Abstract
Despite the current use of some Bacillus spp. as probiotics, looking for and introducing new efficient and safe potential probiotic strains is one of the most important topics in both microbiology and food industry. This study aimed to isolate, identify, and evaluate the probiotic characteristics and safety of some Bacillus spp. from natural sources. Thirty-six spore-forming, Gram-positive, and catalase-positive Bacillus isolates were identified in 54 samples of soil, feces and dairy products. Bacterial identification was performed using 16S rDNA sequencing. To evaluate the probiotic potential of isolates, the resistance of bacterial cells to simulated gastrointestinal tract (GIT) conditions, the presence of enterotoxin genes, their susceptibility to antibiotics, antimicrobial and hemolytic activities and biochemical profiles were investigated. The results revealed that eight sporulating Bacillus spp. isolates fulfilled all tested probiotic criteria. They showed a high growth rate, non-hemolytic and lecithinase activity, and resistance to simulated GIT conditions. These strains exhibited broad-spectrum antibacterial activity against pathogenic bacteria. In addition, they did not exhibit antibacterial resistance to the 12 tested antibiotics. The results of this study suggest that these isolates can be considered as candidates for functional foods and as safe additives to improve diet quality.
Similar content being viewed by others
Introduction
Probiotics are considered to be living and non-pathogenic microorganisms that can enhance the intestinal microbial balance in favor of beneficial microorganisms colonizing in the human intestine. They are used as functional foods and dietary supplements, alternatives to antibiotics in animal farming, growth promoters in feed supplements, and human and aquaculture prophylactics1,2. Previous studies have shown that dietary supplementation with Bacillus-based probiotics could be successfully used in poultry diets with growth-promoting benefits3. Bacillus-based probiotic products are also available for human consumption.
Bacterial probiotics mainly belong to lactic acid bacteria (LAB) and Bifidobacteria4,5. Therefore, Bacillus species have received limited attention compared with LAB and Bifidobacteria. Nevertheless, Enterogermina® (Sanofi-Aventis SpA), a probiotic product containing B. clausii spores was registered in 1958 as an OTC medicinal supplement in Italy. Some strains of rod-shaped, Gram-positive, catalase-positive and spore-forming Bacilli including B. laterosporus, B. subtilis, B. clausii, B. licheniformis, B. cereus, B. coagulans and B. pumilus have been shown to possess probiotic characteristics6,7.
Growth and sporulation rate, the resistance of spores to gastrointestinal tract (GIT) condition, broad antimicrobial activities against pathogenic bacteria, lack of antibiotic resistance, lack of positive hemolytic and lecithinase activity, and lack of enterotoxin genes involved in gastrointestinal disorders are the most important criteria that a microorganism must possess to be considered as a probiotic strain. Bacillus species offer several advantages over lactobacilli and Bifidobacteria as probiotics. Since these bacteria are spore formers, they can survive for a long time at room temperature in a desiccated form. In addition, their endospore structure allows them to withstand harsh conditions in the GIT. They can survive in extreme or stressful environmental conditions such as low pH, high temperatures, high bile salts, nutrient starvation and desiccation8. Furthermore, Bacilli exhibit high metabolic activity mainly associated with the synthesis of a vast array of antimicrobial compounds. For instance, approximately 4–5% of the B. subtilis genome has been devoted to the synthesis of antibiotics, enabling these bacteria to produce more than two dozen antibiotics9. Moreover, several Bacillus species are remarkably important in the industry due to their application in the industrial production of extracellular enzymes such as amylase and protease10,11.
Given the interesting features of Bacillus species as promising probiotic candidates and the limited presence of Bacillus probiotics in the market, this study sought to isolate favorite Bacillus strains from various sources including soil, feces and artisanal dairy products. Following isolation, these strains were identified, and their probiotic properties were systematically investigated and compared with those of some established commercial Bacillus probiotics.
Results
Identification of probiotic isolates
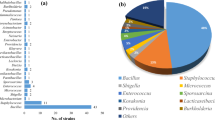
From 54 samples of dairy products, soil, livestock and poultry wastes, in total 36 spore-forming, Gram-positive and catalase-positive strains were isolated (Table 1). The PCR products of the 16S rRNA genes were analyzed using gel electrophoresis, revealing a prominent 1500 bp band. The alignment results for the most of isolates exhibited a high similarity of 98–100% in the 16S rDNA of the Bacilli. The accession numbers of the registered sequences are represented in Table 1. However, the isolates identified as B. cereus, which possess potential pathogenicity, and two strains with limited growth capability were excluded from further testing to ensure the safety and focus of the subsequent investigations.
Hemolytic and lecithinase activity of isolates
As demonstrated in Table 2, it was found that 18 strains out of 28 strains were gamma hemolytic (non-hemolytic), five strains were alpha-hemolytic, and five strains showed beta-hemolytic activity. Also, the lecithinase test results were negative for all strains except one isolate (PR) (Table 2).
Probiotic characteristics
Resistance of the strains to simulated GIT conditions
Survival rates of the 17 isolates were evaluated under simulated gastric fluid (SGF) and simulated intestinal fluid (SIF) conditions. Based on the results, the viability of 13 strains was above 90% under GIT conditions and only 4 strains showed reduced viability of more than 1 log (Table 3).
