Abstract
The recently developed model of the epidemic spread of two virus strains in a closed population is generalized to the situation typical for the couple of strains delta and omicron, when there is a high probability of omicron infection soon enough after recovering from delta infection. This model can be considered as a kind of combination of SIR and SIS models for the case of competition of two strains of the same virus with different contagiousness in a population. The obtained equations and results can be directly implemented for practical calculations of the replacement of strains of the SARS-CoV-2 virus. A comparison between the estimated replacement time and the corresponding statistics shows reasonable agreement.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Existing models for the spread of infection describe the free-running development of an epidemic and all its stages. There are two basic models for such description: the susceptible–infected–susceptible (SIS) and susceptible–infectious–removed, susceptible–exposed–infectious–removed (SIR, SEIR). The SIS model goes back to the pioneering investigations of malaria by Ronald Ross [1] and uses the assumption that the recovered people can immediately get infection again. Existing SIR models assume that the recovered people save immunity during epidemic (see, e.g., [2, 3]). There are many versions of those models [4,5,6,7,8,9,10] (see also references therein). The balance between the susceptible and infected members of population under the various conditions of infection transfer is the subject of research in [8, 11, 12].
Recently, the delayed time-discrete epidemic model (DTDEM) considering the typical long duration of the COVID-19 disease has been developed [13]. In [14, 15], this specific delay has been presented in differential form. The delay discussed in [13,14,15] assumes that the recovered patient is immune, and in this respect fits the SIR model, rather than SIS. The considered delay models do not imply the allocation of a separate category of hidden virus carriers (see, e.g., the SEIR models in [16, 17]). Latent carriers of the virus can infect others without delay and are similar to the infected ones. Currently, the simplest SIS, SIR and SEIR baseline models are being developed taking into account the vaccination process [18,19,20].
Nowadays, the actual problem is the strain appearance and circulation in application to COVID-19 [21,22,23]. In the recent preprint [21], the SIR-type model was considered for the case of the coexistence of two strains of the COVID-19 virus spreading in the same population. In [21], as in the pioneer paper [22] devoted to the circulation of several strains in a population, it was assumed that after being ill with any strain, those who recovered were completely immune. We called this assumption about the properties of strains “strain orthogonality”, bearing in mind a certain analogy with the mathematical orthogonality of functions. An important difference between [21] and [22] is the complete disappearance of strains at long times. This is due to the fact that [21] considers times much shorter than the average time of a human life. It should be noted that in reality, the long-term circulation of strains (see [22]) is mainly not associated with the finite average lifetime and assumptions about the nature of the birth and death of healthy people in the population. The prolonged circulation of strains is due to the emergence of a new, more contagious strain in the patient’s body during a long-term illness with another strain that is easily susceptible to mutations (like SARS-CoV-2, for example) in the absence of a universal vaccine. For infections such as influenza, the main sources of mutations are carriers external to humans (birds, pigs, and other representatives of the animal world). In this paper, we do not include these factors, which differ significantly from those considered in [21, 22] and require separate study.
Due to different contagiousness, a replacement of a less contagious strain to a more contagious one takes place; it was quantitatively described in [21] and, in concrete application to COVID-19, in [22] under the “strain orthogonality” condition. In [22] the characteristic details of such a process were revealed, such as the necessary conditions for the emergence of a maximum in the curves describing the current number of virus carriers, a decrease in the peak incidence of a less contagious strain when a more contagious strain appears, a faster depletion of the part of population that has not affected by any strain. This means a more rapid course of the epidemic when a more contagious strain appears (if a third, even more contagious strain, does not arise) and increase in the required level of recovered patients to achieve collective immunity (if it turns out to be possible) when a less contagious strain of the virus is replaced by a more contagious one, etc.
