Abstract
Background
Gynostemma pentaphyllum is an important perennial medicinal herb belonging to the family Cucurbitaceae. Aerial stem-to-rhizome transition before entering the winter is an adaptive regenerative strategy in G. pentaphyllum that enables it to survive during winter. However, the molecular regulation of aerial stem-to-rhizome transition is unknown in plants. Here, integrated transcriptome and miRNA analysis was conducted to investigate the regulatory network of stem-to-rhizome transition.
Results
Nine transcriptome libraries prepared from stem/rhizome samples collected at three stages of developmental stem-to-rhizome transition were sequenced and a total of 5428 differentially expressed genes (DEGs) were identified. DEGs associated with gravitropism, cell wall biosynthesis, photoperiod, hormone signaling, and carbohydrate metabolism were found to regulate stem-to-rhizome transition. Nine small RNA libraries were parallelly sequenced, and seven significantly differentially expressed miRNAs (DEMs) were identified, including four known and three novel miRNAs. The seven DEMs targeted 123 mRNAs, and six pairs of miRNA-target showed significantly opposite expression trends. The GpmiR166b-GpECH2 module involved in stem-to-rhizome transition probably promotes cell expansion by IBA-to-IAA conversion, and the GpmiR166e-GpSGT-like module probably protects IAA from degradation, thereby promoting rhizome formation. GpmiR156a was found to be involved in stem-to-rhizome transition by inhibiting the expression of GpSPL13A/GpSPL6, which are believed to negatively regulate vegetative phase transition. GpmiR156a and a novel miRNA Co.47071 co-repressed the expression of growth inhibitor GpRAV-like during stem-to-rhizome transition. These miRNAs and their targets were first reported to be involved in the formation of rhizomes. In this study, the expression patterns of DEGs, DEMs and their targets were further validated by quantitative real-time PCR, supporting the reliability of sequencing data.
Conclusions
Our study revealed a comprehensive molecular network regulating the transition of aerial stem to rhizome in G. pentaphyllum. These results broaden our understanding of developmental phase transitions in plants.
Similar content being viewed by others
Background
Gynostemma pentaphyllum (Thunb.) Makino, belonging to the genus Gynostemma in the family Cucurbitaceae, is a perennial herb widely distributed in Asian countries [1]. G. pentaphyllum contains important medicinal components, called gypenosides, which are reportedly effective in the treatment of various illnesses, such as inflammation, cardiovascular diseases, and cancer [2,3,4]. This herb is widely used as tea or functional food [5], and has thus received substantial attention in recent years.
G. pentaphyllum is a dioecious, herbaceous vine with a female-to-male ratio of 1:20, which is not conducive to seed production [6]. Moreover, its seeds contain germination inhibitors and exhibit deep dormancy at maturity, and thus, it propagates mainly vegetatively under natural conditions [7]. The aboveground part of the vine lives only 1 year and dies in winter under natural conditions. Interestingly, before entering the winter, the subapical regions of some aerial stems swell and then drill into the soil to form rhizomes that produce new plants in the next year [6]. This vegetative regeneration is an adaptation of G. pentaphyllum to the natural environment to maintain its population. Aerial stem-to-rhizome transition implies not only morphological changes, but also functional changes in processes ranging from transport and support to storage and reproduction. This developmental phase transition is an interesting research topic in the field of developmental biology.
Accumulating evidence shows that plant developmental phase transitions involve the regulation of a large numbers of genes [8,9,10,11]. For example, transcriptome analysis revealed that genes related to the photoperiod pathway, starch biosynthesis, and hormone signaling are involved in stolon-to-rhizome transition in lotus [9]. miRNAs have also been confirmed to be involved in plant developmental phase transitions [12, 13]. miRNAs are single-stranded small noncoding RNAs of 20–24 nt in length that repress the expression of target genes by transcript cleavage and/or translation inhibition [14]. The identification of miRNA targets is critical for functional investigation of miRNAs. For example, miR156 and miR172 targets SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) and APETALA2 (AP2) regulate juvenile-to-adult and adult-to-reproduction transitions, respectively, in Arabidopsis [8]. miR156 is also involved in the regulation of tuberization in potato, and miR156 abundance increases in stolons under tuber-inductive conditions [15]. The miR159-MYB33 module controls the transition from the vegetative to the reproductive phase, and enhanced miR159 expression delayed flowering time in Arabidopsis [16]. miR166 affects root development by targeting several homeodomain-leucine zipper (HD-ZIP) genes in Medicago truncatula [17], whereas the miR166-PHABULOSA module participates in the embryogenic transition of somatic cells in Arabidopsis [18]. More recently, novel miRNAs involved in potato tuber formation have been identified [13]. To date, little is known about whether and which miRNAs participate in aerial stem-to-rhizome transition in plants. Except for miRNAs and their targets, it is also unknown which other genes are involved in aerial stem-to-rhizome transition.
In this study, we conducted integrated transcriptome and miRNA analyses to investigate the molecular mechanism underlying aerial stem-to-rhizome transition in G. pentaphyllum. We expected our findings to broaden our understanding of developmental transitions in plants.
Results
Morphological and histological traits of aerial stem-to-rhizome transition in G. pentaphyllum
As shown in Fig. 1, aerial stem, aboveground moderately swelling stem, and underground newly formed rhizome were selected as representative stages of developmental aerial stem-to-rhizome transition in G. pentaphyllum and were named stage 1, stage 2, and stage 3, respectively. In the process of stem-to-rhizome transition, the subapical regions of aerial stems swelled and expanded away from the tip, and then grew down into the soil. As swelling intensified, the stem diameter increased by about 1, 3 and 5 mm at the three developmental stages, respectively. Correspondingly, the stem color changed gradually from green to pale green, and finally to white (Fig. 1a-c). Rhizome, as a modified subterranean stem, exhibited anatomical characteristics similar to those of aerial stem (Additional file 1: Figure S1). This result is consistent with a recent report on Oryza longistaminata [19]. Stems at transition stages 1, 2, and 3 were all composed of epidermis, cortex, vascular bundles arranged along the stem circumference, and pith from outside to inside (Additional file 1: Figure S1). It is noteworthy that there is a circle of perivascular fibers composed of several layers of cells outside the vascular bundles (Additional file 1: Figure S1). Histochemical observation revealed that only a small amount of starch grains accumulated in stage 1 and stage 2 stems, whereas more and larger starch grains were present in stems at stage 3 (Additional file 2: Figure S2). The starch grains mainly accumulated in the innermost layer of the cortex, termed the starch sheath, and the pith (Additional file 2: Figure S2). In stage 3, starch grains accumulated even in the phloem parenchyma cells of the vascular bundles (Additional file 2: Figure S2).
Transcriptome analysis of aerial stem-to-rhizome transition in G. pentaphyllum
RNA-Seq and de novo assembly
To explore the molecular basis of aerial stem-to-rhizome transition, RNA-Sequencing (RNA-Seq) was conducted to generate transcriptome profiles. Nine RNA libraries derived from the above-mentioned three developmental stages of aerial stem-to-rhizome transition were sequenced on an Illumina HiSeq X Ten platform. In total, 352,070,555 cleaned reads were generated (Table 1). De-novo assembly of the cleaned reads yielded 207,635 transcripts, which were further assembled into 100,119 unigenes with an N50 length of 1336 bp (Additional file 9: Table S1). E90N50 value was 2658 bp, which represents the N50 of 90% of the total normalized expressed transcripts (Additional file 3: Figure S3b). Bench-marking universal single-copy orthologs (BUSCO) analysis showed a completeness score of 66.4%, a fragmented score of 23.4 and 10.2% as missing BUSCOs (Additional file 10: Table S2). The length distribution of the unigenes is shown in Additional file 3: Figure S3a, and Fig. 2 shows the genes that are similarly and distinctly regulated among the three stages. In total, 46,808 genes were expressed in all three stages, whereas 8616, 8961, and 15,396 genes were uniquely expressed in stage 1, stage 2, and stage 3, respectively (Fig. 2). These stage-specific expressed genes that were primarily assigned to carbon metabolism, amino acid biosynthesis, and ribosomes at each developmental stage, indicating that they exhibited different temporal and spatial expression patterns during the aerial stem-to-rhizome transition of G. pentaphyllum. Because of the lack of a reference genome sequence, the cleaned reads were mapped onto the assembled transcriptome; 81.16% of cleaned reads were aligned (Table 1). Principal component analysis (PCA) revealed that three samples from the same stage were clustered together and nine samples from three stages were clearly assigned to three groups as stage 1, stage 2 and stage 3 (Additional file 3: Figure S3c).