Antibiotic susceptibility
Antibiotic resistance of the strains was evaluated according to the European food safety authority (EFSA) and National Committee for Clinical Laboratory Standards (NCCLS). Due to the lack of a reliable interpretation document for the disc diffusion method for Bacillus spp., the inhibition zones lower than 12 mm were considered resistant to antibiotics as suggested by Hoa et al.12. Table 4 shows the results of the disc diffusion method, indicating that G2a, SHE, GHE, and GUa isolates demonstrated resistance to streptomycin. Fortunately, the resistance gene to this antibiotic is not transferable13. While no inhibition zone was observed for the Gala isolate in relation to chloramphenicol, the microdilution assay confirmed its susceptibility to this antibiotic. Also, the Gua isolate was resistant to erythromycin, clindamycin, chloramphenicol as well as ampicillin based on the microdilution method.
Antimicrobial activity
The ability to inhibit the growth of pathogens is a favorable characteristic of probiotic bacteria. Based on the well-diffusion method, the diameters of the growth inhibition zones of the pathogens treated with the most isolated strains were more than 10 mm (Table 5). The GE1 strain inhibited a broad range of pathogenic bacteria, and the SHE strain displayed the greatest effect on MRSA with a 23 mm diameter inhibition zone. However, none of the isolates exhibited antibacterial activity against the ESBL E. coli.
Enterotoxins and other potential virulence factors
Nhe, hbl and bceT genes were detected using PCR to distinguish virulent strains (Table 6). To evaluate whether these enterotoxin genes are essentially involved in hemolytic activity, ME2a, MA2b, MA1c and Khg2, as β-hemolytic isolates, were investigated for the presence of enterotoxin genes by PCR test. The results showed that ME2a and MA1c possess all enterotoxin genes and were considered as the positive controls for the PCR test of other samples. In four of the γ-hemolytic isolates, namely, GA3a, GHE, SHE and Gua, the presence of enterotoxin genes (bceT, hbl and nhe) was identified.
Biochemical tests
Various carbohydrates were tested to be used as sole carbon sources by the isolated strains and the results are presented in Table 7. It was shown that all strains produced acid from xylose, trehalose, and glucose. None of the strains could ferment lactose, sorbitol, melibiose and raffinose. Only GB isolate fermented rhamnose. Mannitol and fructose were fermented by three and five of the isolates, respectively.
Table 8 presents the results of biochemical tests and the growth characteristics of the strains under harsh conditions. It can be concluded that most isolates were able to survive at high temperatures (55 °C). Notably, the G2a isolate grew even at 65 °C. However, both SHE and Ma1b isolates were unable to withstand 6.5% NaCl. As for resistance to low pH environments, the PUN, G2a, and MA1d isolates were able to survive at pH values ranging from 4.0 to 5.0.
Discussion
Isolation of Bacillus strains
Bacillus species are ubiquitous in nature and can be isolated from a variety of ecological habitats. In this study, several Bacillus strains were isolated from diverse sources, including soil, feces, and artisanal dairy products. While soil serves as a prominent environmental niche for numerous Bacillus species, Bacillus strains displaying promising probiotic properties can also be found in animal feces14 and artisanal dairy products15. Some of isolated Bacillus strains such as B. amyloliquefaciens16 and B. siamensis17, have previously been isolated and identified in similar researches. Sen et al. isolated and identified B. coagulans strains with probiotic potential from soil samples containing dry animal waste. In our investigation, four strains of B. cereus were isolated from dairy samples, which aligns with findings of Montanhini et al. who also isolated 23 B. cereus strains from similar samples18. Eight B. pumilus strains were isolated from livestock and poultry wastes. Guo et al. isolated this strain with probiotic properties from the fresh waste of Tibetan sheep19. Also, five B. amyloliquefaciens strains were isolated from camel and chicken feces and dairy products, while this strain was recently isolated from soil, fruits, and cereals20,21,22. In a recent study, Daneshazari et al. isolated Bacillus strains from camel milk with probiotic potentials23 further contributing to the growing body of knowledge on beneficial Bacillus strains.Furthermore, two strains of B. safensis were isolated from soil and poultry waste samples. Satomi et al. have also isolated 30 strains of Bacillus spp. from spacecraft and assembly-facility surfaces from which 13 strains were identified as B. safensis24. Raja and Omine isolated B. safensis MS11 strain from forest soil samples of Mongolia25.
Additionally, one B. toyonensis strain was identified from poultry waste samples. There are few reports on the isolation of this bacterium. However, recently Okaiyeto et al. have purified B. toyonensis from South African sea sediments26. In Japan, this strain is used in a probiotic product named Toyocerin27.
Probiotic characterization of isolates
Hemolytic activity of isolates
Non-hemolytic (ɤ-hemolytic) strains are generally considered safe for their hosts, while strains with hemolytic activity are considered pathogen. Microorganisms exhibiting hemolytic activity can disrupt red blood cells, leading to the release of hemoglobin. This activity is often associated with the production of hemolysins, which can have cytolytic effects on host cells, and decreasing available hemoglobin content as a Fe source28,29. On the other hand, beta hemolytic activity indicates the presence of cytotoxic phospholipase in microorganisms6.
