium, 70 from CaCO3 medium and 149 from LBm giving a total of 281 colonies to analyse from the initial 3,000 isolates.
The 16S rRNA genes were amplified from these 281 isolates and analysed by ARDRA. Forty distinct ARDRA profiles were obtained. For each profile the 16S rRNA gene was sequenced from one or more randomly chosen isolates (Table 2). The sequences were analysed by BLASTn showing that they originated from 27 bacterial genera. Some genera exhibited identical ARDRA profiles with the two enzymes used. All the genera belonged to three major phyla: Actinobacteria, Firmicutes and Proteobacteria (see Table 2 for details of families, genera and species in each phylum). One isolate was affiliated with the Deinococcus-Thermus phylum.
The distribution of bacterial phyla was significantly different according to mosquito gender (P = 0.0002). Most bacterial isolates from females were Proteobacteria (51.3%) followed by Firmicutes (30.3%) then Actinobacteria (18.4%). Conversely, Actinobacteria was the most abundant phylum in male mosquitoes (48%) followed by Proteobacteria (30.6%) and Firmicutes (20.4%). Some bacterial genera were found in both females and males, namely Curtobacterium flaccumfaciens, Microbacterium, Arthrobacter, Kocuria, Streptomyces, Bacillus, Staphylococcus, Haematobacter massiliensis, Enterobacter, Klebsiella oxytoca, Acinetobacter and Pantoea. Some bacterial genera were only associated with one mosquito gender. For instance, Asaia, Citrobacter freundii and Sphingomonas were only present in females whereas Arseniococcus bolidensis, Cellulosimicrobium, Deinococcus, Leucobacter, Planococcus, Pseudomonas, Skermanella aerolata, Xanthomonas and the Neisseria (referred to as being unculturable in databases) were exclusively isolated from males. According to the number of affiliated sequences, Pantoea was the most abundant genera, representing 25.8% of the total isolates from both male and female mosquitoes (Table 2).
Relative abundance of bacterial isolates differs according to geographic distribution
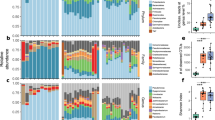
The relative abundance of isolates according to the sampling sites and the isolation media is shown in Figure 1. As expected, the isolation procedure using rich LBm medium gave the most diverse bacterial composition ranging from 3 to 8 distinct families per sampling site. Mosquitoes sampled in Ankazobe harboured only 3 bacterial families (Enterobacteriaceae, Bacillaceae, and Staphylocacceae), whereas mosquitoes from the other three sites (Tsimbazaza Park, Toamasina and Ambohidratrimo) harboured a total of 8 bacterial families per site. However, the abundance and composition of the bacteria from particular families varied between sampling sites. For instance, members of the families Moraxellaceae and Deinococcaceae were only isolated from mosquitoes in Ambohidratrimo, and those of the families Neisseriaceae and Xanthomonadaceae only from mosquitoes in Toamasina and Tsimbazaza park, respectively. While the isolation procedure was initially used to enrich for Asaia, isolates on CaCO3 medium largely belonged to Actinobacteria, irrespective of the origin of mosquitoes. Differences were also observed for members of the family Acetobacteraceae found in mosquitoes from Toamasina. As expected, on Herellea medium Gammaproteobacteria were detected with a majority of Enterobacteriaceae as well as bacteria of the genus Acinetobacter. These bacteria were only noted in mosquitoes from Toamasina and Ankazobe. Overall, the Ambohidratrimo mosquitoes harboured the highest number of distinct bacterial taxa with a total of 10 families in comparison to mosquitoes from other sites, which exhibited no more than 4 families. Members of the families Staphylococcaceae, Rhodobacteraceae, Planoccoccaeae, Intrasporangiaceae, Rhodospirillaceae, Promicromonosporaceae were only present in mosquitoes from Ambohidratrimo.
