Abstract
Biological tissues change their shapes through collective interactions of cells. This coordination sets length and time scales for dynamics where precision is essential, in particular during morphogenetic events. However, how these scales emerge remains unclear. Here, we address this question using the pulsatile domains observed in confluent epithelial MDCK monolayers where cells exhibit synchronous contraction and extension cycles of \(\approx 5\) h duration and \(\approx 200~ \upmu \textrm{m}\) length scale. We report that the monolayer thickness changes gradually in space and time by more than twofold in order to counterbalance the contraction and extension of the incompressible cytoplasm. We recapitulate these pulsatile dynamics using a continuum model and show that incorporation of cell stiffness dependent height variations is critical both for generating temporal pulsations and establishing the domain size. We propose that this feedback between height and mechanics could be important in coordinating the length scales of tissue dynamics.
Graphic abstract




Similar content being viewed by others
References
S. Wong, W.H. Guo, Y.L. Wang, Proc. Natl. Acad. Sci. 111, 17176 (2014)
R. Ananthakrishnan, A. Ehrlicher, Int. J. Biol. Sci. 3, 303 (2007)
J.C. Effler, P.A. Iglesias, D.N. Robinson, Cell Cycle 6, 30 (2007)
D. Ribatti, Exp. Cell Res. 359, 17 (2017)
T. Vicsek, A. Zafeiris, Phys. Rep. 517, 71 (2012)
K. Kruse, D. Riveline, Curr. Top. Dev. Biol. 95, 67–91 (2011)
C. Jouve, T. Iimura, O. Pourquie, Development 129, 1107 (2002)
A. Mara, S.A. Holley, Trends Cell Biol. 17, 593 (2007)
A.C. Martin, M. Kaschube, E.F. Wieschaus, Nature 457, 495 (2009)
M. Roh-Johnson, G. Shemer, C.D. Higgins, J.H. McClellan, A.D. Werts, U.S. Tulu, L. Gao, E. Betzig, D.P. Kiehart, B. Goldstein, Science 335, 1232 (2012)
S. Zhang, X. Teng, Y. Toyama, T.E. Saunders, Curr. Biol. 30, 3364 (2020)
R. Fernandez-Gonzalez, J.A. Zallen, Phys. Biol. 8, 045005 (2011)
G.B. Blanchard, S. Murugesu, R.J. Adams, A. Martinez-Arias, N. Gorfinkiel, Development 137, 2743 (2010)
M. Rauzi, P.F. Lenne, T. Lecuit, Nature 468, 1110 (2010)
J. Solon, A. Kaya-Çopur, J. Colombelli, D. Brunner, Cell 137, 1331 (2009)
H. Zhang, M. Labouesse, J. Cell Sci. 125, 3039 (2012)
R. Thiagarajan, A. Bhat, G. Salbreux, M.M. Inamdar, D. Riveline, bioRxiv (2021)
R. Cheong, A. Levchenko, Curr. Opin. Genet. Dev. 20, 665 (2010)
M. Nomura, M. Okada-Hatakeyama, Front. Physiol. 4, 68 (2013)
S.S. Soumya, A. Gupta, A. Cugno, L. Deseri, K. Dayal, D. Das, S. Sen, M.M. Inamdar, PLoS Comput. Biol. 11, e1004670 (2015)
S.M. Zehnder, M. Suaris, M.M. Bellaire, T.E. Angelini, Biophys. J. 108, 247 (2015)
S.M. Zehnder, M.K. Wiatt, J.M. Uruena, A.C. Dunn, W.G. Sawyer, T.E. Angelini, Phys. Rev. E 92, 032729 (2015)
J. Notbohm, S. Banerjee, K.J.C. Utuje, B. Gweon, H. Jang, Y. Park, J. Shin, J.P. Butler, J.J. Fredberg, M.C. Marchetti, Biophys. J. 110, 2729 (2016)
G. Peyret, R. Mueller, J. d’Alessandro, S. Begnaud, P. Marcq, R.M. Mège, J.M. Yeomans, A. Doostmohammadi, B. Ladoux, Biophys. J. 117, 464 (2019)
