Abstract
Shigellosis remains a common gastrointestinal disease mostly in children < 5 years of age in developing countries. Azithromycin (AZM), a macrolide, is currently the first-line treatment for shigellosis in Bangladesh; ciprofloxacin (CIP) and ceftriaxone (CRO) are also used frequently. We aimed to evaluate the current epidemiology of antimicrobial resistance (AMR) and mechanism(s) of increasing macrolide resistance in Shigella in Bangladesh. A total of 2407 clinical isolates of Shigella from 2009 to 2016 were studied. Over the study period, Shigella sonnei was gradually increasing and become predominant (55%) over Shigella flexneri (36%) by 2016. We used CLSI-guided epidemiological cut-off value (ECV) for AZM in Shigella to set resistance breakpoints (zone-diameter ≤ 15 mm for S. flexneri and ≤ 11 mm for S. sonnei). Between 2009 and 2016, AZM resistance increased from 22% to approximately 60%, CIP resistance increased by 40%, and CRO resistance increased from zero to 15%. The mphA gene was the key macrolide resistance factor in Shigella; a 63MDa conjugative middle-range plasmid was harboring AZM and CRO resistance factors. Our findings show that, especially after 2014, there has been a rapid increase in resistance to the three most effective antibiotics. The rapid spread of macrolide (AZM) resistance genes among Shigella are driven by horizontal gene transfer rather than direct lineage.
Similar content being viewed by others
Introduction
Shigella is the most common pathogen for gastrointestinal infection in developing countries and the leading cause of death among children < 5 years globally1,2,3,4,5. Shigella flexneri is the predominant strain but Shigella sonnei is the uprising strain in low-and-middle-income countries (LMICs) including Bangladesh5,6,7,8,9. The sustained pressure of microbial infection and the tendency to quickly reduce the disease duration and severity has led to indiscriminate use of antimicrobials, therefore, triggering the raise of superbugs in developing countries10,11. World Health Organization (WHO) recommends ciprofloxacin (CIP) as the first-line therapy along with pivmecillinam, ceftriaxone, and azithromycin as alternative options. Due to the high CIP-resistance in Shigella in Bangladesh, the efficacy of CIP is currently in doubt. Recently, Shigella isolates have been reported to acquire resistant genes and plasmid with reduce susceptibility to fluoroquinolones and third-generation cephalosporins12,13. Ceftriaxone resistance is low in Shigella but it is given parentally, therefore not encouraged for children14,15. Therefore, the macrolide AZM is widely used as the most preferred therapy for shigellosis in children16.
The IPC policy in Bangladesh is poorly followed and implemented in practice. Indiscriminate use of antibiotics is one of the key aberrant features of antibiotics usage management in LMICs17,18. It has been recently reported from Bangladesh that over 25% of the antibiotic users self-medicated themselves and azithromycin was the second highest (21%) self-medicated drug17,18.
However, there was no established clinical susceptibility breakpoints of AZM for Shigella before 201619. Therefore, in Bangladesh, few studies reported AZM susceptibility for Shigella but using different breakpoints which was inconsistent with current CLSI guideline7,20,21,22,23. Several studies have reported the emergence of AZM resistance in Shigella spp. globally24,25,26 and described the mechanism for AZM resistance23. To date, different molecular mechanisms involved in the development of resistance to AZM have been described. Shigella confers resistance to macrolides through variety of mechanisms include target site modification by methylases, enzymatic inactivation by esterases or phosphotransferases and through efflux pumps27,28,29,30. Several reports suggested that plasmid-mediated macrolide 2'-phosphotransferase (mphA) mostly and esterase (ermB) for some instances inactivate macrolide through modifying its molecular structure31,32. Furthermore, conjugative R-plasmid mediated horizontal gene transfer (HGT) was demonstrated to be involved in the rapid transfer of genes responsible for resistance33,34,35,36. In 2015, middle range plasmid (50 MDa) mediated transfer of third generation cephalosporin resistance between Escherichia coli and S. sonnei was reported in Bangladesh37. Recently it has been described that a conjugative R-plasmid carrying azithromycin-resistance genes was involved in reduced susceptibility of S. flexneri serotype 3a to AZM35.
Furthermore, given the limited treatment options for children with shigellosis, monitoring resistance rates and studying macrolide resistance mechanisms (AZMs) is not only a necessity but a task. Due to the rapid spread of the MDR phenomenon, advanced studies are always needed to assess and track real-time AMR burden in Shigella. In this study, we reported a trend towards AMR resistance in Shigella and the key mechanism of macrolide resistance in Shigella spp.
Results
Distribution of Shigella strains in Bangladesh between 2009 and 2016
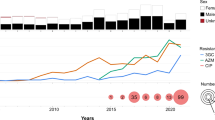
Distribution of Shigella strains (n = 2407) were observed between 2009 and 2016. During this study period, S. flexneri was the dominant species (48%) until 2015. However, the prevalence of S. flexneri decreased by 17% from 2009 (53%; 314/593) to 2016 (36%; 24/66). At the same time, S. sonnei was increased from 20% (119/593) in 2009 to 55% (36/66) in 2016. During this -period, a 10% decrease in the number of S. boydii was observed (from 18 to 8%). The frequency of S. dysenteriae was consistently low in subsequent years, becoming sporadic (2%) in 2016 (Fig. 1A).
