Abstract
In this work, we investigated the oncogenic role of Streptococcus gallolyticus subsp. gallolyticus (SGG), a gut bacterium associated with colorectal cancer (CRC). We showed that SGG UCN34 accelerates colon tumor development in a chemically induced CRC murine model. Full proteome and phosphoproteome analysis of murine colons chronically colonized by SGG UCN34 revealed that 164 proteins and 725 phosphorylation sites were differentially regulated. Ingenuity Pathway Analysis (IPA) indicates a pro-tumoral shift specifically induced by SGG UCN34, as ~ 90% of proteins and phosphoproteins identified were associated with digestive cancer. Comprehensive analysis of the altered phosphoproteins using ROMA software revealed up-regulation of several cancer hallmark pathways such as MAPK, mTOR and integrin/ILK/actin, affecting epithelial and stromal colonic cells. Importantly, an independent analysis of protein arrays of human colon tumors colonized with SGG showed up-regulation of PI3K/Akt/mTOR and MAPK pathways, providing clinical relevance to our findings. To test SGG’s capacity to induce pre-cancerous transformation of the murine colonic epithelium, we grew ex vivo organoids which revealed unusual structures with compact morphology. Taken together, our results demonstrate the oncogenic role of SGG UCN34 in a murine model of CRC associated with activation of multiple cancer-related signaling pathways.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the third most diagnosed cancer worldwide with 1.9 million new cases detected in 20201. CRC develops in patients over many years through the accumulation of genetic and epigenetic alterations. Many intrinsic or extrinsic factors can influence the initiation and/or development of CRC, complicating the study of its etiology.
The colon is composed of several cellular layers, including an epithelial layer and a subepithelial lamina propria containing abundant stromal cells, which are required for intestinal homeostasis2. Both epithelial cells and their stromal microenvironment play a role in CRC development3. The colon is a very singular organ colonized by a vast and complex community of microorganisms known as the gut microbiota. It is composed of over 500 species totaling approximately 1013 bacteria4. Accumulating evidence supports the role of the microbiota in CRC development5,6,7,8,9. A few bacterial species have been identified as playing a role in colorectal carcinogenesis including: Streptococcus gallolyticus subsp. gallolyticus (SGG), Fusobacterium nucleatum, enterotoxigenic Bacteroides fragilis, colibactin-producing Escherichia coli, Parvimonas micra, and Clostridium septicum10. Understanding the role of SGG in CRC is critical to developing novel strategies to improve clinical diagnosis and treatment of this disease.
SGG, formerly known as Streptococcus bovis biotype I, is one of the first bacteria to be associated with CRC11,12. This association, ranging from 47 to 85% depending on the method used for SGG detection has been confirmed by several epidemiological studies13,14,15,16,17,18,19,20. We also studied SGG prevalence in control or CRC-patients and showed that SGG was detected more frequently in the stools of patients with adenocarcinomas (50%) as compared to early-adenoma patients (30.4%) or control ‘tumor-free’ patients (32.5%)21.
Streptococcus gallolyticus has been subdivided into three subspecies, subsp. gallolyticus (SGG), subsp. pasteurianus (SGP), and subsp. macedonicus (SGM) (Fig. S1A). Only SGG causing septicemia and endocarditis in elderly people is associated with CRC. SGP causes bacteremia, endocarditis, urinary tract infection in elderly and immunodeficient people, septicemia and meningitis in newborns and intrauterine infections in pregnant woman22,23,24,25. SGM, the genetically closest SGG relative, is considered as safe and non-pathogenic26,27 and thus used as a control bacterium in this study. Whether SGG is a driver and/or a passenger bacterium in CRC has been an open question for decades (reviewed in28). In favor of the passenger model, we and others previously showed that tumor-associated conditions provide SGG UCN34 with the ideal environment to proliferate29,30. We showed that this colonization advantage was linked to the production of a bacteriocin enabling SGG to kill closely related gut microbiota bacteria30. In favor of the driver model, various molecular mechanisms have been proposed including inflammation31, cell proliferation or blockade of host immunity. Kumar et al. showed that SGG strain TX20005 accelerates tumor growth through activation of the Wnt/β-catenin signaling pathway32,33 leading to increased cell proliferation. The same group reported more recently that a type VII secretion system of SGG TX20005 contributes to the development of colonic tumors34. Increased colonic cell proliferation was recently linked to the modulation of extracellular matrix by SGG TX2000535. Lastly, another group suggested that SGG ATCCBAA-2069 drives tumor formation by modulating anti-tumor-immunity in a colitis-associated CRC model36.
In this work, we tested the oncogenic potential of SGG UCN34 using 2 different murine CRC models: (1) the AOM-induced CRC model in A/J mice and (2) the genetic APCMin/+ model. Azoxymethane (AOM) is an active derivative of dimethyl hydrazine, an azide compound inducing DNA mutations in colon cells37. The AOM-induced mouse model reflects the human sporadic CRC events with adenoma to carcinoma progression which are largely influenced by gut microbiota composition. As a second model, we used the well-known APCMin/+ genetic model. Multiple intestinal neoplasia (Min) mice, containing a point mutation in the Adenomatous polyposis coli (APC) gene in a C57BL/6 background (C57BL/6 APCMin/+ = APCMin/+)38, develop numerous adenomas in the small intestine and was the first model used to demonstrate the involvement of the APC gene in intestinal tumorigenesis39,40.
We show that SGG UCN34 can accelerate tumor development in the AOM-induced CRC model compared to mice colonized with control SGM. Analysis of the global proteome and phosphoproteome from murine colon tissue colonized by SGG UCN34 or control SGM over 12 weeks revealed that SGG UCN34 promote autocrine and paracrine pro-tumor signals. Most of the proteins and phosphoproteins regulated by SGG UCN34 were associated with digestive cancer. In depth bioinformatic analyses using ROMA software revealed that the pro-oncogenic role of SGG UCN34 is associated with activation of several cancer hallmark pathways in tumor cells and their stromal microenvironment. In line with these results, we show here for the first time that SGG UCN34 contributes to pre-cancerous transformation of the colon epithelium using an ex-vivo organoid model. Finally, an independent protein analysis of human colon tumors colonized with SGG revealed up-regulation of PI3K/Akt/mTOR and MAPK pathways, providing clinical relevance to our findings in a murine model of CRC.
Results
SGG UCN34 accelerates tumorigenesis in AOM-induced CRC model
We tested the oncogenic potential of SGG UCN34 in the AOM-induced CRC model in A/J mice as summarized in Fig. 1a. We compared groups of mice infected for 12 weeks with SGG UCN34 or non-pathogenic SGM. A third group of mice, annotated NT, received PBS by oral gavage. We showed that both SGG UCN34 and SGM were able to colonize the murine colon at comparable levels through the duration of the experiment (Fig. 1b). SGM colonization remained very stable over the 12 weeks while SGG UCN34 showed a slight but statistically not significant decrease after week 8. After 12 weeks of colonization, mice were sacrificed, and colons dissected for tumor inspection and counts. We found that mice colonized by SGG UCN34 displayed between 1 and 8 tumors (average of 4 tumors/mouse), mostly in the distal and middle part of the colon, whereas in the control SGM group, only 2 mice exhibited 1 tumor. In the NT group, only 1 mouse showed 1 tumor (Fig. 1c). In addition, the few tumors recorded in the SGM and NT control groups were very small in size compared to those observed with SGG UCN34 (Fig. 1D). Histopathological analyses of the colonic tumors performed in blind indicate that most of the tumors induced by SGG UCN34 were low-grade adenomas (Fig. 1e, Fig. S1a), while low-grade dysplasia and low-grade adenomas were observed in NT and SGM groups respectively (Fig. 1e, Fig. S1a).
SGG UCN34 induces acceleration of tumorigenesis in AOM-induced CRC model. (a) Experimental design used for the development of colonic tumors in an AOM-induced CRC model. (b) CFU (colony forming units) per g of stools. Fecal materials were collected at different time points (1, 5, 8 and 11 weeks), homogenized, and serial dilutions plated onto Enterococcus Selective Agar plates to enumerate SGG UCN34 and SGM bacteria; ns, not significant; Mann–Whitney test. (c) Sum of tumors per mouse. Macroscopic tumors were evaluated by an experimented observer. ns not significant; *p < 0.05; **p < 0.01; Mann–Whitney test. (d) Representative pictures of mouse colon after dissection for SGM and SGG UCN34 groups (T tumor, LA lymphoid aggregate). (e) Representative histological (H&E) sections of colon tumors found in AOM-induced CRC mouse model for each experimental group: NT (low-grade dysplasia), SGM (low-grade dysplasia) and SGG UCN34 (low-grade adenoma).