X. Serra-Picamal, V. Conte, R. Vincent, E. Anon, D.T. Tambe, E. Bazellieres, J.P. Butler, J.J. Fredberg, X. Trepat, Nat. Phys. 8, 628 (2012)
V. Petrolli, M. Le Goff, M. Tadrous, K. Martens, C. Allier, O. Mandula, L. Hervé, S. Henkes, R. Sknepnek, T. Boudou et al., Phys. Rev. Lett. 122, 168101 (2019)
S. Yabunaka, P. Marcq, Soft Matter 13, 7046 (2017)
S. Tlili, E. Gauquelin, B. Li, O. Cardoso, B. Ladoux, H. Delanoë-Ayari, F. Graner, R. Soc. Open Sci. 5, 172421 (2018)
S. Banerjee, K.J.C. Utuje, M.C. Marchetti, Phys. Rev. Lett. 114, 228101 (2015)
V. Hakim, P. Silberzan, Rep. Prog. Phys. 80, 076601 (2017)
L. Kocgozlu, T.B. Saw, A.P. Le, I. Yow, M. Shagirov, E. Wong, R.M. Mège, C.T. Lim, Y. Toyama, B. Ladoux, Curr. Biol. 26, 2942 (2016)
J. Schindelin, I. Arganda-Carreras, E. Frise, V. Kaynig, M. Longair, T. Pietzsch, S. Preibisch, C. Rueden, S. Saalfeld, B. Schmid et al., Nat. Methods 9, 676 (2012)
W. Thielicke, E. Stamhuis, J. Open Res. Softw. 2, e30 (2014)
R. Etournay, M. Popović, M. Merkel, A. Nandi, C. Blasse, B. Aigouy, H. Brandl, G. Myers, G. Salbreux, F. Jülicher et al., eLife 4, e07090 (2015)
S. Curran, C. Strandkvist, J. Bathmann, M. de Gennes, A. Kabla, G. Salbreux, B. Baum, Dev. Cell 43, 480 (2017)
Ã. Fodor, V. Mehandia, J. Comelles, R. Thiagarajan, N.S. Gov, P. Visco, F. van Wijland, D. Riveline, Biophys. J. 114, 939 (2018)
C. Monzel, K. Sengupta, J. Phys. D Appl. Phys. 49, 243002 (2016)
F.Y. Chu, S.C. Haley, A. Zidovska, Proc. Natl. Acad. Sci. 114, 10338 (2017)
A.C. Martin, B. Goldstein, Development 141, 1987 (2014)
D. Pinheiro, E. Hannezo, S. Herszterg, F. Bosveld, I. Gaugue, M. Balakireva, Z. Wang, I. Cristo, S.U. Rigaud, O. Markova et al., Nature 545, 103 (2017)
K. Skouloudaki, D.K. Papadopoulos, T.W. Hurd, Int. J. Mol. Sci. 21, 8804 (2020)
T. Risler, A. Peilloux, J. Prost, Phys. Rev. Lett. 115, 258104 (2015)
A. Taloni, E. Kardash, O.U. Salman, L. Truskinovsky, S. Zapperi, C.A.M. La Porta, Phys. Rev. Lett. 114, 208101 (2015)
L. Saias, J. Swoger, A. D’Angelo, P. Hayes, J. Colombelli, J. Sharpe, G. Salbreux, J. Solon, Dev. Cell 33, 611 (2015)
P. Recho, A. Hallou, E. Hannezo, Proc. Natl. Acad. Sci. 116, 5344 (2019)
A. Janshoff, Biophys. Rep. 1, 100024 (2021)
S.L. Vecchio, R. Thiagarajan, D. Caballero, V. Vigon, L. Navoret, R. Voituriez, D. Riveline, Cell Syst. 10, 535 (2020)
S.P. Palecek, J.C. Loftus, M.H. Ginsberg, D.A. Lauffenburger, A.F. Horwitz, Nature 385, 537 (1997)
J. Brevier, M. Vallade, D. Riveline, Phys. Rev. Lett. 98, 268101 (2007)
Acknowledgements
The MDCK-GFP-E-Cadherin cells were a kind gift from Nelson lab., Stanford. We are grateful to Guillaume Salbreux for discussions and many key insights into the theoretical model. We thank the Riveline lab. for discussions and comments and acknowledge the help of Basile Gurchenkov, Yves Lutz and Pascal Kessler from the IGBMC imaging facility. We thank the late Marcel Boeglin for writing the Fiji plugin to perform height measurements. R.T. was an IGBMC international PhD program fellow supported by LabEx INRT funds. M.M.I acknowledges funding from Science and Engineering Research Board (MTR/2020/000605). D.R. acknowledges support from CNRS (ATIP), ciFRC Strasbourg, the University of Strasbourg, Labex IGBMC. This study has been also supported by a French state fund through the Agence Nationale de la Recherche under the frame program Investissements d’avenir labeled A NR-10-IDEX-0002-02.
