Abstract
This study aimed to develop and validate prediction models to estimate the risk of death and intensive care unit admission in COVID-19 inpatients. All RT-PCR-confirmed adult COVID-19 inpatients admitted to Fujian Provincial Hospital from October 2022 to April 2023 were considered. Elastic Net Regression was used to derive the risk prediction models. Potential risk factors were considered, which included demographic characteristics, clinical symptoms, comorbidities, laboratory results, treatment process, prognosis. A total of 1906 inpatients were included finally by inclusion/exclusion criteria and were divided into derivation and test cohorts in a ratio of 8:2, where 1526 (80%) samples were used to develop prediction models under a repeated cross-validation framework and the remaining 380 (20%) samples were used for performance evaluation. Overall performance, discrimination and calibration were evaluated in the validation set and test cohort and quantified by accuracy, scaled Brier score (SbrS), the area under the ROC curve (AUROC), and Spiegelhalter-Z statistics. The models performed well, with high levels of discrimination (AUROCICU [95%CI]: 0.858 [0.803,0.899]; AUROCdeath [95%CI]: 0.906 [0.850,0.948]); and good calibrations (Spiegelhalter-ZICU: − 0.821 (p-value: 0.412); Spiegelhalter-Zdeath: 0.173) in the test set. We developed and validated prediction models to help clinicians identify high risk patients for death and ICU admission after COVID-19 infection.
Similar content being viewed by others
Explore related subjects
Find the latest articles, discoveries, and news in related topics.Introduction
The coronavirus disease 2019 (COVID-19) pandemic has been a major cause of mortality worldwide and caused an unprecedented global health crisis1,2. A surge in the number of critically ill patients in intensive care units (ICUs) all over the world has placed a burden on ICUs3. Health care systems have been strongly affected and collapsed in most countries worldwide as the pandemic unfolded4. During the peak of the pandemic, rapid diagnosis and assessment of COVID-19 patients at risk for death and ICU admission are very crucial to managing potential capacity challenges. In this scenario, predictive models using readily available data are capable of identifying those at risk of severe disease effectively, greatly contributing to resource allocation and triage5.
Logistic regression models were most common among those prediction models in medicine6. Regression-based scoring systems has been used to assist with triage of patients with COVID-197,8,9. These investigations collectively underscore the utility of regression-based scoring systems in assisting clinicians with timely intervention decisions, crucial for mitigating in-hospital mortality. However, they rely on logistic regression which is one of conventional methods and these static scores may not fully capture patient progression, necessitating a deeper understanding of how to tailor interventions based on individual patient conditions. In recent years, significant progress in predictive modeling, particularly through the application of machine learning (ML) methodologies such as elastic net, has offered an advantage over logistic regression on forecasting capabilities and was preferable for model fitting because it can potentially retain collinear variables10,11. Although prior studies have published prognostic models for COVID-19 patients, the majority of them either rely on logistic regression not an elastic net regression model or develop prediction models for death or ICU admission only on critically ill adult patients admitted to the ICU with no external validation or with a small sample size11,12,13,14,15,16.
To date, taking account of the paucity of validated prognostic models for death and ICU admission for COVID-19 patients with relatively large sample size, we are committed to developing and validating prediction models using elastic net regression to support resource allocation and disease management. We applied a cross-validation framework for model development and carefully evaluated the generalizability of the prediction models on an independent test cohort.
Methods
Study design and patient population
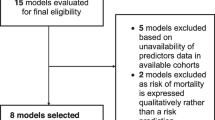
The dataset includes demographic characteristics, clinical symptoms, comorbidities, laboratory results, treatment process and prognosis based on all adult (≥ 18 years of age) COVID-19 patients admitted to Fujian Provincial Hospital between October 2022 and April 2023. Patients with incomplete clinical and laboratory data were excluded. The diagnosis of COVID-19 is based on a positive reverse-transcriptase polymerase-chain reaction (RT-PCR) test. In this retrospective study, patients from the Fujian Provincial Hospital COVID-19 dataset were recruited split into training (n = 1526) and internal test (n = 380) data sets. Besides, external test cohort comes from patients from Fujian Provincial Geriatric Hospital (n = 887) (Fig. 1).