Identification and functional classification of differentially expressed genes
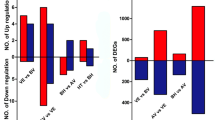
To identify differentially expressed genes (DEGs), pairwise comparisons were conducted among the three stages of G. pentaphyllum aerial stem-to-rhizome transition. In total, 5428 DEGs were filtered out based on FDR < 0.01 and |log2 fold change| ≥1 in each pairwise comparison (Additional file 11: Table S3); 1683 and 792 genes were significantly up- and downregulated, respectively, in the transition from stage 1 to stage 2; and 906 and 763 genes were significantly up- and downregulated, respectively, in that from stage 2 to stage 3 (Additional file 3: Figure S3c). In the transition from stage 1 to stage 3, 2552 and 2075 genes were significantly up- and downregulated, respectively (Additional file 3: Figure S3d).
Five thousand four hundred twenty-eight DEGs were annotated using blastx and the functions of these DEGs were investigated by Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses (Additional file 4: Figure S4 and Additional file 5: Figure S5). Stage 1-to-stage 2 DEGs were predominantly involved in hormone signal transduction, phenylpropanoid biosynthesis, carbon metabolism, ribosome, photosynthesis, and starch and sucrose metabolism (Additional file 5: Figure S5a). Among them, upregulated DEGs were mainly assigned to hormone signal transduction, phenylpropanoid biosynthesis, and starch and sucrose metabolism, whereas downregulated DEGs were mainly involved in photosynthesis, ribosome, and carbon metabolism (Additional file 5: Figure S5b-c). Similar findings were obtained for stage 2-to-stage 3 DEGs (Additional file 5: Figure S54e-f).
DEGs related to the aerial stem-to-rhizome transition
Aerial stem-to-rhizome transition involves a conversion from negative to positive gravitropism. Genes encoding indole-3-pyruvate monooxygenase (YUCCA), LAZY1, and actin-related protein 2/3 (ARP2/3) complex reportedly are involved in gravitropism [20]. Putative homologs of these genes were identified in G. pentaphyllum (Table 2). Among them, two GpYUCCA and GpLAZY1 were respectively well clustered with their homologs whose functions have been reported to be associated with gravitropism (Additional file 8: Figure S8). The expressions of these genes was significantly upregulated during the transition from stage 1 to stage 3 (Table 2).
Phenylpropanoid biosynthesis is involved in rhizome formation [21]. In this study, a large number of DEGs was assigned to the phenylpropanoid pathway, including genes encoding phenylalanine ammonia-lyase (PAL), trans-cinnamate 4-monooxygenase (C4H), caffeic acid 3-O-methyltransferase (COMT), ferulate-5-hydroxylase (F5H), cinnamoyl-CoA reductase (CCR), cinnamyl-alcohol dehydrogenase (CAD), and peroxidase (Px) (Fig. 3). Most of the putative genes encoding these enzymes were significantly upregulated during aerial stem-to-rhizome transition of G. pentaphyllum. Among them, some genes were upregulated at stage 2, whereas others were upregulated at stage 3 when compared with stage 1 (Fig. 3).
Rhizome formation is also controlled by distinct photoperiod-related genes [22]. Some genes encoding phytochrome A (PHYA), CONSTANS-like (COL) protein, cyclic dof factor (CDF), and flavin-binding kelch repeat F-box protein 1 (FKF1) in the photoperiod pathway were identified (Table 3). Among them, GpCOLs and GpCDF were respectively well clustered with their homologs whose functions have been reported to be involved in photoperiod (Additional file 8: Figure S8). Genes encoding PHYA, FKF1, and two out seven of genes encoding COL were significantly upregulated in stage 3 compared to stage 1, whereas genes encoding CDF and five out seven of genes encoding COL were significantly downregulated during aerial stem-to-rhizome transition.
Plant hormones play crucial roles in rhizome formation [10]. Seventy-four genes associated with the biosynthesis, metabolism, and signaling of plant hormones, including gibberellin acid (GA), abscisic acid (ABA), ethylene (ETH), cytokinin (CTK), auxin (IAA), brassinosteroid (BR), jasmonic acid (JA), and salicylic acid (SA), were identified (Fig. 4). It is noteworthy that 30 genes were assigned to the IAA signaling pathway, and their expression was generally significantly upregulated. Genes related to the ETH, CTK, and SA pathways were significantly upregulated in stage 3 compared to stage 1. Most genes involved in the biosynthesis, metabolism, and signaling of GA (3 out of 4), ABA (7 out of 8), IAA (21 out of 30), BR (2 out of 3), and JA (7 out of 8) were also significantly upregulated during aerial stem-to-rhizome transition, except for several downregulated genes, including GA20ox (Fig. 4).
Carbohydrate metabolism-related starch biosynthesis is strongly involved in the development and function of storage organs, including rhizomes, corms, tubers, and bulbs [23]. Several putative genes encoding sucrose synthase (SUS), granule-bound starch synthase (GBSS), cellulose synthase (CESA), and SNF1-related protein kinase regulatory subunit gamma-1 (KING1) were found to be significantly upregulated during aerial stem-to-rhizome transition of G. pentaphyllum (Table 4). These genes have been suggested to be closely related to carbohydrate metabolism [23, 24].
miRNAs and miRNA targets involved in aerial stem-to-rhizome transition in G. pentaphyllum
Sequencing of small RNAs and identification of miRNAs
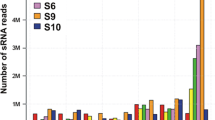
Nine small RNA libraries from three stages in developmental aerial stem-to-rhizome transition of G. pentaphyllum were generated and sequenced. In total, 281,846,013 cleaned reads were obtained (Table 5). Among them, 204,459,222 cleaned reads, accounting for 72.54% of the total cleaned reads, could be mapped to known small RNA databases (Table 5). The mapped reads were categorized into seven classes, including miRNA (4.91%), ribosomal RNA (rRNA, 64.57%), transfer RNA (tRNA, 2.78%), small nucleolar RNA (snoRNA, 0.10%), repeats (0.18%), and unannotated reads (27.46%) (Additional file 12: Table S4). In total, 90 known miRNAs were identified by mapping the cleaned reads to known plant miRNA databases (Additional file 13: Table S5). The remaining unmapped reads were used to predict novel miRNAs; 158 novel miRNAs were identified (Additional file 13: Table S5). These miRNAs were mainly 20–24 nt in length, and the most abundant miRNAs were 21 nt in length (Additional file 6: Figure S6).
Identification of differentially expressed miRNAs
To identify differentially expressed miRNAs (DEMs), pairwise comparisons were performed among the three transition stages based on the criteria of FDR < 0.01 and |log2 fold change| ≥1. Four known and three novel miRNAs were significantly differentially expressed during aerial stem-to-rhizome transition (Table 6). In the transition from stage 1 to stage 2, GpmiR156a, GpmiR159, and Co.47071 were significantly upregulated, whereas Co.25160 and Co.59333 were significantly downregulated; between stages 2 and 3, GpmiR156a and Co.47071 were significantly upregulated, whereas Co.25160 was significantly downregulated. During the transition from stage 1 to stage 3, GpmiR156a, GpmiR159, and Co.47071 were significantly upregulated, whereas GpmiR166b-5p, GpmiR166e-3p, Co.25160, and Co.59333 were significantly downregulated.
Identification and functional annotation of mRNA targets of differentially expressed miRNAs
In total, 123 putative targets for GpmiR166b-5p, GpmiR166e-3p, GpmiR156a, GpmiR159, Co.47071, and Co.59333 were identified, whereas Co.25160 had no predicted target genes (Additional file 14: Table S6). Six miRNA-target pairs exhibiting contrasting expression trends were identified (|log2 fold change| ≥1; FDR < 0.01) (Fig. 5). GpmiR166b-5p and GpmiR166e-3p were significantly downregulated during aerial stem-to-rhizome transition, and their target genes, which encode enoyl-CoA hydratase 2 (ECH2) and scopoletin glucosyltransferase (SGT)-like, respectively, were significantly upregulated (Fig. 5). In contrast, GpmiR156a and miRNA Co.47071 were significantly upregulated during the transition and their targets were significantly downregulated (Fig. 5). Among them, GpmiR156a targeted two SPL transcription factor genes (GpSPL6/GpSPL13A) and a gene encoding related to ABI3/VP1 (RAV)-like factor. Like GpmiR156a, miRNA Co.47071 also targeted the GpRAV-like gene (Fig. 5).