(A) Epidemiological distribution of Shigella spp. from 2009 to 2016. Different patterns presenting the four species of Shigella indicating a clear increasing trend in S. sonnei and decreasing trend in S. boydii and S. dysenteriae. (B) Changing pattern in resistance to AZM, CIP and CRO in the time-period of 2009 to 2016. Bar chart indicating an increasing trend for all three most used drugs to treat shigellosis. Rate of resistance showed sharp increase in 2015 and 2016. Resistance to CIP and CRO increasing gradually. Microsoft Excel 2013 was used in visualization.
Susceptibility breakpoints for azithromycin and Shigella
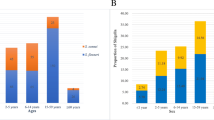
Epidemiological cut-off values (ECVs) are not intended to determine clinical susceptibility cutoffs. Therefore, we performed a non-parametric Spearman rank test between the diameter of the inhibition zone of the azithromycin disk and the available MIC data for 32 S. flexneri and 59 S. sonnei isolates. A significant correlation was observed between MIC and disc diffusion zone size for both S. flexneri (rho, − 0.907; P < 0.0001) and S. sonnei (rho, − 0.862; P < 0.0001). We found no exception to determine the diameter of the disc diffusion zone of S. flexneri and S. sonnei at the respective MIC values; zone diameter ≤ 15 mm in non wild-type (NWT) S. flexneri (MIC ≥ 16 µg/ml) and zone diameter ≤ 11 mm in NWT S. sonnei (MIC ≥ 32 µg/ml) (Fig. 2).
Scatterplot on azithromycin MIC by disc diffusion zone diameter in Shigella spp. (A) Azithromycin MIC (y axis) and inhibition zone diameter (x axis) showing zone diameter ≤ 15 mm can well segregate susceptible (MIC ≤ 8 µg/ml) and resistant (MIC ≥ 16 µg/ml) in Shigella flexneri. (B) Azithromycin MIC (y axis) and inhibition zone diameter (x axis) showing zone diameter ≤ 11 mm can well segregate susceptible (MIC ≤ 16 µg/ml) and resistant (MIC ≥ 32 µg/ml) in Shigella sonnei. IBM SPSS Statistics 26 were used to generate the figure. AZM = azithromycin, NWT = AZM-resistant, WT = AZM-susceptible, MIC = Minimum inhibitory concentration.
Antibiotic resistance pattern in Shigella spp.
A total of 770 Shigella strains (336 S. flexneri, 233 S. sonnei, 162 S. boydii and 39 S. dysenteriae) were subjected to AST. More than 96% (274/284) of the Shigella strains were found resistant to erythromycin and 30% (222/748) to azithromycin (Table 1). In 2014, 27% of Shigella strains were AZM resistant, which was doubled (59%) by 2016 (Fig. 1B). S. flexneri and S. sonnei were found to confer higher resistance (AZMR) than the other two species (Table 1).
Throughout the study, 44% of the Shigella spp. was found CIP-resistant; S. sonnei had significantly higher resistance to CIP (76%) compared to S. flexneri (45%), S. boydii (6%) and S. dysentery (3%). In 2016, more than 70% of Shigella were found to be resistant to CIP, an increase of 40% since 2009 (30%) (Fig. 1B). Before 2014, CRO resistance was less than 5% but, between 2015 and 2016, CRO resistance increased to 15% (Fig. 1B).
Other third generation cephalosporins including cefotaxime (CTX), ceftazidime (CAZ) and cefixime (CFM) was conferred 11%, 3% and 14% resistance in Shigella respectively. In addition, we found 3% resistance to mecillinam (MEL), 42% to ampicillin (AMP) and 58% to trimethoprim-sulfamethoxazole (SXT) in Shigella.
The mphA gene conferring decreasing susceptibility to macrolide in Shigella spp.
We determined macrolide resistance genes among 37 AZM-resistant Shigella spp. which contains 14 S. flexneri, 17 S. sonnei, 4 S. boydii and 2 S. dysenteriae. Out of the 37 AZM-resistant Shigella, 95% were positive for the mphA gene in the PCR test. The remaining 2 isolates did not show a band for any of the macrolide resistance genes studied. The AZM-resistant isolates of S. sonnei with a zone diameter ≤ 11 mm (MIC < 32 µg/ml) and AZM-resistant isolates of the other three species with a zone diameter ≤ 12 mm (MIC < 16 µg/ml) in disc diffusion method found positive for the mphA gene (Table 2).
Prevalence of middle-ranged plasmid (MRP) in macrolide resistant strains
We determined the plasmid profiles of 59 Shigella strains; 42 AZM-resistant and 17 AZM- sensitive isolates. Heterogeneous plasmid patterns were distributed in both resistant and susceptible Shigella strains. The plasmid size was measured between 1.0 and 140.0 MDa. Almost 80% of the Shigella isolates possessed a 140 MDa plasmid and the small plasmid (< 6 MDa) was uniformly distributed in all Shigella isolates. Middle-ranged plasmid (MRP) of approximately 35–90 MDa in size was significantly more prevalent (p < 0.0001) in AZM-resistant Shigella strains (60%, 25/42) compared to susceptible strains (24%, 4/17) (Fig. 3).
Horizontal transfer of AMR
Antimicrobial susceptibility testing confirmed that the transconjugants resistance to azithromycin, erythromycin, ampicillin and ceftriaxone, same as the donor stains (S. flexneri K12582 and S. sonnei K12747) (Table 3). The MIC of azithromycin was ≥ 256 µg/ml for all transconjugants. Plasmid analysis of transconjugants revealed that only 63 MDa plasmid was transferred (Fig. 4A) from both donor Shigella isolates to E. coli K-12 recipient. The mphA gene was confirmed in the transconjugants and their plasmid DNAs by PCR (Fig. 4B).