We compared the oncogenic effect of SGG UCN34 with other SGG clinical isolates distant from UCN34. We chose the USA isolate TX20005 previously shown to accelerate tumor formation in the AOM-induced CRC model32,33,34. Mice colonized over 12 weeks with SGG UCN34 or TX20005 exhibited a higher number of tumors as compared to control groups NT and SGM (Fig. S1b). There was no significant difference in term of tumor count between the two UCN34 and TX20005 SGG isolates (Fig. S1b).
We also tested the oncogenic potential of SGG UCN34 in the APCMin/+ mouse model (Fig. S2a). As shown in Fig. S2b, mice colonized with SGG UCN34 exhibited a slightly increased numbers of adenomas from small intestine that were also larger compared to mice colonized with control SGM bacteria, but these differences were not statistically significant. We hypothesize that this may be due to a lower capacity of colonization of the small intestine by SGG UCN34 (400 × times less) as compared to the colon (Fig. S2c) and to the absence of colonic specific microenvironment such as the gut microbiota.
SGG UCN34 induces multiple pro-tumoral changes in murine colon epithelium
In order to decipher the mechanisms underlying SGG-induced tumorigenic transformation, full proteome and phosphoproteome analyses were carried out on whole protein extracts from macroscopically tumor-free colon tissue of mice colonized by SGG UCN34 or SGM as well as from colonic tumors of mice exposed to SGG UCN34 (Fig. 2a). Phosphoproteome analysis was performed by using proteome samples and phosphopeptide enrichment followed by label-free quantitative mass spectrometry.
Proteome and phosphoproteome analyses on colonic tissue colonized by SGG UCN34 or SGM over 12 weeks reveals a pro-tumoral shift specific to SGG UCN34. (a) Sample preparation for proteome and phosphoproteome LC–MS analysis. All samples were collected at the end of the AOM-induced CRC model. Tissue samples were from macroscopically tumor-free colon sections for SGM and SGG UCN34 groups and from colon tumors found in this experiment only for the SGG UCN34 group. Proteins were extracted by urea lysis and trypsin digestion. For phosphoproteome MS analysis a phosphopeptide enrichment step was added. All samples were processed by label free LC–MS/MS analysis. (b) Principal component analysis (PCA) of proteome samples in 3D representation. (c) PCA of phosphoproteome samples in 3D representation. Proteome and phosphoproteome analyses were done using myProMS web server65. (d) Top disease and bio functions (IPA) of proteome and phosphoproteome changes between SGG UCN34 and SGM. The diseases and functions analysis identified biological functions and/or diseases that were most significant from the data set. Data sets used in this analysis were proteins or phosphoproteins differentially expressed between SGG UCN34 and SGM detected in macroscopically tumor-free colonic tissue. Molecules indicate the number of proteins or phosphoproteins associated with indicated diseases and disorders. A right-tailed Fisher’s Exact Test was used to calculate a p value determining the probability that each biological function and/or disease assigned to that data set is due to chance alone.
The proteome of tumor-free colon segments of mice colonized with SGG UCN34 compared to those colonized with SGM revealed that SGG UCN34 induces significant changes in the levels of 164 proteins as compared to the control SGM group (Fig. 2a, Table S1). Only 35 protein levels were found increased while 129 protein levels were found decreased. Interestingly, stronger changes were observed in the phosphoproteome with 725 phosphosites corresponding to 642 proteins detected at significantly different levels between SGG UCN34 and SGM. The phosphorylation levels of 325 sites (299 proteins were found increased while those of 400 sites (343 proteins) were found decreased (Fig. 2a, Table S2).
Principal Component Analyses (PCA) of all samples based on protein (Fig. 2b) and phosphophosites abundance (Fig. 2c) revealed 3 separate clusters matching the phenotypic observations (SGM tumor-free, SGG tumor-free, SGG tumor). As expected, the protein content of the tumoral tissue (SGG tumor) differs from that of the adjacent macroscopically healthy tissue (SGG tumor-free). Importantly, SGG UCN34 induced specific proteomic and phosphoproteomic changes in macroscopically tumor-free tissue compared to control SGM-treated tumor-free tissue indicating that SGG UCN34 modulates the proteome and phosphoproteome landscapes of the murine colon epithelium before any macroscopically visible changes. These results suggest that SGG UCN34 is not a silent colon inhabitant in sharp contrast with SGM.
To investigate the colonic alterations induced by SGG UCN34, Ingenuity Pathway Analysis (IPA, Qiagen) was applied to our datasets. Most of the differentially expressed proteins and phosphosites in the tumor-free colonic tissue exposed to SGG vs SGM are linked to cancer development (Fig. 2d). As many as (1) 150 proteins out of 163 and (2) 572 phosphoproteins out of 597 were predicted to be associated with cancer development.
This result shows that SGG UCN34 induces a clear pro-tumoral shift at the proteomic and phosphoproteomic levels in mice colons.
SGG UCN34 activates MAPK, mTOR and integrin/ILK/actin cytoskeleton pathways in vivo
To identify the specific signaling pathways altered by SGG UCN34, we first processed our phosphoproteomic data sets with the in-house built ROMA (Representation and quantification Of Module Activities) software, a gene-set-based quantification algorithm41 integrating several databases. As shown in Fig. 3a, the control SGM group clearly differs from SGG group. Secondly, the pro-tumoral shift induced by SGG in the tumor-free region matches that found the tumoral part. Thirdly, 83 pathways were found significantly altered when comparing SGM and SGG groups (evidenced by two-tailed t-tests of activity scores; p value ≤ 0.05) (Fig. 3a, Sup Table S3). Among these 83 pathways, 34 were up-regulated by SGG UCN34 and 49 pathways were down-regulated. The top 10 up-regulated pathways were: epidermal growth factor (EGF), integrin, platelet derived growth factor (PDGF), PYK2, melanoma, p38, mitogen activated protein kinases (MAPK) family signaling cascades, target of rapamycin (mTOR) signaling, regulation of Ras by GTPase activating proteins (GAPs), signaling by fibroblast growth factor receptor 2 (FGFR2) (Table S3). Five of these (EGF, PDGF, p38, GAPs and FGFR2) converge towards MAPK family signaling cascades. Interestingly, PDGFRA (one of the major phosphoproteins identified in the PDGF pathway, Table S5) is exclusively expressed by stromal cells both in normal intestine as well as in colorectal cancer42,43. Through analysis of single cells RNAseq data from human colorectal cancer and normal adjacent tissue42 (Fig. S6), we observed that Pdgfra is mostly expressed by colonic stromal cells positive for podoplanin (Pdpn+). These stromal cells are the main producers of bone morphogenetic protein (BMP) antagonists such as Gremlin 1 (Grem1) involved in stem cells maintenance and tumorigenesis44,45,46 (Fig. S6b). This result shows for the first time that SGG can alter the stromal microenvironment.
ROMA analysis reveals multiple cancer-related pathways altered by SGG UCN34 in comparison to SGM. (a) Heat map view of up and downregulated pathways between three groups: SGM tumor-free, SGG UCN34 tumor-free and SGG UCN34 tumors. (b) Schematic diagram of SGG UCN34-induced activation of MAPK cascades, mTOR signaling and integrin/ILK/actin cytoskeleton pathways. ROMA pathway analysis together with IPA indicates activators of MAPK cascades by UCN34. Apart from the MAPK cascades we can point out activation of the mTOR signaling pathway as well as integrin/ILK/actin cytoskeleton signaling. ROMA: representation and quantification of module activities. Created with Biorender.com.