Molecular characterization of the Pantoeaisolates
As Pantoea was the most prevalent genus isolated from mosquitoes from three of the four sites, it was further characterized by analysing its genomic structure. Nearly complete rrs gene sequences were obtained from 11 isolates that were compared to reference strains (Table 3). PFGE showed that Pantoea contains a high-molecular-weight replicon (>3.13 Mb), which when considered alongside sequenced reference genomes (http://www.ncbi.nlm.nih.gov) probably corresponds to the bacterial chromosome (Figure 2). Two other replicons each less than 1 Mb were also seen in the PFGE pattern which makes it possible to classify isolates into two groups. One group comprises mosquito isolates no. 127 and no. 131 with the reference strain Pantoea stewartii (CFBP 3614), another group included mosquito isolates no. 95 and no. 110 with the reference strain Pantoea agglomerans (CFBP 4740) while all other mosquito isolates have patterns closely related to each other but distinct from the reference strains. When the Eckhardt procedure for plasmid analysis was used, high-molecular-weight plasmids (from 75 kb up to 980 kb) from Pantoea mosquito isolates were detected. The number (from 2 to 6) and size of plasmids were different from those observed in reference strains (Figure 3). If classified according to plasmid content, mosquito isolates no. 127 and no. 131 showed unique patterns that were similar to each other, while the other mosquito isolates clustered into two distinct groups. The first group included 6 isolates (nos. 85, 86, 93, 95, 104 and 124) and the second group contained 3 isolates (nos. 110, 111 and 115) (Figure 3). Using another method to detect lower-molecular-weight plasmids (less than 28 kb), two supplementary plasmids were detected in mosquito isolates no. 127 and no. 131 only, around 8 and 15 kb (data not shown).
PFGE of undigested genomic DNA of Pantoea mosquito isolates and their reference strains. Chromosomal DNA from Hansenula wingei was used as a reference (BioRad). Characteristics of the samples are indicated in Table 3.
One representative sequence of Pantoea isolates according to their ARDRA profile and sequence composition was chosen and the sequences were compared to construct a phylogenetic tree. Sequences of Pantoea isolated from other insect species (stink bug, honeybee, Onion thrip, beetle and the mosquito Culex quinquefasciatus) and the environment (plants and plant nectar) (Figure 4) were also included. The topology of the tree showed that Pantoea isolated from Ae. albopictus clustered with the sequence from Pantoea dispersa from C. quinquefasciatus and the sequence from Pantoea sp. from ant.
Phylogenetic analysis based on partial 16S rRNA gene sequences of Pantoea obtained from this study and some of those available in GenBank. Identification and GenBank accession numbers are indicated for each sample. The phylogenetic tree was constructed using the Hasegawa, Kishino and Yano maximum likelihood method, with bootstrap analysis with 1000 replicates. Numbers on branches indicate support for each clade ≥ 50%. Only one representative sequence of Ae. albopictus Pantoea isolates is listed in the tree corresponding to the classification of the 45 Pantoea isolates according to their ARDRA profile and sequence composition.
Discussion
We found a total of 27 genera of culturable bacteria associated with the mosquito Ae. albopictus caught in different regions of Madagascar. This relatively high number might be partly attributable to the variety of culture media used and provides evidence of the diversity of culturable bacteria present in wild Ae. albopictus. The 16S rDNA sequences from the isolates indicated that they belonged to 19 families from three major phyla, Actinobacteria, Firmicutes and Proteobacteria. Of the 27 bacterial genera identified here, 12 had been previously found in other mosquito species, so this suggests that these bacteria might be in a close relationship with mosquitoes or enable mosquitoes to be better adapted to the environment [8–10, 12, 41, 42].
