Abstract
Background
Carbapenems are the antibiotics of choice to treat infections caused by Acinetobacter baumannii, and resistance to this class can be determined by loss of membrane permeability and enzymatic mechanisms. Here, we analyzed the basis of carbapenem resistance in clinical A. baumannii isolates from different Brazilian regions.
Results
The analyses addressed the carbapenemase activity of OXA-23, CarO expression and alterations in its primary structure. Susceptibility test revealed that the strains presented the COS (Colistin-Only-Sensitive) profile. PCR and sequencing showed the presence of the chromosomally-encoded blaOXA-51 in all isolates. The majority of strains (53%) carried the carbapenemase blaOXA-23 gene associated with ISAba1. The Hodge test indicated that these strains are carbapenemase producers. PFGE revealed 14 genotypes among strains from Rio de Janeiro and Maranhão. The influence of carO on imipenem resistance was evaluated considering two aspects: the composition of the primary amino acid sequence; and the expression level of this porin. Sequencing and in silico analyses showed the occurrence of CarOa, CarOb and undefined CarO types, and Real Time RT-PCR revealed basal and reduced carO transcription levels among isolates.
Conclusions
We concluded that, in general, for these Brazilian isolates, the major carbapenem resistance mechanism was due to OXA-23 carbapenemase activity and that loss of CarO porin plays a minor role in this phenotype. However, it was possible to associate the carO alleles and their expression with imipenem resistance. Therefore, these findings underline the complexity in addressing the role of different mechanisms in carbapenem resistance and highlight the possible influence of CarO type in this phenotype.
Similar content being viewed by others
Background
Acinetobacter baumannii has been considered one of the major nosocomial pathogens worldwide, and it is included in the group of ESKAPE bugs, in which some lineages effectively escape the action of antibacterial drugs [1]. Carbapenems remain the antibiotic of choice to treat A. baumannii and other Gram-negative infections due to both a wider spectrum of antibacterial activity and less frequent side effects. However, their overuse and misuse have selected for nosocomial isolates presenting intrinsic and acquired multidrug resistance determinants [2]. It has been considered that resistance against carbapenems is, in itself, sufficient to define an A. baumannii as highly resistant [3]. The molecular basis of carbapenem resistance in this species has been ascribed to the recruitment and production of carbapenem-hydrolysing class D β-lactamases (CHDLs) and, to a lesser extent, of metallo-β-lactamases (MBLs). In A. baumannii, the CHDLs can be intrinsic (OXA-51-like) or acquired (OXA-23-like, OXA-24-like and OXA-58-like) [3]. Although these enzymes weakly hydrolyze carbapenems, they can confer high resistance when blaOXA genes are overexpressed, as a result of their association with mobile elements, such as ISAba1, which carries a strong promoter [4]. The blaOXA-23 gene, in association with ISAba1, is spread in A. baumannii worldwide, and it is the most prevalent OXA allele in isolates from Brazil [3, 5–7].
In addition to this enzymatic resistance, the loss of membrane permeability, due to alterations in specific porins, is an intrinsic carbapenem resistance mechanism in A. baumannii. The 25/29 kDa heat-modifiable carbapenem-associated outer membrane protein (CarO) is a porin with a β-barrel topology. This channel allows the selective uptake of amino acids and imipenem (but not meropenem) due to its structural conformation and to the presence of an imipenem binding site [8, 9]. This site was not found in any other OMP, and thus, CarO protein is considered a preferred uptake channel for this antibiotic. Based on variable domains of CarO, this channel is classified in two groups, CarOa and CarOb, with high affinity to imipenem. Moreover, CarOb was showed to be twice more specific for imipenem than the CarOa. In this way, carO gene alterations, as disruption by insertion sequences, changes in the primary structure - mainly in the imipenem binding site - or decreased expression, would have a dramatic impact on the entry of imipenem in the cell, thus contributing to resistance to this antibiotic [9]. Concerning the participation of CarO porin in the resistance phenotype, few descriptive studies have only shown the presence/absence of the gene and protein, resulting in a lack of studies showing the association of the carO alleles and their expression with the resistance phenotype.
