Abstract
Issues of the organization of albumin (a protein belonging to the superfamily of albuminoids) are considered using the example of model representatives of lower aquatic vertebrates: jawless Agnatha (Cyclostomata) and jawed bony lobe-finned (Sarcopterygii) and ray-finned (Actinopterygii: Chondrostei, Holostei, Teleostei) fish. A comparative analysis is carried out of the total length and the length of coding and noncoding sequences of the albumin gene; the number of exons, the length of introns, and the presence of repeated DNA elements in them; and the presence/absence of paralogous genes, their chromosomal affiliation, and the composition of syntenic groups. A comparative analysis of the length of the amino acid sequence, the number of domains, and S–S bonds in the albumin structure is also carried out. The issues of the emergence of the ancestral gene for albuminoids, the “expansion” and “narrowing” of the albuminoid repertoire in different groups of vertebrates, the appearance of a cluster of paralogous genes within a single chromosome in the Sarcopterygii branch and paralogous genes on different chromosomes in the Actinopterygii branch in the representatives of cartilaginous ganoids and teleost fish, the elongation of the gene and protein chain of albumin in the lamprey line, and losses of the albumin gene in the largest groups of teleost fish (Ostariophysi and Acanthopterygii) are discussed. The listed scenarios are considered in light of their consistency with the events of whole genome, local, and segment duplications. The problem of compensating for the functions of the “lost” albumin gene due to multiple osmotically active plasma proteins is discussed using the example of the group of teleost fish.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Albumin is a superfamily of albuminoids, which also includes vitamin D binding protein, alpha-fetoprotein, and afamin. Albuminoids are mainly localized in organism fluids and are involved in the transport of several important ligands. An interest in albumin is due to its key contribution to the homeostasis of physiological processes in higher vertebrates, which is achieved in the form of transport and osmotic functions. Out of all albuminoids, albumin transports the widest possible range of ligands, including inorganic cations, fatty acids, bilirubin, vitamins, hormones, peptides, proteins, and other compounds, providing plastic and energy metabolism in vertebrates (Curry et al., 1998; Otterbein et al., 2002; Jerkovic et al., 2005; Ascenzi et al., 2013; Malik et al., 2013). Having a small globule size and a high titer in the blood plasma (~60% of total protein), albumin creates its colloid osmotic pressure, maintaining the vascular volume and osmotic homeostasis of the internal environment of the organism (Dziegielewska et al., 1980; Byrnes and Gannon, 1990; Gray and Doolittle, 1992; Metcalf et al., 1998a, 1998b; Xu and Ding, 2005; Majorek et al., 2012; Anguizola et al., 2013).
Testing 183 species of amniotes, amphibians, and fish for the presence of albumin, a 100% presence of this protein was detected in first two groups of organisms, while only half of the selected species had it in fish (Li et al., 2017). For example, among teleost fish (Teleostei), albumins were found in salmon (Salmoniformes) (Nynca et al., 2017), while they were not found in Cyprinidae (Noel et al., 2010). Such facts of gene loss are associated with the events of whole genome duplications (Noel et al., 2010; Braasch et al., 2014, 2016). In the course of evolution, vertebrates experienced several large-scale and smaller duplication events affecting whole genomes or their limited regions (Ohno, 1970). All multigenic families, including albuminoids, were formed under the influence of these processes (Hoegg et al., 2004; Noel et al., 2010; Ozernyuk and Myuge, 2013; Soshnikova et al., 2013; Braasch et al., 2014, 2016). Simultaneously with the duplication of genes and genomes, “narrowing” of the repertoire in the form of the loss of individual genes occurred in some multigenic families (Braasch et al., 2016; Pasquier et al., 2016). The loss of the albumin gene in the Cyprinidae fish family is an example of the consequences of genomic rearrangements. However, there is still no clarity on the extent of this loss in Teleostei: is it limited to cyprinids or does it cover a wider range of orders?
This review considers several scenarios of evolutionary transformations of albumin in the composition of the albuminoid superfamily using the example of model representative of lower aquatic vertebrates, including jawless Agnatha and jawed bony fish (Gnathostomata: Osteichthyes). The association of these scenarios with the events of whole genome, local, and segment duplications, as well as the problem of albumin gene loss and the compensation for this loss in the largest groups of teleost fish (Ostariophysi and Acanthopterygii), is discussed.
MATERIALS AND METHODS
To search for albumins, 26 species were selected belonging to 20 orders of jawless Agnatha (Cyclostomata, 1 order) and bony jawed fish (19 orders), including lobe-finned and ray-finned (Gnathostomata: Sarcopterygii, Actinopterygii) (Table 1). When selecting objects, we used information about the presence of genomes in DB Genomes NCBI (Genome) and whole genome sequencing (BioProject), as well as the list of model objects from a review (Ravi and Venkatesh, 2018) with BioProject accession identifiers for 60 species of ray-finned fish.
The taxonomy and Latin specific names are given according to “Annotated Catalog of Cyclostomes and Fishes…” (Reshetnikov, 1998). For the rainbow trout, the species name Parasalmo mykiss (Reshetnikov, 1998) and Oncorhynchus mykiss was used. The latter variant (O. mykiss) is used in Proteins, Genes (NCBI), and KEGG databases.
For teleost fish, an extended list of model objects was formed, which includes (1) lower Teleostei from two families (Salmonidae, Esocidae) of the order Salmoniformes, (2) Ostariophysi from five orders (Cypriniformes, Gymnotiformes, Gonorynchiformes, Characiformes, and Siluriformes), and (3) higher teleost Acanthopterygii from ten orders (Gobiiformes, Mugiliformes, Cichliformes, Atheriniformes, Cyprinodontiformes, Carangiformes, Pleuronectiformes, Labriformes, Perciformes, and Tetraodontiformes). Mammals (Homo sapiens) were used as a reference group (Table 1).
Information about amino acid sequences, domain organization, and S–S bonds of the albumin and other albuminoids of tested species were obtained from DB Proteins NCBI. The complete sequences were used for comparison (albumin fragments of lake char and lamprey were an exclusion). Information about the organization of albumin genes (exon–intron structure; complete length; length of coding and non-coding sequences; and paralogous genes, their chromosomal affiliation, and syntenic groups) was obtained from DB Genes and Genome Data Viewer (GDV) NCBI. The SmartBLAST and/or KEGG web resource was used to search for closely related sequence matches in the form of the five best matches and an extended list of homologs, multiple alignment, and quantification of sequence similarity. The alignment and comparison of intron sequences (similar in position in the structure of albumin genes) on the presence of short repeated DNA sequences in them in the tested species was carried out using a MEGA 6 sequence (Tamura et al., 2013).
The names of albuminoid proteins/genes and designations of the parameters of their organization are given according to DB Genes and Proteins NCBI: for albumin and vitamin D binding protein in lower vertebrates, ALB/alb and DBP/dbp, respectively; for albumin, alpha-fetoprotein, afamin, and vitamin D binding protein in humans, ALB/ALB, AFP/AFP, AFM/AFM, and DBP/DBP, respectively; CDS, coding sequence; SINEs, short interspersed nuclear element; LINEs, long interspersed nuclear element; D, domain; L, a.a.r., length of the protein chain in amino acid residues; and L, nt, length of DNA sequences in nucleotides.
Names of the events of whole genome, taxon-specific, and other duplications are given in accordance with the designations accepted in the literature: WGD, whole genome duplication; AsGD, acipenseric-specific WGD; TGD, teleost WGD; CyGD, cyprinidae WGD; SaGD, salmonidae WGD; SD, segment duplication; LD, local duplication; 1R, 2R, 3R, WGD rounds (Ohno, 1970; Freeling and Thomas, 2006; Braash et al., 2016).
Whole Genome, Local, and Segment Duplications in Vertebrates
Most biologists consider the duplication of genes and genomes with subsequent divergence as a source of the evolutionary process, which stimulated the emergence and radiation of large taxa of animals and plants, for example, vertebrates and angiosperms (Freeling and Thomas, 2006). Duplications of genes and genomes are reached using duplications of different scales (WGD, SD, and LD). With WGD, extended DNA segments (ohnologs) are amplified. With SD, less extended genomic segments are amplified under the influence of chromosome rearrangements; during interchromosomal rearrangements, they can be on different chromosomes. LDs usually occur as tandem repeats, as a result of which a cluster of related genes located one after another appears on the chromosome. In all these cases, a duplication of the genome, group of genes, or individual genes occurs, as a result of which homologous copies are obtained from the original (ancestral) genes. Homologous genes in the same genome that occurred by duplication of the ancestral gene are paralogs; homologous genes in the genomes of different organisms occurred as a result of evolution of the same ancestral gene, which was in the genome of the common ancestor of these organisms, are orthologs (Ravi and Venkatesh, 2018).
In 1970, Susumu Ohno (Ohno, 1970)) put forward a 2R hypothesis according to which the ancestral genome of vertebrates went through two rounds of whole genome duplications prior to their diversification (1R (WGD1) and 2R (WGD2)). Later, jawed fish went through the third round of whole genome duplication (3R, WGD3), which gave rise to the Teleostei taxon; for this reason, it was designated as TGD (Ohno, 1970; Noеl et al., 2010; Braasch et al., 2016; Pasquier et al., 2016). Among tetrapods, genome duplications occurred in amphibians and reptiles (Sauria); in birds and mammals, no resistant polyploid forms were found (Vasil’ev, 1985; Uno et al., 2013; Evans et al., 2017; Ravi and Venkatesh, 2018) (Fig. 1).