The top 10 down-regulated pathways were as following: adherens junction, insulin, vascular smooth muscle contraction, protein tyrosine phosphatase 1B (PTP1B), protein tyrosine phosphatase-2 (SHP2), AUF1 (hnRNP D0) which binds and destabilizes mRNA, Feline McDonough Sarcoma-like tyrosine kinase (FLT3), RUNX2 which regulates bone development, TP53, Fc epsilon receptor 1 (FCER1) (Table S3).
Secondly, the same data sets were re-analysed using the commercially available Ingenuity Pathway Analysis software. The top 10 signalling pathways selected by activation prediction score (positive z-score) were: Integrin-linked kinase (ILK), BRCA1, Dilated Cardiomyopathy Signaling Pathway, Protein Kinase A, Actin Cytoskeleton, AMPK signaling, Epithelial Adherens Junction, Oxytocin Signaling Pathway, Integrin Signaling and CNTF Signaling (Fig. S3). The other signaling pathways that were in line with ROMA analysis and were predicted to be activated by SGG UCN34 were ERK/MAPK and mTOR (Fig. S4).
Taken together, ROMA and IPA analyses indicate that SGG UCN34 can interfere with several pro-tumoral signaling pathways, such as MAPK, mTOR and integrin/ILK/actin cytoskeleton pathways (Fig. 3b, Figs. S4, S5) which in turn can contribute to colorectal cancer growth acceleration as well as pre-cancerous transformation of the colon epithelium.
Protein analysis of human colon tumors enriched for SGG reveals up-regulation of PI3K/Akt/mTOR and MAPK pathways
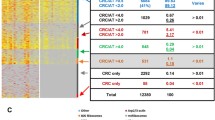
Independently and by a different group of researchers, the protein content of human colon tumor biopsies colonized with SGG (n = 32) vs control colon tumor biopsies negative for SGG (n = 29)32 was analysed using RPPA (Reverse Phase Protein Array)47. In total, 52 proteins and/or phosphorylation sites were differentially regulated between SGG-enriched human colon tumors vs SGG-negative ones. Among them, 16 proteins and 11 phosphosites were up-regulated and 17 proteins and 8 phosphosites were down-regulated (Fig. 4, Table S4). Interestingly the majority of the up-regulated proteins/phosphosites belong to two signaling pathways: (1) PI3K/AKT/mTOR (Akt, Akt pS473, GSK3 p89, GSK3αβ, GSK3αβ PS21, PI3K p85, PTEN, Src pY527, TSC1, PDK1, Lck, Smad2, β-Catenin) and (2) MAPK (p38, p38 pT180pY182, JNK2, Syk, PKCα pS657, PKCβII pS660).
SGG induces up-regulation of PI3K/Akt/mTOR, MAPK and AMPK pathways in human colon tumors. Proteins were extracted from human colon biopsies enriched for SGG (n = 32) vs negative ones (n = 29) and subjected to RPPA. The table displays up-regulated by SGG pathways for which at least 3 RPPA targeted antibody were attributed. RPPA: reverse phase protein array. Created with BioRender.com.
SGG UCN34 induces formation of abnormal ex vivo organoids with compact morphology
We wondered whether the observed pro-tumoral shift in tumor-free sections of murine colon exposed to SGG UCN34 had phenotypical consequences on the colonic tissue and its microenvironment. To test more directly the effect of SGG UCN34 on stromal cells as suggested by activation of PDGFRA (Table S5), a receptor strongly involved in the stroma-cancer cells crosstalk48, we performed quantification of Grem1 producing stromal cells positive for podoplanin (Pdn+) and CD3449 on histological sections. As shown in Fig. 5a, we observed an increased frequency of Pdpn+CD34+ stromal cells in the SGG UCN34 group compared to the SGM group.
SGG UCN34 induces pre-cancerous transformation of murine colon tissue. (a) Immunofluorescence analysis of Pdpn (green) and CD34 (red) in sections of colonic tumor-free regions of mice colonized with SGM (top) or SGG UCN34 (bottom) (DAPI/nuclei in blue and E-cadherin in grey). Scale bar 50 µm. Images were acquired with SP8 microscope (Leica) and a ×40 objective. Graph presents the percentage of colocalization of Pdpn and CD34 signals per field of view (FOV). The quantification was done by Imaris (version8) software on SGM (n = 8) and SGG UCN34 (n = 7) colonized mice, using at least 6 images per mouse and showing 1.1% vs 3.1% of Pdpn + CD34 + signal respectively (*t-test p = 0.01). (b) Experimental design of ex vivo organoid formation originating from macroscopically tumor-free sections of colon tissue colonized by SGG UCN34 or SGM over 12 weeks in the AOM-induced CRC model. (c) The number of organoids with normal or cystic morphology defined as colonospheres or colonoids with a central empty lumen and polarized epithelium (actin staining). (d) The number of organoids with compact morphology defined as colonospheres or colonoids with a lumen full of cells and depolarized, restructured actin filaments. (e). Microscopic pictures showing DAPI/nuclei labeling (blue), Phalloidin/actin (green) and Ki67/proliferation marker (red). Images were acquired with Opterra Multipoint Scanning Confocal microscope (Bruker) at ×63 objective and analyzed by Fiji software. Scale bar 50 µm.
Next, to determine whether intestinal epithelial cells were intrinsically modified in their capacity to proliferate and differentiate, we examined ex vivo organoid formation50. Normally, organoid formation from murine colonic crypts requires supplementation of several niche factors (Wnt, R-Spondin, EGF, Noggin) to maintain their stemness and proliferation status51. We hypothesized that pre-cancerous tissue transformation of the colonic tissue in the group of mice colonized with SGG could allow organoid formation without any supplementation of niche factors.
Development of ex vivo organoids originating from cells obtained from tumor-free colon tissue colonized by SGG UCN34 or SGM over 12 weeks in the AOM-induced CRC model was tested (Fig. 5b). After 6 days of culture in the absence of the four niche factors (Wnt, R-Spondin, EGF, Noggin), we evaluated the presence or absence of organoids and classified them according to their morphology. In both groups, we observed a similar number of organoids that we classified as normal or cystic, characterized by budding crypts or round structures respectively, composed of a monolayer of epithelial cells with a central lumen containing some dead cells50 (Fig. 5c). Strikingly, we found that only SGG UCN34 induced the formation of numerous organoids with a different morphology, defined as compact (Fig. 5d). These compact organoids are characterized by their heterogeneous shape, and a filled lumen. The epithelium of compact organoids appeared depolarized and unstructured compared to the polarized and organized epithelium observed in the normal or cystic organoids, as highlighted by phalloidin staining (yellow arrow, Fig. 5e). Staining of these abnormal organoids induced by SGG with the proliferative Ki-67 marker revealed the presence of proliferative cells within the lumen (red arrow, Fig. 5e). Other examples of organoids with normal and cystic morphology and organoids with compact morphology are shown in Fig. S7.
Taken together these results show for the first time that SGG UCN34 contributes to early (macroscopically not visible) pre-cancerous transformation of murine colon epithelium and its microenvironment (Fig. 6).
Working model of the oncogenic potential of SGG UCN34. Chronic colonization of host colon with SGG UCN34 induces the activation of MAPK cascades, PI3K/AKT/mTOR signaling and integrin/ILK/actin cytoskeleton pathways. SGG UCN34 alters multiple signaling pathways found downstream of EGF and FGF receptors present on colonic epithelial cells, but also downstream of PDGFA receptor present in stromal cells. SGG UCN34 induces the expansion of Pdpn+CD34+ cell in the stromal compartment, which regulates epithelial cells differentiation/proliferation. All these events contribute to pre-cancerous transformation of colon epithelium and acceleration of tumor development. Created with BioRender.com.
Discussion
A better understanding of the contributions of Streptococcus galloyticus subsp. gallolyticus (SGG) to colorectal cancer (CRC) acceleration is critical to developing novel strategies to improve clinical diagnosis and treatment of this disease.
