Abstract
Immune checkpoint inhibitors (ICI) are highly effective in specific cancers where canonical markers of antitumor immunity are used for patient selection. Improved predictors of T cell-inflammation are needed to identify ICI-responsive tumor subsets in additional cancer types. We investigated associations of a 4-chemokine expression signature (c-Score: CCL4, CCL5, CXCL9, CXCL10) with metrics of antitumor immunity across tumor types. Across cancer entities from The Cancer Genome Atlas, subgroups of tumors displayed high expression of the c-Score (c-Scorehi) with increased expression of immune checkpoint (IC) genes and transcriptional hallmarks of the cancer-immunity cycle. There was an incomplete association of the c-Score with high tumor mutation burden (TMB), with only 15% of c-Scorehi tumors displaying ≥10 mutations per megabase. In a heterogeneous pan-cancer cohort of 82 patients, with advanced and previously treated solid cancers, c-Scorehi tumors had a longer median time to progression (103 versus 72 days, P = 0.012) and overall survival (382 versus 196 days, P = 0.038) following ICI therapy initiation, compared to patients with low c-Score expression. We also found c-Score stratification to outperform TMB assignment for overall survival prediction (HR = 0.42 [0.22–0.79], P = 0.008 versus HR = 0.60 [0.29-1.27], P = 0.18, respectively). Assessment of the c-Score using the TIDE and PredictIO databases, which include ICI treatment outcomes from 10 tumor types, provided further support for the c-Score as a predictive ICI therapeutic biomarker. In summary, the c-Score identifies patients with hallmarks of T cell-inflammation and potential response to ICI treatment across cancer types, which is missed by TMB assignment.
Similar content being viewed by others
Introduction
Antitumoral CD8+ T cells mediate immune-checkpoint inhibitor (ICI) response1,2,3. Chemokines have central roles in antitumoral immune cell infiltration, including recruitment of dendritic cells and antigen-specific CD8+ T cells4,5. We have identified a 4-chemokine transcriptomic signature (c-Score: CCL4, CCL5, CXCL9, CXCL10) to associate with T cell-inflammation in pancreatic cancer6. All four chemokines have roles in recruiting immune cells critical to cancer immunity. CCL4 mitigates dendritic cell migration and subsequent T cell activation and tumor infiltration7, while CCL5, CXCL9, and CXCL10 are associated with T cell infiltration across solid tumors8,9,10,11,12,13,14,15.
High mutational burden (TMB) in mismatch repair (MMR) deficient (MMRD) tumors enhances immune antitumor responses16,17,18,19. Similarly, tumors with homologous recombination (HR) repair pathway deficiencies (HRD) exhibit increased mutations and may also harbor a genetically favorable milieu to elicit an antitumor immune response in HRD-associated cancers20,21. In metastatic pancreaticobiliary cancers with HRD, responders to ICI therapy express higher levels of CCL4, CXCL9, and CXCL1022.
Elevated TMB (≥10 mutations/megabase) has emerged as a tumor-agnostic FDA-approved biomarker for ICI therapy23. The transformative treatment benefits of ICIs in a select subset of malignancies has triggered new research to identify improved biomarkers that will identify T cell-inflamed cancer subpopulations within TMB low and broadly immune cold cancer types24. In this study, we investigated the predictiveness of the 4-chemokine c-Score in revealing T cell-inflammation across 25 tumor types by leveraging genomic and transcriptomic data from 6455 patients. Using a real-world clinical cohort of 82 advanced cancer patients with ICI response data, we subsequently evaluated the ability of the 4-chemokine signature to predict ICI treatment response compared to TMB.
Results
Two discovery pan-cancer datasets were used to examine the 4-chemokine signature (CCL4, CCL5, CXCL9, CXCL10; herein referred to as c-Score) as a predictive biomarker for T cell-inflammation and ICI treatment outcomes across different solid tumor types (Fig. 1). The utility of the c-Score in predicting ICI treatment outcomes was then validated using two additional databases (TIDE25, PredictIO26), which include 28 studies with RNA sequencing and ICI response data from 10 tumor types (Fig. 1).
a Consort diagram describing principal cohorts and analysis pipeline. b 4-chemokine signature (c-Score), calculated as a mean expression of CCL4, CCL5, CXCL9, and CXCL10, across 31 tumor types from TCGA. Lowest to highest median c-Score plotted from left to right. Red and blue dashed lines represent top and bottom quartile cut offs, respectively. c Distribution of c-Score, with segregation according to c-Scorehi (n = 1747), c-Scoremed (n = 3493) and c-Scorelo (n = 1747) expression profiles. Expression profile scoring is based on the bottom, middle and top quartiles across the entire dataset. The bottom track displays the reported objective response rate to anti-PD1 or anti-PD-L1 immune checkpoint therapy27. Circles = ORR in MMRD tumors for that tumor subtype. NA = ORR not available. Median, quartiles, minimum and maximum values are represented by the central line, limits of box, and ends of lines of boxplots shown in (b and c).
A wide range of tumor types across TCGA cohort have high expression of the 4-chemokine signature
We analyzed transcriptomic cancer data from The Cancer Genome Atlas (TCGA). The cohort consisted of 31 tumor types from 6987 patients (histological subtypes and abbreviations found in Supplementary Data File 1). Uveal melanoma (UVM), adrenocortical carcinoma (ACC), and lower grade glioma (LGG) had the lowest c-Score expression, while testicular germ cell (TGCT), cervical squamous cell carcinoma (CESC) and head and neck squamous cell carcinoma (HNSC) were the highest expressors of the c-Score (Fig. 1b). To compare differences in T cell-inflammation within and between tumor types, c-Score expression was categorized for each tumor type in quartile distributions based on expression of the 4 chemokines across all 6987 patients in TCGA cohort. Tumors having c-Score expression equal or greater than the third quartile were classified as c-Scorehi (n = 1747), those expressing equal or less than the first quartile were classified as c-Scorelo (n = 1747), and those with scores in between were classified as c-Scoremed (n = 3493). Across these 31 tumor types, only two (pheochromocytoma [PCPG], and kidney chromophobe [KICH]) were devoid of c-Scorehi tumors. An additional four tumor types (UVM, ACC, uterine carcinosarcoma [UCS], glioblastoma multiforme [GBM]) with a paucity of c-Scorehi tumors were excluded, leaving 25 tumor types from 6455 patients for downstream analyses comparing c-Score groups (Methods, Fig. 1). Tumor types with reported objective response rates (ORR) of >11 to anti-PD-1 or PD-L1 therapies27 included top c-Score expressors (mesothelioma [MESO], lung squamous cell carcinoma [LUSC], kidney renal clear cell carcinoma [KIRC], HNSC, CESC), though other modest and strong responders to ICIs were seen throughout the c-Score expression spectrum (Fig. 1c). Interestingly, TGCT, the highest c-Score expressor, had low ORR to ICIs27. For data representation across metrics of T cell-inflammation and antitumor immunity, the c-Scorelo and c-Scoremed cases were grouped together (c-Scorelo+med) and compared to the c-Scorehi group.