Many isolates identified are known to be commensal bacteria for plant and soil environments. In mosquitoes, the origin of commensal bacteria has not yet been fully resolved. Usually bacteria can be acquired in two ways, either by vertical inheritance through generations or through continual acquisition from the environment. Moreover, the mosquito gender is also an important factor that affects bacterial microbiota composition, as already demonstrated [12]. This difference is mainly due to the fact that male and female mosquitoes exhibit different ecological behaviors in terms of nutritional capabilities. Both genders feed on nectar and plant saps, but females are also hematophagous. Consequently, diet regime (sugar and/or blood meals) can significantly affect the bacterial structure. However, information on the sugar feeding of Ae. albopictus in the field is scarce [43]. Recently, a first survey of bacteria in floral nectar from a natural plant community showed that Pantoea was one of the most common bacterial genus recovered [44]. Interestingly, we showed that this bacterium was also the most prevalent in Ae. albopictus mosquitoes, suggesting a potential route of its acquisition through the environment.
A total of eight 16S rDNA sequences identified were similar to those of bacteria encountered in human clinical specimens, including the species Microbacterium, Klebsiella oxytoca and Haematobacter massiliensis[45, 46]. As mosquitoes are mostly known to transmit arboviruses and parasites, it is possible that they also transmit, even on a small scale, opportunistic bacterial pathogens to human and animals.
In our previous study of Ae. albopictus populations from Madagascar, we identified the phyla Proteobacteria and Firmicutes, with Bacillus as a predominant isolated genus [12]. Here the majority of isolates belonged to the Enterobacteriaceae family and Pantoea was the most common genus probably due to the difference in the sampling region as well as the cultural media used. The relatively high prevalence of Pantoea isolates found in the present study emphasizes the need to also consider this bacterium as an intimate partner of the mosquito vector and to better explore its abundance and persistence among field populations, as previously explored in the context of the prevalence study performed on Acinetobacter and Asaia in the same areas. The genus Pantoea is polyphyletic and comprises seven species [47]. Following the results of phylogenetic analyses, sequences of Pantoea isolates from Ae. albopictus tended to cluster together and with those originated from the C. quinquefasciatus species as well as one isolate from ant. A larger number of sequences is thus needed to make conclusions on the presence of well-conserved sequence of Pantoea isolates in mosquitoes. For this purpose, it would be necessary to pursue the global effort to obtain new Pantoea isolates from insects and environment. Members of Pantoea are commonly isolated from the environment, mostly from water and soil, and some isolates have been recovered from human clinical samples or as causative agents of plant diseases. Pantoea agglomerans can establish a symbiotic relationship in western flower thrips (Frankliniella occidentalis) that persists for over 50 generations or about 2 years [48]. Pantoea agglomerans was also the most frequently isolated bacterium from the midgut of Anopheles funestus and An. gambiae species caught in Kenya and Mali [49], and it has been shown to easily adapt to its hosts [50]. This bacterium was also recently detected in Ae. albopictus from North America [51]. Recently, Bisi and Lampe [22] hypothesized that P. agglomerans could be engineered to express and secrete anti-plasmodium effector proteins in Anopheles mosquitoes. As Pantoea was the most prevalent bacterium isolated in our study, it could also be a candidate for paratransgenesis in Ae. albopictus. One strategy in paratransgenesis is to insert the gene of interest into plasmids hosted by the chosen bacterium. We found Pantoea isolates from Ae. albopictus harboured at least two plasmids per genome, so it would be of interest to further determine if plasmids can be laterally transferred between strains and which genes they carry.
Conclusions
This study highlights the diverse culturable bacteria in field populations of Ae. albopictus. Some of them were detected for the first time in this vector and their functions are not known at all. Further studies are needed to investigate the physiological characteristics of the bacterial isolates and their possible interactions with mosquito biology and vector competence. This information could be of great importance in developing new alternative control strategies based on the use of symbiotically modified mosquitoes.
References
Rosenberg E, Zilber-Rosenberg I: Symbiosis and development: the hologenome concept. Birth Defects Res C Embryo Today. 2011, 93 (1): 56-66. 10.1002/bdrc.20196.