Flow diagram of the study. 1906 inpatients were included from Fujian Provincial Hospital finally by inclusion/exclusion criteria and inpatients were divided into derivation cohort (n = 1526) and test cohort (n = 380). 887 inpatients were included from Fujian Provincial Geriatric Hospital finally by inclusion/exclusion criteria for external validation.
Ethics approval and consent to participate
The studies involving human participants were reviewed and approved by the Ethics Review Committee of the Fujian Provincial Hospital (Grant No. K2023-12-016). All clinical investigations were conducted in accordance with the Declaration of Helsinki. We analyzed the data anonymously and written informed consent was waived by ethics committee of the Fujian Provincial Hospital due to the retrospective nature of our study of routine clinical data.
Treatment protocols
Inpatient patients all received appropriate treatment based on their medical conditions. Treatment protocols of patients in the hospital include oxygen therapy, mechanical ventilation, nasal high flow oxygen therapy, invasive ventilation, thymus method, mask oxygen, prone position ventilation, nasal catheter for oxygen, extracorporeal membrane oxygenation (ECMO), antibiotic therapy, anticoagulant therapy glucocorticoid and antipyretic analgesics.
Outcomes
The primary outcomes were in-hospital mortality and ICU admission. We developed and internally validated a prediction model for in-hospital mortality and ICU admission in the dataset form Fujian Provincial Hospital. Besides, we externally validated the prediction model in another dataset from Fujian Provincial Geriatric Hospital.
Predictor variables
We extracted data from electronic medical records based on established risk factors and considered 161 candidate predictors, including demographic information (age, sex), clinical symptoms (cough, fever, fatigue, diarrhea, dry cough, etc.), comorbidities (hypertension, diabetes etc.) and treatments (oxygen therapy, mechanical ventilation, Extracorporeal membrane oxygenation, etc.). A full list of variables is available at Supplemental Materials Table S1.
Statistical methods
Preprocessing
The gathered data from 1906 patients were integrated and the duplicated records were removed. The original dataset consisted of 455 variables. We omitted variables that had more than 30% missing values to minimize bias resulting from missing data and finally a total of 161 variables remained. The missing values were imputed using the K-Nearest Neighbors (KNN) method to generate the analytical dataset.
Model training
We employed elastic net, a regularized regression model17, on complete cases to develop and validate the risk prediction models. Samples were divided into two parts of a ratio of 8:2 using stratified sampling, where 80% of the total study samples were used for model development (derivation cohort). The remaining 20% of the samples were used as a test cohort for validation. The model was applied to an external dataset for external validation.
Risk prediction models were built using the derivation cohort. The derivation cohort was further divided into training and validation sets using a cross-validation framework18 (k = 3, repeats = 2) to allow interval validation. Specifically, the training set was randomly partitioned into three roughly equal-sized parts. We then left out one part as the validation set and the model was built on the remaining parts. The leave-out-modelling process was conducted recursively until each part was treated as a validation set for once. The cross-validation modelling process was repeated twice. Therefore, the number of training samples is double that of the original deviation cohort. We tuned one hundred combinations of hyperparameters, where we specified ten different alphas and ten different lambdas. For each combination of hyperparameters, six (k times number of repeats) prediction models were built for each outcome of interest. The model performance on the validation sets, which was calculated by averaging the model performance of the six models, was used to determine which combination of hyperparameters should be applied to our final prediction model. Finally, hyperparameters that generated the best model performance (highest value of scaled Brier score) were chosen and applied to the whole derivation cohort to obtain our final prediction model.
Assessment of accuracy
We evaluated the model performance (overall performance, discrimination, and calibration) in the validation and test cohort. Accuracy, sensitivity, specificity, negative predictive values, positive predictive values and scaled Brier score were calculated to assess the overall performance of prediction models. The ROC curve was plotted and the area under the ROC curve (AUROC) was calculated to visualize and quantify the calibration19. Spiegelhalter-Z statistics20 were calculated to evaluate calibration. Calibration plots21 which compared the observed risks with the precited risks were provided to visualize the predictive ability of the risk prediction models. The effect coefficients of the included predictors in the risk prediction models.