Tumors in TCGA cohort with high expression of the c-Score display hallmarks of T cell-inflammation
To investigate the immune profiles of c-Scorehi tumors across the 25 TCGA tumor types, we evaluated immunomodulatory genes involved in immune checkpoint blockade response, (CD274 [PD-L1], PDCD1 [PD-1], HAVCR2 [TIM3], LAG3, TIGIT, CTLA4, and FASLG). Across tumor types, these genes were overexpressed in c-Scorehi tumors versus c-Scorelo+med tumors, except for cholangiocarcinoma (CHOL), thymoma (THYM), and kidney papillary renal cell carcinoma (KIRP) that did not meet the false discovery rate (FDR) threshold for CD274, PDCD1, and HAVCR2, respectively (Fig. 2a; Wilcoxon rank-sum, FDR adjusted). CD40, which encodes a key molecule involved in antigen presentation2,28,29,30, was also upregulated in the c-Scorehi tumors across tumor types (Supplementary Fig. 1a; 92%, 23/25).
a Degree of the significance of immunoregulatory gene expression between c-Scorehi versus c-Scorelo+med groups across 25 tumor types from TCGA. Wilcoxon rank-sum test, FDR adjusted. (b) Scores for and (c) degree of significance in eight genesets representing processes involved in the cancer-immunity cycle across c-Scorehi (n = 1735) and c-Scorelo+med (n = 4720) tumors in this dataset. For each pair, c-Scorelo+med tumors are on left, c-Scorehi tumors are on the right, as depicted in the inset. A.U. Arbitrary units, translated onto a positive scale (see Methods). Wilcoxon rank-sum test, FDR adjusted. X represents FDR > 0.05.
To expose higher-dimensional differences in the transcriptomic immune profiles of these tumors, we compared immune cell proportions and gene set differences between the two c-Score groups. Compared to c-Scorelo+med, c-Scorehi tumors had increased transcriptional patterns associated with the presence of M1-type macrophages, CD8+ T cells, and T Regulatory cells in most tumor types (Supplementary Fig. 1b; enriched in 96%, 88%, 64% of tumor types, respectively, Wilcoxon rank-sum, FDR adjusted). We then examined eight genesets representative of processes key to the cancer-immunity cycle, including co-stimulation and inhibition of antigen presenting cells (APCs), tumor antigen cross-presenting BATF3 dendritic cell (DC) expression, co-stimulation and inhibition of T cells, major histocompatibility class I (MHC I) molecule expression, interferon gamma signaling, and cytolytic activity7,31,32 (Fig. 2b). Compared to c-Scorelo+med, c-Scorehi tumors had increased expression of at least six genesets, with 96% (24/25) of tumor types having increased expression of all genesets (Fig. 2c; Wilcoxon rank-sum, FDR adjusted). Similarly, genesets predictive of T cell-inflammation33 and response to ICI34 were upregulated in c-Scorehi tumors across these 25 tumor types (Supplementary Fig. 1c; Wilcoxon rank-sum, FDR adjusted). Together, these results show a correlation between high c-Score expression and transcriptional hallmarks of T cell-inflammation across tumor types.
High proportion of tumors in TCGA cohort are c-Scorehi despite low TMB
Tumors displaying high TMB are presumed to be more immunogenic due to their increased levels of immune-inciting neoantigens27,35,36. However, the association of TMB and markers of T cell-inflammation appears to be specific to certain tumor types and is not observed in others6,37,38,39. Therefore, we evaluated whether the c-Score identifies T cell-inflammation that is missed by the TMB surrogate biomarker. We observed a relationship between median TMB and c-Score expression across tumor types (Fig. 3a; Rho = 0.42, P = 0.020, Spearman correlation, all 31 TCGA tumor types). However, of the 25 tumor types analyzed, only 32% (8/25; CESC, stomach adenocarcinoma [STAD], lung adenocarcinoma [LUAD], breast carcinoma [BRCA], colon adenocarcinoma [COAD], THYM, endometroid endometrial adenocarcinoma [UCEC], and bladder urothelial carcinoma [BLCA]) had higher TMB in the c-Scorehi group versus the c-Scorelo+med group (Fig. 3b; P < 0.05, Wilcoxon rank-sum, FDR adjusted). A similar relationship was also observed for single nucleotide variant (SNV)-derived neoantigens (Supplementary Fig. 2a). Notably, among the 25 histological tumor types, high TMB tumors (TMBhi, n = 530; ≥10 mutations/megabase) were c-Scorehi enriched compared to TMBlo (n = 5925) tumors (Fig. 3c; 48% versus 25%, P < 0.001, Fisher’s exact test). However, TMBhi tumors only represented 15% of the total c-Scorehi tumors across histological types (Fig. 3c), suggesting an incomplete association of TMB and the c-Score.
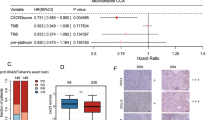
a Spearman correlation of median TMB versus c-Score across all 31 tumor types from TCGA. b Association between TMB and c-Score signature across 25 tumor types from TCGA. Boxplots depict a comparison of TMB in c-Scorehi versus c-Scorelo+med tumors, across tumor types. The left track displays FDR adjusted P values of TMB differences between groups. Right track displays group designation. The dashed line shows TMB threshold of 10 mutations/megabase. Wilcoxon rank-sum test, FDR adjusted. c The proportion (left) and absolute number (right) of c-Scorehi and c-Scorelo+med tumors in TMBlo versus TMBhi (TMB ≥ 10) cases across 25 tumor types. Fisher’s exact test. d Tumor types where c-Scorehi tumors had higher TMB versus c-Scorelo+med were classified as ‘TMB-associated’. TMB (left) and the c-Score (right) in ‘TMB-unassociated’ versus ‘TMB-associated’. Wilcoxon rank-sum test. Median, quartiles, minimum and maximum values are represented by the central line, limits of box, and ends of lines of boxplots shown in b and d.
We, therefore, classified histological tumor types with higher TMB in the c-Scorehi group versus the c-Scorelo-med group as ‘TMB-associated’. Only eight of the 25 histological tumor types were ‘TMB-associated’, whereas the remaining 17 tumor types were ‘TMB-unassociated’ (Fig. 3b). The eight ‘TMB-associated’ tumor types (CESC, STAD, LUAD, BRCA, COAD, THYM, UCEC, BLCA) displayed elevated TMB (P < 0.001) and c-Score (P < 0.001) expression (Fig. 3d; Wilcoxon rank-sum), with a higher proportion of tumors meeting the c-Scorehi designation (Supplementary Fig. 2b; P < 0.001, Fisher’s exact test). Importantly, 52% (n = 902/1735) of all c-Scorehi tumors across histological tumor types were classified as ‘TMB-unassociated’ (Supplementary Fig. 2b).
Next, we compared these tumors across histological types based on their c-Score group and classification as TMBhi or TMBlo. Considering both c-Score and TMB assignments, we found that c-ScorehiTMBhi tumors had increased expression of most cancer-immunity cycle gene sets and marginally increased immune checkpoint (IC) gene expression compared to the c-ScorehiTMBlo group (Supplementary Fig. 2c, d; Wilcoxon rank-sum, FDR adjusted, all 31 TCGA tumor types). These observations suggest that tumors with both high expression of the c-Score and a TMB ≥ 10 mutations/megabase have the strongest antitumor immunity phenotype. However, c-ScorehiTMBhi tumors represent only 3.6% of tumors across the total cohort, with a larger fraction of c-Scorehi tumors having TMBlo scores (Fig. 3c). Therefore, c-Scorehi identifies T cell-inflammation, with potential immune checkpoint treatment implications, that is not captured by TMB assignment.
Tumors with high c-Score expression in the ICI-treated POG cohort respond to immune checkpoint blockade
To explore the prognostic value of the c-Score in immunotherapy response, we used previously published transcriptome sequencing data from the Personalized OncoGenomics (POG570) program40. We first evaluated the c-Score distribution across one cohort of patients from the POG program (POG570, n = 559 patients, 26 tumor types)41. Similar to the distribution in TCGA, c-Scorehi tumors were present in most tumor types (73%, n = 19/26, Supplementary Fig. 3a). Notably, in both POG570 and TCGA, the same tumor types had overall lower c-Score expression values (ACC, UVM, thyroid carcinoma [THCA]) versus higher values (ovarian serous cystadenocarcinoma [OV], CESC and MESO).