Brazil is a continental country and, concerning resistance in A. baumannii, the studies have been concentrated in the Southeast region [5, 7], which could not be representative of the A. baumannii diversity as showed previously [10]. Therefore, the role of CarOa and CarOb in the differential imipenem uptake, and the current diversity of carO alleles available in the GenBank lead us to investigate these aspects in a set of carbapenem-resistant clinical A. baumannii strains from two Brazilian states placed ~3,000 Km apart and with distinct ecological and social contexts. Moreover, the prevalence and participation of carbapenemases in this phenotype was also evaluated.
Results and discussion
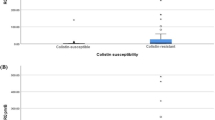
All the 28 strains were resistant to, at least, meropenem. Most of the strains from both Rio de Janeiro and Maranhão were resistant to ceftazidime, cefepime, ciprofloxacin, trimethoprim, amikacin and streptomycin (Table 1). A considerable discrepancy was observed between strains from Rio de Janeiro and Maranhão concerning the resistance to aztreonam, gentamicin, piperacillin/tazobactam, ertapenem, and imipenem (Table 1). Three/thirteen (23%) strains from Rio de Janeiro were resistant to imipenem contrasting with 13/15 (86.6%) resistant strains from Maranhão. All strains from this study were susceptible to tigecycline and colistin (MICs ranging from 0.25 to 0.50 μg/mL), and strains resistant to imipenem presented MICs ≥ 32 μg/mL.
All strains harboured the chromosomally-encoded blaOXA-51 gene, however, the ISAba1 was not associated to this allele in any strain. In fact, the ISAba1-blaOXA-51 combination is not frequent [5], although a previous study had found this association in the Southern region of Brazil [7]. Additionally to blaOXA-51, 13/15 (86.6%) strains from Maranhão carried also the CHDL blaOXA-23 gene, while only 2/13 (15%) strains from Rio de Janeiro (AC2-RJ and AC72-RJ) harboured this allele. Spite the low prevalence of blaOXA-23 in AC-RJ strains (this work) and in São Paulo [6], it is the most prevailing and disseminated allele in Brazil. All blaOXA-23 from both Maranhão and Rio de Janeiro were preceded by ISAba1 as found in previous works from several Brazilian states [5, 7]. Therefore, the high imipenem resistance (MIC, ≥ 32 μg/mL) presented in all strains harbouring blaOXA-23 can be due to its overexpression driven by the ISAba1 promoter [4]. All strains from Rio de Janeiro, except by AC2-RJ and AC72-RJ, carried blaOXA alleles of narrow resistance spectrum (blaOXA-13, -10 or -66), none of them associated with ISAba1, spite its presence in AC-RJ strains (Table 1). The isolates were negative for MBL genes, blaOXA-58-like and blaOXA-24-like alleles. In summary, our results revealed that only OXA with carbapenemase activity (blaOXA-23) was associated with ISAba1 in the strains analyzed here, and that all of them were resistant to carbapenems. These results are summarized in Table 1.
The Hodge test was in agreement with the carbapenemase activity and the consequent carbapenem resistance, where positive results were obtained only for the strains carrying ISAba1-blaOXA-23, while negative results were observed for strains harbouring OXA alleles with no or a weak carbapenemase activity. Thus, these results reflect the OXA-23 overexpression driven by the strong promoter [4, 11] found in ISAba1 (Table 1).
A. baumannii has a clonal structure population [12], however, several unrelated carbapenem-resistant lineages have been reported in Brazil [13, 14]. In the present work, macrorestriction analyses revealed eight genotypes in Rio de Janeiro (A-H) and six genotypes in Maranhão (I-N), totalizing 14 clones, none of them related to the international clones. In Maranhão, the majority of strains (8/15) belonged to genotype I, however in Rio de Janeiro there was no predominant genotype. In this study, the blaOXA-23 gene was distributed in several genotypes, while in São Paulo this allele occurred in only one genotype [6]. These findings show the diversity of clinical A. baumannii circulating in Brazil. One strain of each genotype was selected as a representant to perform further analyses of carO transcription and gene sequencing (Table 1).