Validation of DEGs, DEMs, and their targets
To validate the DEGs, DEMs, and their targets identified by Illumina sequencing, 32 representative genes, five DEMs, and six miRNA-target pairs were investigated by quantitative real-time PCR (qRT-PCR). The qRT-PCR results were consistent with the sequencing data, supporting the reliability of sequencing data (Additional file 7: Figure S7).
Discussion
Transcriptomic analysis reveals the important roles of DEGs involved in G. pentaphyllum aerial stem-to-rhizome transition
RNA-Seq is a powerful and efficient means to discover putative functional genes involved in diverse biological processes, especially for plant species without a reference genome [10]. Using this tool, we found 5428 genes to be differentially expressed during stem-to-rhizome transition in G. pentaphyllum. Among them, DEGs were mostly related to gravitropism, phenylpropanoid biosynthesis, photoperiod, hormone synthesis and signal transduction, and carbohydrate metabolism.
Gravitropism is vital for shaping directional growth of plants in response to gravity [25]. Shoots grow upward (negative gravitropism), whereas roots grow downward (positive gravitropism) due to a gravitropic response, which results in differential growth between upper and lower sides of these organs [26]. Differential growth is thought to be controlled by polar auxin transport and asymmetric auxin distribution in different parts of graviresponding organs [27]. In this study, several homologs of known gravitropism-related genes, including GpYUCCA-a, GpYUCCA-b, GpLAZY1, and GpARP2/3, were significantly upregulated (Table 2). YUCCA genes encode flavin monooxygenases that catalyze a key step in the conversion of tryptophan into IAA [28]. In Arabidopsis, mutation of five YUCCA genes led to IAA deficiency and abnormal gravitropic responses of roots [29]. LAZY1 and ARP2/3 also play essential roles in shoot and root gravitropism by affecting polar auxin transport and asymmetric auxin distribution [20, 30, 31]. Thus, it is suggested that these gravitropism-related DEGs cooperatively control the gravitropic response during aerial stem-to-rhizome transition, probably by promoting auxin biosynthesis and altering auxin polar transport and distribution, thereby enabling the rhizome to acquire a positive gravitropism phenotype and to thus grow into the soil.
The phenylpropanoid pathway generates lignin precursors, which are transported into the cell walls for polymerization into lignin [32]. Lignin is mainly present in sclerenchymatous cells, such as vessel and fiber, whose lignification level is much higher than that in parenchyma cells, such as pith cells [33]. In this study, phenylpropanoid biosynthesis-related DEGs, including GpPAL-a–GpPAL-g, GpC4H-a–GpC4H-b, GpCOMT-a–GpCOMT-b, GpCCR-a–GpCCR-d, GpCAD-a–GpCAD-d, GpF5H-a–GpF5H-b, and GpPx1–GpPx31 genes, were significantly upregulated (Fig. 3). These genes are required for lignin synthesis. In transgenic tobacco, downregulation of PAL or C4H significantly reduced lignin content [34]. In two Vicia sativa varieties, upregulation of COMT, CCR, and CAD led to increased lignin deposition in the cell walls [35]. Overexpression of F5H increased lignin content in transgenic Arabidopsis, tobacco, and poplar [36]. Transgenic tobacco with 10-fold higher Px activity than the wild type exhibited lignin enrichment in the leaves [37]. Given that the enlargement of cells during aerial stem-to-rhizome transition of G. pentaphyllum (Additional file 1: Figure S1), we speculate that the phenylpropanoid biosynthesis-related DEGs are involved in cell-wall expansion to accommodate the enlargement of various cells, in particular vessel and fiber cells, by regulating lignin biosynthesis.
The photoperiod regulates tuber and rhizome formation [9, 38]. The photoperiod-related genes GpPHYA, GpCOL-a–GpCOL-g, GpCDF, and GpFKF1 were identified as DEGs during aerial stem-to-rhizome transition (Table 3). PHYA overexpression increased tuber production in short-day potato [39], which is in line with the upregulation of PHYA in this study. CO family members are involved in tuberization controlled by day length in potato [40]. In lotus, COL members control rhizome development [9]. Based on our findings, we speculate that the COL homologs detected in G. pentaphyllum regulate rhizome formation. CDF belongs to the large DOF transcription factor gene family [41]. In potato, CDF overexpression led to early tuber formation [22]. In our study, two CDF homologs were downregulated. We speculate that the upregulation of CDF expression activates a positive regulator of tuberization to promote tuber formation in potato, whereas the downregulation of GpCDF expression represses a negative regulator for rhizome formation to promote aerial stem-to-rhizome transition in G. pentaphyllum. FKF1, a clock-controlled protein, degrades CDF by ubiquitin-mediated regulation to control photoperiodic flowering in Arabidopsis [42]. The upregulation of GpFKF1 and downregulation of GpCDF observed in our study corroborates an interaction between them and suggests that GpFKF1 might regulate aerial stem-to-rhizome transition in G. pentaphyllum by degrading CDF.
In this study, most of the hormone-related genes were assigned to the IAA signaling pathway, and most of them were upregulated during aerial stem-to-rhizome transition (Fig. 4). In Arabidopsis, cell-wall acidification-triggered root cell expansion is preceded by increase in IAA signaling [43]. Given the enlargement of cells during stem-to-rhizome transition (Additional file 1: Figure S1), we suggest that IAA-related genes are involved in rhizome formation in G. pentaphyllum, probably by promoting cell expansion. ABA, CTK, JA, ETH, and BR reportedly also regulate the formation of plant storage organs [21, 44]. Most genes associated with these five hormones were upregulated in this study, indicating that they also regulate the rhizome formation. GA reportedly inhibits storage organ formation [45, 46]. In transgenic potato, overexpression of the GA-biosynthetic gene GA20ox1 delayed tuberization [46]. In this study, GpGA20ox expression was downregulated, whereas the expression of the GA-catabolic gene GA2ox was upregulated during stem-to-rhizome transition, suggesting that GA levels are reduced during the transition, thereby promoting the rhizome formation in G. pentaphyllum.
Carbohydrate metabolism plays an essential role in plant growth and development as its products are used not only as an energy source, but also for constructing structural cellular components [47]. Starch/sucrose biosynthesis is strongly correlated with the swelling of storage organs [23]. SUS and GBSS encode key enzymes in starch/sucrose synthesis [48, 49]. The upregulated SUS and GBSS was in parallel with rhizome enlargement in lotus [9]. In this study, GpSUS and GpGBSS were significantly upregulated (Table 4), in line with the increased starch accumulation in rhizome cells (Additional file 2: Figure S2). Thus, these two enzymes might promote rhizome formation by providing energy for cell expansion in G. pentaphyllum. Starch is also related to gravitropism. In plants, gravity is perceived by specific starch-containing cells located in the root columella and in the starch sheath of stem endodermis [50]. In this study, starch accumulation was observed in the starch sheath of rhizome (Additional file 2: Figure S2), suggesting that the GpSUS and GpGBSS might indirectly regulate the gravitropic response during stem-to-rhizome transition. SNF1 kinase is a heterotrimer composed of catalytic alpha and regulatory beta and gamma subunits [51] that regulates carbohydrate metabolism [23]. A gene encoding KING1 was significantly upregulated during stem-to-rhizome transition (Table 4), suggesting it is involved in rhizome formation in G. pentaphyllum.