Next, we evaluated the relationship of the c-Score with ICI therapy outcomes in an ICI-treated POG cohort40. This heterogenous pan-cancer cohort of 82 patients, with advanced and previously treated solid cancers, received ICI therapy following tumor biopsy for whole transcriptomic sequencing. Using the transcriptomic data from their tumor biopsies, the 82 patients were stratified into c-Scorehi (n = 28), c-Scoremed (n = 44), and c-Scorelo (n = 10) groups based on the thresholds determined using TCGA data (Fig. 4a; Methods). Patients who exhibited durable clinical benefit (DCB) on treatment had higher c-Score expression (Fig. 4b; Wilcoxon rank-sum), and this observation remained significant after correcting for differences in histological tumor type (P = 0.036, multivariate linear regression). Thirty-six percent (10/28) of patients in the c-Scorehi group had DCB, compared to 16% (7/44) for c-Scoremed and 0% (0/10) for the c-Scorelo group (Supplementary Fig. 3b; P = 0.029, chi-squared). Furthermore, patients with high expression of the c-Score had a longer time to progression (TTP, median 103 versus 72 days, P = 0.012) and overall survival (OS, median 382 versus 196 days, P = 0.038, log-rank) compared to the c-Scorelo+med group (Fig. 4c, d). Comparison of the c-Scorehi, c-Scoremed and c-Scorelo groups also showed parallel TTP (P = 0.012) and OS (P = 0.017, log-rank) clinical outcome differences (Supplementary Fig. 3c). The c-Scorehi association with favorable outcomes was independent of tumor type (Fig. 4d; TTP hazard ratio (HR) = 0.37 [0.18-0.74], P = 0.005; OS HR = 0.42 [0.22-0.79], P = 0.008, Cox proportional hazards).
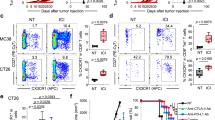
a Distribution of c-Scorehi, c-Scoremed, and c-Scorelo groups across ICI treated patients in the POG program. b c-Score expression in patients with durable clinical benefit (DCB, n = 17) versus no clinical benefit (NCB, n = 65). Wilcoxon rank-sum test. Median, quartiles, minimum and maximum values are represented by the central line, limits of box, and ends of lines of boxplots shown. c Time to progression and overall survival for patients stratified by c-Score group. P values indicate log-rank tests between the c-Scorehi and c-Scorelo+med groups. d Multivariate cox proportional hazards models for time to progression and overall survival. Tumor types with at least 3 patients were included in the models (total of 65 patients). Error bars represent 95% CI.
Since expression of IC genes has been associated with ICI treatment outcomes32,34,35, we examined the ICI treatment response predictiveness of CD274, PDCD1, and CTLA4 in the ICI-treated POG cohort. For each IC gene, expression was stratified into high and low/medium quartiles, similar to c-Score group assignments. Evaluating each IC gene individually in a cox proportional hazards model, along with c-Score, TMB and tumor type, none of the three IC genes were predictive of TTP or OS. The c-Score, however, remained strongly predictive of TTP and OS (Supplementary Fig. 3d).
We then compared the predictiveness of c-Score versus TMB assignments on TTP and OS. The c-Scorehi biomarker performed equally to TMBhi assignments in predicting TTP (HR = 0.37 [0.18–0.74], P = 0.005 versus HR = 0.39 [0.18–0.87], P = 0.022, respectively, Cox proportional hazards) and outperformed TMB when comparing OS (Fig. 4d; HR = 0.42 [0.22-0.79], P = 0.008 versus HR = 0.60 [0.29-1.27], P = 0.18, respectively, Cox proportional hazards). In a subgroup analysis, c-ScorehiTMBhi patients had a longer TTP compared to the other subgroups (P = 0.00092). However, an OS difference between the four groups (c-ScorehiTMBhi, c-ScoreloTMBhi, c-ScorehiTMBlo, c-ScoreloTMBlo) was not observed (Supplementary Fig. 3e; P = 0.094, log-rank). Moreover, no difference in OS between c-Scorehi versus c-Scorelo+med patients was observed across three independent ICI-untreated cohorts (data not shown: PanCuRx20, treatment naïve resectable pancreatic adenocarcinoma (PAAD) cohort, n = 109, P = 0.68; COMPASS42, treatment naïve PAAD metastatic cohort, n = 117, P = 0.48; POG570, entire pan-cancer non-immunotherapy treated cohort, n = 402, P = 0.3). Furthermore, when survival data included time points before patients received ICI (date of advanced disease versus date of ICI initiation), the prognostic ability of the c-Score score was lost (Supplementary Fig. 3f; P = 0.29, log-rank).
Validation of the c-Score as a predictor of immune checkpoint blockade treatment response in additional cohorts
To further validate the c-Score, we first used the TIDE platform25 containing 25 databases from 19 studies spanning 7 tumor types (n = 1290). The c-Score had a positive predictive value of ICI response in 72% (18/25) of the datasets analyzed. Moreover, the c-Score predicted ICI response with an AUC ≥ 0.8 in 24% (6/25) of the datasets, compared to 4% (1/25) using the TIDE computational framework model (Supplementary Fig. 4a). Next, we used the PredictIO platform26, encompassing 15 databases from 15 studies spanning eight tumor types (n = 715). The c-Score predicted treatment response, OS, and progression free survival in a pooled analysis of all datasets (Supplementary Fig. 4b–d, left panels). Furthermore, the c-Score was significantly superior at predicting treatment response, OS, and progression free survival compared to 37 published ICI biomarker signatures, second only to the 100-gene PredictIO gene signature (Supplementary Fig. 4b–d, right panels). These two validation analyses further support the clinical utility of the c-Score for predicting ICI treatment outcomes.
c-Score expression associates with both DNA repair deficient and proficient tumors
We next examined whether the incomplete association between TMB and the c-Score could be explained by genomic differences. Patients with tumors defective in DNA repair and fidelity pathways, including MMRD, HRD, and the POLE/POLD1 genes, have been shown to respond favorably to ICIs and display increased hallmarks of T cell-inflammation20,21,43,44. Mechanisms leading to high TMB may explain an association between TMB and antitumoral immune activity. For example, defects in DNA repair, such as in MMRD, stimulate innate immune signaling pathways45,46. Therefore, we investigated whether tumors that had an association of high TMB with the c-Score (i.e., ‘TMB-associated’) had mutations in genes involved with DNA repair pathways across the 25 TCGA tumor types. Specifically, we considered mutations in DNA mismatch repair (MLH1, MSH2, MSH6, PMS2), homologous recombination repair (BRCA1, BRCA2, PALB2), and DNA replication and proofreading (POLD1, POLE) genes, as well as molecular hallmarks indicative of mutations in these pathways (i.e., for MMRD, microsatellite instability (MSI) inferred using MANTIS scores47; for HRD, HRDScore48,49). Indeed, ‘TMB-associated’ tumor types had a higher frequency of mutations, including protein-truncating mutations, in these DNA integrity pathway genes (Supplementary Fig. 5a–f; Fisher’s exact test) and increased MSI and HRD scores (Fig. 5a, b; Wilcoxon rank-sum). We then stratified tumors into MSI high and low (MANTIShi, n = 205; MANTISlo, n = 4842) and HRD high and low (HRDScorehi, n = 1670; HRDScorelo, n = 4842; Methods, Supplementary Fig. 5g–i). MSIhi tumors displayed increased expression of the c-Score (Fig. 5c; P = 5 × 10−19, Wilcoxon rank-sum), in agreement with previous reports that MSI tumors harbor immunogenicity43. While HRDhi tumors also had increased expression of the c-Score, the association was lost after controlling for tumor type (Fig. 5d; Wilcoxon rank-sum, controlled using multivariate linear regression). Thus, HRD tumors may display increased T cell-inflammation that is both weaker compared to MMRD tumors and more variable among histological tumor types.
a MANTIS scores, excluding HRDScorehi tumors, and (b) HRD scores, excluding MANTIShi tumors, in ‘TMB-unassociated versus ‘TMB-associated’ tumors across 25 tumor types from TCGA. Boxplots depicting the c-Score in (c), MANTISlo versus MANTIShi tumors and (d), HRDScorelo versus HRDScorehi tumors, excluding HRDScorehi and MANTIShi tumors, respectively, from this cohort. Wilcoxon rank-sum test. The proportion (left) and absolute number (right) of c-Scorehi and c-Scorelo+med tumors in (e), MANTISlo (n = 4376) versus MANTIShi (n = 199) and (f), HRDScorelo (n = 4376) versus HRDScorehi (n = 1629) tumors from this cohort. Fisher’s exact test. Median, quartiles, minimum and maximum values are represented by the central line, limits of box, and ends of lines of boxplots shown in a–d.