Considering CarO, imipenem resistance can be influenced in three aspects: i) the occurrence of gene disruption resulting in the lack of porin formation in the outer membrane; ii) amino acid changes in both CarOa and CarOb that could account for an altered porin with low or no affinity for the antibiotic and/or; iii) decrease expression of the carO gene that causes a low density of channels. Therefore, to fully investigate these aspects, it is necessary to assess the carO nucleotide and deduced amino acid sequences and its expression. However, there is a lack of this set of data when the role of carO in imipenem resistance is addressed.
Here, in order to raise all these information concerning carO, we analyzed the expression of this gene by relative quantification, and the nucleotide/predicted amino acid sequences were determined to type the CarO (CarOa or CarOb) and to evaluate gene disruption by IS. The carO sequences from AC-RJ and AC-MA presented substitutions without the presence of frameshifts and/or premature stop codons when compared to CarO of A. baumannii ATCC 19606. Analyses of the carO promoter region showed that all strains harbour the canonical sequence as described previously [15].
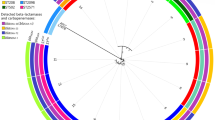
The predicted amino acid sequence showed that CarOa and CarOb were distributed among isolates from Maranhão and Rio de Janeiro. Moreover, blastX analyses revealed CarO sequences with amino acid differences considering both CarOa and CarOb, which was named here undefined CarO (Table 1 and Figure 1). The undefined CarO porins (AC6-RJ, AC-67-MA and AC71-MA) presented one to five amino acid differences among each other, with 70% and 74% identity with CarOa and CarOb, respectively. One of these, from AC6-RJ, is identical to a CarO previously identified in Argentina [GenBank: AAV80243] [15]. AC67-MA and AC71-MA carried two new undefined CarO with 98% and 99% amino acid identity, respectively, with CarO sequences identified in strains from Germany, United States, Leban and China [GenBank: EKL39362, EKL48836, ABG27024, AFH56947, respectively] (Figure 1). Therefore, taking into account the observed CarO diversity, and that the CarO primary structure influence the imipenem affinity and, consequently the imipenem resistance, the impact of other isoforms on this phenotype has to be investigated.
Alignment of A. baumannii CarO amino acid sequences from this study and available in GenBank. The ATCC 19606 [GenBank: ZP_05828783.1] and A6 [GenBank: ADQ27797.1] CarO sequences were selected as the representants of the CarOa and CarOb groups, respectively. Ab244 [GenBank: AAV80243; Argentina], Acb-1 [GenBank: ABG27024; Leban], ZJM007 [GenBank: AFH56947; China], OIFC074 [GenBank: EKL39362; Germany] and Naval-83 [GenBank: EKL48836; United States] were representants of different undefined CarO isoforms. The numbers correspond to amino acid positions. The gray box labels the CarOb group.
Considering the imipenem-susceptible strains, relative quantification assay revealed that AC1-RJ (CarOa), AC3-RJ (CarOa), AC10-RJ (CarOa), AC16-RJ (CarOb), and AC70-MA (CarOa) had a basal transcription level of carO as ATCC 19606 (Table 1). For this group of strains, the carO transcript amount and the type of CarO were in agreement with the imipenem susceptibility phenotype (Table 1). Moreover, these strains did not produce carbapenemases. However, the carO expression did not correlate with this phenotype in the imipenem-susceptible AC60-MA and AC9-RJ strains. In AC60-MA, which is highly susceptible to imipenem (MIC, 0.20 μg/mL), the carO gene was disrupted by the ISAba1 (Table 1). Therefore, the porin loss in this strain was not evoking resistance, indicating the participation of other mechanisms in the imipenem uptake. Similarly, AC9-RJ, susceptible to imipenem (MIC, 0.25 μg/mL), presented a 10-fold decrease in carO expression. It can be suggested that, apparently, this 10-fold expression reduction was not enough to cause the total loss of porin formation. Interestingly, AC9-RJ has a porin from the CarOb group, which is more specific towards imipenem than CarOa [9]. Thus, even in lower amounts of the porin, the imipenem uptake can be occurring due to CarO structural conformation in this strain, explaining its susceptibility to imipenem.