In the present paper, the basic equations [21, 22] are generalized to the case of “strain non-orthogonality”. The respective generalized equation we named “strain-stream” ones. These equations are the development of a general mathematical approach of the “principle of competitive exclusion” (see, e.g., Murray [23]) in epidemiology. In papers [24, 25], the first applications of this principle were considered for the strain transmission. In [26] the model of malaria transmission, which considers the seasonal fluctuations in mosquito population density or spatial heterogeneity with periodic migration, was proposed. It was shown that the strain heterogeneity can generate periodic behavior as a consequence of the interaction between parasite strains and host immunological defences. The model of two strain of dengue circulation in host population due to the transmission from the infected mosquitoes has been developed in detail [27]. In both papers, a vector transmitted disease (caused by different sources of infection: parasites or viruses) is considered on the basis of various differential equations. The extended SIR-type model for COVID-19 waves has been presented in [28] (see also recent the paper [29]) to fit this model calculations to the data in Tonghua City in China (see also references therein). The qualitative consideration of “strain non-orthogonality” has been considered on the basis of simplified model in [30].
It should be stressed that the mathematical structure of various models allows some comparisons in spite of the different processes and conditions which lead to the specific results.
The model under consideration makes it possible, in the presence of a minimum number of parameters, to quantitatively describe various specific situations of coexistence and struggle of two strains for dominance in a population of living organisms. The specific examples considered in this work were based on the choice of initial conditions that correspond to the emergence of a second strain of high contagiousness (for example, omicron) against the background of an already developed epidemic with the dominance of the delta strain. To describe such a situation, it suffices to take into account the initial conditions, bringing them into line with the actual level of delta disease in a certain population, to the time the strain appeared in South Africa, which was subsequently named omicron by WHO.
At the same time, if we are interested in the rivalry of two strains (for specificity, below we designate delta-1 and omicron-2) in any country, region, city or locality, we naturally must use the available statistical data on the incidence, caused by the 1 strain when cases of the disease caused by the 2 strain appear. The detail data concerning COVID-19 disease for different countries, without differentiation on strains are quite fully reflected in [31]. City data are presented on the websites of the respective countries (e.g., the Robert Koch Institute in Germany, Stopcoronavirus in Russia, the Johns Hopkins Institute in the USA, etc.). There are also recent publications on the strain replacement cases and the respective statistics (see, e.g.,[32,33,34] and references therein).
2 Equations for the case of “strain non-orthogonality”
This generalization reflects the observable property to be infected with a high probability by the strain 2 of the COVID-19 disease for those who have already been ill and recovered from infection caused by the strain 1. This means that immunity to strain 2 is not developed (or is only partially developed) after disease caused by strain 1. Obviously, for strains that cause COVID-19 (as well as for influenza viruses), there is only a limited period of immunity but much longer than the average disease duration. In fact, the property of “strain non-orthogonality” means that infection caused by strain 2 (omicron) can appear with some probability even immediately after recovering from the disease caused by strain 1 (e.g., delta). According to our knowledge, the disease COVID-19 caused by two strains which simultaneously coexist in one sick person was not observed (in contrast with the rare cases of COVID-19 and flu). At the same time, according to the existing statistical data infection 1 was not observed after infection by strain 2. The additional reason for this is a fast disappearance of the less contagious strain, as we demonstrate below.
As in [28] we denote S the number of never infected people in a closed population consisting of N people, \(I_1\) and \(I_2\) are the number of strain carriers of type 1 and 2. The simplified SIR-type hooking model equations describing the epidemic spread for the case of two “ non-orthogonal” strains in the close population N can be written in the form using the variables \(I_1(t)/N=y_1(t)\), \(I_2(t)/N=y_2(t)\), \(S(t)/N=u(t)\)
The values \(T_1\) and \(T_2\) are the average durations of the diseases caused by strains 1 an 2. Parameters \(p_1\) and \(p_2\) are the characteristics of the contagiousness for two strains, which are determined as the product of the quantity of dangerous contacts \(n_c\) of the infected people per day and the average susceptibility k of the healthy person on dangerous distance [13, 14]. The new term in Eq. (3) describes the infection process by strain 2 of the people recovered after the disease caused by strain 1.
The coefficient \(0 \le \gamma _2<1\), hereinafter referred to as the Viral Link Attenuation Factor (VLAF), describes a certain decrease in the probability of getting 2 after being infected with 1 (partial increase in immunity) compared to the probability of getting 2 without having been ill before 1 (i.e., directly from the group u). This is due to the production of antibodies after the disease caused by the 1 strain (or after vaccination), which perform some protective function against strain 2.
