Abstract
Diabetes mellitus (DM) is a significant risk factor for peripheral arterial disease (PAD), and PAD is an independent predictor of cardiovascular disorders (CVDs). Growing evidence suggests that long non-coding RNAs (lncRNAs) significantly contribute to disease development and underlying complications, particularly affecting smooth muscle cells (SMCs). So far, no study has focused on transcriptome analysis of lncRNAs in PAD patients with and without DM. Tissue samples were obtained from our Vascular Biobank. Due to the sample’s heterogeneity, expression analysis of lncRNAs in whole tissue detected only ACTA2-AS1 with a 4.9-fold increase in PAD patients with DM. In contrast, transcriptomics of SMCs revealed 28 lncRNAs significantly differentially expressed between PAD with and without DM (FDR < 0.1). Sixteen lncRNAs were of unknown function, six were described in cancer, one connected with macrophages polarisation, and four were associated with CVDs, mainly with SMC function and phenotypic switch (NEAT1, MIR100HG, HIF1A-AS3, and MRI29B2CHG). The enrichment analysis detected additional lncRNAs H19, CARMN, FTX, and MEG3 linked with DM. Our study revealed several lncRNAs in diabetic PAD patients associated with the physiological function of SMCs. These lncRNAs might serve as potential therapeutic targets to improve the function of SMCs within the diseased tissue and, thus, the clinical outcome.
Similar content being viewed by others
Introduction
Peripheral arterial disease (PAD), defined as the narrowing of the peripheral arteries, is primarily caused by atherosclerotic changes within the vessel wall, particularly in the lower extremities. Diabetes mellitus (DM) is a significant risk factor for PAD, and PAD is an independent predictor of cardiovascular and cerebrovascular ischemic events, affecting both the quality and expectancy of life1,2,3. In diabetic patients, atherosclerotic lesions occur earlier with rapid progression and are frequently asymptomatic, thus bearing a high risk of unexpected cardiac and cerebral complications2,3. Multiple metabolic aberrations, such as overproduction of advanced glycation end-products (AGEs), increased oxidative stress, reactive oxygen species (ROS), enhanced inflammation, and dyslipidaemia, have been shown to aggravate PAD in patients with DM3,4,5. The metabolic changes strongly affect the biological function of endothelial cells (ECs) and upregulate the expression of many inflammatory factors6,7,8. These cytokines, in turn, promote atherosclerosis as well as apoptosis of ECs. Increased inflammation, as well as oxidative stress, facilitate the development of atherosclerotic lesions within the arterial wall. In PAD patients, atherosclerosis increases the risk of thrombosis and lower extremity ulceration1,2. Furthermore, individuals with PAD suffering from DM are more susceptible to plaque rupture9,10. In addition, hyperglycaemia in diabetic patients induces the production of ROS and AGEs, leading to vascular damage and diminished bioavailability of nitric oxide (NO)3,6,11,12. NO and prostacyclin PGI2 are the essential vasoactive factors affecting the underlying smooth muscle cells (SMCs)3,13,14,15,16. The impaired ECs, together with other metabolic aberrations due to DM, impair the physiological function of SMCs by changing their phenotype in a proatherogenic manner, leading increasingly also to apoptosis17,18.
Growing evidence suggests that particularly long non-coding RNAs (lncRNAs), participating in many biological processes such as transcriptional regulation of oxidative stress, inflammation, atherosclerosis, as well as insulin sensitivity, may also affect PAD development, especially in patients suffering from DM19,20. Furthermore, many lncRNAs have already been described to significantly affect the phenotype and the physiological behaviour of vascular cells in DM, as well as the crosstalk between ECs and SMCs21,22. Thus, dysregulation of lncRNAs in DM patients suffering from PAD may significantly affect the biological function mainly observed in SMCs and thus aggravate the clinical outcome for these patients. Combining clinical data with omics analyses and bioinformatics is a useful strategy for improving diagnostic precision and pursuing personalised medicine in the treatment of CVDs. Particularly, bioinformatics analysis of big data and the use of artificial intelligence may help to discover novel useful targets or biomarkers with the potential for more accurate diagnosis and therapy23,24,25.
Interestingly, no study has so far focused on the analysis of lncRNAs in diabetic patients with PAD using RNA sequencing of the corresponding tissue. Performing an extended PubMed research, we found no corresponding relevant work dealing with PAD or DM patients and lncRNAs, particularly in humans. Therefore, having the advantage of possessing such tissue samples in our vascular biobank, we performed a detailed transcriptome analysis of PAD patients with and without DM, focusing particularly on lncRNAs and SMCs.
Results
Patient characteristics
The clinical characteristics of the selected study patients included in the transcriptome analysis of lncRNA, associated diseases, and corresponding medication are summarised in Table 1. No significant differences were observed between the study groups of PAD patients with and without DM with regard to age, gender or any other collected clinical data. The mean age of PAD patients with and without DM was 74.9 ± 9.2 and 71.9 ± 11.2 years, 52.9% and 40.0% were of the male sex. Most of the patients suffered from chronic kidney disease (82.4% and 80.0%). Furthermore, 58.8% and 65.0 of the study participants received aspirin or clopidogrel, 64.8% and 40.0% beta-blockers and/or ACE inhibitors, and 64.7 and 75.0% were on statins. The only divergence was observed for diuretic intake, with 52.5% of individuals suffering from DM compared to 15.0% of patients without DM (P = 0.038).
Pathomorphological analysis
At first, we performed histological and immunohistochemical (IHC) staining of the FFPE samples from the diseased iliac artery (n = 79) to obtain an insight into the pathomorphology of the tissue of our study patients. The specimen characterisation revealed highly heterogeneous pathophysiological features (Fig. 1).
Selected histological examples of the tissue samples from PAD patients with and without DM. (a) and (b) Haemalaun-Eosin staining of PAD tissue with and without DM. Most samples were highly atherosclerotic and heavily calcified (arrows). (c) and (d) Smooth muscle actin staining. (e) and (f): Example of MYH10 (synthetic phenotype of SMCs) and MYH11 (contractile phenotype of SMCs) staining. High overlapping was observed. (g) Example of ECs staining using CD31. (h) Example of leukocyte staining using CD45. (i) Example of macrophage staining using CD68. Scale bars: 1 mm (a–d), 100 µm (e–i).
