Abstract
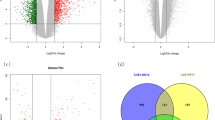
Objective: To identify the key genes and pathways involved in the occurrence and development of gastric cancer (GC). Methods: Differentially expressed genes (DEGs) and differentially expressed miRNAs (DEMs) were obtained using GEO2R. Function and pathway enrichment analysis of DEGs were performed using DAVID database.Protein-protein interaction (PPI) network analysis of DEGs was established by STRING database and visualized by Cytoscape software. Module analysis and hub genes selection of the PPI network was performed using Molecular Complex Detection (MCODE) and cytoHubba plug-in of Cytoscape software, respectively. Overall survival analysis of hub genes was performed by the Kaplan-Meier plotter online tool. Results: Totally, 98 DEGs were picked out, of which, 31 up-regulated genes were mainly involved in extracellular matrix (ECM)-receptor interaction, PI3K-Akt signaling pathway and focal adhesion, while 67 down-regulated DEGs were enriched in gastric acid secretion, collecting duct acid secretion and glycolysis/gluconeogenesis. Top 3 modules and top 10 hub genes with high centrality degree were selected from PPI network. Among these hub genes, high expression of secreted phosphoprotein 1 (SPP1), fibronectin1 (FN1) and collagen type I alpha 1 chain (COL1A1) were significantly associated worse overall survival for gastric cancer patients. Conclusions: The present study identified several key genes and pathways which may play an important role in the initiation and development of gastric cancer and could provide us potential targets for gastric cancer diagnosis and prognostic prediction.



Similar content being viewed by others
REFERENCES
Bray, F., Ferlay, J., Soerjomataram, I., et al., Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries, C.A. Cancer J. Clin., 2018, vol. 68, no. 6, pp. 394—424. https://doi.org/10.3322/caac.21492
Yuan, P., Wu, Z., Li, Z., et al., Impact of postoperative major complications on long-term survival after radical resection of gastric cancer, BMC Cancer, 2019, vol. 19, no. 1, p. 833. https://doi.org/10.1186/s12885-019-6024-3
Song, Z., Wu, Y., Yang, J., et al., Progress in the treatment of advanced gastric cancer, Tumour Biol., 2017, vol. 39, no. 7, p. 1010428317714626. https://doi.org/10.1177/1010428317714626
Guggenheim, D.E., and Shah, M.A., Gastric cancer epidemiology and risk factors, J. Surg. Oncol., 2013, vol. 107, no. 3, pp. 230—236. https://doi.org/10.1002/jso.23262
Su, X., Zhang, J., Yang, W., et al., Identification of the prognosis-related lncRNAs and genes in gastric cancer, Front. Genet., 2020, vol. 11, p. 27. https://doi.org/10.3389/fgene.2020.00027
Jinjia, C., Xiaoyu, W., Hui, S., et al., The use of DNA repair genes as prognostic indicators of gastric cancer, J. Cancer, 2019, vol. 10, no. 20, pp. 4866—4875. https://doi.org/10.7150/jca.31062
Hippo, Y., Taniguchi, H., Tsutsumi, S., et al., Global gene expression analysis of gastric cancer by oligonucleotide microarrays, Cancer Res., 2002, vol. 62, no. 1, pp. 233—240.
Carvalho, J., van Grieken, N.C., Pereira, P.M., et al., Lack of microRNA-101 causes E-cadherin functional deregulation through EZH2 up-regulation in intestinal gastric cancer, J. Pathol., 2012, vol. 228, no. 1, pp. 31—44. https://doi.org/10.1002/path.4032
Barrett, T., Wilhite, S.E., Ledoux, P., et al., NCBI GEO: archive for functional genomics data sets—update, Nucleic Acids Res., 2013, vol. 41, pp. D991—D995. https://doi.org/10.1093/nar/gks1193
Deng, W., Wang, Y., Liu, Z., et al., HemI: a toolkit for illustrating heatmaps, PLoS One, 2014, vol. 9, no. 11. e111988. https://doi.org/10.1371/journal.pone.0111988
Huang, D.W., Sherman, B.T., and Lempicki, R.A., Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources, Nat. Protoc., 2009, vol. 4, no. 1, pp. 44—57. https://doi.org/10.1038/nprot.2008.211
Szklarczyk, D., Franceschini, A., Wyder, S., et al., STRING v10: protein—protein interaction networks, integrated over the tree of life, Nucleic Acids Res., 2015, vol. 43, pp. D447—D452. https://doi.org/10.1093/nar/gku1003
Shannon, P., Markiel, A., Ozier, O., et al., Cytoscape: a software environment for integrated models of biomolecular interaction networks, Genome Res., 2003, vol. 13, no. 11, pp. 2498—2504. https://doi.org/10.1101/gr.1239303
Altaf-Ul-Amin, M., Shinbo, Y., Mihara, K., et al., Development and implementation of an algorithm for detection of protein complexes in large interaction networks, BMC Bioinf., 2006, vol. 7, p. 207. https://doi.org/10.1186/1471-2105-7-207
Bader, G.D., and Hogue, C.W., An automated method for finding molecular complexes in large protein interaction networks, BMC Bioinf., 2003, vol. 4, p. 2. https://doi.org/10.1186/1471-2105-4-2