In this work, we have shown that SGG strain UCN34 can accelerate the development of tumors in the AOM-induced CRC model. To explore the underlying mechanisms, we performed global proteomic and phosphoproteomic analysis of mice colon tissue chronically exposed to SGG UCN34 for 12 weeks. We compared these results to mice colonized with S. gallolyticus subsp. macedonicus (SGM) considered as the closest non-pathogenic relative. Colonization with SGG UCN34 altered the expression of 164 proteins and 725 phosphosites in colon tissue devoid of tumors as compared to the equivalent colon tissue colonized with SGM. Most altered proteins were associated with cancer disease. Bioinformatic analyses using IPA and ROMA software revealed significant up-regulation of multiple signaling pathways previously linked to cancer cells and their stromal microenvironment, including the three major MAPKs (ERK, JNK, p38), the PI3K/AKT/mTOR and integrin/ILK/actin cytoskeleton. These data suggest that SGG UCN34 affects multiple proteins in these pathways altering growth factors signaling cascades such as EGF and FGF receptors, ubiquitously expressed, and PDGFA receptors, which are mostly expressed in the colonic stromal compartment. Importantly, an independent analysis of the protein and phosphoprotein content from human colon tumors enriched for SGG revealed up-regulation of PI3K/Akt/mTOR and MAPK pathways, providing clinical relevance to our findings in the murine model. A working model summarizing our results is depicted in Fig. 6. We hypothesize that this complex activation of multiple signaling pathways in cancer cells and their stromal microenvironment by SGG UCN34 can lead to cell proliferation and transformation, thus accelerating tumor development.
We also showed that SGG UCN34 contributes to early pre-cancerous transformation of murine colon epithelium, with expansion of Pdpn+CD34+ stromal niche, which have been reported to regulate epithelial cells differentiation/proliferation by producing BMP antagonists such as Grem149. Furthermore, using an ex vivo organoid model, we found that SGG UCN34 induces the formation of abnormal organoids with compact morphology. Interestingly, this compact morphology was previously observed in human colon tumor-derived organoids52,53. Staining with phalloidin revealed strong disorganization of the actin cytoskeleton. These SGG- induced cytoskeleton rearrangements may be due to changes in the integrin/ILK/actin cytoskeleton signaling pathways54,55,56,57.
It is worth noting that SGG UCN34 did not significantly increase the number and/or size of adenomas in the APCMin/+ mice, mirroring our previous results in the Notch/APC mice30. These results indicate that the oncogenic effect of SGG UCN34 is influenced by host genetic factors and/or the niche environment. SGG UCN34 is rather an opportunistic pathobiont profiting from a favorable ecological niche (e.g., AOM-induced tumoral environment) and exerting its pro-oncogenic effect only under specific conditions. Consistent with this idea, we did not observe any direct ‘oncogenic’ cellular changes (cell proliferation, cytoskeleton reorganization, cell migration, or induction of double strand DNA breaks) in vitro induced by SGG UCN34 using short-term infection of human colonic cells (Fig. S8). Altogether, our results indicate that SGG UCN34-induced alterations of colon epithelium in vivo are probably not direct and potentially occur through multiple components of this complex system, e.g., gut microbiota, mucus, stroma, immune cells and/or enteric nervous system. Future studies should focus on more complex and integrated systems such as “gut-on-a-chip”58 to unveil the key factors contributing to the acceleration of tumorigenesis by SGG UCN34.
In parallel, the sequencing of several other SGG clinical isolates associated with CRC revealed their highly diverse genetic content (Fig. S10). This result renders the existence of a single toxin-encoded genetic island directly driving colon tumor development unlikely, differing from other CRC-associated bacteria such as pks + E. coli or bft + B. fragilis59,60,61. We thus hypothesize that the oncogenic potential of SGG may differ from one isolate to another. Genetically and geographically distant SGG isolates, namely SGG UCN34 (France) and SGG TX20005 (USA) accelerate the development of tumors in the AOM-induced CRC model at similar levels, but likely through different mechanisms. Indeed, SGG TX20005 was shown to accelerate cell proliferation in vitro and tumor growth in vivo though activation of the β-catenin pathway32,33 which is not the case for SGG UCN34 (Figs. S8a, S9). Interestingly, the genetic island encoding the type VII secretion system of SGG TX20005, shown to contribute to tumor development, exhibits significant differences in term of organization, expression, and secreted effectors with the majority of other SGG isolates exemplified by SGG UCN3462.
Finally, our group and others also previously showed that SGG UCN34 benefits from tumor metabolites29 and is able to colonize the host colon in tumoral conditions by outcompeting phylogenetically-related members of the gut microbiota30. Building on past and present data, together with our recent paper showing significant enrichment of SGG in the stools of patients with adenocarcinoma21, we propose that all SGG isolates have the capacity to take advantage of tumoral conditions in the host colon to multiply. In contrast, SGG’s ability to drive colon tumorigenesis appears strain-specific and will occur at different levels through different mechanisms. Understanding how SGG and other CRC-associated pathobionts increase the risk of developing colonic tumors is key to improving diagnosis and treatment of colorectal cancer.
Material and methods
Patient samples
All procedures were performed according to the Declaration of Helsinki and were approved by the institutional review board at University of Texas M. D. Anderson Cancer Center and Texas A&M Health Science Center and are reported in compliance with the ARRIVE guidelines63. The ethics approval number for analysis of human colon tissues is IRB2012-0507D. Colon biopsy samples were collected from patients at the University of Texas M. D. Anderson Cancer Center (MDACC), Houston, Texas. The cohort contains mostly stage II and III tumors. The patients had previously given written informed consent for use of their samples in future colorectal cancer research. Patient identifiers were anonymized. Collection and handling of patient samples were carried out in strict accordance with relevant guidelines and regulations. Detection of SGG in the colon biopsy samples was described previously32.
Bacterial strains and culture conditions
Streptococcus gallolyticus SGG UCN34, SGG TX2005 and SGM CIP105683T were grown at 37 °C in Todd Hewitt Yeast (THY) broth in standing filled flasks or on THY agar (Difco Laboratories). Starter cultures before mice oral gavage were prepared by growing strains overnight in 50 mL THY broth. Fresh THY broth was then inoculated with the overnight culture at 1:20 ratio. Exponentially growing bacteria were harvested at 0.5 OD600 for the mouse gavage.
AOM-induced CRC model in A/J mice
A/J mice were first imported from Jackson Laboratory (USA) and then bred at the Pasteur Institute animal breeding facility in SOPF condition. Eight-week-old female A/J mice were first treated with AOM (Azoxymethane; A5486; Sigma, France) at a dose of 8 mg/kg by intraperitoneal (i.p.) injection once a week for 4 weeks. After a 2 week-break, mice were treated during 3 days with a broad-spectrum antibiotic mixture including vancomycin (50 μg/g), neomycin (100 μg/g), metronidazole (100 μg/g), and amphotericin B (1 μg/g). Antibiotics were administrated by oral gavage. Additionally, mice were given ampicillin (1 g/L) in drinking water for 1 week and then switched to antibiotic-free water 24 h prior to bacterial inoculation. Mice were orally inoculated with PBS 1× (NT), SGM or SGG using a feeding needle (~ 2 × 109 CFU in 0.2 mL of PBS 1×/mouse) at a frequency of three times per week during the first week of colonization and then once a week for another 11 weeks. Stools were recovered every 2-weeks to monitor SGG and SGM numbers in the colon during the whole experiment. After 12 weeks of bacterial colonization mice were euthanized and colons removed by cutting from the rectal to the cecal end and opened longitudinally for visual evaluation. Tumor numbers were recorded, and tumor sizes measured using a digital caliper. Tumor volumes were calculated by the modified ellipsoidal formula: V = ½ (Length × Width2). For each mouse, all adenomas as well as adjacent tumor-free sections (1 cm2) were dissected and frozen by immersion in liquid nitrogen and stored at − 80 °C until protein extraction. For ex vivo organoid formation, adjacent tumor-free (1 cm2) regions for both groups SGM and SGG UCN34 were dissected and processed immediately for crypt isolation. For further histological analysis, all tissue sections (tumors, tumor-free regions) were fixed for 24 h in 4% paraformaldehyde (PFA).
Determination of SGG/SGM counts in mice stools
Bacterial colonization of murine colons by SGG and SGM was determined by colony forming units (CFU) counts. Briefly, freshly collected stools were weighted and homogenized using a Precellys homogenizer (Bertine) for 2 × 30 s at a frequency of 5000 rpm. Following serial dilutions, samples were plated on Enterococcus agar selective media for counting of SGG/SGM colonies, exhibiting a specific pink color on these plates as described previously31.