Most samples had extended atherosclerotic lesions, inflammation, calcification, and were highly vascularised. Interestingly, staining with MYH10 and MYH11 to distinguish between the synthetic and contractile phenotype of SMCs showed high overlapping characteristics and no clear separation between these two morphologic features was observed (Fig. 1e and f). In order to reduce the broad heterogeneity of our study tissue, samples with a great extent of calcification, inflammation, and a low number of cells, which markedly differed from the average, were excluded. Consequently, from the 79 histologically characterised patients, 37 were finally included in the transcriptomics analysis (DMplus: n = 17, DMminus: n = 20). The semi-quantitative pathomorphological analysis of the included study samples revealed no significant differences between the study groups (Table 2).
Differential expression analysis of lncRNA in the whole PAD tissue samples
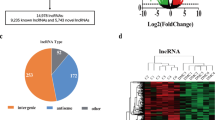
First, we analysed the RNA from the whole tissue samples, focusing on lncRNAs. The volcano plot, comparing PAD patients with and without DM, is shown in Fig. 2. In total, 13,491 lncRNAs were detected, with 6672 above the threshold of 10 counts per million. Nevertheless, only one lncRNA, ACTA2-AS1, demonstrated significantly differential expression between the study groups (4.9-fold increase in DMplus samples, FDR = 0.031, P = 2.7E−06) (Fig. 2a, Table 3).
Volcano plot comparing differentially expressed lncRNAs in the whole tissue (a) and in SMCs (b) from PAD patients with and without DM (DMplus–over–DMminus). The plot shows − log10 transformed FDR as a function of the difference between study groups. The broken lines show 0.5 log2 fold change. Statistically significant differences: FDR < 0.1.
The heatmap of the most differentially expressed lncRNAs in the tissue of PAD patients with DM (DMplus) and without DM (DMminus) revealed broad heterogeneity of the individual samples regarding the expression of lncRNAs (Fig. 3). Even if many samples were clustered, each study group was distributed throughout the whole heatmap.
Heatmap of the most differentially expressed lncRNAs in the whole tissue of PAD patients with (DMplus) and without (DMminus) diabetes mellitus (rows indicate the expression of 241 most expressed lncRNAs, columns indicate the individual samples). Clustering was performed using conditional formatting features. Even if many samples are clustered, high heterogeneity was observed regarding the study groups.
Differential expression analysis of lncRNA in SMCs of the PAD tissue samples
Regardless of the preselection of the study samples according to their histology, wide heterogeneity of the lncRNA expression in the whole tissue was still observed. Therefore, we changed our experimental approach and focused only on the SMCs within the study samples. Consequently, we performed a dissection of the tissue and analysed the lncRNAs only from the microdissected SMC areas. The corresponding volcano plot is shown in Fig. 2b, and the heatmap of the clustering of the differentially expressed lncRNAs is shown in Fig. 4. The expression of lncRNA only from SMCs showed markedly higher homogeneity and better distribution (clustering) of the study samples (Fig. 4) compared to the heatmap of the whole tissue (Fig. 3).
Heatmap of the most differentially expressed lncRNAs in the SMCs from PAD patients with (DMplus) and without (DMminus) diabetes mellitus (rows indicate the expression of 28 most expressed lncRNAs, columns indicate the individual samples). Clustering was performed using conditional formatting features.
In total, 14,499 lncRNAs could be detected, with 761 lncRNAs with counts above the threshold of 10 counts per million. From these 761 lncRNAs, 28 demonstrated significantly differential expression between the study groups (FDR < 0.1) (Table 3). Of these 28 lncRNAs, 14 were novel transcripts not yet described. The function of two other lncRNAs, MIR23AHG and RUFY1-AS1, is also unknown (Table 3). Six of the remaining lncRNAs have been described in different types of cancer (LINC01220, CYTOR, LINC00910, LINC00511, PURPL, and MZF1-AS1), one with macrophage polarisation (MIR222HG), and five of these lncRNAs could be associated with cardiovascular disorders (CVDs), mainly with SMC proliferation, migration, apoptosis, and phenotypic switch (NEAT1, XIST, MIR100HG, HIF1A-AS3, and MIR29B2CHG) (Table 3). The latter lncRNAs were further investigated and RT-PCR was performed to confirm the results from the transcriptome analysis (Fig. 5).
Box plots using RT-PCR to compare the expression of relevant lncRNAs in SMCs between the study groups (with and without DM). The lower part shows subsequent analysis of XIST and MIG222HG separated for sex because these two lncRNAs are expressed only on chromosome X. The expression was normalised for the housekeeping gene GAPDH. Furthermore, for better comparison, the expression of the individual lncRNAs of the study group without DM (w/o DM) was set as references equal 1.
RT-PCR analysis of NEAT1, MIR100HG, HIF1A-AS2, and MIR29B2CHG confirmed the results from the RNA sequencing with a significant reduction of their expression in SMCs of PAD patients suffering from DM (Fig. 5). XIST and MR222HG were expressed only on chromosome X. Therefore, additional subsequent adjustment for sex was performed. Interestingly, the lncRNA XIST was expressed only in females and after the correction for sex, no significant differences were observed (Fig. 5). The same results were found for MIR222HG, even if, in this case, the expression was found in males and females (Fig. 5).
In addition, due to the rather low amount of samples in each study group, we performed a post-hoc power analysis. For NEAT1 (98.1%), HIF1A-AS3 (82.3%), and MIR29B2CHG (83.7%), the statistical power was > 80%. In contrast, MIR100HG showed values < 80% (61.2%). Furthermore, to elucidate a potential clinical significance, receiver operating characteristic (ROC) curve analysis for the significantly differentially expressed lncRNA was performed (Supplementary Fig. 1). For the lncRNAs NEAT1, HIF1AAS3, and MIR29B2CHF, the AUC was significant (P = 0.001, 0.007, and 0.011). For MIR100HG, the AUC was not statistically significant (P = 0.290).
Gene enrichment analysis
In order to identify groups of lncRNA genes that are over-represented in our experimental setup, we performed, in addition, a gene and pathway enrichment analysis with the 761 lncRNAs having counts above the threshold (Fig. 6, Table 4). The gene enrichment pathway analysis (GEPA) revealed the involvement of insulin-like growth factor 2 mRNA binding proteins and diabetes pathways associated with the lncRNA H19. However, only the first pathway showed statistical significance (P-adjusted < 0.1) (Table 4). The gene set enrichment analysis (GSEA) showed significant enrichment for genes involved in miRNA-mediated post-transcriptional gene silencing (GO:0035195), including the RISC complex (GO:0016442) (Fig. 6, Table 4).
Gene set enrichment analysis (GSEA). (a) Biological process (BP): GO:0035195—miRNA-mediated post-transcriptional gene silencing pathway. (b) Cellular component (CC): GO:0016442—RISC complex pathway. The corresponding enrichment data are summarised in Table 2.