Scheme of vertebrate evolution with a mark of the main rounds of whole genome duplications (WGD1, WGD2, WGD3) relative to the stages of Vertebrata divergence into Agnatha and Gnatostomata and the divergence of the latter into Sarcopterygii and Actinopterygii. Esoc, Esocidae; Salm, Salmonidae. Duplications in individual orders in the composition of taxa are noted with small asterisks. Explanations are in the text.
Most authors consider that first two rounds of duplications took place before the separation of Agnatha and Gnathostomata (~500–550 million years ago) (Holland et al., 1994; Kuraku et al., 2009a, 2009b; Van de Peer et al., 2017). Another point of view supports the scenario in which the first round is assigned to the period before the divergence of Agnatha and Gnathostomata, while the second round to the period after their divergence (Nakatani et al., 2021). The third round of duplications occurred presumably ~230–400 (Venkatachalam et al., 2017) or ~300–450 million years ago (Taylor et al., 2001).
The analysis of genomes in cyclostomes indicates a high probability of polyploidization events (hexaploidization) independent of Gnathostomata in the lamprey line (Smith and Keinath, 2015; Nakatani et al., 2021). Independent events of polyploidization occurred in Acipenseriformes in the Acipenseridae and Polyodontidae families (Crow et al., 2012; Du et al., 2020; Cheng et al., 2021). In the branch of ray-finned fish after the TGD event and appearance of Teleostei taxon, the genome duplications occurred in cyprinids (СyGD) (Ma et al., 2014) and salmon fish (SaGD) (Alexandrou et al., 2013; Berthelot et al., 2014; Zhivotovskii, 2015). Polyploids were also found in some orders of Acanthopterygii, for example, in Labriformes, Anabantiformes, and some others (Vasil’ev, 1985; Felip et al., 2009; Cioffi et al., 2015) (Fig. 1).
In addition to 2R hypothesis, models considering the contribution of both whole genome and local and segment duplications in the evolutionary process were proposed (Freeling and Thomas, 2006). For example, the model 1R + SD takes into account the contribution of multiple segment duplications in genomic rearrangements of lampreys before the event 1R and after it (Asrar et al., 2013; Smith and Keinath, 2015; Nakatani et al., 2021). It is assumed that both whole genome duplications TGD and SaGD and SD contributed to the emergence of Salmonidae (Vasil’ev, 1977; Zhivotovskii, 2015).
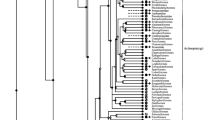
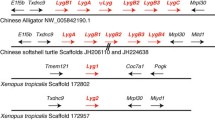
Organization of Albuminoid Genes on Chromosomes
Among vertebrates, only amniotes have a complete set of four albuminoid genes. In mammals, all four genes (DBP, ALB, AFP, and AFM) are located on the same chromosome (Fig. 2). In jawed fish, the genes of two albuminoids (alb and dbp) were found. Agnatha have only a single albuminoid (albumin); they have no dbp gene (Fanali et al., 2012). Like mammals, the alb and dbp genes in gar pike and sterlet are located on the same chromosome in the opposite transcriptional orientation (Fig. 2b). In lower teleost fish, 2–3 albumin paralogous genes were found: 2 genes in Northern pike and 3 in Salmonidae. Moreover, the alb and dbp genes can be located on the same or different chromosomes. In pike, the paralogous genes are located in two different chromosomes. In rainbow trout, 3 albumin paralogous genes are located on chromosomes 1, 11, and 23; both albumin and dbp genes are located on the chromosome 11; only albumin genes in the chromosomes 1 and 23.
In mammals and fish, syntenic groups of genes including albumin are different. They do not match in the gar pike, sterlet, or Salmonidae. At the same time, two stable syntenic groups that include albumin paralogous genes were found in the Salmoniformes order (Figs. 2c, 2d). In Northern pike (which is considered the species closest to the common ancestors of Salmonidae) (Rondeau et al., 2014), the syntenic group on chromosome 13 includes (taking into account albumin) five genes: E3 ubiquitin protein lipase K CMF1 like–solute carrier family 20 member–albumin– methylenetetrahydrofolate dehydrogenase (NADP + dependent) 2 like–epithelial mitogen (epigen). Another syntenic group (also out of five genes) on chromosome 5 includes other genes: poly(A) polymerase gamma–retinal homeobox protein Rx1-like– albumin–v-rel avian reticuloendotheliosis vital oncogene homolog–peroxisomal biogenesis factor 13 (Fig. 2c).
In the Atlantic salmon, the syntenic group on the chromosome ssa15 matches (except for one gene) the sequence of pike genes on chromosome 13, but only in reverse order: epigen–methylenetetrahydrofolate dehydrogenase (NADP + dependent) 2 like–albumin–sodium dependeter phosphate transporter 2– Un–E3 ubiquitin protein lipase K CMF1 like (Fig. 2d). Another syntenic group on an unidentified salmon chromosome (Un) completely matches the order of genes in the pike on chromosome 5 (Fig. 2d).
Both syntenic groups are found on different chromosomes in the brown trout Salmo trutta, lake trout Salvelinus namaycush, and rainbow trout Oncorhynchus mykiss in the form of complete or truncated variants. At the same time, the examples of the transformation of syntenic groups as a result of inversions of single genes, several genes, or complete linkage groups are noted in each case.
Exon–Intron Organization of Albumin Genes
The length of the coding sequence (CDS) of albumin gene in Gnathostomata is relatively constant (Table 2). In all tested species, it is organized in the form of 14–16 exons; albumin of the lake trout Salvelinus namaycush on chromosome 4 has a minimal number of exons (13). Except for two extreme exons, the rest encode a polypeptide chain out of three domains. The lengths of the complete gene sequences vary significantly due to the different length of introns (noncoding DNA regions) (Tables 2, 3; Fig. 3). In the lamprey, all listed indices exceed those in jawed fish.
Graphical representation of exon–intron organization of albumin genes in sea lamprey (Petromyzon marinus), coelacanths (Latimeria chalumnae), sterlet (Acipenser ruthenus), gar pike (Lepisosteus oculatus), Atlantic salmon (Salmo salar), northern pike (Esox luceus), and humans (Homo sapiens). Exons are presented in the form of dark vertical boxes of different thicknesses depending on their length; introns are in the form of horizontal lines between them. D1–D3, domains. Graphical data of DB Genes NCBI were used.
The comparison of lengths of the appropriate introns in albumin genes of jawed fish and human found no matches (Table 3). Most of those presented in DB NCBI intron sequences start with GT dinucleotide and end with AG (splicing sites); we note that these sites are absent in a number of presented sequences (introns between the second and third exons of albumin gene in northern pike Esox lucius on chromosome 5 and albumin gene in gar pike Lepisosteus oculatus).
The comparison of repeated elements in the structure of albumin gene introns in fish and humans located between the exons 2 and 3 and highlighted in pink in Table 3 revealed no matches. There are 11 SINEs and 6 LINEs (short and long dispersed DNA repeats) in human albumin gene introns. They do not follow each other in a linear order, but are scattered throughout the sequence (Nishio and Dugaiczyk, 1996; Nishio et al., 1996). Among SINEs, primate-specific Alu repeats, named after the Alu restriction endonuclease, whose DNA processing led to their discovery, are best known: aattgcttgtgtttctt, gatgtatg, gaatactcattcat, and others. Four Alu sequences were detected in the human albumin gene.
In fish, other short repeats were found in the composition of albumin gene introns. Thus, the intron, located between exons 1 and 2, contains tg repeats of different lengths (from 40 to 124 bp in salmonids and shorter in coelacanth). The longest second intron (7411 bp) in the salmon albumin gene on the chromosome ssa15 contains 13 copies of agagtatggc short repeats, 10 copies of ccttggtaat, and 11 copies of aaccaggtcc. The listed sequences are absent in the paralogous gene of salmon albumin, as well as in albumin genes of rainbow trout, coelacanth, and humans.
The promoter region of the albumin gene in tested species is presented in DB NCBI only by the sequence in Atlantic salmon. It consists of 410 bp (Salmo salar albumin promoter region: GenBank X79487.1). In humans, the promoter region was studied in detail (Kajiyama et al., 2006). It consists of 406 bp (albumin (ALB) 5' regulatory region; RefSeq NC_000004.12). The albumin gene promoters contain ТАТА motifs both in salmon and in humans. We note that, in many vertebrates, the promoters of a number of genes can contain other initiating elements (Carninci et al., 2006). The promoter motif TATA box in the salmon albumin gene consists of six nucleotides (TATAAA) and, in humans, seven (ТАТАААА). Regulatory regions of salmon and human albumin gene are homologous, which is confirmed by a specific binding of salmon HNF1 transcription factor by promoter regions of albumin genes in fish and mammals (Deryckere et al., 1995).
Organization of Albumin Polypeptide Chain
The polypeptide chain of albumin and other albuminoids consists of three albumin domains that are structural and functional units (Li et al., 2017). The domains contain binding sites of various endogenous and exogenous ligands, including seven fatty acid binding sites (Ghuman et al., 2005). Each domain contains ~190 amino acid residues (a.a.r.). In mammals, each domain is stabilized by 5–6 S–S bonds; the complete polypeptide chain has 17 S–S bonds (Saber et al., 1977). Spiralized regions of the polypeptide chain give it a conformational flexibility, while S–S bonds maintain the required rigidity of the spatial structure. Like mammals, fish retain a conservative three-domain organization of albumins, as well as a conservative amount of amino acid residues and S–S bonds. In lampreys, all indices increase proportionally (Table 2, Fig. 4).