Results
Participants
Overall, 1906 COVID-19 confirmed cases were included in our study. Of these, 1526 (event rateICU: 17.3%, event ratedeath: 7.6%) inpatients were selected as the derivation cohort by stratified sampling and the remaining 380 (event rateICU: 17.1%, event ratedeath: 7.4%) inpatients were used as test cohort. Table 1 presents the clinical characteristics of our study samples. In total, 329 (17.3%) patients were transferred to ICU. ICU inpatients were older (median: 76 years, interquartile range [IQR]: 65–83 years) than non-ICU inpatients (median: 72 years, IQR: 56–82 years), and the difference is statistically significant (p-value: 0.001). ICU inpatients more frequently underwent hypertension (62.6% versus 54.1%) and diabetes (42.2% versus 34.9%). Baseline characteristics for patients in our study who died vs survived are displayed in Table 2. 144 (7.6%) patients were non-survivors. The median age in non-survivors (median: 86 years, IQR: 80–91 years was significantly older than that of survivors (median:71 years, IQR: 56–81 years). The non-survivor group has higher proportions of patients with COVID-19 related symptoms including expectoration, cough, malaise fever, shortness of breath and fatigue. Baseline characteristics for patients in the development cohort (Fujian Provincial Hospital) and the external test cohort (Fujian Provincial Geriatric Hospital) are demonstrated in Table 3. The median age in the external test cohort (median: 85 years, IQR: 76–91 years) was significantly older than that in the development cohort (median: 73 years, IQR: 58–82 years). The external test cohort has higher proportions of patients with COVID-19 related symptoms including expectoration and cough than the development cohort (p-value: 0.001).
Model evaluation for COVID-19-associated mortality and ICU admission
The performance of the final prediction models on the validation set and test cohort were demonstrated in Table 4 and Fig. 2. As was shown in Table 4, the prediction model for ICU admission shows good overall performance on the validation set and test cohort, with accuracy of 0.877 and 0.879 and scaled Brier score of 0.327 and 0.330, respectively. The Spiegelhalter-Z statistics with a value of − 0.285 (p-value: 0.776) on the validation set and − 0.821 (p-value: 0.412) on the test set indicate good calibration. ROC plots were provided to visualize the discrimination. AUROC was 0.829 (95% CI: 0.810–0.850) and 0.858 (95% CI: 0.803–0.899) on the validation set and test cohort (Fig. 2A), indicating good discrimination. In Table 5, we provide the sensitivity, specificity, positive predictive values (PPV) and negative predictive values (NPV) on the validation and test sets. We found that the prediction model for ICU admission and death achieved better sensitivity and PPV.
Risk prediction performance indices. risk prediction performance indices. (A) On the validation set and test cohort, the prediction model for ICU admission had an AUROC of 0.829 (95% CI: 0.810–0.850) and 0.858 (95% CI: 0.803–0.899) respectively. The value of test set is higher than validation set. (B) The model performance of risk prediction model for death performed similarly on the validation and test sets. AUROC reached above 0.900 on both validation set (AUROC: 0.923, 95% CI: 0.903–0.943) and test cohort (AUROC: 0.906, 95% CI: 0.850–0.948). The value of validation set is higher than test set. (C) The prediction model for ICU admission showed that AUROC was 0.829 (95% CI: 0.810–0.850) and 0.858 (95% CI: 0.803–0.899) on the validation set and test cohort. The value of test set is higher than validation set. (D) The prediction model for death showed that AUROC reached above 0.900 on both validation set (AUROC: 0.923, 95% CI: 0.903–0.943) and test cohort(AUROC: 0.906, 95% CI: 0.850–0.948). The value of validation set is higher than test set.
The model performance of risk prediction model for death performed similarly on the validation and test sets. The Accuracy was 0.935 for validation set and 0.932 for test cohort, SbrS of 0.328 for validation set and 0.256 for test cohort, respectively. AUROC reached above 0.900 on both validation set (AUROC: 0.923, 95% CI: 0.903–0.943) and test cohort (AUROC: 0.906, 95% CI: 0.850–0.948). The discrimination of risk prediction models for ICU admission and death was further assessed on an external dataset, with AUROC of 0.852 (95% CI: [0.788–0.907]) and 0.882 (95% CI: [0.817–0.931]), respectively.