The silencing complex included, besides the already significantly different lncRNAs (DMplus vs DMminus) such as NEAT1 and MIR29B2CHG, also other relevant lncRNAs involved in SMCs physiology or CVDs, such as CARMN, FTX, and MEG3.
The enrichment score was calculated for each annotation category by screening all the genes from a differential expression analysis and their associated fold changes. GO—gene enrichment; Sub-ontologies of GO hierarchy: BP—biological process, CC—cellular components; NES—normalised enrichment score. The corresponding graphical illustration is shown in Fig. 6.
Discussion
The current study identified a couple of lncRNAs significantly downregulated in SMCs of PAD individuals suffering from DM, which might be of interest for further investigation to elucidate their role in diabetes. Sixteen of them (e.g. ENSG00000289474, ENSG00000288156 or ENSG00000289901) are so far of unknown function. Others, such as LINC01220, CYTOR, LINC00910, LINC00511, PURPL, and MZF1-AS1, have been described so far only in cancer. Four differentially expressed lncRNAs have already been associated with CVDs, mainly with SMC function and phenotypic switch (NEAT1, MIR100HG, HIF1A-AS3, and MRI29B2CHG). These lncRNAs might serve as potential diagnostic markers or therapeutic targets to improve the function of SMCs within the diseased tissue.
Comparing the expression pattern of lncRNAs in the whole tissue of PAD patients with and without DM, only ACTA2-AS1 showed significantly increased expression in the DMplus study group. The inability to detect other differentially expressed lncRNAs in the tissue is mainly due to the broad heterogeneity in the underlying pathomorphology, even after our careful preselection. All samples were highly atherosclerotic, and the additional burden of DM was, therefore, not easy to distinguish. The significantly differentially expressed lncRNA ACTA2-AS1 has been described so far, mainly in the context of cancer development26,27,28. However, Arencibia et al. found that the ACTA2-AS1 might also regulate the phenotypic switch and proliferation of SMCs29. In addition, the downregulation of ACTA2-AS1 shows significant enrichment in collagen biosynthesis and alternation in the structure and composition of the extracellular matrix (ECM). Consequently, ACTA2-AS1 might also play an essential role in PAD patients with DM by affecting the function of SMCs.
Due to the above-mentioned lack of differentially expressed lncRNAs in the whole tissue samples caused by the extended atherosclerotic pathology and wide heterogeneity, we focused on the SMCs within the concerned specimens. In this experimental approach, 28 significantly differential expressed lncRNAs were detected between the study groups. Fourteen of these lncRNAs were novel transcripts. Some of them, for instance, ENSG00000289474, with a 12-fold reduction in its expression in PAD patients with DM, might be of interest to be further investigated. Many of the remaining lncRNAs, such as LINC01220, CYTOR, LINC00910, LINC00511, PURPL, and MZF1-AS1, have been described in association with different types of cancer30,31,32,33,34,35. Interestingly, the expression changes were associated mainly with cell proliferation and migration of the tumour cells. So far, there is no study examining these lncRNAs in CVDs. Consequently, whether these lncRNAs also affect SMCs has not yet been investigated. MIR222HG was also described in association with cancer36,37 but also with macrophage polarisation38. In this context, it is to be noted that SMCs can adopt various phenotypes, including also that of macrophages or foam cells39. Thus, lncRNA MIR222HG might also be involved in the phenotypic switch of SMCs, for example, resembling the macrophage phenotype and taking up lipids and cholesterol derivatives. Various lncRNAs have been described to regulate SMCs plasticity40.
Four significantly differentially expressed lncRNAs between our study groups with and without DM, namely NEAT1, MIR100HG, HIF1A-AS3, and MIR29B2CHG, have already been associated with CVDs32,41,42,43,44,45,46,47,48, mainly with the proliferation, migration, apoptosis, and phenotypic switch of SMCs (Table 3)41,46,47,49,50,51.
LncRNA NEAT1 (nuclear-enriched abundant transcript 1) is involved not only in atherosclerosis44,46 or aneurysm52 but has shortly been mentioned as being also dysregulated in DM, especially in diabetic neuropathy and diabetic kidney disease53,54,55,56. Furthermore, NEAT1 has already been described as having a critical in the phenotypic switching of SMCs by repressing SM-contractile gene expression through an epigenetic regulatory mechanism16. In addition, overexpression of NEAT1 participates in proliferation and can inhibit apoptosis of SMCs41, improving their ability to synthesise ECM components in order to stabilise the arterial wall. Thus, the reduction of NEAT1 expression in PAD patients with DM might further impair the function of SMCs and aggravate the clinical outcome.
Regarding lncRNA HIF1A-AS (hypoxia-inducible factor-1 alpha antisense RNA), only HIF1A-AS1 and -AS2 have been described to modulate the proliferation, migration, and phenotypic switch in SMCs, particularly in aortic aneurysm47,50,59,60. So far, there is no information about the role of HIF1A-AS3 in CVDs. However, because the other isoforms affect the behaviour of SMCs, it can be assumed that HIF1A-AS3, significantly downregulated in the DMplus study group, might also affect the function of SMCs. The lncRNAs MIR100HG and MIR29B2CHG have only been mentioned in the context of cardiomyopathy and heart failure42,48. So far, there is no information about their potential role in PAD, DM or SMCs. Whether or how far these two lncRNAs might be involved in the corresponding pathophysiology is so far unknown and has to be further investigated.
Interestingly, considering the gene enrichment pathway analysis, additional lncRNAs occurred to play an essential role in the pathology of DM in PAD patients. Of particular interest were H19, CARMN, and MEG3. Our group and others have already described the critical role of lncRNA H19 in atherosclerosis and aneurysm61,62,63. Furthermore, H19 has long been proven to be connected with the phenotypic switch of SMCs64 and diabetes, especially in the regulation of glucose metabolism65,66. In our experimental setup, H19 was involved in the binding of RNA by insulin-like growth factor 2 mRNA binding proteins (IGF2BPs). IGF2BPs bind specific sets of RNAs, regulating their translation, stability, and subcellular localisation67. IGF2BPs, belonging to the IGF mRNA binding proteins (IMPs), are highly conserved and believed to play an essential role in cell migration, metabolism, stem cell renewal, and development68. In adults, their expression is either absent or at very low levels in most tissues. However, IMPs, particularly IGF2BPs, appear to resume their physiological functions in tumour cells and exhibit multiple attachments to H19 lncRNA69. Altogether, mRNA binding protein family IMPs/IGF2BPs has been described in a plethora of biological processes, including development, tumorigenesis, and differentiation68. Furthermore, in diabetic patients, IMP genes have been associated with impaired insulin secretion68. Whether and how far vascular cells, particularly SMCs, can be affected has not yet been investigated.