Investigation of the hemolytic activity of 48 Bacillus strains isolated from 50 probiotic products collected from South Korea, Australia, and China revealed that 58% of the isolates displayed hemolytic activity30. Therefore, strains displaying hemolytic activities were excluded from further assessment, as they may pose a risk to the host's health.
Resistance to simulated GIT conditions
The ability to tolerate high acid concentrations in the stomach is a relevant criterion for selecting an appropriate probiotic product31. Some isolated strains demonstrated remarkable resistance to the GIT conditions. Interestingly, in certain cases, these isolated strains exhibited even better viability than commercially available strains like Subtyl32, which indicates their potential as robust candidates for probiotic applications. This increased tolerance to acidic environments enhances their chances of survival and exerting their beneficial effects in the GIT, making them promising candidates for further probiotic investigations.
The microaerophilic conditions prevailing in the digestive tract highlights the importance of probiotic strains' ability to thrive in such environments. In our study, we observed that after 24 h of incubation in microaerophilic conditions, all samples exhibited robust growth. Since gastrointestinal juice imposes stressful conditions against bacteria, tolerance to gastrointestinal juice is a critical property sought in probiotic bacteria33. Sharma et al. suggested that an appropriate probiotic strain should withstand at least pH 3.034, while Hong et al. reported that Bacillus strains exhibit different resistances to gastric acid and not all are resistant to these conditions35. The disparity in resistance might be attributed to the activation of spores exposed to acids and the subsequent destruction of regenerative cells in the acidic environment of the stomach13,32. Understanding these mechanisms is vital for identifying probiotic strains with the highest likelihood of surviving in the GIT.
Probiotic bacteria must possess the property to pass through the small intestine successfully36. However, bile salt and pancreatic enzymes of the small intestine can be detrimental to bacterial cells37. The isolated spore-forming Bacillus spp. showed proper resistance against pepsin and pancreatin, two of the gastrointestinal enzymes. These results suggest that many of the strains could reach the colon without significantly being affected by gastrointestinal conditions. In a recent study, it was shown that the B. coagulans population decreased only by 0.64 log under semi-intestinal conditions. It could be concluded that using Bacillus spp. for probiotic consumption offers functional benefits because of their resistance to food processing temperature, storage, and GIT environment38. Their ability to withstand these challenges suggests their potential as robust probiotic candidates with a high likelihood of providing health benefits to the host.
Antibiotic resistance of isolates
Although no concerning antibiotic resistance was observed regarding the Bacillus spp. isolated in this study, the results of some previous studies already conducted on commercial probiotic products, such as HU36, Natto and PY79, showed the resistance of HU36 strain to clindamycin. The PY79 strain was also resistant to streptomycin and tetracycline. Likewise, the Natto strain showed resistance to streptomycin35. Furthermore, a study on commercial probiotic strains, including Enterogermina and Biosubtyl revealed Enterogermina resistance to penicillin, erythromycin, lincomycin, rifampin, neomycin, and chloramphenicol antibiotics39.
Antibacterial activity of isolates
Antibiotics are among the most important pharmaceuticals, having saved millions of lives since their introduction to the market. However, the excessive use of antibiotics has raised concerns about the development of antibiotic-resistant pathogens and reduced efficacy, posing a significant threat to human life. While ongoing efforts to discover novel antibiotics persist, the consumption of probiotics harboring antimicrobial activity could serve as a potential mitigation strategy for this problem.
As shown in Table 5, several isolates displayed antimicrobial activity. Notably, the GE1 isolate exhibited a broad-spectrum antimicrobial effect against 9 out of the 12 of the examined antibiotics, including MRSA, thereby suggesting this isolate a promising probiotic with potent antimicrobial properties.
The antibacterial activity of the isolates might be due to the production of bacteriocin-like agents and also peptide and non-peptide antibiotics released by bacterial cells40. In recent studies, it was shown that Bacillus strains produce a broad range of antibacterial agents including peptide and lipopeptide compounds against Gram-positive and Gram-negative pathogens41,42,43. Also, the presence of secondary metabolites with antimicrobial potentials, such as bacteriocin, hydrogen peroxide, lactic acid and propionic acid, in bacterial culture supernatant has previously been reported44. Together with the competitive exclusion mechanism, where probiotics compete with harmful bacteria for adhesive receptors and nutrients, these metabolites can effectively destroy and prevent the colonization of pathogens in the body45.
Presence of enterotoxin encoding genes
Although a few numbers of Bacillus spp. have been shown to produce toxins, some Bacillus spp. including B. endophyticus, B. subtilis, B. amyloliquefaciens, B. pumilus and B. licheniformis have been approved as generally recognized as safe (GRAS) probiotic bacteria43,46. Moreover, endospores of certain Bacillus spp. have been studied as Direct Feed Microbials (DFM), and the findings have confirmed their safety and reliability as prophylactic agents in reducing GI disease in livestock and humans47,48. The results of the current study are in good agreement with some similar reports43,46,49,50.