miRNAs and their targets involved in the aerial stem-to-rhizome transition in G. pentaphyllum
GpmiR166b-GpECH2 and GpmiR166e-GpSGT-like modules regulate the aerial stem-to-rhizome transition
miR166 family members modulate various developmental processes by negatively mediating their targets [52]. miR166g and its HD-ZIP targets determine the fate of shoot apical meristem and lateral organ formation in Arabidopsis [53]. Overexpression of miR166a reduced lateral root by targeting several HD-ZIP genes in Medicago truncatula [17]. In Arabidopsis, miR166 is also involved in the embryogenic transition of somatic cells by regulating its target PHABULOSA and the LEC2-mediated auxin-related pathway [18]. However, it is unknown whether miR166 family members are involved in rhizome formation. In this study, GpmiR166b and GpmiR166e were significantly downregulated (Table 6 and Fig. 5), indicating that they are involved in aerial stem-to-rhizome transition in G. pentaphyllum. This is the first report on the potential regulatory functions of miR166 family members in rhizome formation. GpmiR166b was predicted to target GpECH2, which was significantly upregulated (Fig. 5). In Arabidopsis, ECH2 promotes cell enlargement during post-mitotic cell expansion in cotyledon development [54]. ECH2 is a peroxisomal enzyme that is essential for IBA-to-IAA conversion through β-oxidation of IBA [54, 55]. The long-standing acid growth theory postulates that IAA triggers cell-wall acidification, thus activating cell wall-loosening enzymes, which enable cell expansion in shoots [56]. A reduction in the IAA level or signaling abolished both cell wall acidification and cellular expansion in Arabidopsis roots, supporting the acid growth theory [43]. In this study, GpECH2 and IAA signaling-related genes were significantly upregulated during stem-to-rhizome transition (Figs. 4 and 5), suggesting that GpmiR166b and its target GpECH2 promote cell expansion probably by the IBA-to-IAA conversion in the rhizome formation of G. pentaphyllum. IBA β-oxidation also leads to the production of acetyl-CoA [55], which can be converted to glucose via the glyoxylate cycle or gluconeogenesis [57] and further polymerized to cellulose, a major structural cell-wall component [24]. The cellulose synthesis-related genes GpSUS and GpCESA were significantly upregulated in this study (Table 4). Thus, it is possible that GpmiR166b-GpECH2 is involved in cell wall remodeling for cell expansion via conversion of IBA into acetyl-CoA during stem-to-rhizome transition. GpmiR166e was predicted to target an GpSGT-like gene, and they exhibited significantly opposite expression trends (Fig. 5). SGT catalyzes the glucosylation of scopoletin to scopolin [58]. In tobacco, scopolin protects IAA from degradation during seedling development [59]. IAA plays fundamental roles in many aspects of plant growth and development [43], which is supported by the increased expression of GpYUCCA and IAA signaling-related genes (Table 2 and Fig. 4). Therefore, the GpmiR166e-GpSGT-like module might protect IAA from inactivation and thereby promote aerial stem-to-rhizome transition in G. pentaphyllum.
GpmiR156a-GpSPL6/SPL13A modules regulate the aerial stem-to-rhizome transition
miR156 family members are master regulators of various plant developmental traits [15]. Overexpression of miR156/miR156a prolonged the juvenile phase and delayed flowering in Arabidopsis, rice, maize, tomato, and switchgrass [8, 60,61,62,63]. In potato, miR156 overexpression induced the production of aerial tubers and regulated the tuberization [64]. In this study, GpmiR156a was significantly upregulated in the two transition phases (Table 6 and Fig. 5), suggesting that it is involved in stem-to-rhizome transition in G. pentaphyllum. The miR156a upstream sequence contains several light-regulated motifs, indicating a putative light-mediated regulation of this miRNA [15]. In photoperiod-responsive potato species, miR156a accumulation induced tuberization under short-day, but not long-day condition [15]. In late autumn, the photoperiod gradually shortens, implying that the light-mediated regulation of GpmiR156a is probably involved in aerial stem-to-rhizome transition in G. pentaphyllum. Accumulating evidence indicates that miR156 family members can target numerous GpSPL genes [65]. SPL genes encode plant-specific transcription factors that contain a conserved SBP domain, through which they can recognize and bind specifically to the promoters of target genes, thus affecting plant growth and development [66, 67]. In this study, GpmiR156a putatively targeted GpSPL6 and GpSPL13A, whose expression was significantly downregulated in parallel with the upregulation of GpmiR156a during stem-to-rhizome transition in G. pentaphyllum (Fig. 5). Arabidopsis has two SPL13 copies, SPL13A and SPL13B [68]. SPL13 plays a crucial role in vegetative and reproductive plant development. In Arabidopsis, expression of SPL13 with a mutated miR156 site delayed leaf production, whereas a loss-of-function mutant of SPL13 had increased juvenile and rosette leaf numbers [68, 69]. In Medicago sativa, SPL13 overexpression induced severe growth retardation, whereas SPL13 silencing increased branching and delayed flowering [70]. These finding indicate that SPL13 represses vegetative phase transition and promoted reproductive phase transition. SPL6 has a conserved DNA-binding domain similar to that of SPL13 [68] and also functions in developmental phase transition [68, 71]. We suggest that GpmiR156a promotes aerial stem-to-rhizome transition in G. pentaphyllum by repressing the expression of GpSPL13A and GpSPL6, which are negative regulators of vegetative phase transition.
GpmiR156a and a novel miRNA co.47071 regulate GpRAV-like during aerial stem-to-rhizome transition
In addition to GpSPL6 and GpSPL13A, GpmiR156a also targeted GpRAV-like gene, together with a novel miRNA Co.47071 (Fig. 5). All members of the RAV subfamily contain both AP2/ERF and B3 DNA-binding domains and belong to the AP2/ERF family. RAV genes encode transcriptional regulators with various functions in plant developmental and physiological processes [72]. In tobacco, overexpression of a Glycine max RAV gene retarded plant growth and reduced root elongation [73]. In Arabidopsis and soybean, GmRAV1 overexpression induced dwarfism, whereas loss-of-function mutant plants had an opposite phenotype [74]. GmRAV1 promotes root and shoot regeneration by enhancing the expression of cyclins and cyclin-dependent protein kinases to promote cell division [75]. These findings indicate that RAV inhibits plant growth probably by inhibiting cell expansion rather than cell division. Our findings suggest that the downregulation of GpRAV-like through co-repression by GpmiR156a and miRNA Co.47071 promotes stem-to-rhizome transition in G. pentaphyllum, probably by promoting cell expansion.
Conclusion
Our integrated transcriptome and miRNA analysis revealed a comprehensive molecular network regulating the transition of aerial stem to rhizome in G. pentaphyllum. In total, 5428 DEGs were identified, and DEGs associated with gravitropism, cell wall biosynthesis, photoperiod pathway, hormone signaling, and carbohydrate metabolism might largely contribute to aerial stem-to-rhizome transition in this species. Seven DEMs, including four known and three novel miRNAs, were identified. For six DEMs, we were able to predicts targets, which displayed significantly opposite expression trends. The regulatory modules GpmiR166b-GpECH2, GpmiR166e-GpSGT-like, GpmiR156a-GpSPL13A/GpSPL6, and GpmiR156a/Co.47071-GpRAV-like likely play important roles in aerial stem-to-rhizome transition. The qRT-PCR results supported the reliability of sequencing data. These results will help elucidate the molecular mechanisms underlying aerial stem-to-rhizome transition in G. pentaphyllum and broaden our understanding of developmental phase transitions in plants.
Methods
Plant materials
G. pentaphyllum plants grew in the field under normal farming conditions in Jishou, Hunan province, China. As shown in Fig. 1, aboveground moderately swelling stem, underground newly formed rhizome, and aerial stem as a control were collected from G. pentaphyllum plants when subapical regions of some aerial stems swelled and then grew down into the soil to form rhizomes in late autumn 2018. Aerial stem, aboveground moderately swelling stem, and underground newly formed rhizome were named stage 1, stage 2, and stage 3 of aerial stem-to-rhizome transition, respectively. Part of the samples were immediately frozen in liquid nitrogen and stored at − 80 °C for transcriptome and small RNA-Seq analysis, and the remaining samples were used to investigate histological traits.
Histological analysis
To investigate histological traits of the three stages of G. pentaphyllum stem-to-rhizome transition, the samples were cut into small pieces of approximately 0.5 cm in length and fixed in formalin:acetic acid:70% ethanol solution (5:5:90, v/v/v). The fixed samples were dehydrated in a graded ethanol series and then embedded in paraffin. Sections of 8 μm thickness were cut using a microtome (Leica RM2245; Leica, Nussloch, Germany). Sections were stained with safranin O-fast green and periodic acid-Schiff reagent, respectively, and then observed under a light microscope (Leica DM6000B).
RNA extraction
Total RNA was extracted from the samples using the easy-spin Plant RNA Kit (Aidlab Biotech, Beijing, China) following the manufacturer’s instructions. Only RNA samples having an 260/280 ratio of 1.8–2.1, an 260/230 ratio of 2.0–2.5, and an RNA integrity number of 8.0 or higher were used for transcriptome and small RNA-Seq. For each transition stage, three biological replicates were prepared.
Transcriptome sequencing and analysis
cDNA libraries were prepared from 1 μg RNA per sample using a NEBNext® Ultra™ RNA Library Prep Kit for Illumina® (NEB, USA) according to the manufacturer’s recommendations. Nine transcriptome libraries from the above-mentioned three developmental stages were sequenced on an Illumina HiSeq X Ten platform (Biomarker technologies, Beijing, China), generating 150-bp paired-end reads. To obtain high-quality cleaned reads, reads containing adapter and poly-N as well as low-quality reads were removed. The cleaned reads were de-novo assembled into transcripts, which were assembled into unigenes, using Trinity v2.5.1 (run parameters: ‘--seqType fq --bflyHeapSpaceMax 20G --min_contig_length 200 --bflyGCThreads 5 --no_run_butterfly --no_run_quantifygraph) [76]. The completeness of transcriptome assembly was assessed by using BUSCO v.3.0.2 (run parameters: -m tran -c 4 -f [77] . The read count for each gene was obtained by mapping the cleaned reads to the assembled transcriptome using RSEM v1.2.19 with default parameters [78].