Regarding the gene set enrichment analysis, H19 seemed to be involved again, together with NEAT1, CARMN, MIR23AHG, MIR29B2CHG, MIR663AHG, MIRLET7A1HG, LINC-PINT, FTX, and MEG3. Of these lncRNAs, irrespective of the already described NEAT1, MIR29B2CHG, and H19, the lncRNA CARMN and MEG3 have already been described in the context of CVDs and SMCs40,66,70,71,72,73. The lncRNA CARMN has been associated with atherosclerosis40,72 and SMC plasticity70. Dong et al. observed that CARMN overexpression maintains SMCs in the contractile phenotype70. On the contrary, loss of CARMN, as observed in our study, affects the phenotypic plasticity of SMCs and significantly accelerates atherosclerosis40,71. The lncRNA MEG3 has already been considered an essential lncRNA in CVDs and modulation of SMC phenotype64,73. Recently, MEG3, among other lncRNAs such as MIAT, H19, MALAT1, AVRIL, and HOTAIR, has been connected with several diabetic complications, being either upregulated or downregulated depending on the disease context or regulatory partners43,66,74. These lncRNAs are involved particularly in oxidative stress, inflammation, apoptosis, and angiogenesis pathways, mediating either protective function or contributing to the severity of DM66.
Limitations
The limitation of this study is the relatively small number of tissue samples and the broad heterogeneity between the individual specimens. Furthermore, it should be noted that the atherosclerotic patterns in PAD patients with and without DM are usually different, and thus, the results may have also been affected by the sampling site. In order to increase the homogeneity within the study groups, extended histological characterisation of the samples was performed, and tissues with great extent of calcification, inflammation and low amount of cells, which markedly differed from the average, were excluded. In this manner, from the 79 characterised specimens, only 37 were involved in the study. The semi-quantitative analysis of the included samples showed no significant differences between the specimens afterwards (see Table 2). Nevertheless, a more extensive or consecutive study with other larger tissue sample sets is necessary to confirm the results of our present work.
Conclusion
In our current study, we have analysed for the first time lncRNAs in PAD patients suffering from DM, using extended transcriptomics analysis of the corresponding tissue and the underlying SMCs. Particularly in SMCs, 28 significantly differential expressed lncRNAs were detected. Sixteen of them were novel transcripts or so far with an unknown function. Some of them, such as for instance ENSG00000289474 with a 12-fold expression reduction in PAD patients with DM, might be of interest to further investigation. Many of the other lncRNAs, such as LINC01220, CYTOR, LINC00910, LINC00511, PURPL, and MZF1-AS1, have been described in association with different types of cancer, mainly with proliferation and migration of the tumour cells. These lncRNAs have not yet been analysed in the context of CVDs and might affect the pathophysiological behaviour of vascular cells within the diseased tissue. Five of the differentially expressed lncRNAs, namely NEAT1, HIF1A-AS3, MIR100HG, and MIR29B2CHG, have already been proven to play a role in CVDs, especially in proliferation, migration, apoptosis, and phenotypic switch of SMCs. Interestingly, all these lncRNAs were downregulated in PAD patients with DM, having a negative effect on SMCs, thus being able to aggravate the clinical outcome. Furthermore, enrichment analysis revealed additional lncRNAs, such as H19, CARMN, and MEG3, with a possible role in DM and the fate of SMCs. In summary, our study detected a set of relevant lncRNAs affecting the SMCs in the PAD patients suffering from DM, which might serve as potential therapeutic targets to improve the function of SMCs within the diseased tissue and, thus, the clinical outcome for these patients.
Material and methods
All methods were performed in accordance with the corresponding relevant guidelines and regulations.
Tissue samples
Vascular tissue samples of the iliac artery were obtained from patients suffering from peripheral arterial disease (PAD) with and without diabetes mellitus (DM) (n = 79) who underwent open surgical intervention in our Department of Vascular Surgery (USZ/UZH), collected in our Swiss Vascular Biobank (SVB). All patients gave appropriate written informed consent. The local ethics committee (Cantonal Ethics Committee Zurich, Switzerland; BASEC-No 2020-00378 and 2020-01844) approved the tissue sample collection and analysis procedure. All tissue samples were divided for consecutive histological and molecular-biological analyses. For histological studies, the tissue was fixed in 4% formalin and embedded in paraffin (FFPE). Adjacent pieces of the corresponding specimens were immediately frozen and stored at − 80 °C.
Histological evaluation
For a pathomorphological evaluation of the PAD tissue (n = 79), sections of FFPE samples were stained with Haematoxylin–eosin (HE) and Elastica van Gieson (EvG), focusing on cellular composition, inflammatory infiltration, calcification, and distribution of the overall content of collagen and elastin. For immunohistochemistry (IHC), FFPE sections were mounted on pre-coated (0.1% poly-L-lysine; Merck/Sigma Aldrich, Buchs, Switzerland) SuperFrost Plus slides (ThermoFisher Scientific), and antigen retrieval was performed by heat in citrate buffer (pH 6.0). Primary antibodies used in this study were purchased from Lucerna-Chem (Luzern, Switzerland) and Agilent/Dako (Basel, Switzerland): alpha-SM-actin for SMCs (abcam ab5694, 1:100), CD31 for ECs (ab134168, 1:40), CD45 for leukocytes (M0701, 1:2,000), CD68 for macrophages (M0814, 1:2,000), MYH10 for synthetic SMCs (ab230823, 1:200), and MYH11 (ab224804, 1:400) were diluted in DAKO REAL Antibody Diluent (Agilent/Dako). To detect the primary antibody, mouse/rabbit specific HRP/DAB (ABC) detection IHC kit (ab64264, abcam) and Mayer’s haematoxylin (Carl Roth, Switzerland) for the nuclear counterstain was used. All slides were digitalised by Zeiss Axio Scan.Z1 and Zeiss Zen Lite 2.1 software.
RNA extraction and quality determination
For the transcriptome analysis of the whole samples, RNA was isolated from fresh-frozen tissue using the TRIZOL reagent (ThermoFisher), which enables the extraction of total RNA, including non-coding RNAs. The tissue samples were homogenised in a volume of 1 mL TRIZOL, mixed with 200 µL chloroform, and centrifuged. The upper (clear) phase containing the RNA was carefully transferred into a new tube, precipitated with isopropanol, washed with 75% ethanol, the RNA pellet air-dried, suspended in 50 µL RNase-free water and stored at − 80 °C until further use.