Author information
Authors and Affiliations
Contributions
All authors contributed equally to this paper.
Corresponding authors
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file 1 (avi 26082 KB)
Appendices
Appendix A: Derivation of equations for the continuum model and analytical estimates
Here, we systematically develop the continuum model described in the main paper and also provide detailed derivations of the relevant equations.
The theory has 2D cell area a, velocity field of the cells \(v_i\), internal stress \(\sigma _{ij}\) and the contractile matter \(\zeta \) as the field variables [34]. The relevant equations are
In the above equations, \(\alpha \) is the friction coefficient between the monolayer and the substrate, K is the linear area modulus of the cells, \(K_h\) is the height modulus of the tissue that is modeled to depend on the area gradient in the tissue (see “Appendix C”). In Eq. A.4, \(\tau \) is the time required for myosin to reach its homeostatic value \(\zeta _0\), \(\beta \) is the coupling coefficient that relates cellular area with \(\zeta \) modulation, and \(\xi _c\) is time-correlated noise satisfying
In the limit when the correlation time \(\tau _c \rightarrow 0\), the noise becomes uncorrelated or white noise. The parameters in the continuum equations and their units in SI are summarized in Table 1—see also in “Appendix D.”
For small perturbations, \(\delta a = a - a_0\) and \(\delta \zeta = \zeta - \zeta _0\), eliminating the velocity term and linearizing in \(\delta a\) and \(\delta \zeta \), we get
Here, we neglect the advection term of the form \(\zeta _0 \partial _k v_k\) for conceptual simplicity in the weak advection limit. Choosing, \({\bar{\tau }} = {\zeta _0 \over \beta }\) as a reference time scale and \(\ell = {\sqrt{K\zeta _0 \over \beta \alpha }}\) as the length scale, we get the following non-dimensionalized dynamical equations.
We can further rewrite these equations to give
Finally, we present the governing equations as
where the non-dimensional dynamical variables are written as \(a \equiv \left[ {\beta \over \beta _c}{\delta a \over a_0}\right] \) and \(\zeta \equiv \left[ {\beta \over \beta _c}{\delta \zeta \over \zeta _0}\right] \). The four, free, non-dimensional parameters in these equations are \(\mu \equiv {\mu \over \alpha \ell ^2} = {\mu \beta \over K\zeta _0}\), \(K_h \equiv {K_h {\bar{\tau }} \over \alpha K \ell ^4} = {K_h \alpha \beta \over \zeta _0 K^2}\), \({1 \over K} \equiv {\zeta _0 \over K}\) and \(\tau \equiv {\tau \beta \over \zeta _0}\), in terms of the original dimensional parameters. The correlation time of the colored noise \(\xi _c\) is written as \(\tau _c \equiv {\tau _c \beta \over \zeta _0}\).