For functional annotation, assembled unigenes were queried against public databases including NCBI non-redundant protein database (NR, ftp://ftp.ncbi.nih.gov/blast/db/) [79], Swiss-Prot (http://www.uniprot.org/) [80], Gene Ontology (GO, http://www.geneontology.org/) [81], Clusters of Orthologous Groups (COG, http://www.ncbi.nlm.nih.gov/COG/) [82], euKaryotic Orthologous Groups (KOG, http://www.ncbi.nlm.nih.gov/KOG/) [83], orthologous groups of genes (EggNOG, http://eggnogdb.embl.de/) [84] and Kyoto Encyclopedia of Genes and Genomes (KEGG, http://www.genome.jp/kegg/) [85] using BLAST v2.2.31 with an E value cutoff of 1e-5 [86], and Pfam (http://pfam.xfam.org/) database [87] using HMMER v3.1b2 with an E-value cutoff of 1e-10 [88].
Gene expression levels in each sample were normalized as transcripts per million (TPM): TPM = readcount× 1,000,000/ Mapped Reads [89]. Analysis of DEGs was performed using the Bioconductor [90] DESeq2 package v1.6.3 [91] in R v3.1.1 [92] with default parameters, based on a model following a negative binomial distribution [91]. DEGs between three developmental stages were identified based on criteria of FDR < 0.01 and |log2 fold change| ≥1. For functional enrichment analysis of the DEGs, GO and KEGG analyses were carried out.
Small RNA-Seq and data analysis
Nine small RNA libraries were parallelly prepared from isolated total RNA using a NEBNext® Ultra™ small RNA Sample Library Prep Kit for Illumina® (NEB) according to the manufacturer’s protocol. The libraries were sequenced on an Illumina HiSeq X Ten platform (Biomarker technologies, Beijing, China), generating 50-bp paired-end reads. Low-quality reads, contaminating reads with adapters and poly-A tails, and reads without the insert tag were removed. Then, sequences shorter than 18 bp and longer than 35 bp were removed. Finally, rRNAs, tRNAs, snRNAs, snoRNAs, and other noncoding RNAs and repeats were removed by aligning to Rfam (http://rfam.xfam.org/) [93], Silva (http://www.arb-silva.de/) [94], GtRNAdb (http://lowelab.ucsc.edu/GtRNAdb/) [95] and Repbase (http://www.girinst.org/repbase/) [96] databases. Conserved miRNAs were identified by comparing the cleaned small RNA reads with known miRNAs available in miRBase 21 (http://www.mirbase.org/). The alignment was done using Bowtie alignment tool v1.0.0 (run parameters: -v 0) with no mismatch [97]. The unannotated reads were used for prediction of novel miRNAs using miRDeep2 v2.0.5 (run parameters: -g − 1 -l 250 -b 0) [98]. The prediction is based on the biological characteristics of miRNA precursors which have a landmark hairpin-stem-loop structure. Expression levels of miRNAs in each sample were normalized as transcripts per million. DEMs between the transition stages were identified based on criteria of FDR < 0.01 and |log2 fold change| ≥1 by using the Bioconductor [90] DESeq2 package v1.6.3 [91] in R v3.1.1 [92] with default parameters.
miRNA target identification and functional annotation
miRNA-target pairs were predicted using the TargetFinder software v1.6 (run parameters: -c 3) [99]. To predict potential functions of these targets, they were annotated in the NR [79], Swiss-Prot [80], GO [81], COG [82], KEGG [85], KOG [83] and Pfam [87] databases.
Quantitative real-time PCR analysis
To quantify and validate the expression levels of DEGs, DEMs, and their targets, qRT-PCR was used. For the DEMs, stem-loop qRT-PCR was performed as described by Varkonyi-Gasic et al. [100], with 18S rRNA as a reference gene. For the DEGs and DEM targets, qRT-PCR was conducted as previously described by Guo et al. [101], with actin as a reference gene. Expression levels were expressed relative to the corresponding levels in stage 1 and were calculated by the 2−ΔΔCT method [102]. The significance was tested by Duncan’s multiple range test at the 5% level. Each sample included in the analysis was based on three biological replicates. All qRT-PCR primers used are listed in Additional file 15: Table S7.
Availability of data and materials
The Illumina sequencing reads have been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive under the accession number SRP197408, which is associated with BioProject number PRJNA541825 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA541825). The Transcriptome Shotgun Assembly project has been deposited at GenBank under the accession GHVI00000000, and the version described in this paper is the first version (GHVI01000000).
Abbreviations
- ABA:
-
Abscisic acid
- AP2:
-
APETALA2
- ARP 2/3 complex:
-
Actin-related protein 2/3 complex
- BR:
-
Brassinosteroid
- BUSCO:
-
Bench-marking universal single-copy orthologs
- C4H:
-
Trans-cinnamate 4-monooxygenase
- CAD:
-
Cinnamyl-alcohol dehydrogenase
- CCR:
-
Cinnamoyl-CoA reductase
- CDF:
-
Cyclic dof factor
- CESA:
-
Cellulose synthase
- COL:
-
CONSTANS-like
- COMT:
-
Caffeic acid 3-O-methyltransferase
- CTK:
-
Cytokinin
- DEGs:
-
Differentially expressed genes
- DEMs:
-
Differentially expressed miRNAs
- ECH2:
-
Enoyl-CoA hydratase 2
- ETH:
-
Ethylene
- F5H:
-
Ferulate-5-hydroxylase
- FKF1:
-
Flavin-binding kelch repeat F-box protein 1
- GA:
-
Gibberellin acid
- GBSS:
-
Granule-bound starch synthase
- GO:
-
Gene Ontology
- HD-ZIP:
-
Homeodomain-leucine zipper
- IAA:
-
Auxin
- JA:
-
Jasmonic acid
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- KING1:
-
SNF1-related protein kinase regulatory subunit gamma-1
- PAL:
-
Phenylalanine ammonia-lyase
- PCA:
-
Principal component analysis
- PHYA:
-
Phytochrome A
- Px:
-
peroxidase
- qRT-PCR:
-
Quantitative real-time PCR
- RAV:
-
Related to ABI3/VP1
- RNA-Seq:
-
RNA-Sequencing
- rRNA:
-
Ribosomal RNA
- SA:
-
Salicylic acid
- SGT:
-
Scopoletin glucosyltransferase
- snoRNA:
-
Small nucleolar RNA
- SPL:
-
SQUAMOSA PROMOTER BINDING PROTEIN-LIKE
- SUS:
-
Sucrose synthase
- tRNA:
-
Transfer RNA
- YUCCA:
-
Indole-3-pyruvate monooxygenase
References
Subramaniyam S, Mathiyalagan R, Gyo IJ, Bum-Soo L, Sungyoung L, Chun YD. Transcriptome profiling and insilico analysis of Gynostemma pentaphyllum using a next generation sequencer. Plant Cell Rep. 2011;30(11):2075–83.
Circosta C, De-Pasquale R, Occhiuto F. Cardiovascular effects of the aqueous extract of Gynostemma pentaphyllum makino. Phytomedicine. 2005;12(9):638–43.
Xie ZH, Huang HQ, Zhao Y, Shi HM, Wang SK, Wang TTY, Chen P, Yu LL. Chemical composition and anti-proliferative and anti-inflammatory effects of the leaf and whole-plant samples of diploid and tetraploid Gynostemma pentaphyllum (Thunb.) Makino. Food Chem. 2012;132(1):125–33.
Zhang XS, Zhao C, Tang WZ, Wu XJ, Zhao YQ. Gypensapogenin H, a novel dammarane-type triterpene induces cell cycle arrest and apoptosis on prostate cancer cells. Steroids. 2015;104:276–83.
Shi L, Tan DH, Liu YE, Hou MX, Zhao YQ. Two new dammarane-type Triterpenoid saponins from Gynostemma pentaphyllum. Helv Chim Acta. 2014;97(10):1333–9.
Peng XL, Peng XJ, Zhang XZ, Liu SB. Propagation characteristics and seedling breeding techniques of Gynostemma pentaphyllum. Hunan Agr Sci. 2011;17:12–4.
Pan CL, Deng ZJ, Huang YF, Huang XY, Zhang ZJ, Miao JH, Yu LY. Study on seed dormancy mechanism and breaking technique of Gynostemma pentaphyllum (Thunb.) Makino. Acta Bot Boreal –Occident Sin. 2013;33:1658–64.