For the transcriptome analysis of SMCs, FFPE tissue was used in order to be able to perform reliable manual microdissection and to remove only areas containing the cells of interest. Briefly, all samples were stained consecutively with alpha-SM-actin to detect SMCs. Next, a consecutive slide of 25 µm thickness without staining was prepared, and corresponding areas positive for SMCs were excised under the light microscope (Motic AE200) with sterile scalpels. Six to eight slices from the same samples were pooled, and RNA was isolated using the High Pure RNA Paraffin Kit (Roche, Switzerland) according to the manufacturer’s protocol.
The RNA concentration was determined by NanoDrop Lite Plus Spectrophotometer (Witec, Sursee, Switzerland). To evaluate the RNA quality and the degree of degradation in order to select suitable samples for RNA sequencing (atherosclerotic tissue samples have markedly lower RNA quality than healthy tissue75), the RNA integrity numbers (RINs) and DV200 index (percentage of fragments of > 200 nucleotides) were determined (TapeStation 4150; Agilent)) using standard Agilent RNA ScreenTape Assay or Agilent High Sensitivity RNA ScreenTape Assay depending on the RNA concentration in accordance with the manufacturer’s protocols. All measurements were performed in duplicates, and the experiments were repeated in cases of high variability or discrepancies.
Library preparation, RNA sequencing, and RT-PCR analysis
The library preparation and RNA sequencing were performed in collaboration with the Functional Genomic Center Zurich (FGCZ/ETH, Switzerland). In order to be able to analyse non-coding RNAs, the ribosomal RNA depletion method was applied. Briefly, the RNA library was prepared using the SMARTer Stranded Total RNA-Seq Kit from Clontech/Takara Bio (USA) according to the manufacturer’s protocol. The fragmented samples were reversed transcribed into cDNA, end-repaired, polyadenylated, and TruSeq adapters containing the index for multiplexing were ligated at the ends of each fragment. Fragments containing the TruSeq adapters were selectively enriched by PCR. The quality and quantity of the RNA library were validated again by the determination of RIN. The RNA sequencing was performed on the Illumina NovaSeq 6000 (Illumina, Berlin, Germany) with 200 M reads in 100 cycles.
In order to validate the results for the significantly differentially expressed lncRNAs associated with CVD (NEAT1, MIR100HG, MIR29B2CHG, and HIF1A-AS3), additional quantitative real-time PCR (RT-PCR) was performed using TaqMan approach and master kit (ThermoFisher Scientific) on QuantStudio 5.0 (Agilent). The cDNA was synthesised with the RevertAid First Strand cDNA Synthesis Kit (ThermoFisher Scientific). The expression of all lncRNAs was normalised for the housekeeping gene glycerinaldehyde-3-phosphate dehydrogenase (GAPDH).
Data analysis
The data analysis was performed with the SUSHI framework76,77, developed by the FGCZ ETH/UZH, and using the open Galaxy platform. Spliced Transcripts Alignment to a Reference (STAR) software was used to align the human RNA-Seq dataset78. The quality control of the individual reads was proven by FastQC. The read alignment was performed by STAR, and the estimation of the reads abundance by FeatureCounts from the R package subreads79. A minimum of 10 reads in at least one group of replicates was required to consider a gene as detected. Differential expression analysis was assessed using the linear model approach from the Bioconductor package DESeq2 and EdgeR280. The correction for multiple testing was obtained with the Benjamini–Hochberg algorithm calculating the False Discovery Rate (FDR, adjusted P-value)81. The thresholds of FDR < 0.1 was considered significant. All other statistical analyses were performed using IBM’s SPSS software version 29 (SPSS Inc., Chicago, IL, USA). An independent t-test and a Levene test of equality of variances were used. The statistical analyses were two-sided, with P < 0.05 as the significance level.
Data availability
All datasets generated and analysed during the study are stored at the servers of Functional Genomic Server Zurich (FGCZ) (Eidgenoessiche Technische Hochschule Zuerich/Switzerland) (project ID 28383). Datasets used in the current publication are stored at http://www.ncbi.nlm.nih.gov/bioproject/1072644.
References
Mascarenhas, J. V., Albayati, M. A., Shearman, C. P. & Jude, E. B. Peripheral arterial disease. Endocrinol. Metab. Clin. North Am. 43, 149–166. https://doi.org/10.1016/j.ecl.2013.09.003 (2014).
Nativel, M. et al. Lower extremity arterial disease in patients with diabetes: a contemporary narrative review. Cardiovasc. Diabetol. 17, 138. https://doi.org/10.1186/s12933-018-0781-1 (2018).
Yang, S. L. et al. Pathophysiology of peripheral arterial disease in diabetes mellitus. J. Diabetes 9, 133–140. https://doi.org/10.1111/1753-0407.12474 (2017).
Pottecher, J. et al. Diabetes worsens skeletal muscle mitochondrial function, oxidative stress, and apoptosis after lower-limb ischemia-reperfusion: Implication of the RISK and SAFE pathways?. Front. Physiol. 9, 579. https://doi.org/10.3389/fphys.2018.00579 (2018).
Yamagishi, S. I. & Matsui, T. Role of hyperglycemia-induced advanced glycation end product (AGE) accumulation in atherosclerosis. Ann. Vasc. Dis. 11, 253–258. https://doi.org/10.3400/avd.ra.18-00070 (2018).
de la Cruz-Ares, S. et al. endothelial dysfunction and advanced glycation end products in patients with newly diagnosed versus established diabetes: From the CORDIOPREV study. Nutrients https://doi.org/10.3390/nu12010238 (2020).
Demarchi, A., Somaschini, A., Cornara, S. & Androulakis, E. Peripheral Artery disease in diabetes mellitus: Focus on novel treatment options. Curr. Pharm. Des. 26, 5953–5968. https://doi.org/10.2174/1389201021666201126143217 (2020).
Katsiki, N., Kotsa, K., Stoian, A. P. & Mikhailidis, D. P. Hypoglycaemia and cardiovascular disease risk in patients with diabetes. Curr. Pharm. Des. 26, 5637–5649. https://doi.org/10.2174/1381612826666200909142658 (2020).
Batinic, K. et al. YKL-40 is elevated in patients with peripheral arterial disease and diabetes or pre-diabetes. Atherosclerosis 222, 557–563. https://doi.org/10.1016/j.atherosclerosis.2012.03.034 (2012).