Protein extraction and preparation for mass spectrometry
All proteins from previously frozen samples (tumors and tumor-free parts of murine colon) were extracted using AllPrep DNA/RNA/Protein Mini Kit (Qiagen). Protein concentrations were estimated using NanoDrop A280 absorbance. About 300 µg of each sample was incubated overnight at − 20 °C with 80% MethOH, 0.1 M Ammonium Acetate glacial (buffer 1). Proteins were then precipitated by centrifugation at 14,000g for 15 min at 4 °C and the pellets washed twice with 100 µL of buffer 1. Samples were then dried on a SpeedVac (Thermofisher) vacuum concentrator. Proteins were resuspended in 100 µL of 8 M Urea, 200 mM ammonium bicarbonate, reduced in 100 µL of 5 mM dithiotreitol (DTT) at 57 °C for 1 h and then alkylated with 20 µL of 55 mM iodoacetamide for 30 min at room temperature in the dark. The samples were then diluted in 200 mM ammonium bicarbonate to reach a final concentration of 1 M urea. Trypsin/LysC (Promega) was added twice at 1:100 (wt:wt) enzyme:substrate, incubated at 37 °C during 2 h and then overnight. Each sample were then loaded onto a homemade C18 StageTips for desalting. Peptides were eluted using 40/60 MeCN/H2O + 0.1% formic acid and 90% of the starting material was enriched using TitansphereTM Phos-TiO kit centrifuge columns (GL Sciences, 5010-21312) as described by the manufacturer. After elution from the Spin tips, the phospho-peptides and the remaining 10% eluted peptides were vacuum concentrated to dryness and reconstituted in 0.3% trifluoroacetic acid (TFA) prior to LC–MS/MS phosphoproteome and proteome analyses.
Mass spectrometry
Peptides for proteome analyses were separated by reverse phase liquid chromatography (LC) on an RSLCnano system (Ultimate 3000, Thermo Scientific) coupled online to an Orbitrap Exploris 480 mass spectrometer (Thermo Scientific). Peptides were trapped on a C18 column (75 μm inner diameter × 2 cm; NanoViper Acclaim PepMapTM 100, Thermo Scientific) with buffer A (2/98 MeCN/H2O in 0.1% formic acid) at a flow rate of 2.5 µL/min over 4 min. Separation was performed on a 50 cm × 75 μm C18 column (NanoViper Acclaim PepMapTM RSLC, 2 μm, 100 Å, Thermo Scientific) regulated to a temperature of 50 °C with a linear gradient of 2% to 30% buffer B (100% MeCN in 0.1% formic acid) at a flow rate of 300 nL/min over 220 min. MS full scans were performed in the ultrahigh-field Orbitrap mass analyzer in ranges m/z 375–1500 with a resolution of 120,000 (at m/z 200). The top 25 most intense ions were isolated and fragmented via high energy collision dissociation (HCD) activation and a resolution of 15 000 with the AGC target set to 100%. We selected ions with charge state from 2+ to 6+ for screening. Normalized collision energy (NCE) was set at 30 and the dynamic exclusion to 40 s. For phosphoproteome analyses, LC was performed with an RSLCnano system (Ultimate 3000, Thermo Scientific) coupled online to an Orbitrap Fusion mass spectrometer (Thermo Scientific). Peptides were trapped on a C18 column (75 μm inner diameter × 2 cm; NanoViper Acclaim PepMapTM 100, Thermo Scientific) with buffer A (2/98 MeCN/H2O in 0.1% formic acid) at a flow rate of 3 µL/min over 4 min. Separation was performed on a 25 cm × 75 μm C18 column (Aurora Series, AUR2-25075C18A, 1.6 μm, C18, Ionopticks) regulated to a temperature of 55 °C with a linear gradient of 2% to 34% buffer B (100% MeCN in 0.1% formic acid) at a flow rate of 150 nL/min over 100 min. Full-scan MS was acquired using an Orbitrap Analyzer with the resolution set to 120,000, and ions from each full scan were higher-energy C-trap dissociation (HCD) fragmented and analysed in the linear ion trap.
Proteome and phosphoproteome analyses
For identification, the data were compared with the Mus musculus UP000000589 SwissProt database (downloaded 01/2019 and containing 22,266 entries) and a common database of contaminants (245 entries) using Sequest HT through proteome discoverer (version 2.2). Enzyme specificity was set to trypsin and a maximum of two-missed cleavage sites were allowed. Carbamidomethylation of cysteines, oxidized methionine and N-terminal acetylation were set as variable modifications for proteomes. Phospho-serines, -threonines and -tyrosines were also set as variable modifications in phosphoproteome analyses. Maximum allowed mass deviation was set to 10 ppm for monoisotopic precursor ions and 0.02 Da for MS/MS peaks or 0.6 Da for phosphoproteome analyses. FDR calculation used Percolator64 and was set to 1% at the peptide level for the whole study. The resulting files were further processed using myProMS65 v3.9.3 (https://github.com/bioinfo-pf-curie/myproms).
Label free quantification was performed by peptide Extracted Ion Chromatograms (XICs), reextracted across all conditions (healthy/tumor-free, tumor, polyp and mix samples, see PXD038272, PXD038270, PXD038268 and PXD038267) and computed with MassChroQ version 2.2.166. Polyp samples were not used in this study since histological analysis demonstrated that they correspond to lymphoid aggregates (LA, Fig. S1E). For differential analyses, XICs from proteotypic peptides (shared between compared conditions for proteomes, no matching constraints for phosphoproteomes) with at most two-missed cleavages were used. Replicate SGM m8-H which had more than 76% missing values was excluded from the proteome differential analyses. Median and scale normalization was applied on the total signal to correct the XICs of each biological replicate for injection and global variance biases. Phosphosite localization accuracy was estimated in myProMS by using PhosphoRS67 and only phosphopeptide with a localization site probability greater than 95% were quantified. To estimate the significance of the change in protein abundance, a linear model (adjusted on peptides and biological replicates) was performed, and p-values were adjusted with a Benjamini–Hochberg FDR procedure. For proteome analyses, proteins were considered in the analysis only when they were found with at least 5 total peptides. Then, proteins with an adjusted p value ≤ 0.05 were considered significantly enriched in sample comparison. For phosphoproteome analyses, phosphosites were analyzed individually. The threshold was also 5 total phosphopeptides to consider a phosphosite for the downstream analysis. Then, phosphosites with an adjusted p value ≤ 0.05 were considered significantly changed in sample comparisons. Unique phosphosites (only detected in one of the groups in each comparison) were also included when identified in at least 5 biological replicates. In addition, we considered the extent of the change in expression to be significant if higher than or equal to 2 for up-regulated entities or lower or equal to 0.5 for down-regulated entities. For the other bioinformatic analyses, label-free quantification (LFQ) was performed following the algorithm as described68 for each sample after peptide XIC normalization as described above. The resulting LFQ intensities were used as protein (all peptide ≥ 3) or phosphosite (all peptides ≥ 2) abundance. For PCA and ROMA analyses, datasets were further filtered to remove entities with more than 34% of missing values across all samples used. The LFQ values were log10-transmformed and the remaining missing values (around 5% for proteome and 15% for phosphoproteome) were imputed using the R package missMDA69 to produce complete matrices.
Quantification of pathway activity with ROMA
ROMA (representation and quantification of module activities) is a gene-set-based quantification algorithm41 with the MSigDB C2 Canonical pathways sub-collection as pathway database composed of 2232 pathways from Reactome (http://www.reactome.org), KEGG (http://www.pathway.jp), Pathway Interaction Database (http://pid.nci.nih.gov), BioCarta (http://cgap.nci.nih.gov/Pathways/BioCarta_Pathways) and others. ROMA was used to investigate potential changes in pathway activity between the 3 groups (SGG UCN34 tumor-free, SGM tumor-free and SGG UCN34 tumor) based on phosphoproteomic data. In this study, we used an R implementation of ROMA available at https://github.com/Albluca/rRoma. Only the 16 most representative samples of the 3 groups were used for the analysis (samples SGM healthy m6, SGM healthy m9, SGG healthy m10 and SGG tumor m14 were removed due to their outlier behavior as illustrated in Fig. 2C). A data reduction to a single phosphosite per protein/gene was performed to switch from a site-centric to a gene-centric dataset. This reduction was achieved with an ANOVA across groups and selection of the site with the lowest p value (“most informative” site) when multiple sites were quantified for the same protein. 2784 of the 3188 mouse genes implicated were then converted into their human counterparts with the Biomart resource (https://m.ensembl.org/biomart, database version 100) prior to ROMA analysis against the MSigDB (http://www.gsea-msigdb.org) human C2 Canonical pathways sub-collection (version 7.1) as pathway database. 1752 genes from the dataset matched 666 of the 2232 pathways.