However, the last term in Eq. (3) only qualitative describes the infection process by strain 2 of the people recovered after the disease caused by strain 1. People after infection 1 can be infected by the strain 2 not immediately, but after some time. This circumstance should be accounted for the explicit description of the “strain non-orthogonality”. To find the respective equations we divide all virus carriers of strain 2 into two groups—\(y_{2 \leftarrow S}\) and \(y_{2 \leftarrow 1}\) . The proportion of those infected with strain 2 who were not ill with strain 1 (i.e., infected from the set S who were not ill with any of the strains) is designated \(y_{2 \leftarrow S}\) and those infected with strain 2 from the set of those who had previously been sick by strain 1 are designated \(y_{2 \leftarrow 1}\). The entire set of patients with strain 2 is equal to \(y_2\) = \(y_{2 \leftarrow S}\) + \(y_{2 \leftarrow 1}\) . Then, the system of equations reads
Here, the function f(t) is the proportion of those who have or had at moment t strain 1, but did not have strain 2 . To find function f(t) necessary to take into account that only a part from all people who have been ill \(x_1(t)\)
can contribute in f(t). The function \(x_1(t)\) contains all people who have been ill with strain 1 by time t (they are extracted only from S). Among them are those identified as \(\varphi (\tau )\) who obtain strain 2 after being ill with strain 1 at time t (they are taken from the set \(x_1(\tau )-\varphi (\tau ))\). According to the model, they cannot get sick again with strain 2 and therefore must be subtracted from \(x_1(t)\) to find f(t)
Therefore, this subtracted function \(\varphi (t)\) is determined by the integral equation
From (9) and (10), taking into account (8), it follows
Summarizing Eqs. (6), (7), we arrive at equations of the model of “strain non-orthogonality”, which considers that after disease caused by strain 1 one can be infected by strain 2 (but not vice versa)
It is easy to see that all people \(x_2(t)\) diseased by strain 2 at time t can be calculated by integral
All coefficients in Eqs. (12, 13, 14, 15) are positive, \(p_{1,2}>0\), \(T_{1,2}>0\), \(\gamma _2>0\), and values of all variable functions lie in the interval (0, 1). There is only one stationary solution to Eqs. (12, 13, 14, 15), \(y_{1,2}=0\), \(u=u_0\), where \(u_0\) is an arbitrary constant. The stationary state is unstable, that is, in the vicinity of zero one of the functions \(y_{1,2}(t)\) grows in time, if \(u_0>\min [1/(p_1T_1),1/(p_2T_2)]\).
Suppose that initially \(u(0)=u_0\) and \(y_{1,2}(0) \approx 0\). Throughout the present study we are interested in the case when both \(y_{1,2}(t)\) grow in time, that is, \(u_0\) should be large enough, \(u_0>\max [1/(p_1T_1),1/(p_2T_2)]\). According to Eq. (12), u(t) is always a decreasing function of time and sooner or later infected fractions of population start decreasing. Evidently, \(y_{1,2}(t\rightarrow \infty )\rightarrow 0\), however an asymptotic value, \(u(t\rightarrow \infty )\rightarrow u_{\infty }\), depend on initial conditions. It may be only deduced from the qualitative analysis that \(u_{\infty }\) should be small enough, \(u_{\infty }<\min [1/(p_1T_1),1/(p_2T_2)]\).
The two-strain propagation model developed in [21] is the limiting case of the considered general model (12, 13, 14, 15) for \(\gamma _2 =0\). In this case Eq.(15) is split off and Eqs. (12, 13, 14) are closed.
3 Numerical solution for various immunity parameter VLAF
The analysis of the stability of the stationary solution carried out above is similar to one in [21]. It shows that the necessary condition for the development of an epidemic process at \(\gamma _2=0\) is the condition \(p_i T_i u_0> 1\). This condition remains valid for Eqs. (12, 13, 14, 15).
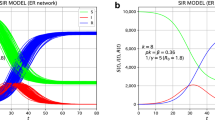
As was revealed in [21] for \(\gamma _2=0\), using the example of specific initial conditions and parameters \(p_i\) and \(T_i\), the coexistence of two viruses of different contagiousness leads over time to the replacement of the less contagious strain by a more contagious one, even if the share of the latter at the beginning of the process was significantly smaller than less contagious. The results of calculations for specific parameters that correspond to the simultaneous emergence of an epidemic with two strains of different contagiousness are shown in Figs. 1 (epidemic process in the presence of strain 1 only, when \(p_2=0\)) and 2 (comparison of the dynamics of the epidemic in the presence of both strains for the model under consideration with \(\gamma _2=0.3\)).