Xiao, F., Zuo, Z., Cai, G., et al., miRecords: an integrated resource for microRNA-target interactions, Nucleic Acids Res., 2009, vol. 37, pp. D105—D110. https://doi.org/10.1093/nar/gkn851
Szasz, A.M., Lanczky, A., Nagy, A., et al., Cross-validation of survival associated biomarkers in gastric cancer using transcriptomic data of 1065 patients, Oncotarget, 2016, vol. 7, no. 31, pp. 49322—49333. https://doi.org/10.18632/oncotarget.10337
Zeng, B., Zhou, M., Wu, H., and Xiong, Z., SPP1 promotes ovarian cancer progression via Integrin beta1/FAK/AKT signaling pathway, OncoTargets Ther., 2018, vol. 11, pp. 1333—1343. https://doi.org/10.2147/OTT.S154215
Choe, E.K., Yi, J.W., Chai, Y.J., and Park, K.J., Upregulation of the adipokine genes ADIPOR1 and SPP1 is related to poor survival outcomes in colorectal cancer, J. Surg. Oncol., 2018, vol. 117, no. 8, pp. 1833—1840. https://doi.org/10.1002/jso.25078
Chen, X., Xiong, D., Ye, L., et al., SPP1 inhibition improves the cisplatin chemo-sensitivity of cervical cancer cell lines, Cancer Chemother. Pharmacol., 2019, vol. 83, no. 4, pp. 603—613. https://doi.org/10.1007/s00280-018-3759-5
Liao, Y.X., Zhang, Z.P., Zhao, J., and Liu, J.P., Effects of fibronectin 1 on cell proliferation, senescence and apoptosis of human glioma cells through the PI3K/AKT signaling pathway, Cell Physiol. Biochem., 2018, vol. 48, no. 3, pp. 1382—1396. https://doi.org/10.1159/000492096
Cai, X., Liu, C., Zhang, T.N., et al., Down-regulation of FN1 inhibits colorectal carcinogenesis by suppressing proliferation, migration, and invasion, J. Cell Biochem., 2018, vol. 119, no. 6, pp. 4717—4728. https://doi.org/10.1002/jcb.26651
Yoshihara, M., Kajiyama, H., Yokoi, A., et al., Ovarian cancer-associated mesothelial cells induce acquired platinum-resistance in peritoneal metastasis via the FN1/Akt signaling pathway, Int. J. Cancer, 2020, vol. 146, no. 8, pp. 2268—2280. https://doi.org/10.1002/ijc.32854
Li, J., Ding, Y., and Li, A., Identification of COL1A1 and COL1A2 as candidate prognostic factors in gastric cancer, World J. Surg. Oncol., 2016, vol. 14, no. 1, p. 297. https://doi.org/10.1186/s12957-016-1056-5
Zhuo, C., Li, X., Zhuang, H., et al., Elevated THBS2, COL1A2, and SPP1 expression levels as predictors of gastric cancer prognosis, Cell Physiol. Biochem., 2016, vol. 40, no. 6, pp. 1316—1324. https://doi.org/10.1159/000453184
Bartel, D.P., MicroRNAs: genomics, biogenesis, mechanism, and function, Cell, 2004, vol. 116, no. 2, pp. 281—297. https://doi.org/10.1016/s0092-8674(04)00045-5
Subramaniam, S., Jeet, V., Clements, J.A., et al., Emergence of microRNAs as key players in cancer cell metabolism, Clin. Chem., 2019, vol. 65, no. 9, pp. 1090—1101. https://doi.org/10.1373/clinchem.2018.299651
Lin, C., Huang, F., Shen, G., and Yiming, A., MicroRNA‑101 regulates the viability and invasion of cervical cancer cells, Int. J. Clin. Exp. Pathol., 2015, vol. 8, no. 9, pp. 10148—10155.
Liu, B., Qu, L., and Yan, S., Cyclooxygenase-2 promotes tumor growth and suppresses tumor immunity, Cancer Cell Int., 2015, vol. 15, p. 106. https://doi.org/10.1186/s12935-015-0260-7
Ma, X., Bai, J., Xie, G., et al., Prognostic significance of microRNA-101 in solid tumor: a meta-analysis, PLoS One, 2017, vol. 12, no. 7. e0180173. https://doi.org/10.1371/journal.pone.0180173
Fan, D., Ren, B., Yang, X., et al., Upregulation of miR-501-5p activates the wnt/beta-catenin signaling pathway and enhances stem cell-like phenotype in gastric cancer, J. Exp. Clin. Cancer Res., 2016, vol. 35, no. 1, p. 177. https://doi.org/10.1186/s13046-016-0432-x
Pan, R., Cai, W., Sun, J., et al., Inhibition of KHSRP sensitizes colorectal cancer to 5-fluoruracil through miR-501-5p-mediated ERRFI1 mRNA degradation, J. Cell Physiol., 2020, vol. 235, no. 2, pp. 1576—1587. https://doi.org/10.1002/jcp.29076
Funding
This work was supported by National Natural Science Foundation of China (no. 30672058); Scientific Research project of Southwest Medical University (no. 2020ZRQNB038).
Author information
Authors and Affiliations
Contributions
ZL and MS made substantial contributions to the conception of the present study. ZL and MS performed the primary bioinformatics analysis and were the major contributor in writing the manuscript. YZ and GT made substantial contributions to data analysis, including the biological significance of hub genes and figure editing. All authors revised and approved the final manuscript.
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest.
This article does not contain any studies involving animals or human participants performed by any of the authors.
Rights and permissions
About this article
Cite this article
Li, Z., Zhou, Y., Tian, G. et al. Identification of Core Genes and Key Pathways in Gastric Cancer using Bioinformatics Analysis. Russ J Genet 57, 963–971 (2021). https://doi.org/10.1134/S1022795421080081
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795421080081