Huijser P, Schmid M. The control of developmental phase transitions in plants. Development. 2011;138(19):4117–29.
Yang M, Zhu L, Pan C, Xu L, Yang P. Transcriptomic analysis of the regulation of rhizome formation in temperate and tropical lotus (Nelumbo nucifera). Sci Rep. 2015;5:13059.
Hu RB, Yu CJ, Wang XY, Jia CL, Pei SQ, He K, He G, Kong YZ, Zhou GK. De novo transcriptome analysis of Miscanthus lutarioriparius identifies candidate genes in rhizome development. Front Plant Sci. 2017;8:492.
Xu Y, Zhang L, Wu G. Epigenetic regulation of juvenile-to-adult transition in plants. Front Plant Sci. 2018;9:1048.
Poethig RS. Vegetative phase change and shoot maturation in plants. Curr Top Dev Biol. 2013;105:125–52.
Kondhare KR, Malankar NN, Devani RS, Banerjee AK. Genome-wide transcriptome analysis reveals small RNA profiles involved in early stages of stolon-to-tuber transitions in potato under photoperiodic conditions. BMC Plant Biol. 2018;18:284.
Voinnet O. Origin, biogenesis, and activity of plant microRNAs. Cell. 2009;136:669–87.
Bhogale S, Mahajan AS, Natarajan B, Rajabhoj M, Thulasiram HV, Banerjee AK. MicroRNA156: a potential graft-transmissible microRNA that modulates plant architecture and tuberization in Solanum tuberosum ssp. andigena. Plant Physiol. 2014;164(2):1011–27.
Achard P, Herr A, Baulocmbe DC, Harberd NP. Modulation of floral development by a gibberellin-regulated microRNA. Development. 2004;131(14):3357–65.
Boualem A, Laporte P, Jovanovic M, Laffont C, Plet J, Combier JP, Niebel A, Crespi M, Frugier F. MicroRNA166 controls root and nodule development in Medicago truncatula. Plant J. 2008;54(5):876–87.
Wójcik AM, Nodine MD, Gaj MD. miR160 and miR166/165 contribute to the LEC2-mediated auxin response involved in the somatic embryogenesis induction in Arabidopsis. Front Plant Sci. 2017;8:2024.
Bessho-Uehara K, Nugroho JE, Kondo H, Angeles-Shim RB, Ashikari M. Sucrose affects the developmental transition of rhizomes in Oryza longistaminata. J Plant Res. 2018;131(4):693–707.
Staiger CJ, Blanchoin L. Actin dynamics: old friends with new stories. Curr Opin Plant Biol. 2006;9(6):554–62.
Huang QQ, Huang X, Deng J, Liu HG, Liu YW, Yu K, Huang BS. Differential gene expression between leaf and rhizome in Atractylodes lancea: a comparative transcriptome analysis. Front Plant Sci. 2016;7:348.
Kloosterman B, Abelenda JA, Gomez MDC, Oortwijn M, de Boer JM, Kowitwanich K, Horvath BM, van Eck HJ, Smaczniak C, Prat S, et al. Naturally occurring allele diversity allows potato cultivation in northern latitudes. Nature. 2013;495(7440):246–50.
Cheng LB, Li SY, Yin JJ, Li LJ, Chen XH. Genome-wide analysis of differentially expressed genes relevant to rhizome formation in lotus root (Nelumbo nucifera gaertn). PLoS One. 2013;8(6):e67116.
Li SD, Bashline L, Lei L, Gu Y. Cellulose synthesis and its regulation. Arabidopsis Book. 2014;12:e0169.
Zou JJ, Zheng ZY, Xue S, Li HH, Wang YR, Le J. The role of Arabidopsis actin-related protein 3 in amyloplast sedimentation and polar auxin transport in root gravitropism. J Exp Bot. 2016;67(18):5325–37.
Went FW. Reflections and speculations. Annu Rev Plant Physiol. 1974;25:1–27.
Morita MT, Tasaka M. Gravity sensing and signaling. Curr Opin Plant Biol. 2004;7(6):712–8.
Zhao Y, Christensen SK, Fankhauser C, Cashman JR, Cohen JD, Weigel D, Chory J. A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science. 2001;291(5502):306–9.
Chen QG, Dai XH, De-paoli H, Cheng YF, Takebayashi Y, Kasahara H, Kamiya Y, Zhao YD. Auxin overproduction in shoots cannot rescue auxin deficiencies in Arabidopsis roots. Plant Cell Physiol. 2014;55(6):1072–9.
Dong ZB, Jiang C, Chen XY, Zhang T, Ding L, Song WB, Luo HB, Lai JS, Chen HB, Liu RY, et al. Maize LAZY1 mediates shoot gravitropism and inflorescence development through regulating auxin transport, auxin signaling, and light response. Plant Physiol. 2013;163(3):1306–22.
Taniguchi M, Furutani M, Nishimura T, Nakamura M, Fushita T, Iijima K, Baba K, Tanaka H, Toyota M, Tasaka M, et al. The Arabidopsis lazy1 family plays a key role in gravity signaling within statocytes and in branch angle control of roots and shoots. Plant Cell. 2017;29(8):1984–99.
Bonawitz ND, Chapple C. The genetics of lignin biosynthesis: connecting genotype to phenotype. Annu Rev Genet. 2010;44(1):337–63.
Siqueira G, Milagres AM, Carvalho W, Koch G, Ferraz A. Topochemical distribution of lignin and hydroxycinnamic acids in sugar-cane cell walls and its correlation with the enzymatic hydrolysis of polysaccharides. Biotechnol Biofuels. 2011;4:7.
Sewalt V, Ni WT, Blount JW, Jung HG, Masoud SA, Howles PA, Lamb C, Dixon RA. Reduced lignin content and altered lignin composition in transgenic tobacco down-regulated in expression of L-phenylalanine ammonia-lyase or cinnamate 4-hydroxylase. Plant Physiol. 1997;115(1):41–50.
Rui HY, Zhang XX, Shinwari KI, Zheng LQ, Shen ZG. Comparative transcriptomic analysis of two Vicia sativa L. varieties with contrasting responses to cadmium stress reveals the important role of metal transporters in cadmium tolerance. Plant Soil. 2018;423(1–2):241–55.
Franke R, McMichael CM, Meyer K, Shirley AM, Cusumano JC, Chapple C. Modified lignin in tobacco and poplar plants overexpressing the Arabidopsis gene encoding ferulate 5-hydroxylase. Plant J. 2000;22(3):223–34.
Lagrimini LM, Bradford S, Rothstein S. Peroxidase induced wilting in transgenic tobacco plants. Plant Cell. 1990;2(1):7–18.
Abelenda JA, Navarro C, Prat S. From the model to the crop: genes controlling tuber formation in potato. Curr Opin Biotechnol. 2011;22(2):287–92.
Yanovsky MJ, Izaguirre M, Wagmaister JA, Gatz C, Jackson SD, Thomas B, Casal JJ. Phytochrome a resets the circadian clock and delays tuber formation under long days in potato. Plant J. 2000;23(2):223–32.
Rodríguez-Falcón M, Bou J, Prat S. Seasonal control of tuberization in potato: conserved elements with the flowering response. Annu Rev Plant Biol. 2006;57:151–80.
Imaizumi T, Schultz TF, Harmon FG, Ho LA, Kay SA. FKF1 F-box protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science. 2005;309(5732):293–7.
Song YH, Smith RW, To BJ, Millar AJ, Imaizumi T. FKF1 conveys timing information for CONSTANS stabilization in photoperiodic flowering. Science. 2012;336(6084):1045–9.
Barbez E, Dünser K, Gaidora A, Lendl T, Busch W. Auxin steers root cell expansion via apoplastic pH regulation in Arabidopsis thaliana. P Natl Acad Sci USA. 2017;114(24):E4884–93.
Peres LE, Carvalho RF, Zsögön A, Bermúdez-Zambrano OD, Robles WGR, Tavares S. Grafting of tomato mutants onto potato rootstocks: an approach to study leaf-derived signaling on tuberization. Plant Sci. 2005;169(4):680–8.
Xu X, van Lammeren AAM, Vermeer E, Vreugdenhil D. The role of gibberellin, abscisic acid, and sucrose in the regulation of potato tuber formation in vitro. Plant Physiol. 1998;117(2):575–84.
Carrera E, Bou J, García-Martínez JL, Prat S. Changes in GA 20-oxidase gene expression strongly affect stem length, tuber induction and tuber yield of potato plants. Plant J. 2000;22(3):247–56.