Thus, Figs. 1 and 2 serve to demonstrate the process of mutual influence of strains during the development of an epidemic for the case of “strain non-orthogonality” based on Eqs. (12, 13, 14, 15).
Functions \(y_1 (t)\) (solid) and \(y_2(t)\) (dashed) of the virus carriers for the case when both strains exist. The parameters are \(p_1 = 0.15\), \(p_2 = 0.4\), \(y_1 (0) = 0.01\), \(y_2 (0) = 10^{-7}\), \(u(0) = 0.8\), the average duration of the virus carrier \(T_1 = T_2 = 15\) days, \(\gamma _2 = 0.3\)
As is easy to see, strain 1 is slightly suppressed by strain 2 since the value of maximum for the solid curve in Fig. 2 is lower than in Fig. 1 for \(u(0)=0.8\). A comparison of these figures shows that the duration of strain 1 circulation is suppressed (\(\simeq 2\) times shorter for the used parameters) due to the appearance of strain 2. A comparison of Figs. 1 and 2 shows that circulation of strain 2 is shorter than circulation of strain 1 in the case of strain 2 absence. It is easy to see that for arbitrary parameters the maximum for strain 1 in Fig. 2 is shifted to earlier time in comparison with Fig. 1. This peculiarity, mentioned in [21], is valid also for the strain-stream model under consideration.
In this paper, we are interested in the impact of a possible infection with strain 2 after recovery from an infection caused by strain 1. This situation corresponds to the epidemic process observed with the appearance of the omicron strain. An important difference from the specific examples considered in [21] is the appearance of strain 2 under conditions of a developed epidemic of strain 1, which is characterized by rather large initial values of u(0) and \(y_1(0)\).
The results of the numerical solution of Eqs. (12, 13, 14, 15) for the initial conditions simulating the situation of the appearance of omicron in already developed epidemic of the delta strain are shown in Figs. 3, 4, 5.
Comparison of the function \(y_1 (t)\) (left) and \(y_2(t)\) (right) for different values \(\gamma _2=0\) (solid), \(\gamma _2=0.2\) (dashed) and \(\gamma _2=0.8\) (dash-dotted) of virus carriers for the case when both virus strains exists. The parameters are \(p_1 = 0.15\), \(p_2 = 0.4\), \(y_1 (0) = 0.01\), \(y_2 (0) = 10^{-7}\), \(u(0) = 0.8\), the average duration of the virus carrier \(T_1 = T_2 = 15\) days
Function u(t) for the various cases: the second strain is absent (\(p_2=0\)), the parameters are \(p_1 = 0.15\), \(y_1 (0) = 0.01\), the average duration of the virus carrier \(T_1 =15\) days for \(u(0) = 0.8\) (solid); both strains coexist, the recovered are immune (the case considered in [21]), the parameters are \(p_1 = 0.15\), \(p_2=0.4\), \(y_1 (0) = 0.01\), \(y_2(0)= 10^{-7}\), \(T_1 =T_2=15\) days for \(u(0) = 0.8\) (dash-dotted), \(\gamma _2=0\); both strains coexist, the parameters are \(p_1 = 0.15\), \(p_2=0.4\), \(y_1 (0) = 0.01\), \(y_2(0)= 10^{-7}\), \(T_1 =T_2=15\) days for \(u(0) = 0.8\), \(\gamma _2=0.2\) (dashed)
Function \(x_1(t)\) (left) of the people affected by the strain 1 and \(x_2(t)\) (right) of the people affected by the strain 2 for different values \(\gamma _2=0\) (solid), \(\gamma _2=0.2\) (dashed) and \(\gamma _2=0.8\) (dash-dotted) of virus carriers for the case when both virus strains exists. The parameters are \(p_1 = 0.15\), \(p_2 = 0.4\), \(y_1 (0) = 0.01\), \(y_2 (0) = 10^{-7}\), \(u(0) = 0.8\), the average duration of the virus carrier \(T_1 = T_2 = 15\) days