Taking Fourier transform of the two coupled equations in space (\(x \rightarrow q_x\) and \(y \rightarrow q_y\)) and time (\(t\rightarrow \omega \)), we get
where \({\tilde{a}}\) and \({\tilde{\zeta }}\) are the space-time Fourier transforms, respectively, of a and \(\zeta \), and \(q^2 = q_x^2 + q_y^2\). Inverting the \({{\textbf{C}}}\) matrix, we get
The power spectrum from \({\tilde{a}}\) and \({\tilde{\zeta }}\) then becomes
We note that since \(q = \sqrt{q_x^2 + q_y^2}\), the power spectrum only in terms of q can now be obtained as \(S_a(q, \omega ) = 2\pi q \times S_a(q_x, q_y, \omega )\). Since the linearized mass balance equation Eq. A.1 in terms of the non-dimensionalized variables and parameters is
the power spectrum for velocity divergence becomes, \(S_d(q, \omega ) = {\beta _c^2 \over \beta ^2}\omega ^2 S_{a}(q, \omega )\). Since we have taken \(\xi _c\) to be correlated noise (Eq. A.4), its power spectrum \(|\tilde{\xi _c}|^2 = 1/(1 + \tau _c^2\omega ^2)\). Hence, the effective power spectrum for the velocity divergence is given as
where
We can perform a scaling near \(q \rightarrow \infty \)
Upon integrating the \(\omega \) terms out, we get
and near \(q \rightarrow 0\) we get
upon integrating the \(\omega \) terms we get
As derived above, the series expansion for \(S_d(q)\) is increasing and decreasing in q for small and large values, respectively, of q. Hence, we could expect the location \(q_m\) of the maxima of \(S_d(q)\) to approximately be at the intersection of these two curves. Hence, we equate them
to give
The wavelength associated with this pattern will be given by \(l_0 = 2 \pi /q_m\).
Following the same procedure as for q, we now check for the asymptotic behavior for \(S_d(q, \omega )\) in terms of \(\omega \).
Similarly,
We find that the location of maxima for \(S_d(\omega )\) is obtained quite well when we equate \(S(q, \omega \rightarrow 0)\) and \(S(q, \omega \rightarrow \infty )\) corresponding \(q = q_m\) as obtained in Eq. A.26. From that, we get that the maxima for \(S_d(\omega )\) occur at
Based on Eqs. A.21-A.29, we can get some insights into the role the non-dimensionalized parameters \(K, K_h, \mu , \tau \) and \(\tau _c\) on the behavior of power spectra \(S_d(\omega )\) and \(S_d(q)\). From Eq. A.26, we find that, as per our model, the three terms \(K_h, \tau \) and \(\tau _c\) are important for the observation of maxima in \(S_d(q)\) as is observed experimentally (Fig. 3e). Hence, the term corresponding to gradients in area in tissue stress (Eqs. 3 and A.3), the active stress homeostatic term (Eqs. 6 and A.14) and the time-correlation component of the noise (Eq. A.6) play key role in setting the length scale of pulsations in the tissue within our model framework. Additionally, from Eqs. A.22 and A.24, we see that the rate at which \(S_d(q)\) grows and decays for small and large values, respectively, of q is governed by the parameters \(\tau , K\), and \(K_h\). From Eq. A.28, it can be seen that although the parameter \(\mu \) does not play an essential role in governing the qualitative nature of \(S_d(q)\) and \(S_d(\omega )\), it modulates the decay behavior of \(S_d(\omega )\) for larger values of \(\omega \).
In Fig. 3, we plot the normalized power spectrum in terms of the wavenumber q and \(\omega \) by defining the following.
Note that for notational simplicity, we use the same symbol \(S_d\) for the different variants of the power spectrum.