Evaluation of DNA showed that the strains carrying the genes coding for two enterotoxins hemolysin (Hbl) and (Nhe) are non-hemolytic enterotoxins51,52. While conclusion about protein function or malfunction solely based on the results of DNA research is not sufficient, this hypothesis still exists that genes encoding for HH and Nhe enterotoxins may not function in some Bacillus strains53. Results of a study on the possible pathogenicity of some commercial probiotic strains including Subtyl, Bactisubtil and BiosubtylDL revealed hemolytic and lecithinase activity in these products and the presence of hbl and nhe enterotoxin genes in these strains was confirmed32.
Physiological characterization of isolates
Some isolates of the present study showed diverse sugar fermentation profiles based on Bergey’s manual of systematic bacteriology book54. One of the main reasons for the discrepancy between the biochemical tests can be attributed to the loss or gain of transferable genes. Since some genes needed for the fermentation of sugars are encoded by plasmids, variation in plasmid content may result in metabolic inconsistencies55.
As expected, all the isolates were able to ferment glucose as the sole carbon source. However, none of the isolates were able to utilize lactose (Table 7). Except for PUN, other strains in Table 7 were isolated from soil or animal feces. These environments typically do not provide a consistent source of lactose, and Bacillus species from these habitats may not have evolved mechanisms for efficient lactose utilization.
Conclusion
In this study, we isolated 36 Bacillus species from soil, animal feces, and artisanal dairy products samples collected from various locations in Iran. After conducting a comprehensive assessment of the probiotic properties of the isolates based on available studies, we identified eight Bacillus strains that met the probiotic criteria. These strains included G2a (B. subtilis strain esf-G2; KX276177), MA1d (B. amyloliquefaciens strain ard-MA1D; KX276185), Khk (B. coagulanse strain NBRC-12583; KX261624), GB (B. endophyticus strain 2DT; KX261623), G1 (B. pumilus strain esf G1; KX270718), Ga1a (B. pumilus strain NBRC-12092; KX261622), Ma1b (B. licheniformis strain ard MA1B; KX270723) and PUN (B. siamensis strain ard-PUN; KX276176). The results of this study suggest that these isolates, especially G2a, Ga1a, G1and Khk, exhibit significant potential for utilization in several sectors, including animal feed additives, promoting gut health, food preservatives, and antibiotic alternatives. However, further in vivo analyses are required to unlock the full spectrum of potential applications for these isolates. Comprehensive in vivo assessments will not only validate the findings but also enhance and broaden the potential applications of these isolates across diverse fields. Moreover, considering the prevalence of probiotic products in the market primarily consisting of LAB and Bifidobacteria, future investigation should focus on exploring the synergistic or antagonistic interactions of our isolated Bacillus strains with these well-established probiotic groups.
Materials and methods
Sample collection and isolation of strains
A total of 54 samples, including soil, animal feces, and artisanal dairy products were taken from Isfahan and Ardabil provinces in Iran. In summary, the collection comprised 5 soil samples (3 from agricultural setting and 2 from greenhouses), 18 samples from animal feces (6 from chickens, 5 from cows, 4 from sheep, 1 from turkeys, 1 from gooses, and 1from camels), and 31 samples from a diverse array of artisanal dairy products. The samples were placed in sterile vials, promptly chilled on ice, and transported to the laboratory.
One gram of each solid or 1 mL of each liquid sample was added to 10 mL of sodium citrate 2% (w/v) and shaken for 4 h. The samples were then placed in a water bath for 15 min at 80 °C to kill vegetative bacteria. Next, 1 mL of the treated samples was inoculated into 10 mL of MRS broth culture medium and incubated with shaking at 37 °C for 24 h. 100 μL of the grown cells were plated on MRS agar and incubated at 37 °C. Colonies with different morphologies were selected and subcultured. Bacillus colonies were screened by Gram staining, spore staining using Schaeffer-Fulton method56, and catalase test57. Finally, the pure Gram-positive, catalase-positive, and spore-forming isolates were stored as glycerol stocks at − 20 and − 80 °C until further use.
Hemolytic and lecithinase activity
Hemolytic activity of the isolates was assayed by culturing the strains on the blood agar medium, followed by the incubation at 30 °C for 24 to 48 hours58. In addition, isolates were cultured on the egg yolk agar and incubated at 37 °C for 24 h to assess their lecithinase (phospholipolytic) activity59.
Molecular identification of the isolate
Molecular identification of the isolates was done by PCR amplification of the corresponding 16s rRNA genes followed by their sequencing. Briefly, the genomic DNA of the pure isolates was extracted using genomic DNA extraction kits (Tali Gene Pars, Iran), and PCR was done using 27F (5′-AGAGTTTGATCCTGGCTCAC-3′) and 1492R (5’-CGGTTACCTTGTTACGACTT-3′) primers as forward and reverse universal 16S rDNA primers, respectively60. After gel purification, the PCR products were sequenced (Gene Fanavaran, Tehran, Iran) and analyzed by the CLC Main Workbench and Bio Edit software61. The National Center for Biotechnology Information (NCBI) database and the BLAST search tool were used to identify the genus and species of the bacteria.