The proportions of \(y_1(t)\) and \(y_2(t)\) infected with strains 1 and 2 are shown in Fig. 3 left and right, respectively, for different parameters \(\gamma _2\). As in Figs. 1 and 2, the initial condition for the proportion of the population that did not encounter either of the two considered strains was chosen at the level of \(u(0)=0.8\), which significantly exceeds the official statistics for, e.g., Germany at the time the omicron strain appeared in the country. By such an overestimation, we take into account a significant number of unreported cases of diseases with the delta strain at the time of the appearance of the omicron strain. The same qualitative picture is observable also in other countries. The initial proportion of those infected with strain 1 is chosen to be rather high \(y_1(0)=0.01\), which also corresponds to the presence of a significant number of hidden virus carriers that can actively infect others. Note, that the purpose of this work is to identify the general patterns of the development of the epidemic in the presence of two strains, and not a calculation based on a detailed analysis of the changing situation from day to day and incomplete statistical data. Nevertheless, the specific time for the essential replacement of strain 1 by strain 2 is in reasonable agreement with statistical data obtained in England [32]. We have stress that the statistical data on strain replacement can be different for different countries and regions due to various conditions.
As follows from Fig. 3, the impact of the appearance of strain 2 capable of infecting those who have been ill with strain 1 depends significantly on the value of VLAF \(\gamma _2\). The larger \(0\le \gamma _2 \le 1\), the faster the process of infection with strain 1 is suppressed, i.e., it is forced out faster than in the case with \(\gamma _2=0\) [21] (see also Fig. 1 ). At the same time, as \(\gamma _2\) grows, the current proportion of strain 2 carriers grows, exceeding by a factor of \(\simeq 3.5\) at the maximum proportion of strain 1 carriers under the chosen parameters.
The effect of a non-zero value \(\gamma _2\) on the fraction u(t) of non-affected by strains at all is shown on Fig. 4. The possibility to become infected with strain 2 soon after disease caused by strain 1 is high. There is much faster and complete depletion of the share of non-affected. This means that with a certain parameter \(\gamma _2\), herd immunity becomes practically unattainable and almost everyone must get sick due to strain 2. It is of interest to determine values \(\gamma _2\) for which the stationary value of the proportion of the population not affected by any of the viruses is reached. It can be considered as a numerical characteristic of herd immunity. The calculation carried out up to 1000 days (not presented in 4) showed that with the selected parameters, the solid curve corresponding to the absence of strain 2 tends to \(u(t = 1000) = 0.205\), the dash-dotted curve (corresponding to the case \(\gamma _2 = 0\) [21] of full lengthy in time immunity after each of the diseases caused by the strain 1 or 2) tends to 0.052 and the dotted curve corresponding to the case under consideration Eqs. (12, 13, 14, 15) for \(\gamma _2 = 0.2\) tends to \(u(1000) = 0.007\). In the latter case the level of collective immunity is only 0.7 percent of the population.
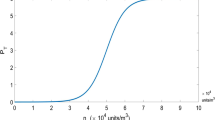
Figure 5 shows the curves for the total part of people (sick plus recovered, or affected) \(x_1(t)\) with strain 1 (left) and strain 2 (right), calculated according Eqs. (8) and (16). All three curves for the function \(x_1(t)\) (left) and for the function \(x_2(t)\) (right) in Fig. 5 correspond to the circulation of two strains, but for different values of \(\gamma _2\). The parameters in Fig. 5 correspond to those selected in Fig. 3. As it follows from Fig. 5 the function \(x_1(t)\) decreases as \(\gamma _2\) increases, while the function \(x_2(t)\) grows. This behavior corresponds to the general pattern of replacement of a less contagious virus by a more contagious one, with the greater efficiency, the greater the VLAF value.