Dillon R, Charnley K: Mutualism between the desert locust Schistocerca gregaria and its gut microbiota. Res Microbiol. 2002, 153: 503-539. 10.1016/S0923-2508(02)01361-X.
Dillon RJ, Dillon VM: The gut bacteria of insects: nonpathogenic interactions. Annu Rev Entomol. 2004, 49: 71-92. 10.1146/annurev.ento.49.061802.123416.
Sharon G, Segal D, Ringo JM, Hefetz A, Zilber-Rosenberg I, Rosenberg E: Commensal bacteria play a role in mating preference of Drosophila melanogaster. Proc Natl Acad Sci USA. 2010, 107 (46): 20051-20056. 10.1073/pnas.1009906107.
Tsuchida T, Koga R, Horikawa M, Tsunoda T, Maoka T, Matsumoto S, Simon JC, Fukatsu T: Symbiotic bacterium modifies aphid body color. Science. 2010, 330: 1102-1104. 10.1126/science.1195463.
Toju H, Fukatsu T: Diversity and infection prevalence of endosymbionts in natural populations of the chestnut weevil: relevance of local climate and host plants. Mol Ecol. 2011, 20: 853-868. 10.1111/j.1365-294X.2010.04980.x.
Pidiyar VJ, Jangid K, Patole MS, Shouche YS: Studies on cultured and uncultured microbiota of wild Culex quinquefasciatus mosquito midgut based on 16 s ribosomal RNA gene analysis. AmJTrop Med Hyg. 2004, 70: 597-603.
Rani A, Sharma A, Rajagopal R, Adak T, Bhatnagar RK: Bacterial diversity analysis of larvae and adult midgut microflora using culture-dependent and culture-independent methods in lab-reared and field-collected Anopheles stephensi-an Asian malarial vector. BMC Microbiol. 2009, 19 (9): 96-
Gusmão DS, Santos AV, Marini DC, Bacci M, Berbert-Molina MA, Lemos FJ: Culture-dependent and culture-independent characterization of microorganisms associated with Aedes aegypti (Diptera: Culicidae) (L.) and dynamics of bacterial colonization in the midgut. Acta Trop. 2010, 115: 275-281. 10.1016/j.actatropica.2010.04.011.
Chavshin AR, Oshaghi MA, Vatandoost H, Pourmand MR, Raeisi A, Enayati AA, Mardani N, Ghoorchian S: Identification of bacterial microflora in the midgut of the larvae and adult of wild caught Anopheles stephensi: a step toward finding suitable paratransgenesis candidates. Acta Trop. 2012, 121: 129-134. 10.1016/j.actatropica.2011.10.015.
Dinparast Djadid N, Jazayeri H, Raz A, Favia G, Ricci I, Zakeri S: Identification of the midgut microbiota of An. stephensi and An. maculipennis for their application as a paratransgenic tool against malaria. PLoS One. 2011, 6: e28484-10.1371/journal.pone.0028484.
Zouache K, Raharimalala FN, Raquin V, Tran-Van V, Raveloson LHR, Ravelonandro P, Mavingui P: Bacterial diversity of field-caught mosquitoes, Aedes albopictus and Aedes aegypti, from different geographic regions of Madagascar. FEMS Microbiol Ecol. 2011, 75: 377-389. 10.1111/j.1574-6941.2010.01012.x.
Streit WR, Schmitz RA: Metagenomics-the key to the uncultured microbes. Curr Opin Microbiol. 2004, 7 (5): 492-498. 10.1016/j.mib.2004.08.002.
Boissière A, Tchioffo MT, Bachar D, Abate L, Marie A, Nsango SE, Shahbazkia HR, Awono-Ambene PH, Levashina EA, Christen R, Morlais I: Midgut microbiota of the malaria mosquito vector Anopheles gambiae and interactions with Plasmodium falciparum infection. PLoS Patho. 2012, 8 (5): e1002742-10.1371/journal.ppat.1002742.