Graphical image of albumin domains (D) of polypeptide chain of human albumin (HSA) (RefSeq NP_000468.1) (a) and Japanese lamprey Lеthenteron camtschaticum (GenBank BAF47283.1) (b); positions of cysteines (С77–С193) and S–S bonds in the domain (D1) HSA (RefSeq NP_000468.1) (c). Explanations are in the text.
The conservatism of the listed parameters is maintained with a relatively high indices of the similarity of amino acid sequences at intragenus and intraorder levels. At the same time, the similarity of sequences in the representatives of different orders is minimal. Thus, the similarity of SDS-1 albumin in sea lamprey with albumins of the same species and Japanese lamprey reaches ~77–99%, while with mammal albumins it is only ~22–24%. The similarity of sterlet albumin (612 a.a.r.) with albumins of Acipenseriformes is ~81–99%; with albumins of Lepisosteidae, Polypteriformes, and Salmonidae, it is ~58, ~49–51, and ~43%, respectively; with AFP and AFM of amniotes, it is ~28–35%; and with DBP of vertebrates, it is <28%. The same pattern is traced for the Atlantic salmon albumin: a high similarity of sequences at intragenus (~78–99%) and intraorder (~64–93%) levels, lower at the interorder level (~41–49% with albumins of gar pike and Acipenseriformes), and minimal with albuminoids of amniotes (~23–27%) and DBP (~20–22%).
For gar pike, a maximal similarity (93%) was also noted at an intraorder level with the albumin of alligator gar Atractosteus spatula; lower indices were noted with albumins of Acipenseriformes (~58–60%) and Salmoniformes (~43–49%); and there was a minimal similarity with albumins of mammals (~31–33%), AFP and AFM of amniotes (~26–32%), and DBP (~21–22%).
The albumins of sea lamprey have a special status due to the large length of the polypeptide chain (1423 a.a.r.), the presence of seven albumin domains, and the number of S–S bonds proportional to the protein size (41), in general, retaining the similarity with albuminoids of Gnathostomata. The high level of similarity of albumin sequences in lampreys at an intraorder level as compared with albumins of other vertebrates is probably explained by their independent origin from one- or two-domain precursor, unlike albumins of other vertebrates starting from three-domain precursor (Gray and Doolittle, 1992). The homology of amino acid sequences of albumins in lampreys and albuminoids in Gnathostomata is preserved throughout the lamprey amino acid chain (Fig. 5).
Multiple alignment of the albumin sequence of Japanese river lamprey Lethenteron camtschaticum (a) and sea lamprey Petromyzon marinus (b) with the five closest homologs, albuminoids ALB, VDB, AFP, and AFM in human, mouse Mus musculus, and zebrafish Danio rerio. The scale of the length of amino acid sequences is on the top, their length is on the right, and the ID identifier is indicated near the name of the protein and species.
Evolutionary Transformations of Albumin and Their Association with Genome/Gene Duplication
Three-domain albuminoids originated from a single-domain precursor. Initially, this conclusion was made based on the analysis of amino acid sequence of bovine serum albumin and, first and foremost, by the location of S–S bridges characterized by a 3-fold repetition in the amino acid sequence (Brown, 1976). Later it became clear that the sequences of albuminoids of other mammal species had a similar architecture of S–S bonds and three-domain structure (Gorin et al., 1981; Jagodzinski et al., 1981; Morinaga et al., 1983; Minghetti et al., 1985, etc.).
The origin of albuminoids was given by an ancient gene encoding a “semi-domain” protein (½D) (Gibbs and Dugaiczyk, 1987). It had five exons in its structure. The gene with seven exons encoding single-domain (1D) protein originated from it. Based on it, the gene encoding a two-domain protein and containing 11 exons appeared, followed by an ancestral gene from 15 exons encoding a three-domain protein (albuminoid precursor, 3D). The elongation of the ancient gene occurred as a result of two events of nonhomologous recombination (unequal crossing over) and one event of homologous recombination. It is assumed that the ancestral gene was duplicated, giving rise to two paralog protogens (proto-ALB and proto-DBP). Proto-ALB evolved into ALB genes, which gave rise to AFP and AFM genes, while vitamin D binding protein genes in vertebrates originated from proto-DBP (Eiferman et al., 1981; Ohno, 1981; Sargent et al., 1981; Gibbs and Dugaiczyk, 1987; Gray and Doolittle, 1992; Noel et al., 2010). The DBP gene at initial stages of evolution lost two exons (the 12th and 13th) (Nishio and Dugaiczyk, 1996; Malik et al., 2013). DBP was not found in lampreys (Gray and Doolittle, 1992); however, it is in almost all other groups of vertebrates, and it preserved its structure well over ~500 million years of evolution (Bouillon et al., 2020). All vertebrates have the albumin gene except for some teleost fish; only amniotes have AFP and AFM (Noel et al., 2010).
The events of gene duplication that resulted in a three-domain structure (common for albuminoids) presumably occurred between the branching of lampreys from the trunk of vertebrates (~450 million years ago) and the emergence of DBP (Gray and Doolittle, 1992). The albumin of lampreys most likely occurred as a result of the duplication of the gene encoding one- or two-domain protein regardless of the events leading to three-domain albumin and before the emergence of the DBP gene (Gray and Doolittle, 1992). The elongation of the albumin polypeptide chain to seven domains occurred in the line of lampreys. In analogy to the Gibbs and Dugaiczyk model (Gibbs and Dugaiczyk, 1987), it can be assumed that this elongation could be a result of nonhomologous recombinations in the form of unequal crossing over. Literature data indicate the presence of two albumins in lampreys (AS and SDS-1) presumably encoded by paralogous genes (Filosa et al., 1998). Recent studies indicate a high probability of polyploidization events independent of Gnathostomata in the branch of lampreys in the form of hexaploidization and segment duplications (Smith and Keinath, 2015; Nakatani et al., 2021), which indirectly supports the point of view that lampreys can have paralogous genes for albumin.
After the 2R even, the evolution of albumin on the basis of three-domain precursor in different branches of Gnathostomata occurred according to different scenarios. The coelacanth (Sarcopterygii) and gar pike (Holostei) had a diploid genome typical for vertebrates similar to the human genome (Ravi and Venkatesh, 2018); their albumins are determined by a single gene. In Tetrapoda (Sarcopterygii), duplication events occurred that affected albuminoids. Thus, a whole genome duplication occurred ~40 million years ago in amphibians, as a result of which frogs have albumin genes on homeologous chromosomes. In amniotes, polyploids appeared in the group Sauria (lizards); no stable polyploids were found in birds and Mammalia (Vasil’ev, 1985). Meanwhile, all amniotes have AFP and AFM genes absent in other Vertebrata. They appeared presumably as a result of the local (tandem) duplication of the ancestral gene of albuminoids and the appearance of the ancestral AFP/AFM gene (Noel et al., 2010). Another point of view considers AFM an intermediate component between ALB and AFP (Fasano et al., 2017). Thus, the evolution of albumin in tetrapods proceeded in the following directions: (a) the appearance of multiple copies of albumin gene in the composition of homeologous chromosomes in allotetraploids (amphibians) and (b) tandem duplication of albumin gene and appearance of paralogous AFP and AFM genes, as a result of which the cluster ALB/AFP/AFM occurred, from which the DBP gene (amniotes) is located at a distance of 1.5 Mb. The appearance of AFP/AFM probably occurred after 2R and the divergence of amphibians and reptiles ~320 million years ago (Noel et al., 2010).
In another branch of Gnathostomata (Actinopterygii), two copies of the albumin gene, which appeared probably as a result of genome duplication specific to Acipenseriformes, were detected in the Acipenseriformes order in the sterlet. A TGD event, which gave rise to the taxon Teleostei and stimulated the diversification of this group, occurred on the time interval ~230–450 million years ago (simultaneously with the divergence of amphibians and reptiles) in the Actinopterygii branch. As a result of the reorganization of the genomes, the largest groups of teleosts (Acanthopterygii and Ostariophysi) lost the albumin gene, while lower teleosts (Salmonidae and Esocidae) received additional copies of this gene.
Based on the data, a scheme of evolution of albumin in the groups Agnatha and Gnathostomata is proposed (Fig. 6), supplemented with the events of the whole genome and segment duplications, recombinations, and duplications of the albumin gene (including in tetrapods in the group Amniota).
Scheme of evolutionary transformations of the precursor of albuminoids and albumin under the influence of WGD: in Agnatha (a) and Gnathostomata before the TGD event (b) and after it (c). The scheme is complemented by the events of segment and local (tandem) duplications, as well as by the event of albumin gene loss. Explanations are in the text.