Chang, Z., Yan, H., Zhen, Y., Zheng, J. & Liu, Z. Lower limb arterial calcification and acute thrombosis risk in patients with peripheral artery disease. Ann. Vasc. Surg. 63, 227–233. https://doi.org/10.1016/j.avsg.2019.06.043 (2020).
Chait, A. & Bornfeldt, K. E. Diabetes and atherosclerosis: Is there a role for hyperglycemia?. J. Lipid Res. 50(Suppl), S335-339. https://doi.org/10.1194/jlr.R800059-JLR200 (2009).
Fiorello, M. L., Treweeke, A. T., Macfarlane, D. P. & Megson, I. L. The impact of glucose exposure on bioenergetics and function in a cultured endothelial cell model and the implications for cardiovascular health in diabetes. Sci. Rep. 10, 19547. https://doi.org/10.1038/s41598-020-76505-4 (2020).
Casella, S., Bielli, A., Mauriello, A. & Orlandi, A. Molecular pathways regulating macrovascular pathology and vascular smooth muscle cells phenotype in type 2 diabetes. Int. J. Mol. Sci. 16, 24353–24368. https://doi.org/10.3390/ijms161024353 (2015).
Cohen, R. A. Role of nitric oxide in diabetic complications. Am. J. Ther. 12, 499–502. https://doi.org/10.1097/01.mjt.0000178776.77267.19 (2005).
Porter, K. E. & Riches, K. The vascular smooth muscle cell: A therapeutic target in Type 2 diabetes?. Clin. Sci. (Lond.) 125, 167–182. https://doi.org/10.1042/CS20120413 (2013).
Son, S. M. Role of vascular reactive oxygen species in development of vascular abnormalities in diabetes. Diabetes Res. Clin. Pract. 77(Suppl 1), S65-70. https://doi.org/10.1016/j.diabres.2007.01.036 (2007).
Lin, S., Lin, R., Zhang, H., Xu, Q. & He, Y. Peripheral vascular remodeling during ischemia. Front. Pharmacol. 13, 1078047. https://doi.org/10.3389/fphar.2022.1078047 (2022).
Shi, J., Yang, Y., Cheng, A., Xu, G. & He, F. Metabolism of vascular Smooth muscle cells in vascular diseases. Am. J. Physiol. Heart Circ. Physiol. 319, H613–H631. https://doi.org/10.1152/ajpheart.00220.2020 (2020).
Perez-Cremades, D., Cheng, H. S. & Feinberg, M. W. Noncoding RNAs in critical limb ischemia. Arterioscler. Thromb. Vasc. Biol. 40, 523–533. https://doi.org/10.1161/ATVBAHA.119.312860 (2020).
Qi, M. et al. Analysis of long non-coding RNA expression of lymphatic endothelial cells in response to type 2 diabetes. Cell. Physiol. Biochem. 41, 466–474. https://doi.org/10.1159/000456599 (2017).
Chen, B. et al. Non-coding RNAs in IGF-1R Signaling regulation: The underlying pathophysiological link between diabetes and cancer. Cells https://doi.org/10.3390/cells8121638 (2019).
Li, M., Qian, M., Kyler, K. & Xu, J. Endothelial-vascular smooth muscle cells interactions in atherosclerosis. Front. Cardiovasc. Med. 5, 151. https://doi.org/10.3389/fcvm.2018.00151 (2018).
Benincasa, G., Marfella, R., Della Mura, N., Schiano, C. & Napoli, C. Strengths and opportunities of network medicine in cardiovascular diseases. Circ. J. 84, 144–152. https://doi.org/10.1253/circj.CJ-19-0879 (2020).
Silverman, E. K. et al. Molecular networks in Network Medicine: Development and applications. Wiley Interdiscip. Rev. Syst. Biol. Med. 12, e1489. https://doi.org/10.1002/wsbm.1489 (2020).
Collaborators, G. B. D. P. A. D. Global burden of peripheral artery disease and its risk factors, 1990–2019: A systematic analysis for the Global Burden of Disease Study 2019. Lancet Glob. Health 11, e1553–e1565. https://doi.org/10.1016/S2214-109X(23)00355-8 (2023).
Hu, Z. et al. Long noncoding RNA ACTA2-AS1 inhibits cell growth and facilitates apoptosis in gastric cancer by binding with miR-6720-5p to regulate ESRRB. Biochem. Genet. 61, 2672–2690. https://doi.org/10.1007/s10528-023-10399-5 (2023).
Peng, Y. et al. Thymic stromal lymphopoietin-induced HOTAIR activation promotes endothelial cell proliferation and migration in atherosclerosis. Biosci. Rep. https://doi.org/10.1042/BSR20170351 (2017).
Wu, X., Pang, Q., Zhi, F., Mao, X. & Hu, Y. Overexpression of long non-coding RNA ACTA2-AS1 inhibits the viability, proliferation, migration and invasion of colorectal cancer cells. Tissue Cell 76, 101769. https://doi.org/10.1016/j.tice.2022.101769 (2022).
Arencibia, A., Lanas, F. & Salazar, L. A. Long non-coding RNAs might regulate phenotypic switch of vascular smooth muscle cells acting as ceRNA: Implications for in-stent restenosis. Int. J. Mol. Sci. https://doi.org/10.3390/ijms23063074 (2022).
Cheng, Z. et al. Long non-coding RNA p53 upregulated regulator of p53 levels (PURPL) promotes the development of gastric cancer. Bioengineered 13, 1359–1376. https://doi.org/10.1080/21655979.2021.2017588 (2022).
Fang, E. et al. Therapeutic targeting of MZF1-AS1/PARP1/E2F1 axis inhibits proline synthesis and neuroblastoma progression. Adv. Sci. (Weinh) 6, 1900581. https://doi.org/10.1002/advs.201900581 (2019).
Ghafouri-Fard, S., Safarzadeh, A., Hussen, B. M., Taheri, M. & Ayatollahi, S. A. A review on the role of LINC00511 in cancer. Front. Genet. 14, 1116445. https://doi.org/10.3389/fgene.2023.1116445 (2023).
Li, Y. et al. Knocking down of LINC01220 inhibits proliferation and induces apoptosis of endometrial carcinoma through silencing MAPK11. Biosci. Rep. https://doi.org/10.1042/BSR20181794 (2019).
Li, Z., Wang, D., Zhang, W., Shi, H. & Zhu, M. Novel PBMC LncRNA signatures as diagnostic biomarkers for colorectal cancer. Pathol. Res. Pract. 253, 154985. https://doi.org/10.1016/j.prp.2023.154985 (2023).