Schäfer A, Konrad R, Kuhnigk T, Kämpfer P, Hertel H, König H: Hemicellulose-degrading bacteria and yeasts from the termite gut. J Appl Bacteriol. 1996, 80 (5): 471-478. 10.1111/j.1365-2672.1996.tb03245.x.
Watanabe Y, Shinzato N, Fukatsu T: Isolation of actinomycetes from termites’ guts. Biosci Biotechnol Biochem. 2003, 7 (8): 1797-1801.
Moran NA, Baumann P: Bacterial endosymbionts in animals. Curr Opin Microbiol. 2000, 3 (3): 270-275. 10.1016/S1369-5274(00)00088-6.
Pinto-Tomás AA, Anderson MA, Suen G, Stevenson DM, Chu FS, Cleland W, Weimer PJ, Currie CR: Symbiotic nitrogen fixation in the fungus gardens of leaf-cutter ants. Science. 2009, 326 (5956): 1120-1123. 10.1126/science.1173036.
Malhotra J, Dua A, Saxena A, Sangwan N, Mukherjee U, Pandey N, Rajagopal R, Khurana P, Khurana JP, Lal R: Genome sequence of Acinetobacter sp. strain HA, isolated from the gut of the polyphagous insect pest Helicoverpa armigera. J Bacteriol. 2012, 194 (18): 5156-10.1128/JB.01194-12.
Coutinho-Abreu IV, Zhu KY, Ramalho-Ortigao M: Transgenesis and paratransgenesis to control insect-borne diseases: current status and future challenges. Parasitol Int. 2010, 59: 1-8. 10.1016/j.parint.2009.10.002.
Favia G, Ricci I, Marzorati M, Negri I, Alma A, Sacchi L, Bandi C, Daffonchio D: Bacteria of the genus Asaia: a potential paratransgenic weapon against malaria. Adv Exp Med Biol. 2008, 27: 49-59.
Bisi DC, Lampe DJ: Secretion of anti-Plasmodium effector proteins from a natural Pantoea agglomerans isolate by using PelB and HlyA secretion signals. Appl Environ Microbiol. 2011, 77: 4669-4675. 10.1128/AEM.00514-11.
Lambrechts L, Scott TW, Gubler DJ: Consequences of the expanding global distribution of Aedes albopictus for dengue virus transmission. PLoS Negl Trop Dis. 2010, 25; 4 (5): e646-
Ratsitorahina M, Harisoa J, Ratovonjato J, Biacabe S, Reynes JM, Zeller H, Raoelina Y, Talarmin A, Richard V, Louis Soares J: Outbreak of dengue and Chikungunya fevers, Toamasina, Madagascar, 2006. Emerging Infect Dis. 2008, 14: 1135-1137. 10.3201/eid1407.071521.
Renault P, Balleydier E, D’Ortenzio E, Bâville M, Filleul L: Epidemiology of chikungunya infection on Reunion Island, Mayotte, and neighboring countries. Med Mal Infect. 2012, 42: 93-101. 10.1016/j.medmal.2011.12.002.
Minard G, Tran FH, Raharimalala FN, Hellard E, Ravelonandro P, Mavingui P, Valiente Moro C: Prevalence, genomic and metabolic profiles of Acinetobacter and Asaia associated with field-caught Aedes albopictus from Madagascar. FEMS Microbiol Ecol. 2013, 83: 63-73. 10.1111/j.1574-6941.2012.01455.x.
Raharimalala FN, Ravaomanarivo LH, Ravelonandro P, Rafarasoa LS, Zouache K, Tran-Van V, Mousson L, Failloux AB, Hellard E, Moro CV, Ralisoa BO, Mavingui P: Biogeography of the two major arbovirus mosquito vectors, Aedes aegypti and Aedes albopictus (Diptera, Culicidae), in Madagascar. Parasit Vectors. 2012, 5: 56-10.1186/1756-3305-5-56.