Plasmid and PCR analysis of transconjugants. (A) Agarose gel electrophoresis of plasmid DNA from conjugation study showing representative patterns of parent donor strains (Lane D = K12582 and Lane G = K12747), recipient strain (Lane B = K-12), transconjugants (Lane C = Tc-K12582 and Lane F = Tc-K12747) and plasmid size markers (Lane A = PDK-9, Lane E & H = Sa + R1 and Lane I = V-517). (B) Gel illusion of mphA gene using plasmid from transconjugant as template. On lane A = Ladder, Lane B = Tc-12582, lane C = Tc-12747, lane D = K-12, lane E = Negative control and lane F = K12747 (PC). *Keynote: Tc = Transconjugant, and PC = Positive control. The original gel images (both A and B) were provided in the Supplementary Fig. S1. Paint application (windows operating system) was used to edit the images.
Plasmid typing and resistome profiling
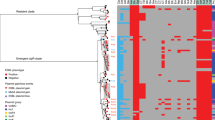
Analyzing the whole genome sequencing based resistome profiles of the three AZM-resistant Shigella, we identified 11, 14 and 15 antimicrobial resistance factors in the genomes of S. flexneri Z12966, S. flexneri Z13032 and S. boydii Z12959 respectively (Table 4). Using PlasmidFinder, we found that all 3 strains were harboring at least a IncFII-type plasmid in their genome. From Mob-suite results, the size of the IncF plasmids were ~ 90 kbp (~ 60 MDa). Separate estimation of AMR-genotype in chromosome and plasmid showed that resistance factors to azithromycin (mphA, ermB, msrE and mphE) and cephalosporins (blaTEM-1, blaDHA-1, blaCTXM-15 and blaOXA-1) were plasmid borne, whereas, ciprofloxacin resistance genes were chromosomal (Table 4). In addition, plasmids were conferring the highest and most potential resistance factors in the three Shigella genomes studied.
Determination of clonal variation of the azithromycin-resistant Shigella isolates
PFGE analysis of Xba-I digested chromosomal DNA of the AzmR and AzmS Shigella strains yielded 21 to 23 reproducible DNA fragments ranging in size approximately from 20 to 690 Kb (Supplementary Fig. S2). Dendrogram on fragment sizes showed no different pulsotype-clustering based on AZM susceptibility. AZM-resistant Shigella was not from any single clone of Shigella spp. (Similarity < 98%). However, species-wise pulsotype-clustering was present. (Fig. 5).
Discussion
Antimicrobial resistance has been a long-persistent major public health issue, particularly in underdeveloped and developing nations where shigellosis is endemic. In this study, we report a rapid increase in the resistance to the first-line antibiotics used to treat shigellosis, especially a 40% increase of AZM resistance among Shigella spp. in just two years (2014–2016) years. In fact, this is the first report in Bangladesh showing a pattern of AZM-resistance in Shigella spp. following the publication of the CSLI defined ECVs for AZM and Shigella. The altered temporal dominance of S. sonnei over S. flexneri has been demonstrated and MRP-mediated HGT is considered to be the main mechanism of AMR spread. Several studies reported emergence of increasing S. sonnei worldwide including Bangladesh6,8. In 2001, 6% of S. sonnei was reported in Bangladesh6,7,38, which was increased to 54% in 2016. This acute temporal alteration of S. sonnei by 48% in just 15 years seems dramatic in geo-environmental timeframe, gives potential massages of weal and woes in parallel. Continuous improvement in the quality of global drinking water, rapid industrialization, improved nutritional status, better sanitation and less immune-cross-reaction have been resulted in reduced less-adaptive Shigella spp. and increased more-adaptive S. sonnei load6,8. Simultaneously, antibiotic driven immense selection pressure and efficient dissemination channels can resonate the emergence of S. sonnei and signs chronic potential problems like spread of MDR S. sonnei39,40.
A decade ago, several drugs were considered to treat Shigellosis e.g. ciprofloxacin, tetracycline, chloramphenicol, ampicillin, trimethoprim-sulfamethoxazole, nalidixic acid etc41. Most of them have long since lost their effectiveness due to low intestinal absorption, cross-reactivity and mainly due to high resistance to Shigella.
After being the most preferred treatment option, CIP is seldom prescribed to treat Shigellosis in countries like Bangladesh currently because of its resistance mediated inefficiency14,42. In recent years, the prevalence of CIP resistance has been about 70% in patients of all ages, especially in Bangladesh43,44. In our study, we also found more than 70% CIP-resistance in 2016. Ceftriaxone is a potential alternative in shigellosis treatment but high cost and route of administration reduces its compatibility14,15. Moreover, a rapid increase of CRO-resistance was found in our study.
Empirically administered AZM offers an attractive option for its low frequent dosage system and high intracellular concentration in the colon of patients with shigellosis. The absence of clinical or epidemiological cutoff values lead to unclear conclusions until 201645,46,47. Previously, Rahman et al. followed Antimicrobial Chemotherapy (BSAC) guidelines (sensitive: ≥ 18 mm and resistant: < 18 mm) for AZM breakpoint22; Bourtchai et al. followed Clinical Laboratory Standards Institute breakpoints recommended for Streptococci (> 1 mg/l, resistant; < 0.25 mg/l, susceptible)20; Murray et al. considered all isolates with an MIC of AZM of > 32 isolates as DSA according to CDC21. In 2016, CLSI suggested ‘epidemiological cutoff values’ (ECVs)46. In 2017, Darton et al. demonstrated S. flexneri (MIC ≥ 16 g/l, zone diameter ≤ 15 mm) and S. sonnei (MIC ≥ 32 g/l, zone diameter ≤ 11 mm) breakpoints for AZM based on ECVs of CLSI guidelines47. However, there is no clinical breakpoint or ECVs for AZM for S. boydii and S. dysenteriae in CLSI and EUCAST46. Therefore, this confusing situation regarding the AZM breakpoint is not over yet.