Paparelli E, Gonzali S, Parlanti S, Novi G, Giorgi FM, Licausi F, Kosmacz M, Feil R, Lunn JE, Brust H, et al. Misexpression of a chloroplast aspartyl protease leads to severe growth defects and alters carbohydrate metabolism in Arabidopsis. Plant Physiol. 2012;160(3):1237–50.
Visser RGF, Somhorst I, Kuipers GJ, Ruys NJ, Feenstra WJ, Jacobsen E. Inhibition of the expression of the gene for granule-bound starch synthase in potato by antisense constructs. Mol Gen Genet. 1991;225(2):289–96.
Wang F, Sanz A, Brenner ML, Smith A. Sucrose synthase, starch accumulation, and tomato fruit sink strength. Plant Physiol. 1993;101(1):321–7.
Morita MT. Directional gravity sensing in gravitropism. Annu Rev Plant Biol. 2010;61:705–20.
Mangat S, Chandrashekarappa D, Mccartney RR, Elbing K, Schmidt MC. Differential roles of the glycogen-binding domains of β subunits in regulation of the Snf1 kinase. Eukaryot Cell. 2010;9(1):173–83.
Li XY, Xie X, Li J, Cui YH, Hou YM, Zhai LL, Wang X, Fu YL, Liu RR, Bian SM. Conservation and diversification of the miR166 family in soybean and potential roles of newly identified miR166s. BMC Plant Biol. 2017;17:32.
Williams L, Grigg SP, Xie M, Christensen S, Fletcher JC. Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHD-ZIP target genes. Development. 2005;132(16):3657–68.
Katano M, Takahashi K, Hirano T, Kazama Y, Abe T, Tsukaya H, Ferjani A. Suppressor screen and phenotype analyses revealed an emerging role of the monofunctional peroxisomal enoyl-CoA hydratase 2 in compensated cell enlargement. Front Plant Sci. 2016;7:132.
Strader LC, Wheeler DL, Christensen SE, Berens JC, Cohen JD, Rampey RA, Bartel B. Multiple facets of Arabidopsis seedling development require indole-3-butyric acid-derived auxin. Plant Cell. 2011;23(3):984–99.
Rayle DL, Cleland R. Enhancement of wall loosening and elongation by acid solutions. Plant Physiol. 1970;46(2):250–3.
Gerhardt B. Fatty acid degradation in plants. Prog Lipid Res. 1992;31(4):417–46.
Hino F, Okazaki M, Miura Y. Effect of 2,4-dichlorophenoxyacetic acid on glucosylation of scopoletin to scopolin in tobacco tissue culture. Plant Physiol. 1982;69(4):810–3.
Gális I, Simek P, Van-Onckelen HA, Kakiuchi Y, Wabiko H. Resistance of transgenic tobacco seedlings expressing the Agrobacterium tumefaciens C58-6b gene, to growth-inhibitory levels of cytokinin is associated with elevated IAA levels and activation of phenylpropanoid metabolism. Plant Cell Physiol. 2002;43(8):939–50.
Xie K, Wu C, Xiong L. Genomic organization, differential expression, and interaction of SQUAMOSA promoter-binding-like transcription factors and microRNA156 in rice. Plant Physiol. 2006;142(1):280–93.
Chuck G, Cigan AM, Saeteurn K, Hake S. The heterochronic maize mutant Corngrass1 results from overexpression of a tandem microRNA. Nat Genet. 2007;39(4):544–9.
Zhang X, Zou Z, Zhang J, Zhang Y, Han Q, Hu T, Xu X, Liu H, Li H, Ye Z. Over-expression of sly-miR156a in tomato results in multiple vegetative and reproductive trait alterations and partial phenocopy of the sft mutant. FEBS Lett. 2011;585(2):435–9.
Fu CX, Sunkar R, Zhou CE, Shen H, Zhang JY, Matts J, Wolf J, Mann DGJ, Stewart CN, Tang YH, et al. Overexpression of miR156 in switchgrass (Panicum virgatum L.) results in various morphological alterations and leads to improved biomass production. Plant Biotechnol J. 2012;10(4):443–52.
Eviatar-Ribak T, Shalit-Kaneh A, Chappell-Maor L, Amsellem Z, Eshed Y, Lifschitz E. A cytokinin-activating enzyme promotes tuber formation in tomato. Curr Biol. 2013;23(12):1057–64.
Aung B, Gruber MY, Amyot L, Omari K, Bertrand A, Hannoufa A. MicroRNA156 as a promising tool for alfalfa improvement. Plant Biotechnol J. 2015;13(6):779–90.
Yamasaki K, Kigawa T, Inoue M, Tateno M, Yamasaki T, Yabuki T, Aoki M, Seki E, Matsuda T, Nunokawa E, et al. A novel zinc-binding motif revealed by solution structures of DNA-binding domains of Arabidopsis SBP-family transcription factors. J Mol Biol. 2004;337(1):49–63.
Chen XB, Zhang ZL, Liu DM, Zhang K, Li AL, Mao L. SQUAMOSA promoter-binding protein-like transcription factors: star players for plant growth and development. J Integr Plant Biol. 2010;52(11):946–51.
Xu M, Hu T, Zhao J, Park MY, Earley KW, Wu G, Yang L, Poethig RS. Developmental functions of mir156-regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) genes in Arabidopsis thaliana. PLoS Genet. 2016;12:e1006263.
Martin RC, Asahina M, Liu PP, Kristof JR, Coppersmith JL, Pluskota WE, Bassel GW, Goloviznina NA, Nguyen TT, Martinez-Andujar C, et al. The regulation of post-germinative transition from the cotyledon- to vegetative-leaf stages by microRNA-targeted SQUAMOSA PROMOTER-BINDING PROTEIN LIKE13 in Arabidopsis. Seed Sci Res. 2010;20(2):89–96.
Gao RM, Gruber MY, Amyot L, Hannoufa A. SPL13 regulates shoot branching and flowering time in Medicago sativa. Plant Mol Biol. 2018;96(1–2):119–33.
Jung I, Kang H, Kim JU, Chang H, Kim S, Jung W. The mRNA and miRNA transcriptomic landscape of Panax ginseng under the high ambient temperature. BMC Syst Biol. 2018;12(2):27.
Zhuang J, Sun CC, Zhou XR, Xiong AS, Zhang J. Isolation and characterization of an AP2/ERF-RAV transcription factor BnaRAV-1-HY15 in Brassica napus L. HuYou15. Mol Biol Rep. 2011;38(6):3921–8.
Zhao L, Luo Q, Yang C, Han Y, Li W. A RAV-like transcription factor controls photosynthesis and senescence in soybean. Planta. 2008;227(6):1389–99.
Zhang K, Zhao L, Yang X, Li M, Sun J, Wang K, Li Y, Zheng Y, Yao Y, Li W. GmRAV1 regulates regeneration of roots and adventitious buds by the cytokinin signaling pathway in Arabidopsis and soybean. Physiol Plant. 2018;165(4):814–29.
Kakimoto T. Cytokinin signaling. Curr Opin Plant Biol. 1998;1(5):399–403.
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, et al. Trinity: reconstructing full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol. 2013;29(7):644–52.
Simao FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 2015;31(19):3210–2.
Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinf. 2011;12:323.
Deng YY, Li JQ, Wu SF, Zhu YP, Chen YW, He FC. Integrated nr database in protein annotation system and its localization. Comput Eng. 2006;32(5):71–4.
Apweiler R, Bairoch A, Wu CH, Barker WC, Boeckmann B, Ferro S, Gasteiger E, Huang H, Lopez R, Magrane M, et al. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 2004;32:D115–9.
Ashburner M, Ball CA, Blake JA, Botstein D, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, et al. Gene ontology: tool for the unification of biology. Gene. 2000;25(1):25–9.
Tatusov RL, Galperin MY, Natale DA, Koonin EV. The COG database a tool for genome scale analysis of protein functions and evolution. Nucleic Acids Res. 2000;28(1):33–6.
Koonin EV, Fedorova ND, Jackson JD, Jacobs AR, Krylov DM, Makarova KS, Mazumder R, Mekhedov SL, Nikolskaya AN, Rao BS, et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 2004;5(2):R7.
Huerta-Cepas J, Szklarczyk D, Forslund K, Cook H, Heller D, Walter MC, Rattei T, Mende DR, Sunagawa S, Kuhn M, et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 2016;44:D286–93.
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 2004;32:D277–80.
Altschul SF, Madden TL, Schäffer AA, Zhang JH, Zhang Z, Miller W, Lipman DG. Gapped BLAST and PSI BLAST a new generation of protein database search programs. Nucleic Acids Res. 1997;25(17):3389–402.