Ou, C., He, X., Liu, Y. & Zhang, X. lncRNA cytoskeleton regulator RNA (CYTOR): Diverse functions in metabolism, inflammation and tumorigenesis, and potential applications in precision oncology. Genes Dis. 10, 415–429. https://doi.org/10.1016/j.gendis.2021.08.012 (2023).
Jia, X. et al. Identification of crucial lncRNAs for luminal A breast cancer through RNA sequencing. Int. J. Endocrinol. 2022, 6577942. https://doi.org/10.1155/2022/6577942 (2022).
Sun, T. et al. Expression of lncRNA MIR222HG co-transcribed from the miR-221/222 gene promoter facilitates the development of castration-resistant prostate cancer. Oncogenesis 7, 30. https://doi.org/10.1038/s41389-018-0039-5 (2018).
Wen, S. et al. MIR222HG attenuates macrophage M2 polarization and allergic inflammation in allergic rhinitis by targeting the miR146a-5p/TRAF6/NF-kappaB axis. Front. Immunol. 14, 1168920. https://doi.org/10.3389/fimmu.2023.1168920 (2023).
Basatemur, G. L., Jorgensen, H. F., Clarke, M. C. H., Bennett, M. R. & Mallat, Z. Vascular smooth muscle cells in atherosclerosis. Nat. Rev. Cardiol. 16, 727–744. https://doi.org/10.1038/s41569-019-0227-9 (2019).
Ni, H. et al. A smooth muscle cell-enriched long noncoding RNA regulates cell plasticity and atherosclerosis by interacting with serum response factor. Arterioscler. Thromb. Vasc. Biol. 41, 2399–2416. https://doi.org/10.1161/ATVBAHA.120.315911 (2021).
Chen, L. et al. Acrolein-induced apoptosis of smooth muscle cells through NEAT1-Bmal1/Clock pathway and a protection from asparagus extract. Environ. Pollut. 258, 113735. https://doi.org/10.1016/j.envpol.2019.113735 (2020).
Chen, X. et al. Evaluation of lncRNA expression pattern and potential role in heart failure pathology. Dis. Markers 2023, 2369352. https://doi.org/10.1155/2023/2369352 (2023).
Sohrabifar, N., Ghaderian, S. M. H., Alipour Parsa, S., Ghaedi, H. & Jafari, H. Variation in the expression level of MALAT1, MIAT and XIST lncRNAs in coronary artery disease patients with and without type 2 diabetes mellitus. Arch. Physiol. Biochem. 128, 1308–1315. https://doi.org/10.1080/13813455.2020.1768410 (2022).
Vlachogiannis, N. I. et al. Adenosine-to-inosine Alu RNA editing controls the stability of the pro-inflammatory long noncoding RNA NEAT1 in atherosclerotic cardiovascular disease. J. Mol. Cell Cardiol. 160, 111–120. https://doi.org/10.1016/j.yjmcc.2021.07.005 (2021).
Yang, K., Xue, Y. & Gao, X. LncRNA XIST Promotes Atherosclerosis by Regulating miR-599/TLR4 Axis. Inflammation 44, 965–973. https://doi.org/10.1007/s10753-020-01391-x (2021).
Zhang, X. et al. NEAT1 knockdown suppresses endothelial cell proliferation and induces apoptosis by regulating miR-638/AKT/mTOR signaling in atherosclerosis. Oncol. Rep. 44, 115–125. https://doi.org/10.3892/or.2020.7605 (2020).
Zhang, X., Li, H., Guo, X., Hu, J. & Li, B. Long noncoding RNA hypoxia-inducible factor-1 alpha-antisense RNA 1 regulates vascular smooth muscle cells to promote the development of thoracic aortic aneurysm by modulating apoptotic protease-activating factor 1 and targeting let-7g. J. Surg. Res. 255, 602–611. https://doi.org/10.1016/j.jss.2020.05.063 (2020).
Ghafouri-Fard, S. et al. A concise review on the role of MIR100HG in human disorders. J. Cell Mol. Med. 27, 2278–2289. https://doi.org/10.1111/jcmm.17875 (2023).
Ahmed, A. S. I. et al. Long noncoding RNA NEAT1 (nuclear paraspeckle assembly transcript 1) is critical for phenotypic switching of vascular smooth muscle cells. Proc. Natl. Acad. Sci. U S A 115, E8660–E8667. https://doi.org/10.1073/pnas.1803725115 (2018).
He, Q., Tan, J., Yu, B., Shi, W. & Liang, K. Long noncoding RNA HIF1A-AS1A reduces apoptosis of vascular smooth muscle cells: Implications for the pathogenesis of thoracoabdominal aorta aneurysm. Pharmazie 70, 310–315 (2015).
Liang, K. et al. LncRNA Xist induces arterial smooth muscle cell apoptosis in thoracic aortic aneurysm through miR-29b-3p/Eln pathway. Biomed. Pharmacother. 137, 111163. https://doi.org/10.1016/j.biopha.2020.111163 (2021).
Shi, C. et al. NEAT1 promotes the repair of abdominal aortic aneurysms of endothelial progenitor cells via regulating miR-204-5p/Ang-1. Am. J. Transl. Res. 13, 2111–2126 (2021).
Alfaifi, M. et al. Circulating long non-coding RNAs NKILA, NEAT1, MALAT1, and MIAT expression and their association in type 2 diabetes mellitus. BMJ Open Diabetes Res. Care https://doi.org/10.1136/bmjdrc-2020-001821 (2021).
Hassani, S. S. et al. Dysregulation of long noncoding RNA NEAT1/miR-199a-5/BiP axis in patients with diabetic neuropathy. Lab. Med. 54, 160–165. https://doi.org/10.1093/labmed/lmac082 (2023).
Jia, D. et al. NEAT1: A novel long non-coding RNA involved in mediating type 2 diabetes and its various complications. Curr. Pharm Des. 28, 1342–1350. https://doi.org/10.2174/1381612828666220428093207 (2022).
Zhao, Y. et al. The impact of lncRNA on diabetic kidney disease: Systematic review and in silico analyses. Comput. Intell. Neurosci. 2022, 8400106. https://doi.org/10.1155/2022/8400106 (2022).
Chen, H., Guo, Y. & Cheng, X. Long non-cording RNA XIST promoted cell proliferation and suppressed apoptosis by miR-423-5p/HMGA2 axis in diabetic nephropathy. Mol. Cell Biochem. 476, 4517–4528. https://doi.org/10.1007/s11010-021-04250-x (2021).