Ingenuity pathway analysis (IPA)
We used IPA (QIAGEN, http://www.ingenuity.com/) to characterize two sets of proteins: (1) one corresponding to the list of proteome changes between the tumor-free SGG UCN34 group and the tumor-free SGM group (164 proteins of which 163 IDs were found in the IPA database) and (2) and one to the list of phosphoproteome changes between the tumor-free SGG UCN34 group and the tumor-free SGM group (597 proteins of which all IDs were found in the IPA database). For the list of phosphoproteins, we first removed phosphosites that were found in conjunction with other phosphosites on the same protein, and in that case, we retained only the site with the lowest p value (in total 725 phosphosites mapped to 597 proteins). Lists containing differentially expressed proteins with their corresponding log2 ratio of expression values for both data sets (proteome and phosphoproteome) were uploaded and analyzed individually. We only considered proteins and phosphoproteins measurements where we had at least 5 non-zero expression measurements in at least one of the conditions. For the cases where we had no expression in one of the conditions, leading to non-finite log2 ratios, we set the log2 ratio values to be equal to + 10 [resp. − 10] in the case of an up-regulation [resp. down-regulation] in the tumor-free SGG UCN34 group. This value corresponded to the upper end (in absolute value) of log2 ratios for comparisons of measurements containing no zero-values. This measurement (log2 ratio) was used by IPA to calculate directionality (z-scores) in the analysis and is displayed in color on pathways and networks (red for up-regulated proteins and green for down-regulated proteins). We used the “Core Analysis” function to relate the expression to known diseases, biological functions, and canonical pathways. This software is based on computer algorithms that analyze the functional connectivity of genes from information obtained from the IPA database. Biological Functions with a corrected p value < 0.05 (Fisher’s exact test) were considered to be statistically significant. The analysis of canonical pathways identified the pathways from the IPA library of canonical pathways which were most relevant to the input data set, based on a test of enrichment (Fisher’s exact test).
Crypt isolation and organoid formation
Half centimeter sections of tumor-free colonic tissue were removed and washed in cold PBS 1×. The tissue was further fragmented in smaller pieces using a scalpel and washed in fresh cold PBS 1× by pipetting 5-times with pre-cut P1000 tips. After sedimentation of the pieces, the supernatant was removed and the fragments were digested in HBSS with calcium and magnesium, 2.5% fetal bovine serum (FBS), 100 U/mL penicillin–streptomycin, 1 mg/mL collagenase from Clostridium histolyticum (Sigma C2139), and 1 µM ROCK1 inhibitor (Y-27632, StemCell technologies), for 60 min in a thermomixer at 37 °C with 800 rpm agitation. After a quick spin, the fragments were further digested using Tryple express (Gibco) with 1 µM ROCK1 inhibitor, for 20 min at 37 °C with 800 rpm agitation. The samples were washed with PBS 1×, left to sediment and the supernatant was collected, centifuged and the pellet resuspended with Matrigel Growth Factor Reduced Basemen Membrane Matrix (Corning) in order to make 2 drops of 30 µL each in a 48-well plate. After 20 min at 37 °C, the Matrigel drops were overlayed with 300 µL of Medium containing Advanced DMEM, P/S, N2 and B27 (all from Gibco), lacking Noggin, R-spondin, Wnt3a, and EGF. The observation of organoids formation was done at day 6.
Immunofluorescence
Tissues and organoids were fixed with 4% paraformaldehyde (Electron Microscopy Science) overnight at 4 °C. Tissues were embedded in OCT compound (Fisher Scientific) and stored at − 80 °C. Frozen blocks were cut at 10 µm thickness, and sections were collected onto Superfrost Plus slides (VWR international). Tissue sections and organoids were permeabilized with 0.5% triton X-100 (Sigma Aldrich) in PBS 1X at RT for 40 min and incubated with a blocking buffer containing 3% bovine serum albumin (BSA, Sigma Aldrich) in PBS 1× RT for 40 min. Samples were incubated overnight at 4 °C with primary antibodies diluted in 0.1% triton X-100, 1% BSA in PBS 1×, washed, incubated with secondary antibodies for 1 h at room temperature, washed, and stained for 20 min with 1 µg/mL DAPI (Invitrogen). Finally, slides were mounted with ProLong™ gold antifade mounting medium (ThermoFisher). The following antibodies were used: anti-CD34 coupled with eF660 (clone RAM34) (eBioscience), anti-E-cadherin (clone ECCD-2) (Takara), goat anti-rat488 (Thermo Fisher), anti-Pdpn (gift from A. Farr, University of Washington, Seattle), and goat-anti-hamster 546 (Thermo Fisher), anti-Ki67 coupled with eF660 (clone SolA15) (eBioscience), phalloidin coupled with AF488 (Thermo Fisher).
Histology and immunohistochemistry
Colon fragments fixed for 48 h in 10% neutral-buffered formalin were embedded in paraffin. Four-µm-thick sections were cut and stained with hematoxylin and eosin (H&E) staining. Histological evaluation was performed by an histopathologist in a blind fashion. Characterization of LA was done by immunostaining performed on Leica Bond RX using anti-CD3 (Dako, Ref. A0452) antibody, and hematoxylin staining. Slides were then scanned using Axioscan Z1 Zeiss slide scanner and images were analyzed with the Zen 2.6 software.
Reverse phase protein array (RPPA)
This was performed in the RPPA core at MDACC using established protocols. Briefly, frozen tumors were lysed and protein extracted. Lysates were serially diluted in 5 two-fold dilutions and printed on nitrocellulose-coated slides using an Aushon Biosystems 2470 arrayer. Slides were probed with primary antibodies selected to represent the breadth of cell signaling and repair pathways70 conditioned on a strict validation process as previously described71, followed by detection with appropriate biotinylated secondary antibodies and streptavidin-conjugated horseradish peroxidase (HRP). The slides were analyzed using Array-Pro Analyzer software (MediaCybernetics) to generate spot intensity, which was adjusted using “control spots” to correct spatial bias72. A fitted curve ("Supercurve") was created for each protein using a non-parametric, monotone increasing B-spline model47. Relative protein levels were estimated using SuperCurve GUI73. Slide quality was assessed using a QC metric74 and only slides greater than 0.8 on a 0–1 scale were included for further processing. Protein measurements were corrected for loading as described73,75 using bidirectional median centering across samples and antibodies.
Statistical analysis
Mann–Whitney nonparametric test was used to test for statistical significance of the differences between the different group parameters. p values of less than 0.05 were considered statistically significant.
Ethical approval
Animals were housed in the Institut Pasteur animal facilities accredited by the French Ministry of Agriculture for performing experiments on live rodents. Work on animals was performed in compliance with French and European regulations on care and protection of laboratory animals (EC Directive 2010/63, French Law 2013-118, February 6th, 2013). All experiments were approved by the Use Ethics Committee of Institut Pasteur #89, registered under the reference dap180064 and were performed in accordance with relevant guidelines and regulations. We declare that these studies are reported in compliance with the ARRIVE guidelines63. Mice were housed in groups up to 7 animals per cage on poplar chips (SAFE, D0736P00Z) and were fed with irradiated food at 25 kGy (SAFE, #150SP-25). The facility has central air conditioning equipment that maintains a constant temperature of 22 ± 2 °C. Air is renewed at least 20 times per hour in animal rooms. Light is provided with a 14:10-h light:dark cycle (06:30–20:30). Mice were kept in polypropylene or polycarbonate cages that comply with European regulations in terms of floor surface per animal. All cages were covered with stainless steel grids and non-woven filter caps.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files. In addition, the raw datasets generated and/or analyzed during the mass spectrometry proteomic and phosphoproteomic study were deposited in the ProteomeXchange Consortium via the PRIDE76 partner repository with the dataset identifiers PXD038272 (reviewer_pxd038272@ebi.ac.uk; Zc74Ei5B); PXD038270 (reviewer_pxd038270@ebi.ac.uk; Ctnt4Ao0); PXD038268 (reviewer_pxd038268@ebi.ac.uk; FW8rInt) and PXD038267 (reviewer_pxd038267@ebi.ac.uk; blfbVkViU). For RPPA analysis, the raw datasets are available from the corresponding author on reasonable request.