In the current study, we found sharp increase of AZM-resistance after 2014 and mphA gene was the key mechanism of resistance in molecular and WGS based approaches. Resistome profiling in individual chromosome and plasmid demonstrated the major contribution of plasmid-borne horizontal gene transfer (HGT) in AMR spread rather than clonal expansion. The predominance of HGT over clonality in the spread of azithromycin resistance is reasonable and well addressed in different studies worldwide35,48,49,50,51,52. HGT is the most energy-efficient way to transfer genetic material for bacterial species. In addition, HGT allows bacteria inter-species dissemination of AMR factors which is not possible for clonal transfer. A IncFII type plasmid pKSR100 is the most crucial genomic entity fostering AZM resistance35,52. We reported for the first time of macrolide resistance pKSR100 when the plasmid carrying IS26–mphA–mrx–mphR(A)–IS6100 in Shigella isolated in Bangladesh the years after 201453.
By correlating the mechanism of AZM resistance, as CRO resistance increased significantly after 2014, the involvement of an emerging R-plasmid carrying CRO-resistant factors in Shigella can be strongly inferred. In addition, the transfer of AZM, CRO and AMP resistance phenomena through one conjugative R-plasmid indicate the chance of rapid inter-species dissemination of resistance factors. The involvement of 63 MDA plasmid in AMR-resistance spread well aligns the WGS-based findings and the global studies on the role of IncF plasmids in AMR spread. This size of plasmid is optimum to harbor multiple resistance genes and mechanisms for HGT but can fit into a tiny bacterial genome54. We do not think that the pathogen responsible for the acute diarrhea acquires resistance genes immediately upon reaching the hospital as patients with severe diarrhea were admitted to the hospital and fecal samples were collected right after patient’s admission in the hospital. In the PFGE study, the AZM-resistant Shigella were not confined in same pulsotype-cluster and they were not from same clone of AZM-resistant bacteria. These findings indicate that horizontal transfer contributes more than direct lineage to spread AMR more rapidly.
Lack of patient data is one of the limitations of the manuscript which might strengthen the schematic illustration of AMR scenario in terms of population features. Although the data presented in the manuscript not from very recent years, it is still appealing and well connected to the current trends of antibiotic resistance in Shigella in Bangladesh. Firstly, it fills the information gap by presenting the AMR scenario during the crucial turnover time period of antibiotic resistance in Shigella in Bangladesh when the dramatic increase in azithromycin and ceftriaxone resistance happened during the time-period of 2014–2015. Secondly, the data presented in the manuscript bridges the azithromycin resistance rates before and after the epidemiological cut-off value (ECV) of azithromycin for Shigella being published in 201646. Lastly but most importantly, azithromycin and third generation cephalosporins (e.g. ceftriaxone) are the most prescribed drugs in case of shigellosis treatment in Bangladesh, therefore, it is crucial to have the clear picture of antibiotic resistance trends.
Overall, the current study provides a clear depiction of the major increase of AMR in Shigella and pragmatically evaluates the azithromycin resistance mechanisms in Bangladesh. It signifies the urgency of robust AMR surveillance, resistome profiling and transmission dynamics study in Shigella at a large and global scale.
Materials and methods
Study population
A total of 2407 Shigella strains were identified and isolated between 2009 and 2016 in the Clinical Microbiology Laboratory from the stool specimen of diarrheal patients admitted in icddr,b hospital unit, Dhaka, Bangladesh. Patients with acute watery or bloody diarrhea were kept under observation for two hours upon arriving at the icddr,b hospital. If situation not improve after the observation period providing normal saline and if the watery discharge remains frequent (> 3 times an hour), they were given admission. Only Bangladeshi patients with no history of recent abroad-visit were asked for consent. Fecal samples were collected from diarrheal patients (with consent) and subjected to culture procedures to identify diarrheal pathogen. The identified diarrheal pathogen was subjected to identification tests using phenotypical and Shigella spp. specific polyvalent antisera. Diarrheal infection was termed as “Shigellosis” once we confirmed the etiological agent as Shigella app. Serotype of all the Shigella strains were confirmed in Laboratory of Gut-Brain Signaling, icddr,b, using standard microbiological and biochemical methods55. Among these strains, 770 isolates were subjected to antibiotic susceptibility test (AST) and further analysis. This study was reviewed and approved by institutional review board (IRB) of icddr,b, Dhaka, Bangladesh. Three multidrug-resistant Shigella (2 Shigella flexneri and 1 Shigella boydii) isolated in 2017 were used for resistome profiling.
Serotyping of Shigella species
Isolated Shigella strains were sub-cultured on MacConkey agar (Difco, Becton Dickinson & Company, Sparks, Md.) plates, and incubated for 16 h for optimum growth. Serotyping was performed by the slide agglutination test56. Two types of commercially available kits were used in this study; (i) antisera specific for all type- and group-factor antigens (Denka Seiken, Tokyo, Japan) (ii) monoclonal antibody reagents (Reagensia AB, Stockholm, Sweden) specific for all S. flexneri type- and group-factor antigens. After serotyping, single colony of the strains was inoculated in Trypticase soy broth containing 0.3% yeast extract (TSBY), incubated for 16 h and stored at − 70 °C with 15% glycerol afterwards.