The calibration plots were demonstrated in Figs. 3 and 4. The X-axis represents the model prediction probability value, and the Y-axis represents the actual observation proportion. By dividing the predicted value into several intervals, we calculated the average prediction probability and the true positive rate for each interval22. In this study, we divided the predicted values into five intervals, which correspond to five levels of risk categories (low-risk, medium–low, medium, medium–high, and high). If the values of Pred and True are close, then the coordinates of (Pred, True) will be close to the diagonal. Therefore, the closer each point is to the diagonal, the better the calibration ability of the model.
Calibration indices of risk prediction model of ICU admission. (A) The calibration of risk prediction model for ICU admission on the validation set. (B) The calibration of ICU admission risk prediction model on the test cohort. Figure 3 indicate that the model tends to overestimate the probability of ICU. In panel A, for instance, when the model predicts more than 0.3 probability of ICU, the observed proportion of ICU is closer to 0.2. This discrepancy suggests that the model is too progressive in its ICU estimates.
Calibration indices of risk prediction model of death. (A) Calibration indices of risk prediction model of death on the validation set. (B) Calibration indices of risk prediction model of death on the test cohort. Both panels in Fig. 4 indicate that the model tends to underestimate the probability of death. In panel A, for instance, when the model predicts a 0.9 probability of death, the observed proportion of death is less than 0.7. This discrepancy suggests that the model is too conservative in its death estimates.
The calibration of risk prediction model for ICU admission on the validation set was showed in Fig. 3A. The data pairs (Pred, True) in the calibration plot for ICU admission were close to the diagonal line, indicating good predictive ability of the prediction model in the different risk groups. The calibration of ICU admission risk prediction model on the test cohort was showed in Fig. 3B. The predicted probabilities and observed risks were close in medium and low-risk subgroups (refers to samples with predicted risks lower than 0.6). However, the predicted probability in the high-risk groups was lower than the observed risks, indicating underestimation of ICU admission risk in high-risk groups.
The calibration of the risk prediction model for death on the validation set and test cohort was presented in Fig. 4. In the high-risk subgroup, the predicted probability was greater than the observed risk, indicating an overestimate of mortality risk. The data pairs in the rest of the subgroups were close to the diagonal line, indicating great calibration. However, we observed poor calibration on the test cohort, with overestimate of death risk in medium and high-risk subgroups and underestimate of death risk in medium–high risk subgroup.
The calibration of the risk prediction models on the external dataset was showed in Fig. 5. We observed good calibration of the risk model for ICU admission (Fig. 5A). However, the risk prediction model for death overestimated risk for high-risk groups (Fig. 5B), indicating poor calibration.
Calibration indices of risk prediction model of ICU admission /death on external dataset. (A) Calibration indices of the risk model for ICU admission. (B) Calibration indices of the risk model for death. Figure 5 indicates that the model overestimated the probability of death and underestimated the probability of ICU in external validation. For example, in panel A, when the model predicts an ICU probability less than 0.8, the observed ICU probability is higher than 0.8. In panel B, when the model predicts a mortality probability close to 0.8, the observed mortality probability is close to 0.6.
Discussion
In this study, we developed and validated risk prediction models for ICU admission and death for COVID-19 inpatients using Elastic Net model. Our study showed that the prediction models had the high discrimination and best calibration of the risk prediction model for ICU admission and death. It could identify patients at high or low risk of ICU admission and mortality. Due to the ongoing impact of the COVID-19 disease on health care systems, no wonder that there are prior studies about prognostic models for COVID-19 patients. In what follows, we outline the main characteristics of the most important studies and compare these works with ours in order to highlight the respective strengths and limitations.
Rahmatinejad et al.7 compared the performance of a variety of regression-based scoring systems, and highlighted the superiority of the APACHE II scoring system in predicting inhospital mortality of patients in the ICU, which was more suitable for critically ill adult patients admitted to the ICU directly from the ED (emergency departments). What sets it apart is the population included in our study, not only the more severe COVID-19 inpatients like in other studies12,23,24 but also all COVID-19 inpatients in our hospital. This broad inclusion criterion makes our study less prone to selection bias and improves the generalizability of our findings, as our study has confirmed that our model had higher discrimination in the external dataset demonstrating the generalizability and robustness of the model.