Long, B., Wan, Y., Zhang, S. & Lv, L. LncRNA XIST protects podocyte from high glucose-induced cell injury in diabetic nephropathy by sponging miR-30 and regulating AVEN expression. Arch. Physiol. Biochem. 129, 610–617. https://doi.org/10.1080/13813455.2020.1854307 (2023).
Wang, S. et al. BRG1 expression is increased in thoracic aortic aneurysms and regulates proliferation and apoptosis of vascular smooth muscle cells through the long non-coding RNA HIF1A-AS1 in vitro. Eur. J. Cardiothorac. Surg. 47, 439–446. https://doi.org/10.1093/ejcts/ezu215 (2015).
Zhang, K., Qi, Y., Wang, M. & Chen, Q. Long non-coding RNA HIF1A-AS2 modulates the proliferation, migration, and phenotypic switch of aortic smooth muscle cells in aortic dissection via sponging microRNA-33b. Bioengineered 13, 6383–6395. https://doi.org/10.1080/21655979.2022.2041868 (2022).
Busscher, D., Boon, R. A. & Juni, R. P. The multifaceted actions of the lncRNA H19 in cardiovascular biology and diseases. Clin. Sci. (Lond) 136, 1157–1178. https://doi.org/10.1042/CS20210994 (2022).
Li, D. Y. et al. H19 induces abdominal aortic aneurysm development and progression. Circulation 138, 1551–1568. https://doi.org/10.1161/CIRCULATIONAHA.117.032184 (2018).
Shi, X. et al. Long non-coding RNA H19 in atherosclerosis: what role?. Mol. Med. 26, 72. https://doi.org/10.1186/s10020-020-00196-w (2020).
Simion, V., Haemmig, S. & Feinberg, M. W. LncRNAs in vascular biology and disease. Vascul. Pharmacol. 114, 145–156. https://doi.org/10.1016/j.vph.2018.01.003 (2019).
Bi, Y., Wang, Y. & Sun, X. Recent advances of LncRNA H19 in diabetes LncRNA H19 in diabetes. Horm. Metab. Res. 54, 212–219. https://doi.org/10.1055/a-1769-5032 (2022).
Hussein, R. M. Long non-coding RNAs: The hidden players in diabetes mellitus-related complications. Diabetes Metab. Syndr. 17, 102872. https://doi.org/10.1016/j.dsx.2023.102872 (2023).
Bergman, D., Halje, M., Nordin, M. & Engstrom, W. Insulin-like growth factor 2 in development and disease: A mini-review. Gerontology 59, 240–249. https://doi.org/10.1159/000343995 (2013).
Dai, N. The diverse functions of IMP2/IGF2BP2 in metabolism. Trends Endocrinol. Metab. 31, 670–679. https://doi.org/10.1016/j.tem.2020.05.007 (2020).
Nielsen, F. C., Nielsen, J. & Christiansen, J. A family of IGF-II mRNA binding proteins (IMP) involved in RNA trafficking. Scand. J. Clin. Lab. Invest. Suppl. 234, 93–99 (2001).
Dong, K. et al. CARMN is an evolutionarily conserved smooth muscle cell-specific LncRNA that maintains contractile phenotype by binding myocardin. Circulation 144, 1856–1875. https://doi.org/10.1161/CIRCULATIONAHA.121.055949 (2021).
Vacante, F. et al. CARMN loss regulates smooth muscle cells and accelerates atherosclerosis in mice. Circ. Res. 128, 1258–1275. https://doi.org/10.1161/CIRCRESAHA.120.318688 (2021).
Wu, Z., Yin, H., Guo, Y., Yin, H. & Li, Y. Detection of cell-type-enriched long noncoding RNAs in atherosclerosis using single-cell techniques: A brief review. Life Sci. 333, 122138. https://doi.org/10.1016/j.lfs.2023.122138 (2023).
Zhou, H. et al. Long noncoding RNAs in pathological cardiac remodeling: A review of the update literature. Biomed. Res. Int. 2019, 7159592. https://doi.org/10.1155/2019/7159592 (2019).
Rajabinejad, M. et al. The MALAT1-H19/miR-19b-3p axis can be a fingerprint for diabetic neuropathy. Immunol. Lett. 245, 69–78. https://doi.org/10.1016/j.imlet.2022.03.004 (2022).
Pelisek, J. et al. Swiss vascular biobank: Evaluation of optimal extraction method and admission solution for preserving RNA from human vascular tissue. J. Clin. Med. https://doi.org/10.3390/jcm12155109 (2023).
Hatakeyama, M. et al. SUSHI: an exquisite recipe for fully documented, reproducible and reusable NGS data analysis. BMC Bioinform. 17, 228. https://doi.org/10.1186/s12859-016-1104-8 (2016).
Qi, W., Schlapbach, R. & Rehrauer, H. RNA-seq data analysis: From raw data quality control to differential expression analysis. Methods Mol. Biol. 1669, 295–307. https://doi.org/10.1007/978-1-4939-7286-9_23 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: Fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108. https://doi.org/10.1093/nar/gkt214 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Reiner, A., Yekutieli, D. & Benjamini, Y. Identifying differentially expressed genes using false discovery rate controlling procedures. Bioinformatics 19, 368–375. https://doi.org/10.1093/bioinformatics/btf877 (2003).
Acknowledgements
The authors gratefully acknowledge the Functional Genomics Center Zurich (FGCZ) of the University of Zurich and ETH Zurich, particularly Dr Peter Leary, Dr Hubert Rehrauer, and Dr Anna Bratus-Neuenschwander, for their support in the RNA sequencing and analysis.
Funding
This research project was founded by the Swiss Heart Foundation (FF21065). The Swiss Vascular Biobank was supported by the Department of Vascular Surgery (Director Professor Alexander Zimmermann, University of Zurich, Switzerland).
Author information
Authors and Affiliations
Contributions
J.P. and Y.Y. wrote and designed the manuscript; J.P. and Y.Y. processed the tissue samples and performed all the corresponding experiments. S.M. contributed through statistical analysis of the RNA sequencing data sets. B.R., F.P., and A.Z. contributed by providing the vascular tissue samples and critically revising the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yundung, Y., Mohammed, S., Paneni, F. et al. Transcriptomics analysis of long non-coding RNAs in smooth muscle cells from patients with peripheral artery disease and diabetes mellitus. Sci Rep 14, 8615 (2024). https://doi.org/10.1038/s41598-024-59164-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-59164-7
- Springer Nature Limited