In order to quantitatively obtain the low and high q and \(\omega \) behavior of \(S_d(q)\) and \(S_d(\omega )\), respectively, we show in Fig. 4 log–log plot of the data presented in Fig. 3a, c. Since, as opposed to the theoretical calculation, \(S_d(\omega = 0) \ne 0\) for the experimental data, we fit a power law of the form \(\omega ^{m}\) while neglecting a few data points at the beginning (Fig. 4a). The exponents \(m \approx 1.4\) and \(m \approx -1.7\) (Fig. 4a) are not the same as \(m \approx 2\) (Eq. A.27) and \(m \approx -4\) (Eq. A.28), respectively, as is predicted by the model. Similarly, the exponents \(m \approx 1.5\) and \(m \approx -4.8\) (Fig. 4b) for \(S_d(q)\) also do not match with the theoretical estimate of \(m = 5\) (Eq. A.24) and \(m = -3\) (Eq. A.22).
Log–log plot for (a) \(S_d(\omega )\) and (b) \(S_d(q)\) presented earlier in Fig. 3c, e, respectively. The behavior of the power spectrum at small and large values of \(\omega \) and q, respectively, is shown as blue lines in (a, b). a For small values of \(\omega \), \(S_d(\omega ) \sim \omega ^{1.4}\) and for larger values of \(S_d(\omega ) \sim \omega ^{-1.7}\). The theoretical values of these exponents, 2 (Eq. A.27) and \(-4\) (Eq. A.28), are not the same as those experimentally observed. b For small values of q, \(S_d(q) \sim q^{1.5}\) and for large values of q, \(S_d(q) \sim q^{-4.8}\). The theoretical values of these exponents 5 (Eq. A.24) and \(-3\) (Eq. A.22) are different than their experimentally obtained counterparts
Appendix B: Power spectrum for cellular pulsations on non-patterned surface
Power spectra a \(S_d(\omega )\), c \(S_d(q)\) and the corresponding log–log plots (b) and (d) for pulsatory cellular movements in MDCK epithelial monolayers on non-patterned surfaces. The nature of the power spectra is similar to that obtained for patterned surfaces in Fig. 3c, e. The blue lines and the gray-shaded region in (a) and (c) correspond to the mean and standard deviation, respectively, of \(S_d(\omega )\) and \(S_d(q)\) obtained over five separate experimental repeats. The power-law fits for low and high values of (b) \(\omega \) and (d) q are represented with the blue lines, and the corresponding exponents are indicated
In our previous work [17], we had studied pulsations on epithelial monolayers. Here, similar to the power spectrum \(S_d(\omega )\) and \(S_d(q)\) for the divergence of pulsatory cellular movements on patterned surfaces discussed in Sect. 3.3 and plotted in Fig. 3c, e, respectively, we do the equivalent for the non-patterned surfaces in Fig. 5a, c. As shown in Fig. 5, the blue curves and the gray-shaded region correspond to the mean and standard deviation of the quantities over five experimental repeats. The peak for \(S_d(\omega )\) happens at \(\omega /2\pi \approx 0.15~\textrm{h}^{-1}\) corresponding to oscillation period of approximately 6.5 h. Although the peak is not as well defined for \(S_d(q)\), its maxima occur at around \(q/2\pi \approx 0.004~{ \upmu \textrm{m}}^{-1}\) corresponding to a periodic pattern of approximately \(250~{ \upmu \textrm{m}}\). The values are not significantly different from that obtained for the patterned surface. We also check the values of the power spectra at larger and smaller values of \(\omega \) (Fig. 5b) and q (Fig. 5d) and obtain the power-law exponents as shown in Fig. 4 for patterned substrate. Although the qualitative rise and decay in q and \(\omega \) for the power spectra is similar for the patterned and non-patterned surfaces, the power-law exponents are not the same for these two conditions. However, the exponents for \(S_d(q)\) do exhibit a better match with each other. As for the patterned surface, the exponents also do not match the values predicted by the model.