Among the full 31 TCGA tumor type dataset, the MSI score was positively associated with the c-Score only in COAD (Supplementary Fig. 6a; Rho = 0.35, P < 0.001, Spearman correlation, FDR adjusted). The HRDScore was positively correlated with the c-Score in THYM (Rho = 0.33, P < 0.01) and negatively correlated in HNSC (Supplementary Fig. 6b; Rho = −0.36, P < 0.01, Spearman correlation, FDR adjusted). These results suggest continuous metrics of MSI or HRD correlate linearly with the c-Score only in a subset of tumor types, and analyses using dichotomous classifications of these pathway defects may highlight further correlations with the c-Score across tumors.
Similar to their correlation with TMB, TCGA tumors harboring DNA repair pathway defects were enriched for c-Scorehi expression (Fig. 5e, f; 49% versus 24%, P = 3 × 10−14 for MSI; 32% versus 24%, P = 2 × 10−10 for HRD, Fisher’s exact test). However, the majority of c-Scorehi tumors had low MSI and HRD scores (Fig. 5e, f; 91% and 67%, respectively). HRDetecthi (n = 8) and MSIsensorhi (n = 3) tumors in POG570 cohort had higher median c-Score expression than their low counterparts, which was not significant in this small patient cohort (Supplementary Fig. 6c, d; Wilcoxon rank-sum). Together, these analyses show an association between the c-Score and DNA repair deficiency. Potentially more clinically relevant, these observations also suggest that additional mechanisms beyond DNA repair deficiency underlie the T cell-inflammation predicted by the c-Score and highlight the potential complementary clinical utility of the c-Score with TMB and markers of DNA repair deficiency.
Relationship of c-Score, somatic mutations and TMB
Considering the potential association of oncogenic mutations, beyond DNA repair genes, with cancer immunity, we also investigated the relationship of somatic mutations with T cell-inflammation, as predicted by the c-Score, across TCGA cohort. Among the entire 31 tumor types, the top 10 genes with somatic mutations were TTN, TP53, MUC16, CSMD3, RYR2, LRP1B, SYNE1, USH2A, FLG, and PIK3CA (Fig. 6a). We compared the expression of the c-Score in wild-type versus mutated tumors for the top three mutated genes. For all three genes, tumors harboring mutations in these genes had increased c-Score expression, including after controlling for tumor type (Fig. 6b). Tumors harboring these mutations also had increased TMB (Fig. 6c).
a Histogram of the top 10 somatic gene mutations across the 31 TCGA tumor types. Comparison of (b), c-Score and (c), TMB in tumors with wild-type (wt) versus mutated (mut) TTN, TP53, or MUC16 across the 25 tumor types from TCGA. d Overview of somatic KRAS mutation status across the cohort (n = 6386; KRAS-mutated tumors were not identified in MESO), shown as percentage (top) and absolute counts (bottom), with missense representing the majority of mutations. e c-Score, and (f), TMB in KRAS-mutated versus wild-type tumors across the top five KRAS-mutated tumors across the cohort. Wilcoxon rank-sum test. Median, quartiles, minimum and maximum values are represented by the central line, limits of box, and ends of lines of boxplots shown in (b, c, e, and f).
Relationship of c-Score, KRAS mutations and TMB
Oncogenic KRAS mutations commonly occur across cancer types50, have been associated with immunosuppressive phenotypes, and may represent a potential mechanism of ICI treatment evasion24,51,52. Therefore, we investigated the relationship of KRAS-mutated tumors with the c-Score. Pancreatic ductal adenocarcinoma (PAAD) harboured the most KRAS mutations (73%), followed by COAD (48%), rectum adenocarcinoma [READ] (38%), LUAD (30%), and UCEC (18%, Fig. 6d). A tumor-agnostic analysis across the 25 TCGA tumor types did not identify a c-Score expression difference between KRAS wild-type versus mutated tumors (Supplementary Fig. 7a; Wilcoxon rank-sum). Further analysis of the five tumor types with highest prevalence of KRAS mutations revealed an association in PAAD and READ, whereby KRAS-mutated tumors displayed lower expression of the c-Score versus KRAS wild-type tumors (Fig. 6e; MSI status was controlled using MANTIS for READ (P = 0.016, multivariate linear regression), while PAAD did not include cases with MSI). A c-Score association with KRAS-mutated tumors was not observed in COAD (P = 0.085) and LUAD (P = 0.43), while there was a trend to a lower c-Score in KRAS-mutated UCEC (P = 0.053, Wilcoxon rank-sum).
KRAS-mutated PAAD and LUAD were found to harbor higher TMB (Fig. 6f; P = 2 × 10−9, P = 0.029, respectively, Wilcoxon rank-sum). However, this association was lost in LUAD when controlling for MANTIS designations. Across tumor types, KRAS-mutated tumors had a higher TMB versus KRAS wild-type tumors (Supplementary Fig. 7b; P = 3 × 10−22, Wilcoxon rank-sum; P = 0.013 after controlling for MSI status using MANTIS) but was also lost after controlling for tumor type (P = 0.58, multivariate linear regression). Although there were more c-Scorehi tumors classified as KRAS wild-type versus KRAS-mutated, this correlation did not reach significance (Supplementary Fig. 7c; P = 0.26, Fisher’s exact test). The absence of a significant c-Scorehi association with KRAS wild-type tumors may be, at least in part, due to a countereffect exerted by a higher mutational load in KRAS-mutated tumors, supported by the higher proportion of tumors with TMB ≥ 10 in the KRAS-mutated group (Supplementary Fig. 7d; P = 1 × 10−9, Fisher’s exact test). Taken together, these results support a paradigm whereby genomic drivers, such as mutated KRAS, and defects in DNA repair pathways/TMB may have counteracting effects on tumor immunogenicity, where contributions from each process are dictated by additional underlying mechanisms specific to tumor types and their microenvironments.
Discussion
The c-Score, derived from expression of CCL4, CCL5, CXCL9, and CXCL10, identified subpopulations of tumors across cancer types with hallmarks of T cell-inflammation. Tumors with high expression of the c-Score displayed increased transcriptional hallmarks of a T cell-inflamed phenotype, suggesting that c-Scorehi tumors have properties of an activated immune response. We found that most histological tumor types had a subpopulation of tumors displaying a high expression of the c-Score, including histological tumor types that have been broadly classified as immune cold. Across cancer types, the proportion of c-Scorehi tumors ranged from 0% in pheochromocytoma and kidney chromophobe tumors to 61% in testicular germ cell tumors. T cell-inflammation stratification of tumor types according to c-Score correlated with previously reported antitumor immunity phenotypes and ICI response classifications27,31,38,48. Notably, two of the four chemokines (CXCL9, CXCL10) in the c-Score were present in a 10 gene M1-M2 macrophage signature recently shown to be predictive of ICI therapy outcomes41. These observations align with evidence implicating chemokines as putative mediators of tumor-homing in adoptive T cell therapy53.