Ravaonjanahary C: Les Aedes de Madagascar. 1978, France: Travaux et documents de 1′ORSTOM
Bouvet PJM, Joly-Guillou ML: Acinetobacter. Précis de bactériologie Clinique. Edited by: Freney J, Renaud F, Hansen et W, Bollet C. 2000, Paris: Editions ESKA, 1239-1258.
Mandel AD, Wright K, McKinnon JM: Selective medium for isolation of Mima and Herellea organisms. J Bacteriol. 1964, 88: 1524-1525.
Listiyanti P, Kawasaki H, Seki T, Yamoda Y, Chimura T, Komagata K: Identification of Acetobacter Strains isolated from Indonesian sources, and proposals of Acetobacter syzygii sp. nov., Acetobacter Cibinongensis sp.nov. Acetobacter cibinongensis sp. nov., and Acetobacter orientalis sp. J Gen Appl Microbiol. 2001, 47: 119-131. 10.2323/jgam.47.119.
Chouaia B, Rossi P, Montagna M, Ricci I, Crotti E, Damiani C, Epis S, Faye I, Sagnon N, Alma A, Favia G, Daffonchio D, Bandi C: Molecular evidence for multiple infections as revealed by typing of Asaia bacterial symbionts of four mosquito species. Appl Environ Microbiol. 2010, 76: 7444-7450. 10.1128/AEM.01747-10.
Hall TA: BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser. 1999, 41: 95-98.
Schwartz DC, Cantor CR: Separation of yeast chromosome-sized DNAs by pulsed field gradient gel electrophoresis. Cell. 1984, 37: 67-75. 10.1016/0092-8674(84)90301-5.
Eckhardt T: A rapid method for the identification of plasmid desoxyribonucleic acid in bacteria. Plasmid. 1978, 1: 584-588. 10.1016/0147-619X(78)90016-1.
Mavingui P, Flores M, Guo X, Dávila G, Perret X, Broughton WJ, Palacios R: Dynamics of genome architecture in Rhizobium sp. strain NGR234. J Bacteriol. 2002, 184: 171-176. 10.1128/JB.184.1.171-176.2002.
Seifert H, Boullion B, Schulze A, Pulverer G: Plasmid DNA profiles of Acinetobacter baumannii: clinical application in a complex endemic setting. Infect Control Hosp Epidemiol. 1994, 15: 520-528. 10.1086/646970.
Caballero-Mellado J, López-Reyes L, Bustillos R: Presence of 16S rRNA Genes in Multiple Replicons in Azospirillum brasilense. FEMS Microbial Lett. 1999, 178: 283-288. 10.1111/j.1574-6968.1999.tb08689.x.
Wisniewski-Dyé F, Borziak K, Khalsa-Moyers G, Alexandre G, Sukharnikov LO, Wuichet K, Hurst GB, McDonald WH, Robertson JS, Barbe V, Calteau A, Rouy Z, Mangenot S, Prigent-Combaret C, Normand P, Boyer M, Siguier P, Dessaux Y, Elmerich C, Condemine G, Krishnen G, Kennedy I, Paterson AH, González V, Mavingui P, Zhulin IB: Azospirillum genomes reveal transition of bacteria from aquatic to terrestrial environments. PLoS Genet. 2011, 7: e1002430-10.1371/journal.pgen.1002430.
R Development Core Team: R: A Language and Environment for Statistical computing. R Foundation for Statistical Computing, Vienna. 2009, Available at: http://www.R-project.org
Lindh JM, Terenius O, Faye I: 16S rRNA gene-based identification of midgut bacteria from field-caught Anopheles gambiae sensu lato and A. funestus mosquitoes reveals new species related to known insect symbionts. Appl Environ Microbiol. 2005, 71: 7217-7223. 10.1128/AEM.71.11.7217-7223.2005.