Antibiotic susceptibility tests (AST)
We determined the bacterial susceptibility to antimicrobial agents by the disc diffusion method according to the guidelines of CLSI46 using Muller-Hinton agar and commercially available antimicrobial discs (Oxoid, Basingstoke, United Kingdom)57. We used E. coli (ATCC 25922) strain as negative control in AST. As per CLSI guideline, S. flexneri with azithromycin disc diffusion zone diameter ≤ 15 mm and MIC ≥ 16 µg/ml respectively can be considered as NWT. In case of S. sonnei, only MIC (WT, ≤ 16 µg/ml and NWT, ≥ 32 µg/ml) was asserted in CLSI guideline58. In 2018, Thomas C. Darton et al. suggested disc diameter ≤ 11 mm as a cutoff value for S. sonnei. Thus, we aimed to confirm that disc diffusion zone diameter ≤ 11 mm for S. sonnei and ≤ 15 mm for S. flexneri can be used to well discriminate AZM-resistant and sensitive isolates in our population. We followed Centers for Disease Control and Prevention (CDC) guided methodology for S. boydii and S. dysenteriae to define the susceptibility to AZM, where MIC ≥ 32 was defined AZM-resistant (NWT) isolates)59. Four different groups of antibiotic discs were used to perform AST: (i) azithromycin (AZM, 15 μg), erythromycin (ERY, 15 μg) from macrolide, (ii) ampicillin (AMP, 10 μg) and amoxicillin/clavulanate (AMC, 10/20 μg) form penicillin, (iii) ciprofloxacin (CIP, 5 μg), and nalidixic acid (NAL, 30 μg) from quinolone and (iv) ceftriaxone (CRO, 30 μg), ceftazidime (CAZ, 30 μg), cefotaxime (CTX, 30 μg) and cefixime (CFM, 30 μg) from cephalosporins and sulfamethoxazole-trimethoprim (SXT, 25 μg). The MIC was determined by the epsilometer test (E-test; AB Biodisk, Solna, Sweden) as per manufacture’s guideline.
Isolation of plasmid DNA
Plasmid DNA was extracted using modified alkaline lysis method of Kado and Liu60,61. Gel electrophoresis was performed in 0.5% agarose gel at 100 V current for 3 h. Gel was stained with ethidium bromide and visualized under ultraviolet rays. The mobility and size of plasmids present in previously described strains E. coli PDK- 9 (140, 105, 2.7 and 2.1 MDa), R1 (62 MDa), RP- 4 (36 MDa), Sa (23 MDa) and V517 (35.8, 4.8, 3.7, 3.4, 3.1, 2.0, 1.8 and 1.4 MDa) were used as molecular mass standards to scale the unknown plasmid DNA62.
Molecular detection of macrolide resistance genes in Shigella species
A total of 37 AZM-resistant Shigella isolates for macrolide were selected to extracted the DNA and determine macrolide resistance genes. Polymerase chain reaction (PCR) was performed to determine phosphotransferase genes (mphA and mphB), esterase genes (ereA and ereB), rRNA methylase genes (ermA and ermB) and efflux pump mediating factors (mefA and msrA)63 (Supplementary Table S1). The primers used for this study were taken from previously published article20,45.
Conjugation and transfer of R-plasmid
We used two multidrug resistant Shigella strains as donor strain and E. coli K-12 (NALR, Lac+, F-) as the recipient, to conjugate described previously64. In our study, both of the donor strains had an MIC of ≥ 256 µg/ml to azithromycin and were positive for macrolide resistance factor mphA gene. Transconjugants were selected on MacConkey agar containing azithromycin (32 μg/ml: Sigma-Aldrich Corporation, St. Louis, Mo.) and nalidixic acid (32 μg/ml: Sigma-Aldrich Corporation, St. Louis, Mo.) that produce lactose-fermenting pink colonies of E. coli in contrast to non-lactose-fermenting pale colonies of Shigella isolate. As the recipient K-12 was lactose fermenting and susceptible to azithromycin, it can grow only if it receives the AZM resistance factor(s) from donor. The transconjugants were cultured onto MacConkey agar plates, and their identities were reconfirmed as E. coli using with API 20E. The selected and confirmed transconjugants were then subjected to plasmid analysis and PCR confirmation for mphA gene.
Resistome profiling
Three MDR-Shigella having similar pattern of AMR and conferring middle-range plasmid were subjected to whole genome sequencing using Illumina technologies. Sequence data was processed and assembled using the bio-informatics techniques described previously57. Resistome profiling was performed using the AMRFinderPlus v3.10.565 and PlasmidFinder v2.1 (https://cge.cbs.dtu.dk/services/PlasmidFinder/) and MOB-suite v3.1.0 was used to identify plasmids and segregate plasmid-borne AMR genes66,67,68.
Pulsed-field gel electrophoresis (PFGE)
To observe the clonal relationship between azithromycin resistant and sensitive Shigella strains, a total of 11 Shigella strains (7 S. sonnei, 2 S. boydii, 2 S. flexneri type 6) were compared using PFGE typing. Genomic DNA of Shigella strains was embedded in intact agarose gel and digested for 4 h at 37 °C with XbaI (New England Biolabs) restriction enzyme according to the PulseNet program69,70. CHEF-MAPPER system apparatus (Bio-Rad Laboratories) was used to separate the restriction fragments under suitable condition (switching time from 5 to 35 s at 6 V cm−1 for 18 h at 14 °C). TIFF image of PFGE were analyzed using BioNumerics version 4.5 (Applied Maths, Kortrijk, Belgium) fingerprinting software. The dendrogram was generated by the UPGMA algorithm with the Dice-predicted similarity value of two PFGE patterns at 1.0% pattern optimization and 1.5% band position tolerance.