We also show that the c-Score has complementary clinical utility to TMB. In a cohort of 82 patients treated with ICIs40, the c-Scorehi group exhibited a longer median time to progression and overall survival. We observed a correlation between c-Scorehi and TMBhi assignments, with c-ScorehiTMBhi tumors showing the longest time to progression following ICI treatment initiation. However, the correlation between c-Scorehi and TMBhi was imperfect, identifying patients that were c-Scorehi while TMBlo with responses to ICI therapy. In addition, the c-Score outperformed TMB for overall survival prediction. While a limitation of this real-world cohort is that patients received different types of ICIs, combined ICIs, or ICIs with chemotherapy, our observations support a role for ICI therapy in patients with tumors characterized by high expression of the c-Score that would otherwise be ineligible for ICI therapy based on TMB stratification alone. The clinical utility of the c-Score as a predictor of ICI response was further validated using the TIDE25 and PredictIO26 databases.
Since genome-wide tumor mutagenesis may promote T cell-inflammation20,54, we evaluated the c-Score relationship with DNA repair mutations. We examined tumors with MSI as a proof of principle since those with MMRD, the pathway resulting in MSI, respond favorably to ICIs43,54, and explored whether tumors harboring HRD are also c-Scorehi. We found tumors with high MSI and HRD scores harbored increased expression of the c-Score, and that a higher proportion of patients with ‘TMB-associated’ tumors (i.e., c-Scorehi and elevated TMB) had a mutation in a DNA damage repair gene versus ‘TMB-unassociated’ tumors. MSI tumors had elevated c-Score expression, and the HRD associations support a role for DNA repair pathway defects in cancer-immunity cycle that extends beyond MMRD. However, the c-Scorehi association in HRD tumors was lost when controlling for histological tumor type, suggesting T cell-inflamed phenotypes vary across HRD tumor types.
To examine the relationship of somatic mutations on T cell-inflammation as predicted by the c-Score, we also applied an agnostic approach. We identified the most common mutated genes in TCGA solid tumors and evaluated the c-Score relationship with tumors harboring mutations in these genes. When considering the three most mutated genes (TTN, TP53, and MUC16), we found that tumors with mutations in these genes had elevated c-Score and TMB measurements. Elevated TMB may reflect increased immune activation through a mutational load triggered by genomic instability, rather than an effect of these mutated genes on TMB. Deciphering the mechanisms underlying these observations was beyond the scope of this study and merits further investigation.
The relationship between KRAS-mutated tumor genomes and tumor inflammation predicted by the c-Score was also examined. We evaluated this relationship since oncogenic KRAS has been implicated as a prognostic and predictive cancer biomarker. When considering cancer immunity, oncogenic KRAS may promote immunosuppressive tumor microenvironments51,52. Moreover, KRAS is mutated in ~25% of all cancers50, and in >90% of cases in certain cancers such as pancreatic ductal adenocarcinoma55. The immunosuppressive role of oncogenic KRAS is multifaceted, directly and indirectly affecting various steps of the cancer-immunity cycle56,57,58. Our observations suggest that, although KRAS mutations may contribute to immunosuppressive properties, they do not fully predict tumor inflammation. We propose that tumor T cell-inflammation predicted by increased expression of the c-Score is mediated by mechanisms specific to histological tumor types and their microenvironments, which include the interplay between the immunosuppressive properties of KRAS mutations with the immune stimuli resulting from DNA repair deficiencies characterized by elevated TMB and neoantigen tumor loads. In support of this paradigm, KRAS mutations did not impart lower c-Scores in lung adenocarcinoma. That is, the immunosuppressive effects of mutated KRAS may be offset by a higher TMB in these KRAS-mutated tumors. In fact, KRAS mutational status has been associated with increased PD-L1 expression in non-small-cell lung cancers (NSCLC)59, and anti-PD-1 therapy responses in NSCLC have been correlated with KRAS mutations occurring alone or together with TP53 loss60. Contrary to NSCLC, KRAS mutations in pancreatic adenocarcinoma are understood to generate an immunosuppressive milieu61. To this end, we found increased expression of the c-Score in KRAS wild-type versus mutated PAAD. Although a higher TMB was observed in KRAS-mutated compared to wild-type PAAD, the TMB levels in PAAD rarely meet the ≥10 mutations/megabase threshold predictive of tumor inflammation and ICI response, suggesting the immunosuppressive effects of oncogenic KRAS signaling overpower the immune-stimulating features of increased TMB.
In summary, we show that the c-Score identifies tumor subsets with predicted T cell-inflammation and ICI treatment benefits across a wide spectrum of cancers. Our findings suggest that the c-Score is complementary to TMB in predicting ICI therapy clinical outcomes, providing clinical equipoise for its validation in clinical trial settings.
Methods
Cohort and description of publicly available TCGA datasets
Datasets were downloaded as described in the DATA AVAILABILITY STATEMENT. Samples with matched RNA (EBPlusPlusAdjustPANCAN_IlluminaHiSeq_RNASeqV2.geneExp.tsv) and MAF data were then filtered (i.e., those not passing quality control, or with other annotations advising removal). Only primary tumors were included. Patients with a prior history of cancer or treatments were excluded. Except for CHOL, esophageal carcinoma (ESCA), mesothelioma (MESO), sarcoma (SARC), STAD, TGCT, and thymoma THYM, the prominent histological subtype was selected. Hematological cancers (diffuse large B-cell lymphoma, [DLBC], acute myeloid leukemia [LAML]) were omitted from downstream analyses due to their immunological properties. In cases of multiple aliquots for primary tumors, the first aliquot was selected. In cases of multiple samples per patient, the aliquot with the highest median expression was selected. A final cohort of 6987 patients was assembled for analyses. In Fig. 1, tumor types with clinical response data to ICI27 were matched to TCGA nomenclature, with renal cells classified as KIRC and NSCLC classified as lung adenocarcinoma LUAD. The barcodes, c-Scores, and associator designations for all 6987 patients are listed in Supplementary Data File 2.
Patient enrollment, data collection, processing, and sequencing for the POG570 cohort
This work was approved by and conducted under the University of British Columbia BC Cancer Research Ethics Board (H12-00137, H14-00681), in compliance with the Tri-Council Policy Statement and the FDA regulations (Belmont) and the Good Clinical Practice principles (Helsinki) and approved by the institutional review board. Patients with advanced or metastatic disease gave informed written consent and were enrolled into the POG (NCT02155621) as described previously40,41,62. Treatment histories, response, and survival data for the POG cohort were collected retrospectively using the BC Cancer Pharmacy database and chart review. Durable clinical benefit was defined as treatment for greater than 6 months without disease progression40. Time to progression was defined as the time from ICI initiation to the date of discontinuation due to progression, and overall survival as the time from ICI initiation to death. Tumor specimens were collected using needle core biopsies, endobronchial ultrasound biopsies, or tissue resection. Solid tumor specimens were snap frozen, while liquid biopsies were spun down into a cell pellet and resuspended. RNA extraction was performed as previously described41. Briefly, following pathological review of samples for tumor cellularity, four 50 um sections were pooled in 2.0 mL tubes with 420-600 uL of RLT Plus lysis buffer (Qiagen) and tris (2-carboxyethyl) phosphine, and co-extraction of DNA and RNA was done from 3–11 tubes using an Aline EvoPure kit (Aline Biosciences, R-907-400-C5). Transcriptomes were sequenced to a target of 150–200 million 75-bp end reads on Illumina HiSeq2500 or NextSeq500. Sequencing statistics were performed as previously described40. The barcodes and c-Scores for the POG570 and ICI-treated cohorts are listed in Supplementary Data File 3 and Supplementary Data File 4, respectively.