Evaluation of the probiotic characteristics of the isolates
Resistance of spores and vegetative cells to simulated GIT conditions
The acidic liquid of the stomach was simulated by adding 1 mg/mL pepsin to the sterile normal saline (0.85%) at pH 2. The bacterial suspension (0.5 McFarland) was then exposed to this solution and incubated at 37 °C. After 3, 30 and 60 min of incubation, 100 µL of each treated spore solution was inoculated into Tryptic Soy Agar (TSA) culture medium and incubated at 37 °C for 24 hours32.
The intestine condition was simulated by adding 1 mg/mL pancreatin and 0.2% of bile salts into the sterile isotonic buffer (pH 7.4), followed by exposing the bacterial cells to this condition. After 3, 60, 120 and 180 min, the resistance of the bacterial cells to these conditions was evaluated.
Antibiotic resistance of the isolates
To assess the antibiotic resistance of the isolates, the disc diffusion method (DDM) and Micro dilution methods were applied62. The sensitivities of the bacteria to antibiotics were interpreted as sensitive (S), intermediate (I), and resistant (R) based on the Clinical & Laboratory Standards Institute (CLSI) guidelines63. The Antibiotic discs used were chloramphenicol (30 µg/disc), gentamicin (10 µg/disc), erythromycin (15 µg/disc), clindamycin (2 µg/disc), kanamycin (30 µg/disc), vancomycin (30 µg/disc), penicillin (10 IU/disc), streptomycin (10 µg/disc), tetracycline (30 µg/disc) and ampicillin (10 µg/disc) that were purchased from MAST Group (Merseyside, UK). To evaluate the minimum inhibitory concentrations (MICs) of these ten antibiotics, different amounts of each antibiotic were diluted in broth medium to prepare the following concentrations: 0.5 to 256 μg/mL (chloramphenicol), 0.125 to 64 μg/mL (clindamycin, erythromycin, and ampicillin) and 0.25 to 128 μg/mL (vancomycin, tetracycline). The strains treated with these antibiotics were incubated at 35 °C for 24 h. The minimum bactericidal concentrations (MBCs) were determined as described by CLSI63.
Antimicrobial activities of the isolated strains
Antimicrobial activities of the isolated strains were assessed by spotting and agar well diffusion method against several pathogens including Salmonella typhimurium PTCC1609 (Persian Type Culture Collection), Escherichia coli ATCC11775, Listeria monocytogenes ATCC13932, Shigella sonnei ATCC9290, Staphylococcus aureus ATCC33591, Klebsiella pneumoniae, Enterococcus faecium, Acinetobacter baumannii, Streptococcus pyogenes, MRSA, VRE, and Extended-Spectrum Beta-Lactamase (ESBL)-producing E. coli provided by Alzahra hospital (Isfahan, Iran). To assess the antimicrobial activity through the agar well diffusion method, 106 CFU/mL of the pathogenic bacteria suspensions were prepared and cultured on the Mueller–Hinton Agar (MHA). Wells with 0.5 cm diameter were made on MHA plates using a sterile Pasteur pipette. Two milliliters of Bacillus isolate medium was centrifuged at 9000×g, and 4 °C for 10 min5 and the supernatant was filtered. Next, 60 µL of the cell-free supernatant of each bacterial culture was poured into the wells and the plates were incubated at 37 °C. The diameter of the growth inhibition zoon appeared around the wells was recorded after 24 h of incubation64,65,66.
Presence of genes encoding virulence factors
The presence of enterotoxin genes, which are responsible for pathogenicity and food toxicity in Bacillus spp. was investigated. The presence of common enterotoxins coding genes in Bacillus bacteria including nhe, hbl, cytk, and bceT. nhe, hbl, and bceT was examined by multiplex PCR method using the corresponding primers for each gene as described in the literature67,68,69. A positive control and a negative control were included in the experiment.
Growth in microaerophilic condition
The ability of the isolates to grow in microaerophilic conditions was investigated by culturing on the Luria Bertani (LB) agar culture medium followed by incubating anaerobically at 37 °C for 24 hours70.
Biochemical tests
The isolates were also characterized with several biochemical tests including the starch hydrolysis, Voges Proskauer test, citrate test, carbohydrate fermentation and production of acid test, NaCl tolerance test, growth at high temperatures, and acid resistance test as described by Vos et al.54.
Statistical analysis
Data were statistically analyzed with a significance level set at p < 0.05 using GraphPad Prism version 9.5.1.733 and one-way analysis of variance (ANOVA) was performed to identify significant differences between the means.
Data availability
The datasets generated and/or analysed during the current study are available in the GenBank repository, under the accession numbers stated in Table 1.
References
Nithya, V. & Halami, P. M. Evaluation of the probiotic characteristics of Bacillus species isolated from different food sources. Ann. Microbiol. 63, 129–137 (2013).
Rowland, I. Probiotics and benefits to human health—The evidence in favour. Environ. Microbiol. 1, 375–376 (1999).
Ciurescu, G., Dumitru, M., Gheorghe, A., Untea, A. E. & Drăghici, R. Effect of Bacillus subtilis on growth performance, bone mineralization, and bacterial population of broilers fed with different protein sources. Poult. Sci. 99, 5960–5971 (2020).