Ethical declaration
All experiments were performed in accordance with relevant guidelines and regulations, and all participants gave their written informed consent prior enrollment. The study was reviewed and approved by the Institutional Review Board (IRB) and the Ethical Committee of icddr,b, Dhaka, Bangladesh.
Data availability
All data and analysis results generated during this study are included in this article and its supplementary information files; raw data are available from the corresponding author on reasonable request. Sequence data are available under the accession numbers mentioned in the Table 4.
References
de Paula, C. M. D., Geimba, M. P., do Amaral, P. H. & Tondo, E. C. Antimicrobial resistance and PCR-ribotyping of Shigella responsible for foodborne outbreaks occurred in Southern Brazil. Braz. J. Microbiol. 41, 966–977 (2010).
Kotloff, K. L. et al. Global burden of diarrheal diseases among children in developing countries: Incidence, etiology, and insights from new molecular diagnostic techniques. Vaccine 35, 6783–6789 (2017).
Kotloff, K. L. et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): A prospective, case-control study. The Lancet 382, 209–222 (2013).
Kotloff, K. L. The burden and etiology of diarrheal illness in developing countries. Pediatr. Clin. N. Am. 64, 799–814. https://doi.org/10.1016/j.pcl.2017.03.006 (2017).
Kotloff, K. L. et al. Global burden of Shigella infections: Implications for vaccine development and implementation of control strategies. Bull. World Health Organ. 77, 651–666 (1999).
Ud-Din, A. I. M. S. et al. Changing trends in the prevalence of Shigella species: Emergence of multi-drug resistant Shigella sonnei biotype g in Bangladesh. PLoS ONE 8, e82601 (2013).
Rahman, M. et al. Emergence of extensively drug-resistant Shigella sonnei in Bangladesh. Immunol. Infect. Dis. 5, 1–9 (2017).
Thompson, C. N., Duy, P. T. & Baker, S. The rising dominance of Shigella sonnei: An intercontinental shift in the etiology of bacillary dysentery. PLoS Negl. Trop. Dis. https://doi.org/10.1371/journal.pntd.0003708 (2015).
World Health Organization. Guidelines for the Control of Shigellosis, Including Epidemics due to Shigella Dysenteriae Type 1 437–445 (WHO, 2005). http://apps.who.int/iris/bitstream/10665/43252/1/924159330X.pdf?ua=1&ua=1 (2005).
Weissman, J. B., Schmerler, A., Gangarosa, E. J., Marier, R. L. & Lewis, J. N. Shigellosis in day-care centres. The Lancet 305, 88–90 (1975).
Ranjbar, R. & Farahani, A. Shigella: Antibiotic-resistance mechanisms and new horizons for treatment. Infect. Drug Resist. 12, 3137–3167. https://doi.org/10.2147/IDR.S219755 (2019).
Nüesch-Inderbinen, M. et al. Shigella antimicrobial drug resistance mechanisms, 2004–2014. Emerg. Infect. Dis. 22, 1083 (2016).
Aggarwal, P. et al. Multi drug resistance and Extended Spectrum Beta Lactamases in clinical isolates of Shigella: A study from New Delhi, India. Travel Med. Infect. Dis. 14, 407–413 (2016).
Tribble, D. R. Antibiotic therapy for acute watery diarrhea and dysentery. Mil. Med. 182, 17–25 (2017).
Poramathikul, K. et al. Multidrug-resistant Shigella infections in patients with diarrhea, Cambodia, 2014–2015. Emerg. Infect. Dis. 22, 1640–1643 (2016).
Erdman, S. M., Buckner, E. E. & Hindler, J. F. Options for treating resistant Shigella species infections in children. J. Pediatr. Pharmacol. Ther. 13, 29–43 (2008).
Hossain, M. M., Glass, R. I. & Khan, M. R. Antibiotic use in a rural community in Bangladesh. Int. J. Epidemiol. 11, 402–405 (1982).
Biswas, M. et al. Self medicated antibiotics in Bangladesh: A cross-sectional health survey conducted in the Rajshahi City. BMC Public Health 14, 847 (2014).
Salah, M. et al. Evaluation of Shigella species azithromycin CLSI epidemiological cutoff values and macrolide resistance genes. J. Clin. Microbiol. 57, 10–1128 (2019).
Boumghar-Bourtchai, L. et al. Macrolide-resistant Shigella sonnei. Emerg. Infect. Dis. 14, 1297–1299 (2008).
Murray, K. et al. Increasing antibiotic resistance in Shigella spp. from infected New York city residents, New York, USA. Emerg. Infect. Dis. 23, 332–335 (2017).
Rahman, M. et al. Increasing spectrum in antimicrobial resistance of Shigella isolates in Bangladesh: Resistance to azithromycin and ceftriaxone and decreased susceptibility to ciprofloxacin. J. Health Popul. Nutr. 25, 158–167 (2007).
Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing M100-S27. M100, 27th ed. (2017).
Heiman, K. E., Grass, J. E., Sjölund-Karlsson, M. & Bowen, A. Shigellosis with decreased susceptibility to azithromycin. Pediatr. Infect. Dis. J. 33, 1204–1205. https://doi.org/10.1097/INF.0000000000000397 (2014).