Change history
17 October 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41598-023-44834-9
References
Cancer today. http://gco.iarc.fr/today/home.
Sylvestre, M., Di Carlo, S. E. & Peduto, L. Stromal regulation of the intestinal barrier. Mucosal Immunol. S1933–0219(23), 00006–00015. https://doi.org/10.1016/j.mucimm.2023.01.006 (2023).
Kamali Zonouzi, S., Pezeshki, P. S., Razi, S. & Rezaei, N. Cancer-associated fibroblasts in colorectal cancer. Clin. Transl. Oncol. 24, 757–769 (2022).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Bråten, L. S., Sødring, M., Paulsen, J. E., Snipen, L. G. & Rudi, K. Cecal microbiota association with tumor load in a colorectal cancer mouse model. Microb. Ecol. Health Dis. 28, 1352433 (2017).
Flemer, B. et al. Tumour-associated and non-tumour-associated microbiota in colorectal cancer. Gut 66, 633–643 (2017).
Gagnière, J. et al. Gut microbiota imbalance and colorectal cancer. World J. Gastroenterol. 22, 501–518 (2016).
Lucas, C., Barnich, N. & Nguyen, H. T. T. Microbiota, inflammation and colorectal cancer. Int. J. Mol. Sci. 18, E1310 (2017).
Rebersek, M. Gut microbiome and its role in colorectal cancer. BMC Cancer 21, 1325 (2021).
Janney, A., Powrie, F. & Mann, E. H. Host-microbiota maladaptation in colorectal cancer. Nature 585, 509–517 (2020).
McCOY, W. C. & Mason, J. M. Enterococcal endocarditis associated with carcinoma of the sigmoid; report of a case. J. Med. Assoc. State Ala. 21, 162–166 (1951).
Hoppes, W. L. & Lerner, P. I. Nonenterococcal group-D streptococcal endocarditis caused by Streptococcus bovis. Ann. Intern. Med. 81, 588–593 (1974).
Klein, R. S. et al. Association of Streptococcus bovis with carcinoma of the colon. N. Engl. J. Med. 297, 800–802 (1977).
Waisberg, J., de Matheus, C. O. & Pimenta, J. Infectious endocarditis from Streptococcus bovis associated with colonic carcinoma: Case report and literature review. Arq. Gastroenterol. 39, 177–180 (2002).
Kok, H., Jureen, R., Soon, C. Y. & Tey, B. H. Colon cancer presenting as Streptococcus gallolyticus infective endocarditis. Singap. Med. J. 48, e43-45 (2007).
Gupta, A., Madani, R. & Mukhtar, H. Streptococcus bovis endocarditis, a silent sign for colonic tumour. Colorectal Dis. 12, 164–171 (2010).
Corredoira-Sánchez, J. et al. Association between bacteremia due to Streptococcus gallolyticus subsp. gallolyticus (Streptococcus bovis I) and colorectal neoplasia: A case–control study. Clin. Infect. Dis. 55, 491–496 (2012).
Chand, G., Shamban, L., Forman, A. & Sinha, P. The association of Streptococcus gallolyticus subspecies pasteurianus bacteremia with the detection of premalignant and malignant colonic lesions. Case Rep. Gastrointest. Med. 2016, 7815843 (2016).
Boleij, A., van Gelder, M. M. H. J., Swinkels, D. W. & Tjalsma, H. Clinical importance of Streptococcus gallolyticus infection among colorectal cancer patients: Systematic review and meta-analysis. Clin. Infect. Dis. 53, 870–878 (2011).
Kwong, T. N. Y. et al. Association between bacteremia from specific microbes and subsequent diagnosis of colorectal cancer. Gastroenterology 155, 383-390.e8 (2018).
Périchon, B. et al. Detection of Streptococcus gallolyticus and four other CRC-associated bacteria in patient stools reveals a potential ‘driver’ role for enterotoxigenic Bacteroides fragilis. Front. Cell. Infect. Microbiol. 12, 794391 (2022).
Li, Y. et al. Microbiological and clinical characteristics of Streptococcus gallolyticus subsp. pasteurianus infection in China. BMC Infect. Dis. 19, 791 (2019).
Matesanz, M. et al. Is Streptococcus bovis a urinary pathogen?. Eur. J. Clin. Microbiol. Infect. Dis. 34, 719–725 (2015).
Late-onset Streptococcus pasteurianus sepsis in a preterm baby in a neonatal intensive care unit. http://turkarchpediatr.org/en/late-onset-streptococcus-pasteurianus-sepsis-in-a-preterm-baby-in-a-neonatal-intensive-care-unit-13428.
Binghuai, L., Wenjun, S. & Xinxin, L. Intrauterine infection and post-partum bacteraemia due to Streptococcus gallolyticus subsp. pasteurianus. J. Med. Microbiol. 62, 1617–1619 (2013).
De Vuyst, L. & Tsakalidou, E. Streptococcus macedonicus, a multi-functional and promising species for dairy fermentations. Int. Dairy J. 18, 476–485 (2008).
Maragkoudakis, P. A. et al. In vitro and in vivo safety evaluation of the bacteriocin producer Streptococcus macedonicus ACA-DC 198. Int. J. Food Microbiol. 133, 141–147 (2009).
Pasquereau-Kotula, E., Martins, M., Aymeric, L. & Dramsi, S. Significance of Streptococcus gallolyticus subsp. gallolyticus association with colorectal cancer. Front. Microbiol. 9, 614 (2018).
Boleij, A. et al. Bacterial responses to a simulated colon tumor microenvironment. Mol. Cell. Proteom. 11, 851–862 (2012).
Aymeric, L. et al. Colorectal cancer specific conditions promote Streptococcus gallolyticus gut colonization. Proc. Natl. Acad. Sci. USA 115, E283–E291 (2018).
Abdulamir, A. S., Hafidh, R. R. & Bakar, F. A. Molecular detection, quantification, and isolation of Streptococcus gallolyticus bacteria colonizing colorectal tumors: Inflammation-driven potential of carcinogenesis via IL-1, COX-2, and IL-8. Mol. Cancer 9, 249 (2010).
Kumar, R. et al. Streptococcus gallolyticus subsp. gallolyticus promotes colorectal tumor development. PLoS Pathog. 13, e1006440 (2017).
Kumar, R., Herold, J. L., Taylor, J., Xu, J. & Xu, Y. Variations among Streptococcus gallolyticus subsp. gallolyticus strains in connection with colorectal cancer. Sci. Rep. 8, 1514 (2018).
Taylor, J. C. et al. A type VII secretion system of Streptococcus gallolyticus subsp gallolyticus contributes to gut colonization and the development of colon tumors. PLoS Pathog. 17, e1009182 (2021).
Kumar, R. et al. Modulation of the extracellular matrix by Streptococcus gallolyticus subsp. gallolyticus and importance in cell proliferation. PLoS Pathog. 18, e1010894 (2022).
Zhang, Y., Weng, Y., Gan, H., Zhao, X. & Zhi, F. Streptococcus gallolyticus conspires myeloid cells to promote tumorigenesis of inflammatory bowel disease. Biochem. Biophys. Res. Commun. 506, 907–911 (2018).
Rosenberg, D. W., Giardina, C. & Tanaka, T. Mouse models for the study of colon carcinogenesis. Carcinogenesis 30, 183–196 (2009).
Moser, A. R., Pitot, H. C. & Dove, W. F. A dominant mutation that predisposes to multiple intestinal neoplasia in the mouse. Science 247, 322–324 (1990).
Ren, J., Sui, H., Fang, F., Li, Q. & Li, B. The application of ApcMin/+ mouse model in colorectal tumor researches. J. Cancer Res. Clin. Oncol. 145, 1111–1122 (2019).