Rahmatinejad et al.8 compared the prognostic accuracy of six different regression-based scoring systems for predicting in-hospital mortality among Covid-19 patients with AUC values ranging between 70 and 80% and revealed the WPS, REMS, and NEWS have a fair discriminatory performance, which was reported relatively low performance compared with our study. In our study, the model achieved AUROC of 0.852 (95% CI: [0.788–0.907]) and 0.882 (95% CI: [0.817–0.931]) for ICU admission and death respectively, which were classified as a good performance on an external dataset comparable to or better than alternative COVID-19 prediction models3,12,23,25,26,27,28,29,30,31. Our models showed that a reliable prediction can be made for ICU admission and death in COVID-19 inpatients. We note that the authors report that multiple imputations within bootstrap samples were used. Thus, the reported results may be impacted by an over-estimation.
Rahmatinejad et al.13 developed logistic regression and ensemble learning models with the highest AUROC of 83.9% for predicting in-hospital mortality in emergency departments. While the reported results are comparable with our own, they validated the model only on the test dataset and limited themselves to three levels of ESI acuity, making it prone to overfitting and unclear as to what extent these models can be generalized to a broader ED population.
Famiglini et al.27 introduced a robust and parsimonious machine learning method to predict ICU admission in COVID-19 patients but the generalizability of the developed models was not externally validated. The difference is that our prediction models were applied to an external dataset and had higher discrimination in the external dataset, demonstrating the generalizability and robustness of our model.
Regarding ICU admission, Sabetian et al.11 reported that machine learning offered an advantage over logistic regression for predicting patients requiring intensive care. However, this study was based on a small sample of 506 patients.
It was encouraging to note that we incorporated four categories of clinical characteristics, symptoms, comorbidities and treatment information based on all patients’ data in our prediction models when making determinations related to ICU admission and death for COVID-19 inpatients. Our model finally identified 162 variables as being predictive of ICU admission and death, having more variables than other studies23,31. Some variables in our study have been reported as strong contributors to COVID mortality and ICU admission prediction in other studies32,33.Our prediction models were derived using these variables and were exported to an external dataset.
Since clinical judgment is subjective, models can be helpful to physicians, especially those who are not adequately experienced. Providing these models to physicians can help them better estimate patients’ prognoses. Rahmatinejad et al.9 revealed that regression-based mSOFA predicted a better-calibrated mortality risk than emergency residents’ judgment. Models can augment the physician’s clinical judgment. Elastic net regression is interpretable and can provide better prediction accuracy compared to standard statistical analysis3,10,31. As proved in this study, applying elastic net regression indeed achieved a good performance. It is noteworthy that we conducted risk stratification to five levels of risk categories. However, the calibration of the risk prediction model indicated underestimation of ICU admission risk in high-risk groups and death risk in medium–high risk subgroup and an overestimate of death risk in medium and high-risk subgroups. The possible reasons for the above situations were as follows: 1) Model Complexity: we employed elastic net, a regularized regression model in our study, which has a certain degree of complexity. Complex models may overfit the training data, leading to poor generalization and calibration performance on new data. 2) Data Quality: there are inconsistencies and biases in the data between the derivation cohort and the external test cohort, which can adversely affect model calibration.3) Missing Variables: the elastic net models in our study cannot adequately capture all relevant variables that influence the outcomes. Missing variables or unaccounted-for confounding factors can introduce bias and lead to calibration discrepancies.
This study has several strengths. First, we conducted this study with a larger sample size relative to certain studies to develop elastic net regression models for not only mortality but also ICU admission in all COVID-19 patients. Second, we presented external validation results across patients admitted to Fujian Provincial Geriatric Hospital. Although the performance degraded with external validation as opposed to internal validation, we could estimate the expected model performance by this approach when used on new patients from the same or other centers.
There are also several limitations to this work. First, we left out the test cohort as an artificial “external data” to evaluate the generalizability of our prediction models. However, the prediction models are prone to overestimate the performance since the test and derivation cohorts are of the same source. Our perdition model could be used for new patients whose characteristics are similar to those of our study samples. However, when applying it to new data with different features, the prediction results could be inaccurate. Second, this study was conducted during the COVID-19 pandemic, so our findings may vary in non-pandemic conditions. Third, we could only analyze those parameters that had been recorded and documented in our hospital's clinical electronic record system because this was a retrospective study. Various parameters, such as specific leukocyte/lymphocyte subgroups, tumor necrosis factor or interleukins, may be overlooked although they might prove to be better predictors than the parameters included in this study and might carry additional independent information. Fourth, considering that imputing missing values within each bootstrap sample can introduce additional variability into the analysis and increase the risk of model overfitting, these variables were removed from the analysis in our study. However, excluding variables with high missingness can introduce bias and limit generalizability.