Mook, P. et al. ESBL-producing and macrolide-resistant Shigella sonnei infections among men who have sex with men, England, 2015. Emerg. Infect. Dis. 22, 1948–1952 (2016).
Brown, J. D. et al. Shigella species epidemiology and antimicrobial susceptibility: The implications of emerging azithromycin resistance for guiding treatment, guidelines and breakpoints. J. Antimicrob. Chemother. 72, 3181–3186 (2017).
Leclercq, R. Mechanisms of resistance to macrolides and lincosamides: Nature of the resistance elements and their clinical implications. Clin. Infect. Dis. 34, 482–492 (2002).
Ojo, K. K. et al. The mef(A) gene predominates among seven macrolide resistance genes identified in gram-negative strains representing 13 genera, isolated from healthy Portuguese children. Antimicrob. Agents Chemother. 48, 3451–3456 (2004).
O’Hara, K., Kanda, T., Ohmiya, K., Ebisu, T. & Kono, M. Purification and characterization of macrolide 2’-phosphotransferase from a strain of Escherichia coli that is highly resistant to erythromycin. Antimicrob. Agents Chemother. 33, 1354–1357 (1989).
Arthur, M., Andremont, A. & Courvalin, P. Distribution of erythromycin esterase and rRNA methylase genes in members of the family Enterobacteriaceae highly resistant to erythromycin. Antimicrob. Agents Chemother. 31, 404–409 (1987).
Gaudreau, C. et al. Shigella spp with reduced azithromycin susceptibility, Quebec, Canada, 2012–2013. Emerg. Infect. Dis. 20, 854–856 (2014).
Liao, Y. S., Liu, Y. Y., Lo, Y. C. & Chiou, C. S. Azithromycin-nonsusceptible Shigella flexneri 3a in men who have sex with men, Taiwan, 2015–2016. Emerg. Infect. Dis. 23, 345–346. https://doi.org/10.3201/eid2302.161260 (2017).
Ke, X., Gu, B., Pan, S. & Tong, M. Epidemiology and molecular mechanism of integron-mediated antibiotic resistance in Shigella. Arch. Microbiol. 193, 767–774. https://doi.org/10.1007/s00203-011-0744-3 (2011).
Partridge, S. R. Analysis of antibiotic resistance regions in Gram-negative bacteria. FEMS Microbiol. Rev. 35, 820–855. https://doi.org/10.1111/j.1574-6976.2011.00277.x (2011).
Baker, K. S. et al. Intercontinental dissemination of azithromycin-resistant shigellosis through sexual transmission: A cross-sectional study. Lancet Infect. Dis. 15, 913–921 (2015).
Mannion, A. J. et al. Plasmid-mediated quinolone resistance in Shigella flexneri isolated from macaques. Front. Microbiol. 9, 311 (2018).
Rashid, H. & Rahman, M. Possible transfer of plasmid mediated third generation cephalosporin resistance between Escherichia coli and Shigella sonnei in the human gut. Infect. Genet. Evol. 30, 15–18 (2015).
Talukder, K. A. et al. Temporal shifts in the dominance of serotypes of Shigella dysenteriae from 1999 to 2002 in Dhaka, Bangladesh. J. Clin. Microbiol. 41, 5053–5058 (2003).
Okeke, I. N. et al. Antimicrobial resistance in developing countries. Part I: Recent trends and current status. Lancet Infect. Dis. 5, 481–493. https://doi.org/10.1016/S1473-3099(05)70189-4 (2005).
Bhattacharya, D. et al. Changing patterns and widening of antibiotic resistance in Shigella spp. over a decade (2000–2011) Andaman Islands, India. Epidemiol. Infect. 143, 470–477 (2015).
Williams, P. C. M. & Berkley, J. A. Guidelines for the treatment of dysentery (shigellosis): A systematic review of the evidence. Pediatric. Int. Child Health 38, S50–S65. https://doi.org/10.1080/20469047.2017.1409454 (2018).
Christopher, P. R., David, K. V., John, S. M. & Sankarapandian, V. Antibiotic therapy for Shigella dysentery. Cochrane Database Syst. Rev. https://doi.org/10.1002/14651858.CD006784.PUB4/ABSTRACT (2010).
Gruninger, R. J. et al. Socioeconomic determinants of cipro-floxacin-resistant Shigella infections in Bangladeshi children. Pathog. Immun. 2, 89–101 (2017).
Nuzhat, S. et al. Antimicrobial resistance in shigellosis: A surveillance study among urban and rural children over 20 years in Bangladesh. PLoS ONE 17, e0277574 (2022).
Nguyen, M. C. P. et al. Escherichia coli as reservoir for macrolide resistance genes. Emerg. Infect. Dis. 15, 1648–1650 (2009).
Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing M100-S26. M100, 26th ed. (2016).
European Committee on Antimicrobial Susceptibility Testing. Breakpoint Tables for Interpretation of MICs and Zone Diameters Version 9.0, 2019. (2019).
Darton, T. C. et al. Azithromycin resistance in Shigella spp. in Southeast Asia. Antimicrob. Agents Chemother. 62, 10–1128 (2018).
Moreno-Mingorance, A. et al. Circulation of multi-drug-resistant Shigella sonnei and Shigella flexneri among men who have sex with men in Barcelona, Spain, 2015–2019. Int. J. Antimicrob. Agents 58, 106378 (2021).
Stenhouse, G. E. et al. The genomic epidemiology of shigellosis in South Africa. Nat. Commun. 14, 7715 (2023).