Zangeneh, M., Khorrami, S. & Khaleghi, M. Bacteriostatic activity and partial characterization of the bacteriocin produced by L. plantarum sp. isolated from traditional sourdough. Food Sci. Nutr. https://doi.org/10.1002/fsn3.1890 (2020).
Jafari-Nasab, T., Khaleghi, M., Farsinejad, A. & Khorrami, S. Probiotic potential and anticancer properties of Pediococcus sp. isolated from traditional dairy products. Biotechnol. Rep. 29, e00593 (2021).
Sorokulova, I. B. et al. The safety of two Bacillus probiotic strains for human use. Dig. Dis. Sci. 53, 954–963 (2008).
Urdaci, M. C. & Pinchuk, I. Antimicrobial activity of Bacillus probiotics. Bact. Spore Formers Probiotics Emerg. Appl. Norfolk UK Horiz. Biosci. 17, 171–182 (2004).
Al-Thubiani, A. S. A. et al. Identification and characterization of a novel antimicrobial peptide compound produced by Bacillus megaterium strain isolated from oral microflora. Saudi Pharm. J. 26, 1089–1097 (2018).
Stein, T. Bacillus subtilis antibiotics: Structures, syntheses and specific functions. Mol. Microbiol. 56, 845–857 (2005).
Rai, A. K. et al. Production of bioactive hydrolysate using protease, β-glucosidase and α-amylase of Bacillus spp. isolated from kinema. Bioresour. Technol. 235, 358–365 (2017).
Cai, D., Rao, Y., Zhan, Y., Wang, Q. & Chen, S. Engineering Bacillus for efficient production of heterologous protein: Current progress, challenge and prospect. J. Appl. Microbiol. 126, 1632–1642 (2019).
Hoa, N. T. et al. Characterization of Bacillus species used for oral bacteriotherapy and bacterioprophylaxis of gastrointestinal disorders. Appl. Environ. Microbiol. 66, 5241–5247 (2000).
Hong, H. A., Duc, L. H. & Cutting, S. M. The use of bacterial spore formers as probiotics. FEMS Microbiol. Rev. 29, 813–835 (2005).
Gu, S.-B. et al. Potential probiotic attributes of a new strain of Bacillus coagulans CGMCC 9951 isolated from healthy piglet feces. World J. Microbiol. Biotechnol. 31, 851–863 (2015).
Khan, M. N., Bashir, S. & Imran, M. Probiotic characterization of Bacillus species strains isolated from an artisanal fermented milk product Dahi. Folia Microbiol. (Praha) 68, 1–13 (2023).
Niazi, A., Manzoor, S., Bejai, S., Meijer, J. & Bongcam-rudloff, E. Complete genome sequence of a plant associated bacterium Bacillus amyloliquefaciens subsp. plantarum UCMB5033. Stand. Genomic Sci. 9, 718–725 (2014).
Sumpavapol, P. et al. Bacillus siamensis sp. nov., isolated from salted crab (poo-khem) in Thailand. Int. J. Syst. Evol. Microbiol. 60, 2364–2370 (2010).
Montanhini, M. T. M., Montanhini Neto, R. & Bersot, L. S. Enterotoxigenic potential of Bacillus cereus strains isolated from dairy products at different incubation temperatures. Int. Food Res. J. 22(3), 1315–1317 (2015).
Guo, Z., Zhou, B. & Yan, H. Study on the isolation, identification and biological characteristics of a strain of Bacillus pumilus isolated from Tibetan Sheep. China Anim. Husb. Vet. Med., 6, 1610–1617 (2016).
Chen, X., Zhang, Y., Fu, X., Li, Y. & Wang, Q. Isolation and characterization of Bacillus amyloliquefaciens PG12 for the biological control of apple ring rot. Postharvest Biol. Technol. 115, 113–121 (2016).
Lee, A., Cheng, K.-C. & Liu, J.-R. Isolation and characterization of a Bacillus amyloliquefaciens strain with zearalenone removal ability and its probiotic potential. PLoS One 12, e0182220 (2017).
Jamal, Q., Lee, Y. S., Jeon, H. D., Park, Y. S. & Kim, K. Y. Isolation and biocontrol potential of Bacillus amyloliquefaciens Y1 against fungal plant pathogens. Korean J. Soil Sci. Fertil. 48, 485–491 (2015).
Daneshazari, R., Rabbani Khorasgani, M., Hosseini-Abari, A. & Kim, J.-H. Bacillus subtilis isolates from camel milk as probiotic candidates. Sci. Rep. 13, 3387 (2023).
Satomi, M., La Duc, M. T. & Venkateswaran, K. Bacillus safensis sp. nov., isolated from spacecraft and assembly-facility surfaces. Int. J. Syst. Evol. Microbiol. 56, 1735–1740 (2006).
Raja, C. E. & Omine, K. Arsenic, boron and salt resistant Bacillus safensis MS11 isolated from Mongolia desert soil. Afr. J. Biotechnol. 11, 2267–2275 (2012).