The formulated equations and the model under consideration can be easily extended to the consideration of vaccination and different quarantine measures, accounting the government and personal restrictions, vaccination process, etc. In general we accounted, after recovering from strain 1, a person may not immediately become infected with strain 2. However, and taking into account the more elaborated equations which takes into account the delay factor associated with time shifts is beyond the scope of this work. Also the cases of death, re-infection with the same strain long time after recovery, limited time for vaccination efficiency and other known factors can be included in a more elaborated models. Above, we restricted our consideration to the case of the free-running epidemic under two “non-orthogonal” strains of a same virus. This assumption can be considered as realistic for fast developing epidemic caused by, e.g., the omicron strain (or another highly contagious virus strain) appeared in a population affected earlier by a less contagious virus strain. The considered model clarifies the main specific features of competition of two “non-orthogonal” viruses in population.
4 Conclusions
The principal picture of the replacement of one strain by another has already been revealed in the recently considered mathematical model [21], where the basic equations were proposed that describe the replacement of a less contagious strain by a more contagious one. Further development of the theory is connected with taking into account the incomplete “orthogonality” of the strains under consideration. This is manifested in the fact that with a significant mutation of the virus, leading to a different molecular structure, a different virulence, and a different clinical picture of the disease, both strains, spreading in the population, are more interdependent. Immunity to one of them (for example, due to a previous disease), generally speaking, does not means the presence of immunity in relation to another. So, for example, omicron can infect those who have recovered from the delta strain, but not vice versa.
Thus, the situation cannot be described in the framework of SIR and similar models, where all recovered patients have a long immunity, nor within the SIS model, where immunity disappears immediately after recovery. This important property is taken into account by transferring to the “strain-stream” equations by formulation of Eqs. (12, 13, 14, 15). The additional term includes the new VLAF parameter \(\gamma _2 \le 1\), due to the development of partial immunity to strain 2 as a result of the disease caused by strain 1, or to the effective vaccination against strain 1, giving partial protection also against strain 2.
Data Availability
This manuscript has no associated data or the data will not be deposited. [Authors’ comment: This is a theoretical study with the reference on the open statistical data [31].]
References
R. Ross, The Prevention of Malaria (Dutton, New York, 1910)
W.O. Kermack, A.G. McKendrick, A contribution to the mathematical theory of epidemics. Proc. R. Soc. A 115, 700–721 (1927). https://doi.org/10.1098/rspa.1927.0118
F. Brauer, C. Castillo-Chavez, Mathematical Models in Population Biology and Epidemiology (Springer-Verlag, Cham, 2000)
F. Ball, Stochastic and deterministic models for SIS epidemics among a population partitioned into households. Math. Biosci. 156, 41 (1999)
M. Keeling, K. Eames, Networks and epidemic models. J. R. Soc. Interface 2, 295–307 (2005)
G. Chowell, L. Sattenspiel, S. Bansald, C. Viboud, Mathematical models to characterize early epidemic growth: a review. Phys. Life Rev. 18, 66–97 (2016)
J. Bedford, J. Farrar, C. Ihekweazu, G. Kang, M. Koopmans, J. Nkengasong, A new twenty-first century science for effective epidemic response. Nature 575, 130–136 (2019)
G. Ghoshal, L.M. Sander, I.M. Sokolov, SIS epidemics with household structure: the self-consistent field method (2003) arXiv:cond-mat/0304301 v1 [cond-mat.stat-mech]. Accessed 12 Apr 2003
E.B. Postnikov, Estimation of COVID-19 dynamics on a back-of-envelope: does the simplest SIR model provide quantitative parameters and predictions? Chaos Solitons Fractals 135, 109841 (2020)