Evolutionary Fate of Albumin Paralogous Genes after Duplication Events
Susumo Ohno (1970) suggested that, most frequently, one of the two paralogous genes loses its functionality after duplication. The accumulation of harmful mutations that lead to degeneration, pseudogenization, and the nonfunctional state of the gene can be one reason for the loss. Pseudogenes are characterized by a certain degree of homology with functional genes and changes in organization, preventing their normal transcription and translation. Most copies lose their functionality during rediploidization following the duplication event. The analysis of TGD consequences on the example of multigenic Teleostei families indicates a high percentage of the loss of their genes after the duplication of genomes (Brunet et al., 2006; Braash et al., 2016). Thus, ~20% of all duplicated genes in Danio rerio are retained; the rest lose their functionality (Postlethwait et al., 2000) and, in general, this index reaches ~17% for the group of teleosts (Braasch and Postlethwait, 2012). The absence of the albumin gene in Acanthopterygii and Ostariophysi fish is also probably an example of the loss of the gene as a result of TGD. This event is not related to the absence of DBP in lampreys. In the first case, we are talking about a possible loss of albumin genes, since albumins were in all groups of bony Gnathostomata before the TGD event. In the second case, the absence of DBP is explained by the fact that the branching of Agnatha occurred before the emergence of the three-domain precursor of all albuminoids, and hence before the appearance of proto-DBP.
At the same time, some groups of fish (for example, Salmonidae) demonstrate the inclusion of almost half of all duplicated genes in the processes of sub- and neo-functionalization (Berthelot et al., 2014). These two scenarios are achieved in the evolution of paralogous genes that are preserved after duplication events. During subfunctionalization, the functions of the original gene are divided between paralogs. A duplication–degeneration–complementation (DDC) model (Force et al., 1999) assumes a possible scenario of distribution of the functions of ancestral gene between two paralogs. According to it, the accumulation of mutations occurs in both copies. This leads to a decrease in their functionality, as a result of which both copies together reproduce the effect of a single (original) gene. If mutations concern regulatory regions, then this will be reflected in the expression patterns of paralogs. If mutations concern protein-coding regions, then they can lead to the distribution of functions of the original gene between its daughter copies. The examples of these scenarios were described in a number of experimental works and reviews (Ozernyuk and Myuge, 2013; Kamenskaya and Brykov, 2020; Bayramov et al., 2021; Gu and Xia, 2019; etc.).
Trying on the listed scenarios for albuminoids, we note that initially three-domain ancestral protein encoded by the ancestral gene probably performed a transport function. Its duplication led to the appearance of two paralogs (proto-ALB and proto-DBP) that gave rise to albumin and vitamin D binding protein genes. Their subsequent subfunctionalization could occur in the form of the specialization of each of them in the transport of a certain range of ligands: wide (in the case of albumin) and narrow (in the case of DBP). It is possible that the narrowing of the functionality of DBP occurred due to the loss of two exons by it in the early stages of evolution. The acquisition by one of the two paralogs (namely, albumin) of a new (osmotic) function could be an example of neofunctionalization. This could occur due to an increase in the level of expression of the gene encoding it (probably due to mutations in its regulatory region). A high expression provided a high albumin titer in the plasma, which turned out to be sufficient to create a colloid–osmotic pressure of the plasma. Since only albumin has the highest titer in the plasma of vertebrates (among all albuminoids), it was fixed by selection as an ideal protein factor controlling the homeostasis of extracellular fluid in the organism of vertebrates.
Another example of subfunctionalization concerns albumin in lampreys and tetrapods. The paralogs of albumin gene SDS-1 and AS divided the function of ligand transport between different stages of ontogenesis (Filosa et al., 1998), like ALB and AFP paralogous genes in mammals that divided the function of transport between the embryonic and postnatal stages of development.
One more example of sub- and neofunctionalization is demonstrated by Salmonidae, the genomes of which underwent whole genome and segment duplications and experienced at least six multiple chromosome fusion events (Makhrov, 2017). Almost half of ancestral genes (48%) were preserved in Salmonidae as copies, a significant part of which has divergent profiles (levels) of expression (Berthelot et al., 2014). This circumstance indicates a high probability of mutations in the regulatory regions of paralogs and that almost half of the paralogous genes in Salmonidae are involved in the processes of sub- and neofunctionalization. It is possible that albumin paralogous genes are also involved in these processes.
After duplication events, genomes go through a period of rediploidization or a return to a diploid state (Ravi and Venkatesh, 2018). This probably explains the reason for the discrepancy in the numbers of real and expected paralogous genes in polyploids. It was established (Ramberg et al., 2021) that the average number of paralogs for a number of genes encoding the enzymatic proteins is three in Salmo salar. The same number of paralogs was also established for albumins in S. salar. Apparently, this number is more than two due to the fact that Salmonidae went through (in addition to 1R and 2R) two whole genome duplications (TGD and SaGD). At the same time, it is less than the expected value of four, probably due to the process of rediploidization of the polyploid genome, which Salmonidae went through in the late Cretaceous and Eocene (Gundappa et al., 2021). Meanwhile, only two, but not three, albumin paralogous genes are found in northern pike, which is probably explained by only one (in addition to 1R and 2R) duplication event in the form of TGD in the evolution of Esocidae.
After the given examples of one or another strategy in the evolution of paralogous genes, a natural question arises: what factors determine its choice?
Factors Affecting the Evolutionary Fate of Paralogous Genes
The evolutionary strategy of paralogous genes depends on the way of duplication (WGD or SD) and type of amplified DNA fragments (Freeling and Thomas, 2006). In whole genome duplications, DNA is copied as long copies (ohnologs) uniting multiple groups of genes. In this form, the genes involved in the processes of regulation of ontogenesis, oogenesis, cell cycle, signaling cascades, and others are preserved. In the composition of ohnologs, the genes are often combined into so-called “functional modules.” The proteins encoded by the genes of such modules are, as a rule, organized in supramolecular functional complexes. It is considered that the duplication of such modules leads to a jump in the adaptability of organisms. At the same time, individual genes that are not combined into functional modules are preserved in the form of short DNA copies under segment duplications. Such a method of amplification does not lead to a jump in adaptability, but makes a certain contribution to the modulation of biological processes. The genes responsible for the conservative functions, for example, DNA metabolism, activity of nucleases, and others, are preserved in the form of short copies (Bowers et al., 2003; Papp et al., 2003; Yang et al., 2003; Maere et al., 2005; Freeling and Thomas, 2006). The ideas about the role of gene balance in the composition of long and short DNA copies are based on the fact that the balance is observed in ohnologs, while local duplications (for example, tandem), due to a risk of imbalance, preferably cover the genes encoding monomeric proteins or the genes weakly bound on the chromosome (Freeling and Thomas, 2006).
Returning to albumins of Salmoniformes (Esocidae and Salmonidae fish), we note that there is no information whether they are included or not included in functional modules. However, it was found that they are stably present in the composition of two syntenic groups out of five–six genes. One such “pike” group of genes on chromosome 5 is completely reproduced on one of the chromosomes of Atlantic salmon, brown trout, and rainbow trout. Another pike group of genes on chromosome 13 is completely reproduced on chromosome 18 of the lake char and in the form of an inverted gene sequence on salmon chromosome ssa15; at the same time, one of the five pike genes in salmon is “lost” and replaced by another. Inversions of individual or several genes from these two syntenic groups of the pike, substitutions of one gene for another, also occur in other Salmoniformes. Neither polyploidization nor the intense chromosomal rearrangements typical for Salmoniformes (which were also found in the precursors of teleosts) (Braash et al., 2016) “broke” these groups, but only rearranged them.
A comparative analysis of the genomes of teleost fish, gar pike, Australian ghostshark Callorhinchus milii, and other Gnathostomata demonstrated that short conservative syntenic groups or blocks are typical for teleosts (Ravi and Venkatesh, 2018). For single genes encoding monomeric proteins not organized in clusters in the form of tandem repeats and not included in the composition of functional modules, another regularity was established: they are lost more often than related genes in the composition of functional modules of ohnologs (Freeling and Thomas, 2006). It is possible that a similar scenario for the loss of the albumin gene could be realized in the groups Ostariophysi and Acanthopterygii.
Due to WGD, teleost fish made a global jump in evolutionary development, while a “finer” adjustment of the mechanisms aimed at the maintenance of a balanced work of the doubled genome (and, ultimately, on the homeostasis of physiological functions) can be reached due to segment duplications and the duplications of individual genes.
Problem of Compensating Albumin Function in the Groups Acanthopterygii and Ostariophysi
Taking into account the role of serum albumin as one of the key factors of physiological homeostasis in higher vertebrates, it could be assumed that its loss in the representatives of lower vertebrates should have unfavorable consequences. However, Acanthopterygii and Ostariophysi belong to the largest and most evolutionary advanced groups of vertebrates; from this, it follows that some other plasma proteins took over the osmotic and transport functions of albumin in them.
The analysis of the protein composition of the blood plasma in teleost fish, including Acanthopterygii and Ostariophysi, revealed a wide spectrum of proteins with a high electronegative potential capable of binding inorganic cations and water dipoles and thus exhibiting osmotic activity. It was found that not one, but several proteins in the composition of the osmotically active protein fraction are present in the blood plasma of albumin-containing and “albumin-free” teleost fish: (1) albumin, (2) warm temperature acclimated 65 kDa protein exhibiting the properties of hemopexin, (3) inhibitors of serine and cysteine proteinases, (4) apolipoproteins ApoA-I and Apo-14 in the composition of high-density lipoproteins, and (5) multiple additional anode fraction out of low molecular weight proteins with highly negative surface potential (Andreeva, 2019, 2020, 2021). The latter has not only “true” plasma proteins (extracellular proteins with specific functions in the circulatory system), but also so called “transit” proteins, whose pool is constantly filled with intracellular proteins entering the blood due to cell destruction (Andreeva, 2021).