RNA-seq analyses
Gene identifiers were converted using the mapIds function of the AnnotationDbi package (v.1.54.1) and org.Hs.eg.db (v.3.13.0). In cases where multiple matches were found, the first match was used. The c-Score was calculated by taking the mean log2(TPM + 0.001) expression of CCL4, CCL5, CXCL9, and CXCL10 for each patient. c-Scorehi and c-Scorelo tumors were defined based on samples with ≥3rd (4.835088) or ≤1st (1.940425) quartile expression, respectively. For downstream analyses, tumor types with at least 5 patients per c-Score group were included, resulting in 25 tumor types for subsequent evaluation (n = 6455). CIBERSORT scores for leukocyte fractions of TCGA patients were obtained from Thorsson et al.48. Duplicate scores for neutrophils and eosinophils were identified, and the first score was used. For single sample Gene Set Enrichment Analysis (ssGSEA), log2(TPM + 0.001) data were filtered to remove genes with 0 expressions in greater than 50% of the samples. Genesets used for ssGSEA were chosen based on their representation of processes in the cancer-immunity cycle, including co-stimulation and co-inhibition of antigen presenting cells, major histocompatibility complex I presence, co-stimulation and co-inhibition of T cells, and cytolytic activity31, BATF3 dendritic cell presence7, and interferon gamma signaling associated with response to ICI32. When present, CCL4, CCL5, CXCL9, and CXCL10 were removed from these genesets prior to analysis by ssGSEA. ssGSEA was calculated using the gsva function of the GSVA package (v.1.40.1) as previously described6. To display ssGSEA scores on a positive scale in Fig. 2b, arbitrary units (AU) were created as follows: a pseudo-value equal to 1 minus the median value of the ssGSEA score for genesetj for the c-Scorelo+med group was added to the median value of the ssGSEA score for genesetj for the c-Scorelo+med and c-Scorehi groups, such that the lowest transformed value was 1. This is shown in equations 1 and 2 below:
For genesetj and groupi, where i = c-Scorehi or c-Scorelo+med,
where y = c-Scorelo+med specifically, such that the lowest ssGSEA score value is 1:
Statistics were calculated on non-transformed values. For analysis of the POG570 dataset, RNA-seq reads were aligned using STAR73 (v.2.5.2b) and expression was quantified using RSEM74 (v.1.3.0) as TPMs. All final RNA-seq count data are expressed as log2(TPM + 0.001). Input indexed files for STAR and RSEM were generated from the hg38 reference genome (http://hgdownload.cse.ucsc.edu/goldenPath/hg38/bigZips/) and gene annotations were based on Ensembl v.85.
Mutational data processing
For TCGA analysis, MAF files were processed using the maftools package (v.2.8.0). Tumor mutation burden was calculated using the tmb function, with a capture size of 35.8 megabases used, as outlined by the tcgaCompare function. SNV-derived neoantigen load was obtained for TCGA patients48 and was used to corroborate TMB analyses. To compare mutation rates of specific mutation types in DNA repair and fidelity pathways (MMR: MLH1, MSH2, MSH6, PMS2; HR: BRCA1, BRCA2, PALB2; DNA replication: POLE, POLD1), the number of respective mutations, divided by the total number of patients in that group, were used. To compare overall rates of mutations in DNA repair and fidelity pathways, the total number of patients with any mutation in these genes was tallied and compared between groups. Protein-truncating mutations were defined as frameshift deletions, frameshift insertions, nonsense, splice site, translation start site, and nonstop mutations.
MSI and HRD subtyping and analyses
For TCGA analysis, MSI status was inferred from MANTIS scores as previously calculated47, with MANTIShi patients having a score greater than 0.4. The HRD score was obtained for TCGA patients48,49, which represents a combined metric of HRD-Loss of heterozygosity63, large-scale state transitions64, and a number of telomeric allelic imbalances65, and was used to infer an HRD phenotype. HRDScorelo (a proxy for HR proficiency [HRP]) and HRDScorehi (a proxy for HRD) tumor designations were assigned based on third-quartile thresholds of HRDScore values across all 31 tumor types. Given the high prevalence of HRD in BRCA and OV compared to the rest of the cancers, distributions, and quartiles for these tumors were calculated separately. HRDScorehi tumors were defined as those with an HRDScore ≥46 in BRCA and OV, and ≥27 in the rest of the cohort. Following designation as c-Scorelo, c-Scoremed, and c-Scorehi, any tumors with high MANTIS or high HRDScores were further classified. Tumors with both high MANTIS and HRDScores were classified as MANTIShi. The final subclassifications were c-Scorelo+med (n = 3797), c-Scorehi (n = 1045), HRDScorehi (n = 1670), MANTIShi (n = 205), with HRDScorelo and MANTISlo comprised of both c-Scorelo+med and c-Scorehi patients. For KRAS mutational status analyses, tumors with missing MANTIS/HRDScore values were classified as MANTISlo/HRDScorelo. For POG570 dataset, MSI sensor scores were computed as previously described66. Scores above 0.2 were considered high. HRDetect scores were computed using a logistic regression model previously described by Davies and colleagues67, and implementation can be found at https://github.com/eyzhao/hrdtools68.
Survival and regression analyses
Kaplan–Meier survival analysis was performed for TTP and OS using the R packages survival (v.2.42.3) and survminer (v.0.4.2). Differences in nonparametric survival functions were assessed across groups using log-rank tests. To ensure sample sizes were large enough for comparison of the chemokine groups, the c-Scorelo and c-Scoremed groups were combined. Cox proportional hazards models were performed using the R packages survival (v.2.42.3) and forest model (v0.5.0). Log-rank tests were used to calculate P values, and in the case of ties, the Efron approximation was used. Linear regression models were used to account for histology in the durable clinical benefit group versus no clinical benefit group (NCB). In the case of multivariate analyses using histology, only tumor types with at least three samples were included. Tumor types for the ICI treated cohort are as previously described40. For analysis of PanCuRx20 and COMPASS datasets42, c-Score designations were calculated relative to the sample sets as opposed to cut offs used for TCGA and POG analysis.
c-Score validation in pan-cancer ICI-treated datasets
The c-Score (CCL4, CCL5, CXCL9, CXCL10) was input into the TIDE25 and PredictIO26 biomarker query (http://tide.dfci.harvard.edu/setquery/, https://predictio.ca/explore/biomarker/request, respectively) using default settings. For PredictIO, both male and female sexes were included, as well as patients that received anti-CTLA4, anti-PD-1, anti-PD-L1, or a combination. Both FPKM and TPM sequencing results were included.
Statistical analyses
All statistical analyses were done using the R Statistical programming language (v.4.0.4) and select packages from the CRAN and Bioconductor repositories. Specific statistical tests were performed as outlined in the text. Two-tailed P values are shown. P values were adjusted for multiple tests when necessary, using FDR correction via the Benjamini-Hochberg procedure. Controlling for variables was performed using a multivariate linear regression model.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Gene expression for TCGA pan-cancer datasets (TCGA Pan-Cancer (PANCAN)/gene expression RNA-sequencing (RNA-seq)/TOIL RSEM tpm/tcga_RSEM_gene_tpm), containing the log2(TPM + 0.001) expression matrix of 10,535 samples across 60,499 features, was downloaded from the University of California Santa Cruz data repository (https://xenabrowser.net/datapages/). Mutation (mc3.v0.2.8.PUBLIC.maf), annotation (merged_sample_quality_annotations.tsv), and clinical data files (TCGA-CDR-SupplementalTableS1.xlsx, clinical_PANCAN_patient_with_followup.tsv) were downloaded from respective manifests and GDC transfer tool (https://gdc.cancer.gov/about-data/publications/pancanatlas, https://gdc.cancer.gov/access-data/gdc-data-transfer-tool). The genomic and transcriptomic sequence datasets, including metadata with library construction and sequencing approaches for the POG570 cohort, have been deposited at the European Genome-phenome Archive (http://www.ebi.ac.uk/ega/) as part of the study EGAS00001001159, and can be downloaded from: http://bcgsc.ca/downloads/POG570/. Specific analysis files for the full cohort and ICI treated cohorts have been previously reported40,41.