Appendix C: Reasoning behind the term involving \(K_h\) in tissue stress
We provide an intuitive reasoning for including the term \(K_h \nabla ^2 \left( {a - a_0 \over a_0} \right) \) in tissue stress \(\sigma _{ij}\) in Eq. 3. We first note that a cell has at least two modes of deformation. First is the in-plane area change \(\delta a = a - a_0\) without any gradients in tissue height h. For an incompressible tissue, \(a h = a_0 h_0\) due to which \(\delta a/a_0 = - \delta h/h_0\). For the area deformation mode, the energy density of the tissue is \(k_1 (a - a_0)^2\). The other mode of deformation in which there is no change in cell area would necessarily involve cell height gradients, and the energy density, to the lowest order, can be written as \(k_2 |\nabla h|^2\). Thus, the total energy of the tissue due to these two modes of deformation is
over the total region \(\varOmega \) of the tissue. We note that the apical and basal layers of the epithelium can have distinct mechanical properties [46]. Since spatial variation in tissue height h can lead to different deformations of the apical and basal layers, the difference in mechanical properties can further contribute to the tissue energy via the \(|\nabla h|^2\) term in Eq. C.33. For small deformation of the tissue, the isotropic stress \(\sigma \) in the tissue will be obtained by taking the functional derivative of the tissue \(\sigma = \delta U/ \delta a\). After noting that \(\nabla h = -h_0/a_0\nabla a\), and renormalizing the values of the material coefficients \(k_1\) and \(k_2\), we get
as written in Eq. 3. We note that in the absence of volume incompressibility of the monolayer, we cannot rigorously replace the \(\nabla h\) term with \(\nabla a\) as described above. Although, in such a case, we do not expect major qualitative changes in our model except for re-scaling of \(K_h\), the use of \(\nabla a\) term is, strictly speaking, a model approximation.
Appendix D: Estimate for pattern length scale \(l_0\)
We estimate the typical length \(l_0\) given in Eq .10 by proposing the following experimental values for the parameters : substrate friction \(\eta \approx 30~{ \textrm{N}\, \textrm{s}\, \textrm{m}^{-1}}\) between the cells and the surface [47, 48], cell stiffness \(Y \approx 100~{\textrm{pN} \, \upmu \textrm{m}^{-2}}\) in terms of the typical force per unit area for cell–cell contacts [49], myosin homeostasis time \(\tau \approx 60~\textrm{min}\) [23] and myosin fluctuation correlation time along cell–cell junctions \(\tau _c \approx 2~\textrm{min}\) [35]. Taking for cell area \(a_0 \approx 100~{ \upmu \textrm{m}}^2\) and for cell height \(h_0 \approx 10~{ \upmu \textrm{m}}\), we can express \(\alpha \approx \eta \times a_0\) and \(K_h \approx Y\times a_o\times h_0\). Using these numbers, we obtain an approximate length scale associated with the pulsating pattern \(l_0 \approx 100~{ \upmu \textrm{m}}\), which is consistent with the typical length scales measured experimentally.
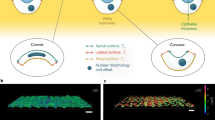
Supplementary movie caption: Movie 1: Height fluctuations during monolayer pulsations. In the top, domain of MDCK-E-Cadherin-GFP cells on FN grid is shown at two z-planes: 2.5 Â\(\mu \)m and 8 Â\(\mu \)m. Scale bar, 100 Â\(\mu \)m. In the bottom, 3D surface view of cells where height is encoded as intensity is shown. Color bar shows the height values. Time in hh:mm.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thiagarajan, R., Inamdar, M.M. & Riveline, D. Interplay between cell height variations and planar pulsations in epithelial monolayers. Eur. Phys. J. E 45, 49 (2022). https://doi.org/10.1140/epje/s10189-022-00201-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epje/s10189-022-00201-6