I. Cooper, A. Argha Mondal, C.G. Antonopoulos, A SIR model assumption for the spread of COVID-19 in different communities. Chaos Solitons Fractals 139, 110057 (2020)
F. Frey, F. Ziebert, U. Schwarz, Stochastic dynamics of nanoparticle and virus uptake. Phys. Rev. Lett. 122, 088102 (2019)
G.M. Nakamura, A.S. Martinez, Hamiltonian dynamics of the SIS epidemic model with stochastic fluctuations. Sci. Rep. 9, 15841 (2019)
S.A. Trigger, E.B. Czerniawski, Equation for epidemic spread with the quarantine measures: application to COVID-19. Phys. Scr. 95, 105001 (2020)
S.A. Trigger, E.B. Czerniawski, A.M. Ignatov, Epidemic transmission with quarantine measures: application to COVID-19. MedRxiv (2021). https://doi.org/10.1101/2021.02.09.21251288
A.M. Ignatov, S.A. Trigger, E.B. Czerniawski, Delay influence on epidemic evolution. High Temp. 59(6), 960 (2021). (in Russian)
Jose M. Carcione, Juan E. Santos, Claudio Bagaini, Jing Ba, A simulation of a COVID-19 epidemic based on a deterministic SEIR model. Front. Public Health 8, 230 (2020)
M.J. Keeling, P. Rohani, Modeling Infectious Diseases in Humans and Animals (Princeton University Press, Princeton, 2008)
Xiao-Jie. Li, Xiang Li, Vaccinating SIS epidemics under evolving perception in heterogeneous networks. Eur. Phys. J. B 93, 185 (2020)
A. Shnip, Epidemic Dynamics Kinetic Model and Its Testing on the COVID-19 Epidemic Spread Data. J. Eng. Phys. Thermophys. 94(1), 6 (2021)
S.A. Rella, Y.A. Kulikova, E.T. Dermitzakis et al., Rates of SARS-CoV-2 transmission and vaccination impact the fate vaccine-resistant strains. Sci. Rep. 11, 15729 (2021)
A.M. Ignatov, S.A. Trigger, Two viruses competition in the SIR model of epidemic spread: application to COVID-19. MedRxiv (2022). https://doi.org/10.1101/2022.01.11.22269046
H.J. Bremermann, H.R. Thieme, A competitive exclusion principle for pathogen virulence. J. Math. Biol. 27, 179 (1989)
J.D. Murray, Mathematical Biology I, An Introduction (Springer-Verlag, Berlin, 2002)
M. Fudolig, R. Howard, The local stability of a modified multi-strain SIR model for emerging viral strains. PLoS One 15(12), e0243408 (2020)
E.F. Arruda, S.S. Das, S.M. Dias, D.H. Pastore, Modelling and optimal control of multi strain epidemics, with application to COVID-19. PLoS One 16(9), e0257512 (2020)
S. Gupta, J. Swinton, R.M. Anderson, Theoretical studies of the effects of heterogeneity in the parasite population on the transmission dynamics of malaria. Proc. R. Soc. Lond. B256, 231 (1994)
Z. Feng, J.X. Velasco-Hernandez, Competitive exclusion in a vector-host model for the dengue fever. J. Math. Biol. 35, 523 (1977)
Youming Guo, L. Tingting, Modeling the transmission of second-wave COVID-19 caused by imported cases: a case study. Math. Methods Appl. Sci. (2021). https://doi.org/10.1002/mma.8041
F.J. Schwarzendahl, J. Grauer, B. Liebchen, H. Löwen, Mutation induced infection waves in diseases like COVID -19. Sci. Rep. 12, 9641 (2022)
S.A. Trigger, A.M. Ignatov, Strain-stream model of epidemic spread in application to COVID-19. MedRxiv (2022). https://doi.org/10.1101/2022.03.26.22272973
Worldometer counter. https://www.worldometers.info/coronavirus/ (2022)
A.E. Samoilov et al., Case report: change of dominant strain during dual SARS-CoV-2 infection. BMC Infect Dis. 21, 959 (2021)
K. Khan et al., Omicron infection enhances Delta antibody immunity in vaccinated persons. Nature 607, 356 (2022)
R.S. Paton, E.O. Christopher, T. Ward, The rapid replacement of the SARS-CoV-2 Delta variant by Omicron (B.1.1.529). Sci. Transl. Med. 14(652), 55 (2022)
Acknowledgements
S.T. is thankful to infectious disease physician Dr. M. Karavaeva, Prof. Dr. F. Onufrieva and colleagues from the clinic Charité (Berlin) for many useful discussions.
Author information
Authors and Affiliations
Contributions
Both authors listed have made a substantial, direct and intellectual contribution to the work, and approved it for publication.
Corresponding author
Ethics declarations
Conflict of interest
There are no any competing interests.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Trigger, S.A., Ignatov, A.M. Strain-stream model of epidemic spread in application to COVID-19. Eur. Phys. J. B 95, 194 (2022). https://doi.org/10.1140/epjb/s10051-022-00457-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epjb/s10051-022-00457-z