The portion of osmotically active plasma proteins in teleost fish from the abovementioned groups reaches ~50–60% of the total protein, which is comparable with the content of albumin in mammalian plasma. This suggests that the contribution of these proteins in a colloid–osmotic pressure of the plasma in “albumin-free” teleost fish is comparable with Mammalia. The transport function of blood plasma was not affected in Acanthopterygii and Ostariophysi. Thus, the comparison of fatty acid transport functions between albumins and high density lipoproteins suggests their more efficient implementation with the involvement of lipoprotein particles (Andreeva, 2019).
Reasons for the Evolutionary Success of Polyploids Using the Example of Teleostei, or Why the Loss of Albumin Was Unnoticed in Teleost Fish
The main changes that emerged in Teleostei when compared with more ancient ray-finned fish (and apparently determined their evolutionary success) affected the mechanisms of locomotion, nutrition, and fertility (Romer and Parsons, 1986). Teleostei differ from other vertebrates by the highest diversity of body shape, way of life, type of nutrition, and high fertility. Since the genomes of the representative of teleost fish passed through an additional round of WGD and taxon-specific events SaGD, CyGD, etc., the progressive traits of this group are largely associated with genome duplication events. Among duplicated genes, HOX genes occupy a special place. After the separation of Acipenseriformes and Semionotiformes, the duplication of four clusters of HOX genes occurred in the precursors of teleosts. It is assumed that it is this event that affected shape-building processes; organism morphology; and caused the radiation of the ancestral group, which gave rise to a new taxon (Ozernyuk and Myuge, 2013; Soshnikova et al., 2013).
The genes encoding transcription factors are another group of genes whose duplications had important consequences for the evolution of Teleostei. They are represented by different multigenic families (Sox, Bmp, Gtf3a, Hif, HNF1, etc.) involved in the regulation of the expression of hundreds of other genes; signaling cascades; and control of the processes of embryogenesis, ontogenesis, oogenesis, cell cycle, etc. (Ozernyuk and Myuge, 2013; Pelster and Egg, 2018; Rojo-Bartolome et al., 2020). The genes encoding proteins of the intermediate metabolism undoubtedly contributed to the evolutionary success of Teleostei. First and foremost, these are proteins involved in lipid metabolism, which is the basis of the energy of fish. A special role is given to transport proteins responsible for providing tissue metabolic processes with all required ligands, including albuminoids (Sharma et al., 2006).
The state of rRNA gene clusters is important for the survival of polyploid organisms, since the expression of duplicated genes is limited by a protein-synthesizing ability of ribosomes depending on the efficiency of replenishment of pools of different rRNA fractions. No clear correlation was detected when comparing the expression of the clusters of these genes in allotetraploid (4n = 200) and diploid (2n = 100) goldfish Carassius auratus, although differences in the expression of 5S and 45S rRNA were noted (Zhao et al., 2021). The effect of duplication events and chromosome rearrangements on redistribution, accumulation, and homeostasis of rRNA, providing the required level of expression of in-demand duplicated genes, was demonstrated using the example of Danio rerio and polyploid Perciformes (Cioffi et al., 2015; Rojo-Bartolome et al., 2020). Unlike teleost fish and other eukaryotes, the multiplicity of alleles of the 18S rRNA gene (associated presumably with the polyploidization of this group) was demonstrated in the Acipenseridae family using the example of nine species of North American Acipenseridae (Krieger et al., 2000, 2006; Krieger and Fuerst, 2002). A significant individual variation of 18S rRNA gene is absent in most other species, for which its sequences were established. The authors suggest that a high polymorphism of these sequences in Acipenseridae can be associated with a low rate of coordinated evolution in this group, slowing down the loss of any resulting polymorphic variation. This circumstance allows us to explain the absence of similar variability in teleost fish by the rapid pace of evolutionary transformation in this group (Ravi and Venkatesh, 2018).
CONCLUSIONS
This search suggests differences in the scenarios of the evolution of albumin genes in jawless and jawed vertebrates. In the line of lampreys, the evolution of albumin occurred under the influence of one or two rounds of WGD and independent events of hexaploidization and segment duplications. Taking the origin not from the three-domain precursor, but from shorter one- or two-domain proteins, albumins of lampreys reached a two-fold superiority in the length of the amino acid sequence over albumins of Gnathostomata. Two paralogous genes, sharing the function of ligand transport in juvenile and mature lamprey individuals, are very similar to paralogs (albumin and alpha-fetoprotein) in amniotes sharing the transport function of the original gene between the embryonic and postnatal stages. Jawed fish unaffected by the third whole genome duplication (coelacanth and gar spotted pike) have, like mammals, one albumin gene. At the same time, two albumin paralogous genes were found in sterlet, in the evolution of which, in addition to the first rounds of WGD, whole genome duplication specific for Aciprnseriformes was present. In the group of teleost fish, the third round of WGD and taxon-specific whole genome duplications led, on the one hand, to the appearance of albumin paralogous genes in lower Teleostei and, on the other, to the loss of albumin in the largest and evolutionary advanced Acanthopterygii and Ostariophysi groups; as a result, they had a “narrowing” of the albuminoid repertoire to a single protein (DBP).
The presence of sequenced genomes in the absence of annotations or weak support in the form of sequence annotation of some analyzed species do not allow one to consider the search results final. However, the extensive literature data on the identification of plasma proteins and tissue filtrates in model Acanthopterygii and Ostariophysi species assure the absence of albumins in the composition of plasma proteomes (Dietrich et al., 2014, 2021; Vilchez et al., 2016; Banerjee et al., 2017; Schrama et al., 2017; etc.). Albumins were also not found in a number of nonmodel objects demonstrating a high level of identity with the proteins of model Acanthopterygii and Ostariophysi (Andreeva et al., 2015, 2017, 2019). These facts allow us to consider the loss of albumin genes as an event that goes beyond one family of Cyprinidae fish.
The conservativism of the organization of albumin polypeptide chains in all groups of Gnathostomata (with the preservation of their length, number of domains, and S–S-bonds, as well as the proportional preservation of these peculiarities with an increase in the length of albumin in lampreys) is probably explained by the fact that the protein organized in this way successfully performs the functions of transport and osmotic activity assigned to it. All these conservative traits are supported by selection with a relatively low similarity of amino acid sequences at an interorder level rarely exceeding 40%. At the same time, the support of the conservative organization of the polypeptide chain is reached by a high turnover of introns. We did not identify any examples of the coincidence of the lengths of the corresponding introns in paralogous genes and albumin orthologs. A dynamic turnover of introns is demonstrated by albumin paralogs in the sterlet and lower teleost fish (they did not have coincidences in the lengths of the corresponding introns and DNA repeats). Meanwhile, a deficiency in intron acquisition and turnover was noted for mammal genes with the conservative protein-coding regions (Roy et al., 2003). Taking into account that introns also include regulatory elements, their high turnover in the albumin genes of teleost fish probably contributed to the processes of their sub- and neofunctionalization.
Another evolutionary scenario of albumin transformations in teleost fish (in the form of its loss due to WGD) did not prevent the Teleostei taxon from developing and conquering new ecological niches. Against the background of dynamic transformations of genomes, a high level of intra- and interchromosomal rearrangements, and increased rate of evolution of protein-coding sequences, as well as a higher (compared with other vertebrates) rate of the turnover of introns and regulatory elements (Ravi and Venkatesh, 2018), the loss of albumin in Teleostei went unnoticed and other osmotically active proteins took over the function of albumin.