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, et al. Pfam: the protein families database. Nucleic Acids Res. 2014;42:D222–30.
Eddy SR. Profile hidden Markov models. Bioinformatics. 1998;14(9):755–63.
Fahlgren N, Howell MD, Kasschau KD, Chapman EJ, Sullivan CM, Cumbie JS, Givan SA, Law TF, Grant SR, Dangl JL, et al. High-throughput sequencing of Arabidopsis microRNAs: evidence for frequent birth and death of MIRNA genes. PLoS One. 2007;2(2):e219.
Huber W, Carey VJ, Gentleman R, Anders S, Carlson M, Carvalho BS, Bravo HC, Davis S, Gatto L, Girke T, et al. Orchestrating high-throughput genomic analysis with bioconductor. Nat Methods. 2015;12(2):115–21.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550.
R Core Team (2019). R: a language and environment for statistics computing. R Foundation for statistical computing, Vienna, Austria. URL https://www.R-project.org/.
Griffiths-Jones S, Bateman A, Marshall M, Khanna A, Eddy SR. Rfam: an RNA family database. Nucleic Acids Res. 2003;31(1):439–41.
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Glockner FO. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007;35(21):7188–96.
Chan PP, Lowe TM. GtRNAdb: a database of transfer RNA genes detected in genomic sequence. Nucleic Acids Res. 2009;37:D93–7.
Jurka J, Kapitonov VV, Pavlicek A, Klonowski P, Kohany O, Walichiewicz J. Repbase update, a database of eukaryotic repetitive elements. Cytogenet Genome Res. 2005;110(1–4):462–7.
Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10(3):R25.
Friedlander MR, Mackowiak SD, Li N, Chen W, Rajewsky N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012;40(1):37–52.
Allen E, Xie Z, Gustafson AM, Carrington JC. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell. 2005;121(2):207–21.
Varkonyi-Gasic E, Wu R, Wood M, Walton EF, Hellens RP. Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods. 2007;3:12.
Guo HH, Li RF, Liu SB, Zhao N, Han S, Lu MM, Liu XM, Xia XL. Molecular characterization, expression, and regulation of Gynostemma pentaphyllum squalene epoxidase gene 1. Plant Physiol Biochem. 2016;109:230–9.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods. 2001;25(4):402–8.
Acknowledgements
Not applicable.
Funding
This work was supported by the National Natural Science Foundation of China (Grant Nos. 31760044, 31870650). The funding bodies played no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript.
Author information
Authors and Affiliations
Contributions
HG designed, supervised the research, and made revision of the manuscript. QY and SL performed most of the experiments and wrote the paper, with assistance from XH, JM, WD, XW and XX. All authors contributed in the final version of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 1: Figure S1.
Anatomical characteristics at different stages in aerial stem-to-rhizome transition in Gynostemma pentaphyllum. (a, d) Aerial stem (stage 1). (b, e) Aboveground moderately swelling stem (stage 2). (c, f) Underground newly formed rhizome (stage 3). E: epidermis; Co: cortex; Pe: perivascular fiber; V: vascular bundle; Pi: pith. Bar = 300 μm (a-c); Bar = 50 μm (d-f).
Additional file 2: Figure S2.
Starch deposition at different stages in aerial stem-to-rhizome transition of Gynostemma pentaphyllum. (a, d, g) Aerial stem (stage 1). (b, e, h) Aboveground moderately swelling stem (stage 2). (c, f, i) Underground newly formed rhizome (stage 3). Red granules in the cells indicated by black arrows are starch grains stained with periodic acid-Schiff reagent. E: epidermis; Co: cortex; Pe: perivascular fiber; Ss: starch sheath; V: vascular bundle; Pi: pith. Bar = 50 μm.
Additional file 3: Figure S3.
(a) Length distribution of assembled unigenes. (b) unigene N50 by expression level. (c) Principal component analysis of the RNA-Seq data. (d) Numbers of differentially expressed genes (DEGs) from pairwise comparisons among different stages of aerial stem-to-rhizome transition in Gynostemma pentaphyllum.
Additional file 4: Figure S4.
Gene Ontology (GO) functional classification of differentially expressed genes (DEGs) for the aerial stem-to-rhizome transition in Gynostemma pentaphyllum. (a, d) All DEGs for stage 1 vs stage 2, stage 2 vs stage 3, respectively. (b, e) Up-regulated DEGs for stage 1 vs stage 2, stage 2 vs stage 3, respectively. (c, f) Down-regulated DEGs for stage 1 vs stage 2, stage 2 vs stage 3, respectively.
Additional file 5: Figure S5.
Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment of differentially expressed genes (DEGs) for the aerial stem-to-rhizome transition in Gynostemma pentaphyllum. (a, d) All DEGs for stage 1 vs stage 2, stage 2 vs stage 3, respectively. (b, e) Up-regulated DEGs for stage 1 vs stage 2, stage 2 vs stage 3, respectively (c, f) Down-regulated DEGs for stage 1 vs stage 2, stage 2 vs stage 3, respectively.
Additional file 6: Figure S6.
Length distribution of miRNAs.
Additional file 7: Figure S7.
Validation of selected differentially expressed genes (DEGs), as well as differentially expressed miRNAs (DEMs) and their targets by qRT-PCR. (a) DEGs. (b) DEMs. (c) DEMs and their targets. All data in the figure represents the mean values of three independent experiments ± standard deviation (SD) (n = 3). Different letters above the columns indicate significant differences at P < 0.05.
Additional file 8: Figure S8.
(a) Phylogenetic relationship between the deduced amino acid sequences of GpYUCCAs and AtYUCCAs. (b) Phylogenetic relationship between the deduced amino acid sequences of GpLAZY1 and AtLAZYs. (c) Phylogenetic relationship between the deduced amino acid sequences of GpCOLs and AtCOLs. (d) Phylogenetic relationship between the deduced amino acid sequences of GpCDF and other plant CDFs. Notes: Gp: Gynostemma pentaphyllum; At: Arabidopsis thaliana; St: Solanum tuberosum. Black arrows indicate protein associated with photoperiod and gravitropism in Arabidopsis or Solanum tuberosum, and red arrows indicate putative proteins in G. pentaphyllum. Accession numbers: AtYUCCA1, number: NP_194980; AtYUCCA2, number: NP_193062; AtYUCCA3, number: NP_171955; AtYUCCA4, number: NP_196693; AtYUCCA5, number: NP_199202; AtYUCCA6, number: NP_001190399; AtYUCCA7, number: NP_180881; AtYUCCA8, number: NP_194601; AtYUCCA9, number: NP_171914; AtYUCCA10, number: NP_175321; AtYUCCA11, number: NP_173564; AtLAZY1, number: NP_196913; AtLAZY2, number: NP_173183; AtLAZY3, number: NP_001117313; AtLAZY4, number: NP_177393; AtLAZY5, number: NP_189119; AtLAZY6, number: NP_850639; AtCOL1, number: NP_197089; AtCOL2, number: NP_186887; AtCOL3, number: NP_180052; AtCOL4, number: NP_197875; AtCOL5, number: NP_568863; AtCOL7, number: NP_177528; AtCOL9, number: NP_187422; AtCOL12, number: NP_188826; AtCOL16, number: NP_173915; AtCDF1, number: NP_197695; AtCDF2, number: NP_851106; AtCDF3, number: NP_190334; AtCDF4, number: NP_180961; AtCDF5, number: NP_177116; AtCDF6, number: NP_174001; StCDF1, number: NP_001305611.
Additional file 9: Table S1.
Length distribution of assembled transcripts and unigenes.
Additional file 10: Table S2.
BUSCO analysis of transcriptome assembly in Gynostemma pentaphyllum.
Additional file 11: Table S3.
Detailed list of differentially expressed genes in during aerial stem-to-rhizome transition Gynostemma pentaphyllum.
Additional file 12: Table S4.
Classification of small RNAs.
Additional file 13: Table S5.
Detailed information of known and novel miRNAs in Gynostemma pentaphyllum.
Additional file 14: Table S6.
Predicted targets of differentially expressed miRNAs during the aerial stem-to-rhizome transition of Gynostemma pentaphyllum.
Additional file 15: Table S7.
Primers used for quantitative RT-PCR in the study.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Yang, Q., Liu, S., Han, X. et al. Integrated transcriptome and miRNA analysis uncovers molecular regulators of aerial stem-to-rhizome transition in the medical herb Gynostemma pentaphyllum. BMC Genomics 20, 865 (2019). https://doi.org/10.1186/s12864-019-6250-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-019-6250-8