Okaiyeto, K., Nwodo, U. U., Mabinya, L. V. & Okoh, A. I. Bacillus toyonensis strain AEMREG6, a bacterium isolated from South African marine environment sediment samples produces a glycoprotein bioflocculant. Molecules 20, 5239–5259 (2015).
Kyriakis, S. C. et al. Evaluation of Toyocerin, a probiotic containing Bacillus toyoi spores, on health status and productivity of weaned, growing and finishing pigs. Asian Australas. J. Anim. Sci. 16, 1326–1331 (2003).
Shin, H.-J. et al. Probiotic potential of Pediococcus pentosaceus BCNU 9070. J. Life Sci. 22, 1194–1200 (2012).
Şeker, E. Identification of Candida species isolated from bovine mastitic milk and their in vitro hemolytic activity in Western Turkey. Mycopathologia 169, 303–308 (2010).
Deng, F. et al. Antimicrobial resistance, virulence characteristics and genotypes of Bacillus spp. from probiotic products of diverse origins. Food Res. Int. 139, 109949 (2021).
Dumitru, M., Lefter, N., Idriceanu, L., Ciurescu, G. & Habeanu, M. Identification and characterization of Bacillus megaterium as probiotic bacteria in poultry feed. Sci. Pap. Anim. Sci. Biotechnol. 54, 124–133 (2021).
Duc, L. H., Hong, H. A., Barbosa, T. M., Henriques, A. O. & Cutting, S. M. Characterization of Bacillus probiotics available for human use. Appl. Environ. Microbiol. 70, 2161–2171 (2004).
Jena, P. K. et al. Isolation and characterization of probiotic properties of lactobacilli isolated from rat fecal microbiota. Microbiol. Immunol. 57, 407–416 (2013).
Sharma, K., Sharma, N. & Sharma, R. Identification and evaluation of in vitro probiotic attributes of novel and potential strains of lactic acid bacteria isolated from traditional dairy products of North-West Himalayas. J. Clin. Microbiol. Biochem. Technol. 2, 18–25 (2016).
Hong, H. A. et al. The safety of Bacillus subtilis and Bacillus indicus as food probiotics. J. Appl. Microbiol. 105, 510–520 (2008).
Huang, Y. & Adams, M. C. In vitro assessment of the upper gastrointestinal tolerance of potential probiotic dairy propionibacteria. Int. J. Food Microbiol. 91, 253–260 (2004).
Floch, M. H., Binder, H. J., Filburn, B. & Gershengoren, W. The effect of bile acids on intestinal microflora. Am. J. Clin. Nutr. 25, 1418–1426 (1972).
Shinde, T. et al. Probiotic Bacillus coagulans mtcc 5856 spores exhibit excellent in-vitro functional efficacy in simulated gastric survival, mucosal adhesion and immunomodulation. J. Funct. Foods 52, 100–108 (2019).
Green, D. H. et al. Characterization of two Bacillus probiotics. Appl. Environ. Microbiol. 65, 4288–4291 (1999).
Zangeneh, M., Khaleghi, M. & Khorrami, S. Isolation of Lactobacillus plantarum strains with robust antagonistic activity, qualified probiotic properties, and without antibiotic-resistance from traditional sourdough. Avicenna J. Clin. Microbiol. Infect. 6, 66–74 (2019).
Abriouel, H., Franz, C. M. A. P., Omar, N. B. & Gálvez, A. Diversity and applications of Bacillus bacteriocins. FEMS Microbiol. Rev. 35, 201–232 (2011).
Larsen, N. et al. Characterization of Bacillus spp. strains for use as probiotic additives in pig feed. Appl. Microbiol. Biotechnol. 98, 1105–1118 (2014).
Manhar, A. K. et al. In vitro evaluation of celluloytic Bacillus amyloliquefaciens AMS1 isolated from traditional fermented soybean (Churpi) as an animal probiotic. Res. Vet. Sci. 99, 149–156 (2015).
El-Mabrok, A. S. W., Hassan, Z., Mokhtar, A. M., Hussain, K. M. A. & Kahar, F. Screening of lactic acid bacteria as biocontrol against (Collectotrichum capsici) on chilli Bangi. Res. J. Appl. Sci. 7, 446–473 (2012).
Barzegar, H., Alizadeh Behbahani, B. & Falah, F. Safety, probiotic properties, antimicrobial activity, and technological performance of Lactobacillus strains isolated from Iranian raw milk cheeses. Food Sci. Nutr. https://doi.org/10.1002/fsn3.2365 (2021).
Lee, S. et al. Probiotic characteristics of Bacillus strains isolated from Korean traditional soy sauce. LWT-Food Sci. Technol. 79, 518–524 (2017).
Guo, X., Li, D., Lu, W., Piao, X. & Chen, X. Screening of Bacillus strains as potential probiotics and subsequent confirmation of the in vivo effectiveness of Bacillus subtilis MA139 in pigs. Antonie Van Leeuwenhoek 90, 139–146 (2006).
La Ragione, R. M. & Woodward, M. J. Competitive exclusion by Bacillus subtilis spores of Salmonella enterica serotype Enteritidis and Clostridium perfringens in young chickens. Vet. Microbiol. 94, 245–256 (2003).