Change history
17 December 2022
An Erratum to this paper has been published: https://doi.org/10.1134/S1995082922090012
REFERENCES
Alexandrou, M., Swartz, B., Matzke, N., and Oakley, T., Genome duplication and multiple evolutionary origins of complex migratory behavior in Salmonidae, Mol. Phylogenet. Evol., 2013, vol. 69, no. 3, pp. 514–523. https://doi.org/10.1016/j.ympev.2013.07.026
Andreeva, A.M., The Strategies of Organization of the Fish Plasma Proteome: with and without Albumin, Russ. J. Mar. Biol., 2019, vol. 45, no. 4, pp. 263–274. https://doi.org/10.1134/S1063074019040023
Andreeva, A.M., Structural organization of plasma proteins as a factor of capillary filtration in pisces (Review), Inland Water Biol., 2020, vol. 13, no. 4, pp. 664–673. https://doi.org/10.1134/S1995082920060036
Andreeva, A.M., Organization and function of osmotically active fraction of fish (Pisces) plasma proteome, Inland Water Biol., 2021, vol. 14, no. 4, pp. 449–460.https://doi.org/10.1134/S1995082921040039
Andreeva, A.M., Lamash, N.E., Serebryakova, M.V., et al., Reorganization of low-molecular-weight fraction of plasma proteins in the annual cycle of cyprinidae, Biochemistry (Moscow), 2015, vol. 80, no. 2, pp. 208–218. https://doi.org/10.1134/S0006297915020078
Andreeva, A.M., Serebryakova, M.V., and Lamash, N.E., Oligomeric protein complexes of apolipoproteins stabilize the internal fluid environment of organism in redfins of the Tribolodon genus [Pisces; Cypriniformes, Cyprinidae], Comp. Biochem. Physiol., Part D: Genomics Proteomics, 2017, vol. 22, pp. 90–97. https://doi.org/10.1016/j.cbd.2017.02.007
Andreeva, A.M., Vasiliev, A.S., Toropygin, I.Y., et al., Involvement of apolipoprotein A in maintaining tissue fluid balance in goldfish Carassius auratus, Fish Physiol. Biochem., 2019, vol. 45, no. 5, pp. 1717–1730. https://doi.org/10.1007/s10695-019-00662-1
Anguizola, J., Matsuda, R., Barnaby, O.S., et al., Review: Glycation of human serum albumin, Clin. Chim. Acta, 2013, vol. 425, pp. 64–76. https://doi.org/10.1016/j.cca.2013.07.013
Ascenzi, P., di Masi, A., Leboffe, L., et al., Molecular phylogenetic analyses of albuminoids reveal the molecular evolution of allosteric properties, IUBMB Life, 2013, vol. 65, no. 6, pp. 544–549. https://doi.org/10.1002/iub.1164
Asrar, Z., Haq, F., and Abbasi, A.A., Fourfold paralogy regions on human HOX-bearing chromosomes: Role of ancient segmental duplications in the evolution of vertebrate genome, Mol. Phylogenet. Evol., 2013, vol. 66, no. 3, pp. 737–747. https://doi.org/10.1016/j.ympev.2012.10.024
Bayramov, A.V., Ermakova, G.V., Kucheryavyy, A.V., and Zaraisky, A.G., Genome duplications as the basis of vertebrates’ evolutionary success, Russ. J. Dev. Biol., 2021, vol. 52, no. 3, art. ID 170. https://doi.org/10.1134/S1062360421030024
Banerjee, S., Mahanty, A., Mohanty, S., et al., Identification of potential biomarkers of hepatotoxicity by plasma proteome analysis of arsenic-exposed carp Labeo rohita, J. Hazard. Mater., 2017, vol. 336, pp. 71–80. https://doi.org/10.1016/j.jhazmat.2017.04.054
Berthelot, C., Brunet, F., Chalopin, D., et al., The rainbow trout genome provides novel insights into evolution after whole-genome duplication in vertebrates, Nat. Commun., 2014, vol. 5, art. ID 3657. https://doi.org/10.1038/ncomms4657
Bouillon, R., Schuit, F., Antonio, L., and Rastinejad, F., Vitamin D binding protein: a historic overview, Front. Endocrinol. (Lausanne), 2020, vol. 10, art. ID 910. https://doi.org/10.3389/fendo.2019.00910
Bowers, J.E., Chapman, B.A., Ron, J., and Paterson, A.H., Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events, Nature, 2003, vol. 422, pp. 433–438. https://doi.org/10.1038/nature01521
Braasch, I. and Postlethwait, J.H., Polyploidy in fish and the teleost genome duplication, in Polyploidy and Genome Evolution, Berlin: Springer-Verlag, 2012. https://doi.org/10.1007/978-3-642-31442-1_17
Braasch, I., Guiguen, Y., Loker, R., et al., Connectivity of vertebrate genomes: Paired-related homeobox (Prrx) genes in spotted gar, basal teleosts, and tetrapods, Comp. Biochem. Physiol., Part C, 2014, vol. 163, pp. 24. https://doi.org/10.1016/j.cbpc.2014.01.005
Braasch, I., Gehrke, A.R., Smith, J.J., et al., The spotted gar genome illuminates vertebrate evolution and facilitates human-teleost comparisons, Nat. Genet., 2016, vol. 48, no. 4, pp. 427–437. https://doi.org/10.1038/ng.3526
Brown, J.D., Structural origins of mammalian albumin, Fed. Proc., 1976, vol. 35, no. 10, pp. 2141–2144.
Brunet, F.G., Crollius, H., Paris, M., et al., Gene loss and evolutionary rates following whole-genome duplication in teleost fishes, Mol. Biol. Evol., 2006, vol. 23, no. 9, pp. 1808–1816. https://doi.org/10.1093/molbev/msl049
Byrnes, L. and Gannon, F., Atlantic salmon (Salmo salar) serum albumin: cDNA sequence, evolution, and tissue expression, DNA Cell Biol., 1990, vol. 9, no. 9, pp. 647–655. https://doi.org/10.1089/dna.1990.9.647
Carninci, P., Sandelin, A., Lenhard, B., et al., Genome-wide analysis of mammalian promoter architecture and evolution, Nat. Genet., 2006, vol. 38, no. 6, pp. 626–635. https://doi.org/10.1038/ng1789
Cheng, P., Huang, Y., Lv, Y., et al., The American paddlefish genome provides novel insights into chromosomal evolution and bone mineralization in early vertebrates, Mol. Biol. Evol., 2021, vol. 38, no. 4, pp. 1595–1607. https://doi.org/10.1093/molbev/msaa326
Cioffi, M., Bertollo, L.A., Villa, M.A., et al., Genomic organization of repetitive DNA Elements and its implications for the chromosomal evolution of channid fishes (Actinopterygii, Perciformes), PLoS One, 2015, vol. 10, no. 6, art. ID e0130199. https://doi.org/10.1371/journal.pone.0130199
Crow, K.D., Smith, C.D., Cheng, J.F. et al., An independent genome duplication inferred from Hox paralogs in the American paddlefish—a representative basal ray-finned fish and important comparative reference, Genome Biol. Evol., 2012, vol. 4, no. 9, pp. 937–953. https://doi.org/10.1093/gbe/evs067
Curry, S., Mandelkow, H., Brick, P., and Franks, N., Crystal structure of human serum albumin complexed with fatty acid reveals an asymmetric distribution of binding sites, Nat. Struct. Biol., 1998, vol. 5, no. 9, pp. 827–835. https://doi.org/10.1038/1869
Deryckere, F., Byrnes, L., Wagner, A., et al., Salmon HNF1: cDNA sequence, evolution, tissue specificity and binding to the salmon serum albumin promoter, J. Mol. Biol., 1995, vol. 247, no. 1, pp. 1–10. https://doi.org/10.1006/jmbi.1994.0115
Dietrich, M.A., Arnold, G.J., Nynca, J., et al., Characterization of carp seminal plasma proteome in relation to blood plasma, J. Proteomics, 2014, vol. 98, pp. 218–232. https://doi.org/10.1016/j.jprot.2014.01.005
Dietrich, M.A., Irnazarow, I., Adamek, M., et al., 2D-DIGE proteomic analysis of blood plasma reveals changes in immune- and stress-associated proteins following hormonal stimulation of carp males, Fish Shellfish Immunol., 2021, vol. 118, pp. 354–368. https://doi.org/10.1016/j.fsi.2021.09.018
Du, K., Stöck, M., Kneitz, S., et al., The sterlet sturgeon genome sequence and the mechanisms of segmental rediploidization, Nat. Ecol. Evol., 2020, vol. 4, no. 6, pp. 841–852. https://doi.org/10.1038/s41559-020-1166-x
Dziegielewska, K.M., Evans, C.A., Fossan, G., et al., Proteins in cerebrospinal fluid and plasma of fetal sheep during development, J. Physiol., 1980, vol. 300, no. 1, pp. 441–455. https://doi.org/10.1113/jphysiol.1980.sp013171
Eiferman, F.A., Young, P.R., Scott, R.W., and Tilghman, S.M., Intragenic amplification and divergence in the mouse α-fetoprotein gene, Nature, 1981, vol. 294, no. 5843, pp. 713–718. https://doi.org/10.1038/294713a0
Evans, B., Upham, N., Golding, G., et al., 2017. Evolution of the largest mammalian genome, Genome Biol. Evol., vol. 9, no. 6, pp. 1711–1724. https://doi.org/10.1093/gbe/evx113
Fanali, G., Ascenzi, P., Bernardi, G., and Fasano, M.J., Sequence analysis of serum albumins reveals the molecular evolution of ligand recognition properties, J. Biomol. Struct. Dyn., 2012, vol. 29, no. 6, pp. 1195–1205. https://doi.org/10.1080/07391102.2011.672632
Fasano, M., Fanali, G., Leboffe, L., and Ascenzi, P., Heme binding to albuminoid proteins is the result of recent evolution, IUBMB Life, 2007, vol. 59, no. 7, pp. 436–440. https://doi.org/10.1080/15216540701474523
Felip, A., Carrillo, M., and Zanuy, S., Older triploid fish retain impaired reproductive endocrinology in the European sea bass Dicentrarchus labrax, J. Fish Biol., 2009, vol. 75, no. 10, pp. 2657–2669. https://doi.org/10.1111/j.1095-8649.2009.02458.x
Filosa, M.F., Adam, I., Robson, P., et al., Partial clone of the gene for AS protein of the lamprey Petromyzon marinus, a member of the albumin supergene family whose expression is restricted to the larval and metamorphic phases of the life cycle, J. Exp. Zool., 1998, vol. 282, no. 3, pp. 301–309. https://doi.org/10.1002/(sici)1097-010x(19981015)282:3<301::aid-jez3>3.0.co;2-q
Force, A., Lynch, M., Pickett, F.B., et al., Preservation of duplicate genes by complementary, degenerative mutations, Genetics, 1999, vol. 151, pp. 1531–1545. https://doi.org/10.1093/genetics/151.4.1531
Freeling, M. and Thomas, B.C., Gene-balanced duplications, like tetraploidy, provide predictable drive to increase morphological complexity, Genome Res., 2006, vol. 16, no. 7, pp. 805–814. https://doi.org/10.1101/gr.3681406
Ghuman, J., Zunszain, P.A., Petitpas, I., et al., Structural basis of the drug-binding specificity of human serum albumin, J. Mol. Biol., 2005, vol. 353, no. 1, pp. 38–52. https://doi.org/10.1016/j.jmb.2005.07.075
Gibbs, P.E. and Dugaiczyk, A., Origin of structural domains of the serum-albumin gene family and a predicted structure of the gene for vitamin D-binding protein, Mol. Biol. Evol., 1987, vol. 4, no. 4, pp. 364–379. https://doi.org/10.1093/oxfordjournals.molbev.a040447
Gorin, M.B., Cooper, D.L., Eiferman, F., et al., The evolution of alpha-fetoprotein and albumin, I. A comparison of the primary amino acid sequences of mammalian alpha-fetoprotein and albumin, J. Biol. Chem., 1981, vol. 256, no. 4, pp. 1954–1959.