Code availability
All bioinformatic analyses were done using the R Statistical programming language (v.4.0.4). The specific packages, parameters, and versions used are outlined in their respective sections found in Methods. The code used in this study, including pipelines for filtering, processing, and analysis, is available upon request.
References
Palucka, A. K. & Coussens, L. M. The basis of oncoimmunology. Cell 164, 1233–1247 (2016).
Chen, D. S. & Mellman, I. Oncology meets immunology: the cancer-immunity cycle. Immunity 39, 1–10 (2013).
Trujillo, J. A., Sweis, R. F., Bao, R. & Luke, J. J. T cell–inflamed versus non-T cell–inflamed tumors: a conceptual framework for cancer immunotherapy drug development and combination therapy selection. Cancer Immunol. Res. 6, 990–1000 (2018).
van der Woude, L. L., Gorris, M. A. J., Halilovic, A., Figdor, C. G. & de Vries, I. J. M. Migrating into the tumor: a roadmap for T cells. Trends Cancer 3, 797–808 (2017).
Nagarsheth, N., Wicha, M. S. & Zou, W. Chemokines in the cancer microenvironment and their relevance in cancer immunotherapy. Nat. Rev. Immunol. 17, 559–572 (2017).
Romero, J. M. et al. A four-chemokine signature is associated with a T cell-inflamed phenotype in primary and metastatic pancreatic cancer. Clin. Cancer Res. 2803, 2019 (2020).
Spranger, S., Dai, D., Horton, B. & Gajewski, T. F. Tumor-residing Batf3 dendritic cells are required for effector T cell trafficking and adoptive T cell therapy. Cancer Cell 31, 711–723.e4 (2017).
Liu, J. et al. Local production of the chemokines CCL5 and CXCL10 attracts CD8+ T lymphocytes into esophageal squamous cell carcinoma. Oncotarget 6, 24978–26489 (2015).
Muthuswamy, R., Corman, J. M., Dahl, K., Chatta, G. S. & Kalinski, P. Functional reprogramming of human prostate cancer to promote local attraction of effector CD8+T cells. Prostate 76, 1095–1105 (2016).
Mlecnik, B. et al. Biomolecular network reconstruction identifies T-cell homing factors associated with survival in colorectal cancer. Gastroenterology 138, 1429–1440 (2010).
Berghuis, D. et al. Pro-inflammatory chemokine-chemokine receptor interactions within the Ewing sarcoma microenvironment determine CD8+ T-lymphocyte infiltration and affect tumour progression. J. Pathol. 223, 347–357 (2011).
Harlin, H. et al. Chemokine expression in melanoma metastases associated with CD8 + T-CeII recruitment. Cancer Res 69, 3077–3085 (2009).
Andersson, A. et al. IL-7 promotes CXCR3 ligand-dependent T cell antitumor reactivity in lung cancer. J. Immunol. 182, 6951–6958 (2009).
Kryczek, I. et al. Phenotype, distribution, generation, and functional and clinical relevance of Th17 cells in the human tumor environments. Blood 114, 1141–1150 (2009).
Zhang, L. et al. Intratumoral T cells, recurrence, and survival in epithelial ovarian cancer. N. Engl. J. Med. 348, 203–213 (2003).
Spranger, S. & Gajewski, T. F. Impact of oncogenic pathways on evasion of antitumour immune responses. Nat. Rev. Cancer 18, 139–147 (2018).
Jardim, D. L., Goodman, A., de Melo Gagliato, D. & Kurzrock, R. The challenges of tumor mutational burden as an immunotherapy biomarker. Cancer Cell 39, 154–173 (2021).
Wellenstein, M. D. & de Visser, K. E. Cancer-cell-intrinsic mechanisms shaping the tumor immune landscape. Immunity 48, 399–416 (2018).
Mandal, R. et al. Genetic diversity of tumors with mismatch repair deficiency influences anti–PD-1 immunotherapy response. Sci. (80-.) 364, 485–491 (2019).
Connor, A. A. et al. Association of distinct mutational signatures with correlates of increased immune activity in pancreatic ductal adenocarcinoma. JAMA Oncol. 1–10 https://doi.org/10.1001/jamaoncol.2016.3916 (2016).
Mouw, K. W., Goldberg, M. S., Konstantinopoulos, P. A. & D’Andrea, A. D. DNA damage and repair biomarkers of immunotherapy response. Cancer Discov. 7, 675–693 (2017).
Terrero, G. et al. Ipilimumab/nivolumab therapy in patients with metastatic pancreatic or biliary cancer with homologous recombination deficiency pathogenic germline variants. JAMA Oncol. 1–3 https://doi.org/10.1001/jamaoncol.2022.0611 (2022).
Marabelle, A. et al. Association of tumour mutational burden with outcomes in patients with advanced solid tumours treated with pembrolizumab: prospective biomarker analysis of the multicohort, open-label, phase 2 KEYNOTE-158 study. Lancet Oncol. 21, 1353–1365 (2020).
Chen, D. S. & Mellman, I. Elements of cancer immunity and the cancer-immune set point. Nature 541, 321–330 (2017).
Jiang, P. et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24, 1550–1558 (2018).
Bareche, Y. et al. Leveraging Big Data of Immune Checkpoint Blockade Response Identifies Novel Potential Targets. Ann. Oncol. https://doi.org/10.1016/j.annonc.2022.08.084 (2022).
Yarchoan, M., Hopkins, A. & Jaffee, E. M. Tumor mutational burden and response rate to PD-1 inhibition. N. Engl. J. Med. 377, 2500–2501 (2017).
Gajewski, T. F., Schreiber, H. & Fu, Y.-X. Innate and adaptive immune cells in the tumor microenvironment. Nat. Immunol. 14, 1014–1022 (2013).
Melero, I. et al. Evolving synergistic combinations of targeted immunotherapies to combat cancer. Nat. Rev. Cancer 15, 457–472 (2015).
Ma, D. Y. & Clark, E. A. The role of CD40 and CD40L in dendritic cells. Semin. Immunol. 21, 265–272 (2009).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Ayers, M. et al. IFN-γ–related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest 127, 2930–2940 (2017).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic β-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Herbst, R. S. et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 515, 563–567 (2014).
Havel, J. J., Chowell, D. & Chan, T. A. The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat. Rev. Cancer 19, 133–150 (2019).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 2189–2199 https://doi.org/10.1056/NEJMoa1406498 (2014).
Danilova, L. et al. Association of PD-1/PD-L axis expression with cytolytic activity, mutational load, and prognosis in melanoma and other solid tumors. Proc. Natl Acad. Sci. USA. 113, E7769–E7777 (2016).
Spranger, S. et al. Density of immunogenic antigens does not explain the presence or absence of the T-cell–inflamed tumor microenvironment in melanoma. Proc. Natl Acad. Sci. 113, E7759–E7768 (2016).
Balli, D., Rech, A. J., Stanger, B. Z. & Vonderheide, R. H. Immune cytolytic activity stratifies molecular subsets of human pancreatic cancer. Clin. Cancer Res. 23, 3129–3138 (2017).
Pender, A. et al. Genome and transcriptome biomarkers of response to immune checkpoint inhibitors in advanced solid tumors. Clin. Cancer Res. 27, 202–212 (2021).