Seo, W.-T., Nam, S.-H., Lee, C.-K. & Cho, K.-M. Identification of potential Bacillus subtilis probiotics from Korean soybean paste and their antimicrobial and immune activities. Prev. Nutr. Food Sci. 16, 37–44 (2011).
Talebi, S., Makhdoumi, A., Bahreini, M., Matin, M. M. & Moradi, H. S. Three novel Bacillus strains from a traditional lacto-fermented pickle as potential probiotics. J. Appl. Microbiol. 125, 888–896 (2018).
Ehling-Schulz, M. et al. Toxin gene profiling of enterotoxic and emetic Bacillus cereus. FEMS Microbiol. Lett. 260, 232–240 (2006).
Kim, J. et al. Toxin genes profiles and toxin production ability of Bacillus cereus isolated from clinical and food samples. J. Food Sci. 76, T25–T29 (2011).
Jiménez, G., Blanch, A. R., Tamames, J. & Rosselló-Mora, R. Complete genome sequence of Bacillus toyonensis BCT-7112T, the active ingredient of the feed additive preparation Toyocerin. Genome Announc. 1, e01080-e1113 (2013).
Vos, P. et al. Bergey’s Manual of Systematic Bacteriology: Volume 3: The Firmicutes Vol. 3 (Springer Science & Business Media, 2011).
Ahrné, S., Molin, G. & StÅhl, S. Plasmids in Lactobacillus strains isolated from meat and meat products. Syst. Appl. Microbiol. 11, 320–325 (1989).
Schaeffer, A. B. & Fulton, M. D. A simplified method of staining endospores. Science 77, 194 (1933).
Patel, A. K., Deshattiwar, M. K., Chaudhari, B. L. & Chincholkar, S. B. Production, purification and chemical characterization of the catecholate siderophore from potent probiotic strains of Bacillus spp. Bioresour. Technol. 100, 368–373 (2009).
Angmo, K., Kumari, A. & Bhalla, T. C. Probiotic characterization of lactic acid bacteria isolated from fermented foods and beverage of Ladakh. LWT-food Sci. Technol. 66, 428–435 (2016).
Chon, J.-W. & Seo, K.-H. Development of a new chromogenic medium for the enumeration of Bacillus cereus in various ready-to-eat foods. Food Control 128, 108188 (2021).
Weisburg, W. G., Barns, S. M., Pelletier, D. A. & Lane, D. J. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 173, 697–703 (1991).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Bauer, A. W., Kirby, W. M. M., Sherris, J. C. & Turck, M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 45, 493 (1966).
Jorgensen, J. H., Institute, C. & L. S.,. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of Infrequently Isolated or Fastidious Bacteria: Approved Guideline (National Committee for Clinical Laboratory Standards, 2010).
Sen, R., Pal, D., Kodali, V. P., Das, S. & Ghosh, S. K. Molecular characterization and in vitro analyses of a sporogenous bacterium with potential probiotic properties. Probiotics Antimicrob. Proteins 2, 152–161 (2010).
Yilmaz, M., Soran, H. & Beyatli, Y. Antimicrobial activities of some Bacillus spp. strains isolated from the soil. Microbiol. Res. 161, 127–131 (2006).
Balouiri, M., Sadiki, M. & Ibnsouda, S. K. Methods for in vitro evaluating antimicrobial activity: A review. J. Pharm. Anal. 6, 71–79 (2016).
Oltuszak-Walczak, E., Walczak, P. & Modrak, R. Detection of enterotoxic Bacillus cereus producing hemolytic and non hemolytic enterotoxins by PCR test. Polish J. Microbiol. 55, 113–118 (2006).
Abbas, B. A., Khudor, M. H. & Saeed, B. M. S. Detection of hbl, nhe and bceT toxin genes in Bacillus cereus isolates by multiplex PCR. Int. J. Curr. Microbiol. App. Sci 3, 1009–1016 (2014).
Wiwat, C. & Thiramanas, R. Detection of hemolysin BL gene of Bacillus cereus isolates. Mahidol Univ. J. Pharm. Sci. 41, 22–30 (2014).
Salim, S. M., Mandal, J. & Parija, S. C. Isolation of Campylobacter from human stool samples. Indian J. Med. Microbiol. 32, 35–38 (2014).
Acknowledgements
Financial support received from Ehyagaran Ghalb Sanat Asia through the contract number 94/146295 is appreciated.
Funding
This work was partially supported by Ehyagaran Ghalb Sanat Asia through the contract No. 94/146295.
Author information
Authors and Affiliations
Contributions
M.G., N.B., and Z.M. isolated the microorganisms, characterized the isolates, analyzed the results, and prepared the manuscript draft. M.R.K., M.A.A., S.S.-A.F., and R.S. supervised the work, provided the resources, validated the results, administered the project, and reviewed and edited the manuscript. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Golnari, M., Bahrami, N., Milanian, Z. et al. Isolation and characterization of novel Bacillus strains with superior probiotic potential: comparative analysis and safety evaluation. Sci Rep 14, 1457 (2024). https://doi.org/10.1038/s41598-024-51823-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-51823-z
- Springer Nature Limited