In future works, we aim to further investigate our models by refining model variables, improving calibration, or validating the models in diverse populations and combine our methods with models predicting the worsening of health status over time34, thus providing clinicians with more information about patients’ health status and better risk stratification indications. Furthermore, it is necessary to conduct multicentric studies to further investigate the performance of these models in the COVID-19 inpatients.
In conclusion, the prediction models in this study can predict the risk of ICU admission and death for COVID-19 inpatients with good performance on an external dataset. COVID-19 ICU admission and death risk stratification tools can help physicians in patient stratification during the stages of the pandemic, ultimately improving care, informing accurate and rapid triage decisions and facilitating better resource allocation.
Data availability
The datasets analyzed during the current study are available from the corresponding author on reasonable request.
References
Arshad Ali, S. et al. The outbreak of Coronavirus disease 2019 (COVID-19)-an emerging global health threat. J. Infect. Public Health 13(4), 644–646 (2020).
Gupta, S. et al. Factors associated with death in critically ill patients with Coronavirus disease 2019 in the US. JAMA Intern. Med. 180(11), 1436–1447 (2020).
Islam, K. R. et al. Prognostic model of ICU admission risk in patients with COVID-19 infection using machine learning. Diagnostics 12(9), 2144 (2022).
De OliveiraAndrade, R. Covid-19 is causing the collapse of Brazil’s national health service. Bmj 370, m3032 (2020).
Supady, A. et al. Allocating scarce intensive care resources during the COVID-19 pandemic: Practical challenges to theoretical frameworks. Lancet Respir. Med. 9(4), 430–434 (2021).
Goldstein, B. A. et al. Opportunities and challenges in developing risk prediction models with electronic health records data: A systematic review. J. Am. Med. Inform. Assoc. JAMIA 24(1), 198–208 (2017).
Rahmatinejad, Z. et al. Internal validation of the predictive performance of models based on three ED and ICU scoring systems to predict in hospital mortality for intensive care patients referred from the emergency department. BioMed Res. Int. 20, 3964063 (2022).
Rahmatinejad, Z. et al. Comparison of six scoring systems for predicting in-hospital mortality among patients with SARS-COV2 presenting to the emergency department. Indian J. Crit. Care Med. Peer Rev. Off. Publ. Indian Soc. Crit. Care Med. 27(6), 416–425 (2023).
Rahmatinejad, Z. et al. Comparing In-hospital mortality prediction by senior emergency resident’s judgment and prognostic models in the emergency department. BioMed Res. Int. 2023, 6042762 (2023).
Moxley, T. A. et al. Application of elastic net regression for modeling COVID-19 sociodemographic risk factors. PLoS ONE 19(1), e0297065 (2024).
Sabetian, G. et al. Prediction of patients with COVID-19 requiring intensive care: A cross-sectional study based on machine-learning approach from Iran. Indian J. Crit. Care Med. Peer Rev. Off. Publ. Indian Soc. Crit. Care Med. 26(6), 688–695 (2022).
Churpek, M. M. et al. Machine learning prediction of death in critically Ill patients with Coronavirus disease 2019. Crit. Care Explor. 3(8), e0515 (2021).
Rahmatinejad, Z. et al. A comparative study of explainable ensemble learning and logistic regression for predicting in-hospital mortality in the emergency department. Sci. Rep. 14(1), 3406 (2024).
Vaid, A. et al. Machine learning to predict mortality and critical events in a cohort of patients with COVID-19 in New York city: Model development and validation. J. Med. Internet Res. 22(11), e24018 (2020).
Yadaw, A. S. et al. Clinical features of COVID-19 mortality: Development and validation of a clinical prediction model. Lancet Digital Health 2(10), e516–e525 (2020).
Yan, L. et al. An interpretable mortality prediction model for COVID-19 patients. Nat. Mach. Intell. 2(5), 283–288 (2020).