Considering the imipenem-resistant strains, AC2-RJ (CarOa), AC62-MA (CarOa), AC65-MA (CarOb), AC67-MA (undefined CarO) and AC71-MA (undefined CarO) had low carO transcript amounts (Table 1). The exception was AC72-RJ, which presented basal carO transcription. All of them produce the CHDL OXA-23, and therefore, it was not possible to establish the contribution of CarO in this resistance phenotype (Table 1). Conversely, the imipenem-resistant AC6-RJ strain (MIC, 32 μg/mL), presented basal carO transcription level (1.07-fold), and no carbapenemase production was detected (Table 1). As showed above, the AC6-RJ harbours an undefined CarO type, presenting 70% and 74% amino acid identity with CarOa and CarOb, respectively. Therefore, these CarO could have an altered imipinem binding site, influencing the antibiotic affinity, resulting in the observed resistance. In fact, this undefined CarO from AC6-RJ was identical to that from Ab244 strain from Argentina, which has been shown to play an essential role in the selective uptake of L-ornithine, but not of imipenem [8, 9].
Therefore, to correlate CarO with imipenem resistance, not only the expression has to be considered, but also the CarO primary sequence. The results obtained for AC6-RJ and AC9-RJ provide evidences of a possible correlation between CarO isoforms and imipenem resistance/susceptibility phenotype (Table 1). Moreover, PROVEAN analyses revealed that the amino acid substitutions (approximately 67 residues) found in the different undefined CarO identified in this study had a neutral impact in protein function. These results indicate that even with this high proportion of substitutions they are not interfering with CarO function and formation.
The particular role of some CarO (CarOa and CarOb) in the imipenem affinity had been experimentally determined by Catel-Ferreira and colleagues [9]. Vashist et al. [16] indirectly showed a correlation between expression, the CarO isoform and imipenem resistance based on proteomic approaches. Here, we directly determined the CarO isoforms by assessing the substitutions in the amino acid primary sequence and the transcription level of CarO. Also, Vashist and colleagues showed that altered CarO isoforms were upregulated while the native CarO (CarOa and CarOb) were downregulated in carbapenem-resistant A. baumannii. These results obtained at the post-translational level contrast with ours since no correlation between the CarO (native or undefined) and the transcription level (up and downregulation) was observed in this study (Table 1). This reflects that post-transcriptional events may be occurring during porin formation in the membrane, since in that work the authors analyzed the protein. Therefore, our study is the first to infer the role of CarO on imipenem resistance considering its expression and allelic diversity.
Conclusions
Studies in distinct Brazilian geographic regions have already shown the prevalence of OXA-23 among the several CHDLs in imipenem resistant A. baumannii strains [5, 7]. However, there is a lack of data concerning the influence of membrane permeability loss in this phenotype. We concluded that, spite of the carbapenemase production being the major carbapenem resistance mechanism in A. baumannii analyzed here, the amino acid composition of CarO porins can affects more drastically the imipenem resistance than the transcription level itself. Therefore, these findings highlight the possible role of CarO isoforms in this phenotype.
Methods
Bacterial strains and detection of carbapenem-resistance genes
The study included 28 clinical carbapenem-resistant A. baumannii belonging to the Bacterial Culture Collection of Environment and Health (CBAS) from the Oswaldo Cruz Institute (FIOCRUZ). The isolates were recovered from Maranhão in 2006 (n = 15) and Rio de Janeiro in 2008 (n = 13), and presented a COS phenotype, with the exception of AC60-MA strain. PCR and sequencing reactions were performed targeting the MBLs genes blaIMP and blaVIM alleles, blaSPM-1, and blaNDM-1; and the CHDLs blaOXA-58-like, blaOXA-51, blaOXA-24-like, blaOXA-23, blaOXA-143 and other OXA alleles (Table 2), alone and downstream ISAba1. Also, the entire carO gene and its promoter region were evaluated. Primer list is detailed in Table 2.
Antimicrobial susceptibility testing
The resistance phenotype was obtained by the disc-diffusion method [17] for the following antibiotics: aztreonam, cefepime, ceftazidime, piperacillin-tazobactam, ciprofloxacin, trimethoprim, imipenem, meropenem, ertapenem, gentamicin, amikacin, streptomycin, colistin and tigecycline. The MIC was determined by E-Test method (Biomerieux) for imipenem and colistin.