Gray, J.E. and Doolittle, R.F., Characterization, primary structure, and evolution of lamprey plasma albumin, Prot. Sci., 1992, vol. 1, no. 2, pp. 289–302. https://doi.org/10.1002/pro.5560010211
Gu, L. and Xia, C., Cluster expansion of apolipoprotein D (ApoD) genes in teleost fishes, BMC Evol. Biol., 2019, vol. 19, art. ID 9.
Gundappa, M.K., To, T.H., Grønvold, L., et al., Genome-wide reconstruction of rediploidization following autopolyploidization across one hundred million years of salmonid evolution, Mol. Biol. Evol., 2021, vol. 39, no. 1, art. ID msab310. https://doi.org/10.1093/molbev/msab310
Hoegg, S., Brinkmann, H., Naylor, J.S., et al., Phylogenetic timing of the fish specific duplication correlates with the diversification of teleost fish, J. Mol. Evol., 2004, vol. 59, pp. 190–203.
Holland, P.W., Garcia-Fernàndez, J., Williams, N.A., and Sidow, A., Gene duplications and the origins of vertebrate development, Dev Suppl., 1994, pp. 125–133.
Jagodzinski, L.L., Sargent, T.D., Yang, M., et al., Sequence homology between RNAs encoding rat alpha-fetoprotein and rat serum albumin, Proc. Natl. Acad. Sci. U. S. A., 1981, vol. 78, no. 6, pp. 3521–3525. https://doi.org/10.1073/pnas.78.6.3521
Jerkovic, L., Voegele, A.F., Chwatal, S., et al., Afamin is a novel human vitamin E-binding glycoprotein characterization and in vitro expression, J. Proteome Res., 2005, vol. 4, no. 3, pp. 889–899. https://doi.org/10.1021/pr0500105
Kajiyama, Y., Tian, J., and Locker, J., Characterization of distant enhancers and promoters in the albumin-alpha-fetoprotein locus during active and silenced expression, J. Biol. Chem., 2006, vol. 281, no. 40, pp. 30122–30131. https://doi.org/10.1074/jbc.M603491200
Kamenskaya, D.N. and Brykov, V.A., Fish growth hormone genes: structure and divergence, Russ. J. Mar. Biol., 2020, vol. 46, pp. 233–242. https://doi.org/10.31857/S0134347520040038
Krieger, J. and Fuerst, P.A., Evidence of multiple alleles of the nuclear 18S ribosomal RNA gene in sturgeon (Family: Acipenseridae), J. Appl. Ichthyol., 2002, vol. 18, pp. 290–297. https://doi.org/10.1046/J,1439-0426.2002.00377.X
Krieger, J., Fuerst, P.A., and Cavender, T.M., Phylogenetic relationships of the North American sturgeons (order Acipenseriformes) based on mitochondrial DNA sequences, Mol. Phylogenet. Evol., 2000, vol. 16, no. 1, pp. 64–72. https://doi.org/10.1006/mpev.1999.0743
Krieger, J., Hett, A.K., Fuerst, P.A., et al., Unusual intraindividual variation of the nuclear 18S rRNA gene is widespread within the Acipenseridae, J. Hered., 2006, vol. 97, no. 3, pp. 218–225. https://doi.org/10.1093/jhered/esj035
Kuraku, S. and Meyer, A., The evolution and maintenance of Hox gene clusters in vertebrates and the teleost-specific genome duplication, Int. J. Dev. Biol., 2009b, vol. 53, nos. 5–6, pp. 765–773. https://doi.org/10.1387/ijdb.072533km
Kuraku, S., Meyer, A., and Kuratani, S., Timing of genome duplications relative to the origin of the vertebrates: did cyclostomes diverge before or after?, Mol. Biol. Evol., 2009a, vol. 26, no. 1, pp. 47–59. https://doi.org/10.1093/molbev/msn222
Li, S., Cao, Y., and Geng, F., Genome-wide identification and comparative analysis of albumin family in vertebrates, Evol. Bioinf. Online, 2017, vol. 13, pp. 1–6. https://doi.org/10.1177/1176934317716089
Ma, W., Zhu, Z., Bi, X., et al., Allopolyploidization is not so simple: evidence from the origin of the tribe Cyprinini (Teleostei: Cypriniformes), Curr. Mol. Med., 2014, vol. 14, no. 10, pp. 1331–1338. https://doi.org/10.2174/1566524014666141203101543
Maere, S., De Bodt, S., Raes, J., et al., Modeling gene and genome duplications in eukaryotes, Proc. Natl. Acad. Sci., 2005, vol. 102, no. 15, pp. 5454–5459. https://doi.org/10.1073/pnas.0501102102
Majorek, K.A., Porebski, P.J., Dayal, A., et al., Structural and immunologic characterization of bovine, horse, and rabbit serum albumins, Mol. Immunol., 2012, vol. 52, nos. 3–4, pp. 174–182. https://doi.org/10.1016/j.molimm.2012.05.011
Makhrov, A.A., A narrowing of the phenotypic diversity range after large rearrangements of the karyotype in Salmonidae: The relationship between saltational genome rearrangements and gradual adaptive evolution, Genes, 2017, vol. 8, pp. 297–306. https://doi.org/10.3390/genes8110297
Malik, S., Fu, L., Juras, D.J., et al., Common variants of the vitamin D binding protein gene and adverse health outcomes, Crit. Rev. Clin. Lab. Sci., 2013, vol. 50, no. 1, pp. 1–22. https://doi.org/10.3109/10408363.2012.750262
Metcalf, V., Brennan, S., Chambers, G., and George, P., The albumins of Chinook salmon (Oncorhynchus tshawytscha) and brown trout (Salmo trutta) appear to lack a propeptide, Arch. Biochem. Biophys., 1998a, vol. 350, no. 2, pp. 239–244. https://doi.org/10.1006/abbi.1997.0509
Metcalf, V.J., Brennan, S.O., Chambers, G.K., and George, P.M., The albumin of the brown trout (Salmo trutta) is a glycoprotein, Biochim. Biophys. Acta, 1998b, vol. 1386, no. 1, pp. 90–96.
Minghetti, P.P., Law, S.W., and Dugaiczyk, A., The rate of molecular evolution of alpha-fetoprotein approaches that of pseudogenes, Mol. Biol. Evol., 1985, vol. 2, no. 4, pp. 347–358. https://doi.org/10.1093/oxfordjournals.molbev.a040350
Morinaga, T., Sakai, M., Wegmann, T.G., and Tamaoki T., Primary structures of human alpha-fetoprotein and its mRNA, Proc. Natl. Acad. Sci. U. S. A., 1983, vol. 80, no. 15, pp. 4604–4608. https://doi.org/10.1073/pnas.80.15.4604
Nakatani, Y., Shingate, P., Ravi, V., et al., Publisher Correction: Reconstruction of proto-vertebrate, protocyclostome and proto-gnathostome genomes provides new insights into early vertebrate evolution, Nat. Commun., 2021, vol. 12, no. 1, art. ID 4704. https://doi.org/10.1038/s41467-021-25110-8
Nishio, H. and Dugaiczyk, A., Complete structure of the human alpha-albumin gene, a new member of the serum albumin multigene family, Proc. Natl. Acad. Sci. U. S. A., 1996, vol. 93, no. 15, P. 7557–7561. https://doi.org/10.1073/pnas.93.15.7557
Nishio, H., Heiskanen, M., Palotie, A., et al., Tandem arrangement of the human serum albumin multigene family in the sub-centromeric region of 4q: Evolution and chromosomal direction of transcription, J. Mol. Biol., 1996, vol. 259, no. 1, pp. 113–119. https://doi.org/10.1006/jmbi.1996.0306
Noel, E.S., Reis, M., Arai, Z., and Ober, E.A., Analysis of the Albumin/α-Fetoprotein/Afamin/Group specific component gene family in the context of zebrafish liver differentiation, Gene Exp. Patterns, 2010, vol. 10, no. 6, pp. 237–243. https://doi.org/10.1016/j.gep.2010.05.002
Nynca, J., Arnold, G., Fröhlich, T., and Ciereszko, A., Proteomic identification of rainbow trout blood plasma proteins and their relationship to seminal plasma proteins, Proteomics, 2017, vol. 17, no. 11, p. 1. https://doi.org/10.1002/pmic.201600460
Ohno, S., Evolution by Gene Duplication, Berlin: Springer-Verlag, 1970.