Terenius O, Lindh JM, Eriksson-Gonzales K, Bussière L, Laugen AT, Bergquist H, Titanji K, Faye I: Midgut bacterial dynamics in Aedes aegypti. FEMS Microbiol Ecol. 2012, 80: 556-565. 10.1111/j.1574-6941.2012.01317.x.
Müller GC, Xue RD, Schlein Y: Differential attraction of Aedes albopictus in the field to flowers, fruits and honeydew. Acta Trop. 2011, 118: 45-49. 10.1016/j.actatropica.2011.01.009.
Alvarez-Pérez S, Herrera CM, de Vega C: Zooming-in on floral nectar: a first exploration of nectar-associated bacteria in wild plant communities. FEMS Microbiol Ecol. 2012, 80: 591-602. 10.1111/j.1574-6941.2012.01329.x.
Gneiding K, Frodl R, Funke G: Identities of Microbacterium spp. encountered in human clinical specimens. J Clin Microbiol. 2008, 46: 3646-3652. 10.1128/JCM.01202-08.
Helsel LO, Hollis D, Steigerwalt AG, Morey RE, Jordan J, Aye T, Radosevic J, Jannat-Khah D, Thiry D, Lonsway DR, Patel JB, Daneshvar MI, Levett PN: Identification of “Haematobacter” a new genus of aerobic Gram-negative rods isolated from clinical specimens, and reclassification of Rhodobacter massiliensis as “Haematobacter massiliensis comb. nov.”. J Clin Microbiol. 2007, 45: 1238-1243. 10.1128/JCM.01188-06.
Brady C, Cleenwerck I, Venter S, Vancanneyt M, Swings J, Coutinho T: Phylogeny and identification of Pantoea species associated with plants, humans and the natural environment based on multilocus sequence analysis (MLSA). Syst Appl Microbiol. 2008, 31 (6–8): 447-460.
de Vries EJ, Jacobs G, Breeuwer JA: Growth and transmission of gut bacteria in the Western flower thrips. Frankliniella occidentalis. J Invertebr Pathol. 2001, 77 (2): 129-137. 10.1006/jipa.2001.5010.
Straif SC, Mbogo CN, Toure AM, Walker ED, Kaufman M, Toure YT, Beier JC: Midgut bacteria in Anopheles gambiae and An. funestus (Diptera: Culicidae) from Kenya and Mali. J Med Entomol. 1998, 35: 222-226.
Riehle MA, Moreira CK, Lampe D, Lauzon C, Jacobs-Lorena M: Using bacteria to express and display anti-Plasmodium molecules in the mosquito midgut. Int J Parasitol. 2007, 37: 595-603. 10.1016/j.ijpara.2006.12.002.
Joyce JD, Nogueira JR, Bales AA, Pittman KE, Anderson JR: Interactions between La Crosse virus and bacteria isolated from the digestive tract of Aedes albopictus (Diptera: Culicidae). J Med Entomol. 2011, 48 (2): 389-94. 10.1603/ME09268.
Acknowledgments
We are grateful to Madagascar National Parks for authorizing the collection of wild mosquitoes under ethical approval. This work was carried out within the frameworks of GDRI “Biodiversité et Développement Durable à Madagascar” and COST action F0701 ‘Arthropod Symbioses: from fundamental to pest disease management’.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
CVM conceived the design of the study, participated in all the tasks and performed sequence analysis. FHT carried out the molecular identification of bacteria (ARDRA, PFGE, plasmid profiles). FNR participated in the sampling of mosquitoes and the isolation of bacteria. PR participated in the design of the study. PM conceived of the study and drafted the manuscript. All authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Valiente Moro, C., Tran, F.H., Nantenaina Raharimalala, F. et al. Diversity of culturable bacteria including Pantoea in wild mosquito Aedes albopictus. BMC Microbiol 13, 70 (2013). https://doi.org/10.1186/1471-2180-13-70
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2180-13-70