Evaluation of carbapenemase production
The carbapenem-resistant isolates were submitted to the modified Hodge test [18] in order to detect the carbapenemase production. In summary, the assay was carried out on Müeller-Hinton agar plates using ertapenem, meropenem and imipenem disks. The blaSPM-1 MBL-producing Pseudomonas aeruginosa (PS600) from CBAS and the Escherichia coli strain ATCC 25922 were used as positive and negative controls, respectively.
PFGE genotyping
PFGE analyses were performed using ApaI (New England Biolabs) restriction enzyme in order to determine the genetic relationship among A. baumannii isolates. The genetic relatedness was attributed when the macrorestriction DNA patterns differed by fewer than three bands [19].
Relative quantification of CarO porin
The carO transcription was evaluated by real time RT-PCR using Power-SYBR Green PCR Master Mix (Applied Biosystems). The single-copy housekeeping cpn60 gene from A. baumannii, coding for a 60-kDa chaperonin, was used as endogenous gene for normalization. The relative quantification (RQ) results were presented as ratios of normalized target gene transcription between the A. baumannii clinical isolates and the Type Strain ATCC 19606 (calibrator), which were obtained according to the following equation: RQ = 2-∆∆CT, where CT is the value corresponding to the crossing point of the amplification curve with the threshold; ∆CT = target CT or calibrator CT – endogenous CT; and ∆∆CT = target ∆CT – calibrator ∆CT. carO differential transcription of clinical strains relative to that of ATCC 19606 was considered significant when the ratios obtained between RQ values (RQ value of calibrator/RQ value of clinical strains) were ≥2.0, taking into account the standard deviation intervals.
CarO amino acid sequence alignment and in silico analyses
The deduced amino acid sequences from the carO obtained in this study and from the CarOa (ATCC 19606 strain), CarOb (A6 strain) and several undefined CarO sequences available in GenBank were aligned using ClustalW. In order to verify whether the amino acid changes observed in the undefined CarO isoforms relative to the canonical CarOa are deleterious, in silico analyses was performed with the software PROVEAN (http://provean.jcvi.org/), which predicts if an amino acid substitution or indel has any impact on the biological function of a protein.
References
Boucher HW, Talbot GH, Bradley JS, Edwards JE, Gilbert D, Rice LB, Scheld M, Spellberg B, Bartlett J: Bad bugs, No drugs: No ESKAPE! an update from the infectious diseases society of America. Clin Infect Dis. 2009, 48: 1-12. 10.1086/595011.
Kuo HY, Chang KC, Kuo JW, Yueha HW, Liouf ML: Imipenem: a potent inducer of multidrug resistance in Acinetobacter baumannii. Int J Antimicrob Agents. 2012, 39: 33-38. 10.1016/j.ijantimicag.2011.08.016.
Poirel L, Nordmann P: Carbapenem resistance in Acinetobacter baumannii: mechanisms and epidemiology. Clin Microbiol Infect. 2006, 12: 826-836. 10.1111/j.1469-0691.2006.01456.x.
Turton JF, Ward ME, Woodford N, Kaufmann ME, Pike R, Livermore DM, Pitt TL: The role of ISAba1 in expression of OXA carbapenemase genes in Acinetobacter baumannii. FEMS Microbiol Lett. 2006, 258: 72-77. 10.1111/j.1574-6968.2006.00195.x.
Werneck JS, Picão RC, Girardello R, Cayô R, Marguti V, Dalla-Costa L, Gales AC, Antonio CS, Neves PR, Medeiros M, Mamizuka EM, Elmor de Araújo MR, Lincopan N: Low prevalence of blaOXA-143 in private hospitals in Brazil. Antimicrob Agents Chemother. 2011, 55: 4494-4495. 10.1128/AAC.00295-11.
Mostachio AK, Levin AS, Rizek C, Rossi F, Zerbini J, Costa SF: High prevalence of OXA-143 and alteration of outer membrane proteins in carbapenem-resistant Acinetobacter spp. isolates in Brazil. Int J Antimicrob Agents. 2012, 39: 396-401. 10.1016/j.ijantimicag.2012.01.021.
Pagano M, Martins AF, Machado AB, Barin J, Barth AL: Carbapenem-susceptible Acinetobacter baumannii carrying the ISAba1 upstream blaOXA-51-like gene in Porto Alegre, southern Brazil. Epidemiol Infect. 2012, 19: 1-4.