Pleasance, E. et al. Pan-cancer analysis of advanced patient tumors reveals interactions between therapy and genomic landscapes. Nat. Cancer 1, 452–468 (2020).
Aung, K. L. et al. Genomics-driven precision medicine for advanced pancreatic cancer: Early results from the COMPASS trial. Clin. Cancer Res 24, 1344–1354 (2018).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Wang, F. et al. Evaluation of POLE and POLD1 mutations as biomarkers for immunotherapy outcomes across multiple cancer types. JAMA Oncol. 5, 1504 (2019).
Lu, C. et al. DNA sensing in mismatch repair-deficient tumor cells is essential for anti-tumor immunity. Cancer Cell 39, 96–108.e6 (2021).
Guan, J. et al. MLH1 deficiency-triggered DNA hyperexcision by exonuclease 1 activates the cGAS-STING pathway. Cancer Cell 39, 109–121.e5 (2021).
Bonneville, R. et al. Landscape of microsatellite instability across 39 cancer types. JCO Precis. Oncol. 1–15 https://doi.org/10.1200/po.17.00073 (2017).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 1–19 (2018).
Knijnenburg, T. A. et al. Genomic and molecular landscape of DNA damage repair deficiency across the cancer genome atlas. Cell Rep. 23, 239–254.e6 (2018).
Li, S., Balmain, A. & Counter, C. M. A model for RAS mutation patterns in cancers: finding the sweet spot. Nat. Rev. Cancer 18, 767–777 (2018).
Pylayeva-Gupta, Y., Lee, K. E., Hajdu, C. H., Miller, G. & Bar-Sagi, D. Oncogenic kras-induced GM-CSF production promotes the development of pancreatic neoplasia. Cancer Cell 21, 836–847 (2012).
Carvalho, P. D. et al. KRAS oncogenic signaling extends beyond cancer cells to orchestrate the microenvironment. Cancer Res 78, 7–14 (2018).
Foeng, J., Comerford, I. & McColl, S. R. Harnessing the chemokine system to home CAR-T cells into solid tumors. Cell Rep. Med 3, 100543 (2022).
Le, D. T. et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Sci. (80-.) 357, 409–413 (2017).
Mizrahi, J. D., Surana, R., Valle, J. W. & Shroff, R. T. Pancreatic cancer. Lancet 395, 2008–2020 (2020).
Zdanov, S. et al. Mutant KRAS conversion of conventional T cells into regulatory T cells. Cancer Immunol. Res 4, 354–365 (2016).
El-Jawhari, J. J. et al. Blocking oncogenic RAS enhances tumour cell surface MHC class I expression but does not alter susceptibility to cytotoxic lymphocytes. Mol. Immunol. 58, 160–168 (2013).
Coelho, M. A. et al. Oncogenic RAS signaling promotes tumor immunoresistance by stabilizing PD-L1 mRNA. Immunity 47, 1083–1099.e6 (2017).
Garon, E. B. et al. Pembrolizumab for the treatment of non–small-cell lung cancer. N. Engl. J. Med. 372, 2018–2028 (2015).
Dong, Z. Y. et al. Potential predictive value of TP53 and KRAS mutation status for response to PD-1 blockade immunotherapy in lung adenocarcinoma. Clin. Cancer Res 23, 3012–3024 (2017).
Bear, A. S., Vonderheide, R. H. & O’Hara, M. H. Challenges and Opportunities for Pancreatic Cancer Immunotherapy. Cancer Cell https://doi.org/10.1016/j.ccell.2020.08.004 (2020).
Laskin, J. et al. Lessons learned from the application of whole-genome analysis to the treatment of patients with advanced cancers. Mol. Case Stud. 1, a000570 (2015).
Abkevich, V. et al. Patterns of genomic loss of heterozygosity predict homologous recombination repair defects in epithelial ovarian cancer. Br. J. Cancer 107, 1776–1782 (2012).
Popova, T. et al. Ploidy and large-scale genomic instability consistently identify basal-like breast carcinomas with BRCA1/2 inactivation. Cancer Res 72, 5454–5462 (2012).
Birkbak, N. J. et al. Telomeric allelic imbalance indicates defective DNA repair and sensitivity to DNA-damaging agents. Cancer Discov. 2, 366–375 (2012).
Niu, B. et al. MSIsensor: Microsatellite instability detection using paired tumor-normal sequence data. Bioinformatics 30, 1015–1016 (2014).
Davies, H. et al. HRDetect is a predictor of BRCA1 and BRCA2 deficiency based on mutational signatures. Nat. Med. 23, 517–525 (2017).
Zhao, E. Y. et al. Homologous recombination deficiency and platinum-based therapy outcomes in advanced breast cancer. Clin. Cancer Res 23, 7521–7530 (2017).
Acknowledgements
This work was supported by the Terry Fox Research Institute (project no. 1078), the Québec Cancer Consortium with funding from the Ministère de l’Économie et de l’Innovation du Québec through the Fonds d’accélération des collaborations en santé, and the Ontario Institute for Cancer Research (PanCuRx Translational Research Initiative) through funding provided by the Government of Ontario, the Wallace McCain Centre for Pancreatic Cancer supported by the Princess Margaret Cancer Foundation, the Canadian Cancer Society Research Institute, and the Pancreatic Cancer Canada Foundation. The study was also supported by a charitable donation from the Canadian Friends of the Hebrew University (Alex U. Soyka). This work includes data generated from the Personalized OncoGenomics program at BC Cancer supported by the BC Cancer Foundation and their donors and Genome British Columbia. J.M. Romero is supported by a Fonds de recherche du Québec–Santé MD-PhD scholarship and a Vanier Canada Graduate Scholarship. Y. Wang was supported by a Vanier Canada Graduate Scholarship, the Fonds de recherche du Québec–Santé/Ministère de la Santé et des Services sociaux training program, and the McGill University Surgical-Scientist Program. S. Gallinger is the recipient of an Investigator Award from the Ontario Institute for Cancer Research. G. Zogopoulos is a clinical research scholar of the Fonds de recherche du Québec–Santé and recipient of the Michal & Renata Hornstein Career Award from McGill University. Funding sources for this study had no role in the design, collection, analysis, interpretation or writing of this manuscript.
Author information
Authors and Affiliations
Contributions
J.M.R.: Conceptualization, data curation, formal analysis, investigation, visualization, methodology, writing-original draft, writing-review and editing. E.T.: Data curation, formal analysis, visualization, methodology, writing-original draft, writing-review and editing. Y.W.: Formal analysis, resources. A.P.: Resources, methodology. G.H.J.: Data curation, methodology, formal analysis. A.Z.: Data curation, methodology. B.G., T.L., J.V.: Formal analysis. L.W.: Methodology, writing-review and editing. B.G.: Writing-review and editing. G.M.O.: Writing-review and editing, resources. S.J.M.J., M.A.M.: Resources. J.M.W.: Resources, project administration. S.G.: Resources, funding acquisition, writing-review and editing. J.L.: Resources, funding acquisition. G.Z.: Conceptualization, resources, supervision, funding acquisition, writing-original draft, project administration, writing-review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
This work included investigators throughout the research progress, with roles and responsibilities as outlined in CONTRIBUTIONS. The clinical POG dataset used in this study has been approved by the local ethics review committee, as outlined in Patient enrollment, data collection, processing, and sequencing for the POG570 cohort section in METHODS. The anonymized nature of these datasets used does not result in stigmatization, incrimination, discrimination, or otherwise personal risk to participan.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Romero, J.M., Titmuss, E., Wang, Y. et al. Chemokine expression predicts T cell-inflammation and improved survival with checkpoint inhibition across solid cancers. npj Precis. Onc. 7, 73 (2023). https://doi.org/10.1038/s41698-023-00428-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-023-00428-2
- Springer Nature Limited