McCart, A. E., Vickaryous, N. K. & Silver, A. Apc mice: Models, modifiers and mutants. Pathol. Res. Pract. 204, 479–490 (2008).
Martignetti, L., Calzone, L., Bonnet, E., Barillot, E. & Zinovyev, A. ROMA: Representation and quantification of module activity from target expression data. Front. Genet. 7, 18 (2016).
Pelka, K. et al. Spatially organized multicellular immune hubs in human colorectal cancer. Cell 184, 4734-4752.e20 (2021).
Jacob, J.-M. et al. PDGFRα-induced stromal maturation is required to restrain postnatal intestinal epithelial stemness and promote defense mechanisms. Cell Stem Cell 29, 856-868.e5 (2022).
Davis, H. et al. Aberrant epithelial GREM1 expression initiates colonic tumorigenesis from cells outside the stem cell niche. Nat. Med. 21, 62–70 (2015).
Liu, Y., Li, Y., Hou, R. & Shu, Z. Knockdown GREM1 suppresses cell growth, angiogenesis, and epithelial-mesenchymal transition in colon cancer. J. Cell. Biochem. 120, 5583–5596 (2019).
Kobayashi, H. et al. The balance of stromal BMP signaling mediated by GREM1 and ISLR drives colorectal carcinogenesis. Gastroenterology 160, 1224-1239.e30 (2021).
Gallagher, R. I. & Espina, V. Reverse phase protein arrays: Mapping the path towards personalized medicine. Mol. Diagn. Ther. 18, 619–630 (2014).
Kim, T. W. et al. The role of PDGFRA as a therapeutic target in young colorectal cancer patients. J. Transl. Med. 19, 446 (2021).
Stzepourginski, I. et al. CD34+ mesenchymal cells are a major component of the intestinal stem cells niche at homeostasis and after injury. Proc. Natl. Acad. Sci. USA 114, E506–E513 (2017).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Fujii, M. et al. A colorectal tumor organoid library demonstrates progressive loss of niche factor requirements during tumorigenesis. Cell Stem Cell 18, 827–838 (2016).
van de Wetering, M. et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 161, 933–945 (2015).
d’Aldebert, E. et al. Characterization of human colon organoids from inflammatory bowel disease patients. Front. Cell Dev. Biol. 8, 25 (2020).
Górska, A. & Mazur, A. J. Integrin-linked kinase (ILK): The known vs. the unknown and perspectives. Cell. Mol. Life Sci. 79, 100 (2022).
McDonald, P. C. & Dedhar, S. New perspectives on the role of integrin-linked kinase (ILK) SIGNALING IN CANCER METAStasis. Cancers 14, 3209 (2022).
Dotan, I. et al. The role of integrins in the pathogenesis of inflammatory bowel disease: Approved and investigational anti-integrin therapies. Med. Res. Rev. 40, 245–262 (2020).
Brakebusch, C. & Fässler, R. The integrin-actin connection, an eternal love affair. EMBO J. 22, 2324–2333 (2003).
Marrero, D. et al. Gut-on-a-chip: Mimicking and monitoring the human intestine. Biosens. Bioelectron. 181, 113156 (2021).
Rhee, K.-J. et al. Induction of persistent colitis by a human commensal, enterotoxigenic Bacteroides fragilis, in wild-type C57BL/6 mice. Infect. Immun. 77, 1708–1718 (2009).
Cougnoux, A. et al. Bacterial genotoxin colibactin promotes colon tumour growth by inducing a senescence-associated secretory phenotype. Gut 63, 1932–1942 (2014).
Cuevas-Ramos, G. et al. Escherichia coli induces DNA damage in vivo and triggers genomic instability in mammalian cells. Proc. Natl. Acad. Sci. USA 107, 11537–11542 (2010).
Teh, W. K. et al. Characterization of TelE, an LXG effector of Streptococcus gallolyticus, antagonized by a non-canonical immunity protein. Preprint at https://doi.org/10.1101/2022.09.07.506920 (2022).
Kilkenny, C., Browne, W. J., Cuthill, I. C., Emerson, M. & Altman, D. G. Improving bioscience research reporting: The ARRIVE guidelines for reporting animal research. PLoS Biol. 8, e1000412 (2010).
The, M., MacCoss, M. J., Noble, W. S. & Käll, L. Fast and accurate protein false discovery rates on large-scale proteomics data sets with percolator 3.0. J. Am. Soc. Mass Spectrom. 27, 1719–1727 (2016).
Poullet, P., Carpentier, S. & Barillot, E. myProMS, a web server for management and validation of mass spectrometry-based proteomic data. Proteomics 7, 2553–2556 (2007).
Valot, B., Langella, O., Nano, E. & Zivy, M. MassChroQ: A versatile tool for mass spectrometry quantification. Proteomics 11, 3572–3577 (2011).
Taus, T. et al. Universal and confident phosphorylation site localization using phosphoRS. J. Proteome Res. 10, 5354–5362 (2011).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteom. 13, 2513–2526 (2014).
Josse, J. & Husson, F. missMDA: A package for handling missing values in multivariate data analysis. J. Stat. Softw. 70, 25 (2016).
Akbani, R. et al. A pan-cancer proteomic perspective on The Cancer Genome Atlas. Nat. Commun. 5, 3887 (2014).
Hennessy, B. T. et al. A technical assessment of the utility of reverse phase protein arrays for the study of the functional proteome in non-microdissected human breast cancers. Clin. Proteom. 6, 129–151 (2010).
Neeley, E. S., Baggerly, K. A. & Kornblau, S. M. Surface adjustment of reverse phase protein arrays using positive control spots. Cancer Inform. 11, 77–86 (2012).
Hu, J. et al. Non-parametric quantification of protein lysate arrays. Bioinform. Oxf. Engl. 23, 1986–1994 (2007).
Ju, Z. et al. Development of a robust classifier for quality control of reverse-phase protein arrays. Bioinform. Oxf. Engl. 31, 912–918 (2015).
Gonzalez-Angulo, A. M. et al. Functional proteomics can define prognosis and predict pathologic complete response in patients with breast cancer. Clin. Proteom. 8, 11 (2011).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: A hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).
Acknowledgements
This work was supported by the Institut National contre le Cancer (INCA, Grant PLBIO16-025) and from the French Government’s Investissement d’Avenir program, Laboratoire d’Excellence Integrative Biology of Emerging Infectious Diseases (Grant no. ANR-10-LABX-62-IBEID). EPK received a 2-year Roux-Cantarini post-doctoral fellowship. We are grateful to Johan Bedel and Magali Tichit of the Institut Pasteur Histological Platform for paraffin embedding, sectioning, histochemistry and immunohistochemistry, Karim Sebastien and Marion Bérard for technical help with animal experimentation and all the zootechnicians from the Animal Breeding Facility from Pasteur Institute. We thank Dmitry Ershov for image analysis for DNA damage (yH2AX) quantifications, Adriana Mihalache for her expertise and generous help in tumor grades scoring, Bruno Périchon and Thomas Cokelaer for SGG isolate sequencing, Laurence du Merle for technical assistance with bacterial cultures and CFU counts as well as Valentin Sabatet from the Institut Curie Proteomics Platform for protein quantification and Olivier Mirabeau for bioinformatic help.
Author information
Authors and Affiliations
Contributions
E.P.-K. and S.D. conceived the project and designed the experiments; E.P.-K., G.N., F.D., S.K. and J.D. performed the experiments; E.P.-K., G.N., L.D., P.P., Y.X., L.P. analyzed the data; L.D. and F.D. performed the proteomics and phosphoproteomics analyses; P.P. performed bioinformatic analyses; C.R.-M., P.S., P.T.-C., Y.X. and S.D. were involved in funding acquisition. E.P.-K. and S.D., wrote the paper. All authors edited and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: In the original version of this Article the authors’ given names and surnames were reversed.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pasquereau-Kotula, E., Nigro, G., Dingli, F. et al. Global proteomic identifies multiple cancer-related signaling pathways altered by a gut pathobiont associated with colorectal cancer. Sci Rep 13, 14960 (2023). https://doi.org/10.1038/s41598-023-41951-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-41951-3
- Springer Nature Limited