Mussi MA, Relling VM, Limansky AS, Viale AM: CarO, an Acinetobacter baumannii outer membrane protein involved in carbapenem resistance, is essential for L-ornithine uptake. FEBS Lett. 2007, 581: 5573-5578. 10.1016/j.febslet.2007.10.063.
Catel-Ferreira M, Coadou G, Molle V, Mugnier P, Nordmann P, Siroy A, Jouenne T, Dé E: Structure-function relationships of CarO, the carbapenem resistance-associated outer membrane protein of Acinetobacter baumannii. J Antimicrob Chemother. 2011, 66: 2053-2056. 10.1093/jac/dkr267.
Fonseca EL, Fdos Freitas S, Scheidegger ÉM, Jacinto T, Vicente AC: Class 2 integrons in multidrug-resistant Acinetobacter baumannii circulating in different Brazilian geographic regions. Int J Antimicrob Agents. 2011, 38: 95-96. 10.1016/j.ijantimicag.2011.03.013.
Corvec S, Poirel L, Naas T, Drugeon H, Nordmann P: Genetics and expression of the carbapenem-hydrolyzing oxacillinase gene blaOXA-23 in Acinetobacter baumannii. Antimicrob Agents Chemother. 2007, 51: 1530-1533. 10.1128/AAC.01132-06.
Higgins PG, Dammhayn C, Hackel M, Seifert H: Global spread of carbapenem-resistant Acinetobacter baumannii. J Antimicrob Chemother. 2010, 65: 233-238. 10.1093/jac/dkp428.
Mugnier P, Poirel L, Naas T, Nordmann P: Worldwide dissemination of the blaOXA-23 carbapenemase gene of Acinetobacter baumannii. Emerg Infect Dis. 2010, 16: 35-40.
Zarrilli R, Pournaras S, Giannouli M, Tsakris A: Global evolution of multidrug-resistant Acinetobacter baumannii clonal lineages. Int J Antimicrob Agents. 2013, 41: 11-19. 10.1016/j.ijantimicag.2012.09.008.
Mussi MA, Limansky AS, Viale AM: Acquisition of resistance to carbapenems in multidrug-resistant clinical strains of Acinetobacter baumannii: natural insertional inactivation of a gene encoding a member of a novel family of beta-barrel outer membrane proteins. Antimicrob Agents Chemother. 2005, 49: 1432-1440. 10.1128/AAC.49.4.1432-1440.2005.
Vashist J, Tiwari V, Kapil A, Rajeswari MR: Quantitative profiling and identification of outer membrane proteins of beta-lactam resistant strain of Acinetobacter baumannii. J Proteome Res. 2010, 9: 1121-1128. 10.1021/pr9011188.
CLSI: Performance standards for antimicrobial susceptibility testing; twenty-first informational supplement. 2011, Wayne, PA: M100-S21. Clinical and Laboratory Standards Institute
Lee K, Kim CK, Yong D, Jeong SH, Yum JH, Seo YH, Docquier JD, Chong Y: Improved performance of the modified hodge test with MacConkey agar for screening carbapenemase-producing gram-negative bacilli. J Microbiol Methods. 2010, 83: 149-152. 10.1016/j.mimet.2010.08.010.
Tenover FC, Arbeit RD, Goering RV, Mickelsen PA, Murray BE, Persing DH, Swaminathan B: Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995, 33: 2233-2239.
Acknowledgements
This work was supported by FAPERJ (Fundação de Amparo à Pesquisa do Estado do Rio de Janeiro), CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) and Oswaldo Cruz Institute grants.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
EL conceived and designed the study, carried out the experimental assays, analyzed and interpreted the data and drafted the manuscript. EMDS and FSF performed the experimental assays. RC provided the isolates and performed the preliminary microbiological tests. ACPV conceived, designed and supervised the study, and edited the manuscript. All authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an open access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Fonseca, E.L., Scheidegger, E., Freitas, F.S. et al. Carbapenem-resistant Acinetobacter baumannii from Brazil: role of carO alleles expression and blaOXA-23 gene. BMC Microbiol 13, 245 (2013). https://doi.org/10.1186/1471-2180-13-245
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2180-13-245