Ohno, S., Original domain for the serum albumin family arose from repeated sequences, Proc. Natl. Acad. Sci. U. S. A., 1981, vol. 78, no. 12, pp. 7657–7661. https://doi.org/10.1073/pnas.78.12.7657
Otterbein, L.R., Cosio, C., Graceffa, P., and Dominguez, R., Crystal structures of the vitamin D-binding protein and its complex with actin: Structural basis of the actin-scavenger system Proc. Natl. Acad. Sci. U. S. A., 2002, vol. 99, no. 12, pp. 8003–8008. https://doi.org/10.1073/pnas.122126299
Ozernyuk, N.D. and Myuge, N.S., Large-scale genome duplications and paralog divergence in fish, Russ. J. Genet., 2013, vol. 49, no. 1, pp. 63–69. https://doi.org/10.1134/S1022795413010080
Papp, B., Pál, C., and Hurst, L.D., Dosage sensitivity and the evolution of gene families in yeast, Nature, 2003, vol. 424, pp. 194–197. https://doi.org/10.1038/nature01771
Pasquier, J., Cabau, C., Nguyen, T., et al., Gene evolution and gene expression after whole genome duplication in fish: the PhyloFish database, BMC Genomics, 2016, vol. 17, art. ID 368. https://doi.org/10.1186/s12864-016-2709-z
Pelster, B. and Egg, M., Hypoxia-inducible transcription factors in fish: expression, function and interconnection with the circadian clock, J. Exp. Biol., 2018, vol. 221, no. 13, art. ID jeb163709. https://doi.org/10.1242/jeb.163709
Postlethwait, J.H., Woods, I.G., Ngo-Hazelet, P., et al., Zebrafish comparative genomics and the origins of vertebrate chromosomes, Genome Res., 2000, vol. 10, no. 12, pp. 1890–1902. https://doi.org/10.1101/gr.164800
Ramberg, S., Hoyheim, B., Ostbye, T., and Andreassen, R., A de novo full-length mRNA transcriptome generated from hybrid- corrected PacBio long- reads improves the transcript annotation and identifies thousands of novel splice variants in Atlantic Salmon, Front. Genet., 2021, vol. 12, pp. 1‒22. https://doi.org/10.3389/fgene.2021.656334
Ravi, V. and Venkatesh, B., The Divergent Genomes of Teleosts, Annu. Rev. Anim. Biosci., 2018, vol. 6, pp. 47–68. https://doi.org/10.1146/annurev-animal-030117-014821
Reshetnikov, Yu.S., Annotirovannyi katalog kruglorotykh i ryb kontinental’nykh vod Rossii (Annotated Check-List of Cyclostomata and Fishes of the Continental Waters of Russia), Moskva: Nauka, 1998.
Rojo-Bartolomé, I., de Souza, J., de Cerio, O., and Cancio, I., Duplication and subfunctionalisation of the general transcription factor IIIA (gtf3a) gene in teleost genomes, with ovarian specific transcription of gtf3ab, PLoS One, 2020, vol. 15, no. 1, art. ID e0227690. https://doi.org/10.1371/journal.pone.0227690
Romer, A.S. and Parsons, T.S., The Vertebrate Body, Philadelphia: Saunders College Publ., 1986.
Rondeau, E.B., Minkley, D.R., Leong, J.S., et al., The genome and linkage map of the northern pike (Esox lucius): conserved synteny revealed between the salmonid sister group and the Neoteleostei, PLoS One, 2014, vol. 9, no. 7, art. ID e102089. https://doi.org/10.1371/journal.pone.0102089
Roy, S.W., Fedorov, A., and Gilbert, W., Large-scale comparison of intron positions in mammalian genes shows intron loss but no gain, Proc. Natl. Acad. Sci. U. S. A., 2003, vol. 100, no. 12, pp. 7158–7162. https://doi.org/10.1073/pnas.1232297100
Saber, M.A., Stöckbauer, P., Morávek, L., and Meloun, B., Disulfide bonds in human serum albumin, Collect. Czech. Chem. Commun., 1977, vol. 42, no. 2, pp. 564–579. https://doi.org/10.1135/cccc19770564
Sargent, T.D., Jagodzinski, L.L., Yang, M., and Bonner, J., Fine structure and evolution of the rat serum albumin gene, Mol. Cell. Biol., 1981, vol. 1, no. 10, pp. 871–883. https://doi.org/10.1128/mcb.1.10.871-883
Schrama, D., Richard, N., Silva, T.S., et al., Enhanced dietary formulation to mitigate winter thermal stress in gilthead sea bream (Sparus aurata): a 2D-DIGE plasma proteome study, Fish Physiol. Biochem., 2017, vol. 43, no. 2, pp. 603–617. https://doi.org/10.1007/s10695-016-0315-2
Sharma, M.K., Liu, R.Z., Thisse, C., et al., Hierarchical subfunctionalization of fabp1a, fabp1b and fabp10 tissue-specific expression may account for retention of these duplicated genes in the zebrafish (Danio rerio) genome, FEBS J., 2006, vol. 273, no. 14, pp. 3216–3229. https://doi.org/10.1111/j.1742-4658.2006.05330.x
Smith, J.J. and Keinath, M.C., The sea lamprey meiotic map improves resolution of ancient vertebrate genome duplications, Genome Res., 2015, vol. 25, no. 8, pp. 1081. https://doi.org/10.1101/gr.184135.114
Soshnikova, N., Dewaele, R., Janvier, P., et al., Duplications of HOX gene clusters and the emergence of vertebrates, Dev. Biol., 2013, vol. 378, no. 2, pp. 194–199. https://doi.org/10.1016/j.ydbio.2013.03.004
Tamura, K., Stecher, G., Peterson, D., Filipski, A., and Kumar, S., MEGA6: Molecular Evolutionary Genetics Analysis version 6.0, Mol. Biol. Evol., 2013, vol. 30, pp. 2725–2729. https://doi.org/10.1093/molbev/mst197
Taylor, J.S., Van de Peer, Y., Braasch, I., and Meyer, A., Comparative genomics provides evidence for an ancient genome duplication event in fish, Philos. Trans. R. Soc., B, 2001, vol. 356, no. 1414, pp. 1661–1679. https://doi.org/10.1098/rstb.2001.0975
Uno, Y., Nishida, C., Takaqi, C., et al., Homoeologous chromosomes of Xenopus laevis are highly conserved after whole-genome duplication, Heredity (Edinb), 2013, vol. 111, no. 5, pp. 430–436. https://doi.org/10.1038/hdy.2013.65
Van de Peer, Y., Mizrachi, E., and Marchal, K., The evolutionary significance of polyploidy, Nat. Rev. Genet., 2017, vol. 18, no. 7, pp. 411–424. https://doi.org/10.1038/nrg.2017.26
Vasil’ev, V.P., Polyploidy in fishes and some problems concerning the evolution of karyotypes in the Salmonidae, Zh. Obshch. Biol., 1977, vol. 38, no. 3, pp. 380–392.
Vasil’ev, V.P., Evolyutsionnaya kariologiya ryb (Evolutionary Kariology of Fishes), Moskva: Nauka, 1985.
Venkatachalam, A.B., Parmar, M.B., and Wright, J.M., Evolution of the duplicated intracellular lipid-binding protein genes of teleost fishes, Mol. Genet. Genomics, 2017, vol. 292, no. 4, pp. 699–727. https://doi.org/10.1007/s00438-017-1313-5
Vílchez, M.C., Pla, D., Gallego, V., et al., Identification of the major proteins present in the seminal plasma of European eel, and how hormonal treatment affects their evolution. Correlation with sperm quality, Comp. Biochem. Physiol., Part A: Mol. Integr. Physiol., 2016, vol. 201, pp. 37–45. https://doi.org/10.1016/j.cbpa.2016.06.025
Xu, Y. and Ding, Z., N-terminal sequence and main characteristics of Atlantic salmon (Salmo salar) albumin, Prep. Biochem. Biotechnol., 2005, vol. 35, no. 4, pp. 283–290. https://doi.org/10.1080/10826060500218081
Yang, J., Lusk, R., and Li, W.-H., Organismal complexity, protein complexity, and gene duplicability, Proc. Natl. Acad. Sci. U. S. A., 2003, vol. 100, no. 26, pp. 15661–15665. https://doi.org/10.1073/pnas.2536672100
Zhao, C., Zhang, Y., Qin, H., et al., Organization and expression analysis of 5S and 45S ribosomal DNA clusters in autotetraploid Carassius auratus, BMC Ecol. Evol., 2021, vol. 21, art. ID 201. https://doi.org/10.1186/s12862-021-01918-2
Zhivotovskii, L.A., Evolutionary history of Pacific salmon and trout, Tr. Vseross. Nauchno-Issled. Inst. Rybn. Khoz. Okeanogr., 2015, vol. 157, pp. 4–23.
Funding
This work was performed as part of State Task no. 121050500046-8.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests. The author declares that she has no conflicts of interest.
Statement on the welfare of animals. This article does not contain any studies involving animals or human participants performed by the author.
Additional information
Translated by A. Barkhash
Abbreviations: a.a.r., amino acid residues; bp, base pair; nt, nucleotides; WGD, whole genome duplication; SD, segment duplication; LD, local duplication; 1R, 2R, 3R, WGD rounds.
The original online version of this article was revised: Due to a retrospective Open Access order.
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Andreeva, A.M. Evolutionary Transformations of Albumin Using the Example of Model Species of Jawless Agnatha and Bony Jawed Fish (Review). Inland Water Biol 15, 641–658 (2022). https://doi.org/10.1134/S1995082922050029
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1995082922050029