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net. J. Royal Stat. Soc. Series B Stat. Methodol. 67(2), 301–320 (2005).
Valente, G. et al. Cross-validation and permutations in MVPA: Validity of permutation strategies and power of cross-validation schemes. NeuroImage 238, 118145 (2021).
Fawcett, T. An introduction to ROC analysis. Pattern Recognit. Lett. 27(8), 861–874 (2006).
Huang, Y. et al. A tutorial on calibration measurements and calibration models for clinical prediction models. J. Am. Med. Inform. Assoc. JAMIA 27(4), 621–633 (2020).
Fenlon, C. et al. A discussion of calibration techniques for evaluating binary and categorical predictive models. Prev. Vet. Med. 149, 107–114 (2018).
Bateman, R. M. et al. 36th international symposium on intensive care and emergency medicine: Brussels, Belgium. 15-18 March 2016. Crit. Care 20(Suppl 2), 94 (2016).
Lichtner, G. et al. Predicting lethal courses in critically ill COVID-19 patients using a machine learning model trained on patients with non-COVID-19 viral pneumonia. Sci. Rep. 11(1), 13205 (2021).
Sakagianni, A. et al. Automated ML techniques for predicting COVID-19 mortality in the ICU. Stud. Health Technol. Inform. 305, 517–520 (2023).
Castro, V. M., McCoy, T. H. & Perlis, R. H. Laboratory findings associated with severe illness and mortality among hospitalized individuals with Coronavirus disease 2019 in Eastern Massachusetts. JAMA Network Open 3(10), e2023934 (2020).
di Napoli, A. et al. 3D CT-inclusive deep-learning model to predict mortality, ICU admittance, and intubation in COVID-19 patients. J. Digit. Imag. 36(2), 603–616 (2023).
Famiglini, L. et al. A robust and parsimonious machine learning method to predict ICU admission of COVID-19 patients. Med. Biol. Eng. Comput. 1–13 (2022).
Ottenhoff, M. C. et al. Predicting mortality of individual patients with COVID-19: A multicentre dutch cohort. BMJ open 11(7), e047347 (2021).
Rodriguez-Nava, G. et al. Performance of the quick COVID-19 severity index and the Brescia-COVID respiratory severity scale in hospitalized patients with COVID-19 in a community hospital setting. Int. J. Infect. Dis. 102, 571–576 (2021).
van Dam, P. et al. Performance of prediction models for short-term outcome in COVID-19 patients in the emergency department: A retrospective study. Ann. Med. 53(1), 402–409 (2021).
Xu, Y. et al. Machine learning-based derivation and external validation of a tool to predict death and development of organ failure in hospitalized patients with COVID-19. Sci. Rep. 12(1), 16913 (2022).
Kukoč, A. et al. Clinical and laboratory predictors at ICU admission affecting course of illness and mortality rates in a tertiary COVID-19 center. Heart Lung J. Crit. Care 53, 1–10 (2022).
Meijs, D. A. M. et al. Predicting COVID-19 prognosis in the ICU remained challenging: External validation in a multinational regional cohort. J. Clin. Epidemiol. 152, 257–268 (2022).
Montomoli, J. et al. Machine learning using the extreme gradient boosting (XGBoost) algorithm predicts 5-day delta of SOFA score at ICU admission in COVID-19 patients. J. Intensiv. Med. 1(2), 110–116 (2021).
Acknowledgements
The authors thank all study participants for support and cooperation during the study.
Funding
This work was supported by the National Key Clinical Specialty (Grant No. 2023002).
Author information
Authors and Affiliations
Contributions
B.X. contributed substance to ideas and design of the study. W.Z. drafted the paper and contributed to critical revision of the manuscript for important intellectual content. X.Y., Y.C., X.L., J.H., Y.Z., L.Y. contributed to acquisition of data and analysis, interpretation of data. X.Y. gave a lot of assistance and revises manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zou, W., Yao, X., Chen, Y. et al. An elastic net regression model for predicting the risk of ICU admission and death for hospitalized patients with COVID-19. Sci Rep 14, 14404 (2024). https://doi.org/10.1038/s41598-024-64776-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-64776-0
- Springer Nature Limited