Mitchell, H. D. et al. Linkage of whole genome sequencing, epidemiological, and clinical data to understand the genetic diversity and clinical outcomes of Shigella flexneri among men who have sex with men in England. Microbiol. Spectr. 9, e01213 (2021).
Baker, K. S. et al. Horizontal antimicrobial resistance transfer drives epidemics of multiple Shigella species. Nat. Commun. 9, 949 (2018).
Nusrin, S. et al. Multiple mechanisms confer resistance to azithromycin in Shigella in Bangladesh: A comprehensive whole genome-based approach. Microbiol. Spectr. 10, e0074122 (2022).
Darphorn, T. S., Bel, K., Koenders-Van Sint Anneland, B. B., Brul, S. & Ter Kuile, B. H. Antibiotic resistance plasmid composition and architecture in Escherichia coli isolates from meat. Sci. Rep. 11, 2136 (2021).
World Health Organization. Programme for control of diarrhoeal disease. In Manual for Laboratory Investigation of Acute Enteric Infections Infections 9–20 CDD/93.3 (World Health Organization, 1987).
Talukder, K. A. et al. Altering trends in the dominance of Shigella flexneri serotypes and emergence of serologically atypical S. flexneri strains in Dhaka, Bangladesh. J. Clin. Microbiol. 39, 3757–3759 (2001).
Asad, A. et al. Draft genome sequences of multidrug-resistant shigella strains isolated from diarrheal patients in Bangladesh. Microbiol. Resour. Announc. 10, 10–1128 (2021).
Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Disk Susceptibility Tests-Tenth Edition M02-A10 vol. 29 (2009).
Heiman, K. E. et al. Shigella with decreased susceptibility to azithromycin among men who have sex with men - united states, 2002–2013. Morb. Mortal. Wkly. Rep. 63, 132–133 (2014).
Kado, C. I. & Liu, S. T. Rapid procedure for detection and isolation of large and small plasmids. J. Bacteriol. 145, 1365–1373 (1981).
Talukder, K. A. et al. Phenotypic and genotypic characterization of serologically atypical strains of Shigella flexneri type 4 isolated in Dhaka, Bangladesh. J. Clin. Microbiol. 40, 2490–2497 (2002).
Haider, K., Huq, M. I., Talukder, K. A. & Ahmad, Q. S. Electropherotyping of plasmid DNA of different serotypes of Shigella flexneri isolated in Bangladesh. Epidemiol. Infect. 102, 421–428 (1989).
Islam, D. & Lindberg, A. A. Detection of Shigella dysenteriae type 1 and Shigella flexneri in feces by immunomagnetic isolation and polymerase chain reaction. J. Clin. Microbiol. 30, 2801 (1992).
Munshi, M. H. et al. Plasmid-mediated resistance to nalidixic acid in shigella dysenteriae type 1. The Lancet 330, 419–421 (1987).
Feldgarden, M. et al. Validating the AMRFINder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrob. Agents Chemother. 63, 10–1128 (2019).
Carattoli, A. et al. In Silico detection and typing of plasmids using plasmidfinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 58, 3895–3903 (2014).
Robertson, J. & Nash, J. H. E. MOB-suite: software tools for clustering, reconstruction and typing of plasmids from draft assemblies. Microb. Genom. 4, e000206 (2018).
Robertson, J., Bessonov, K., Schonfeld, J. & Nash, J. H. E. Universal whole-sequence-based plasmid typing and its utility to prediction of host range and epidemiological surveillance. Microb. Genom. 6, 1–12 (2020).
Okada, N. et al. Construction of a physical map of the chromosome of Shigella flexneri 2a and the direct assignment of nine virulence-associated loci identified by Tn5 insertions. Mol. Microbiol. 5, 2171–2180 (1991).
Talukder, K. A., Dutta, D. K. & Albert, M. J. Evaluation of pulsed-field gel electrophoresis for typing of Shigella dysenteriae type 1. J. Med. Microbiol. 48, 781–784 (1999).
Acknowledgements
This research activity was funded by the icddr,b, Dhaka, Bangladesh. ZI has received grant support from the Fogarty International Center, National Institute of Neurological Disorders and Stroke of the National Institutes of Health, USA, under Award Number K43TW011447. SH has received grant support from “Global Health Equity Scholars NIH FIC TW010540”, USA. icddr,b acknowledges with gratitude the commitment of the Government of Bangladesh to its research efforts, and also gratefully acknowledges all donors who provide unrestricted support, including the icddr,b is grateful to the governments of Bangladesh and Canada for providing core/unrestricted support. The authors are indebted to the neurologists who encouraged their patients to take part in this study.
Author information
Authors and Affiliations
Contributions
Z.I. and A.A. conceptualized the study. Research methodology and execution plan was theorized by A.A. Material acquisition, technique optimization, laboratory experiments, and data acquisition were performed by A.A., M.A.M., R.B., M.A.M. (2), K.S., S.N.F., I.J., and S.H. Data presentation and interpretations were accomplished by A.A. which was further checked by I.J. A.A. drafted the primary manuscript which was exclusively scrutinized by Z.I., I.J. and S.H. Proofreading and reviews were done by all other authors. The final manuscript was read and approved by all authors before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Asad, A., Jahan, I., Munni, M.A. et al. Multidrug-resistant conjugative plasmid carrying mphA confers increased antimicrobial resistance in Shigella. Sci Rep 14, 6947 (2024). https://doi.org/10.1038/s41598-024-57423-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-57423-1
- Springer Nature Limited