Abstract
Malaria is a major health problem in Peru despite substantial progress achieved by the ongoing malaria elimination program. This study explored the population genetics of 63 Plasmodium falciparum and 170 P. vivax cases collected in the Peruvian Amazon Basin between 2015 and 2019. Microscopy and PCR were used for malaria detection and positive samples were genotyped at neutral and drug resistance-associated regions. The P. falciparum population exhibited a low nucleotide diversity (π = 0.02) whereas the P. vivax population presented a higher genetic diversity (π = 0.34). All P. falciparum samples (n = 63) carried chloroquine (CQ) resistant mutations on Pfcrt. Most P. falciparum samples (53 out of 54) carried sulfadoxine (SD) resistant mutations on Pfdhfr and Pfdhps. No evidence was found of artemisinin resistance mutations on kelch13. Population structure showed that a single cluster accounted for 93.4% of the P. falciparum samples whereas three clusters were found for P. vivax. Our study shows a low genetic diversity for both species with significant differences in genetic sub-structuring. The high prevalence of CQ-resistance mutations could be a result of indirect selection pressures driven by the P. vivax treatment scheme. These results could be useful for public health authorities to safeguard the progress that Peru has achieved towards malaria elimination.
Similar content being viewed by others
Introduction
Malaria remains a major health problem in tropical and subtropical regions causing more than 241 million cases and over 627,000 deaths during 20201. Despite major progress achieved towards malaria elimination in the Americas, this disease still puts at risk of infection to nearly 138 million people leading to 889,000 new cases each year. Furthermore, up to 90% of these cases occur in Amazonian regions of Venezuela, Brazil, Colombia and Peru2.
In Peru, the Amazonian region of Loreto accounts for 84% of the 15,519 malaria cases reported in 2020 with most of those caused by Plasmodium vivax followed by P. falciparum (ratio Pv/Pf = 4)3. Malaria transmission is perennial in this region, with seasonal increases of cases between February and July4.
The malaria treatment policy for uncomplicated P. falciparum infections consists of mefloquine (MQ, 12.5 mg/kg/day for 2 days) plus artesunate (AS, 4 mg/kg/day for 3 days). In contrast, treatment for P. vivax comprises chloroquine for three days (CQ, 10 mg/kg/day the first two days and 5 mg/kg/day on the third day) plus primaquine (PQ, 0.5 mg/kg/day for 7 days). The change to MQ-AS was implemented nearly two decades ago due to the widespread prevalence of CQ resistance in P. falciparum5.
In Peru, malaria incidence has presented several fluctuations over the last two decades. One of these fluctuations occurred during the implementation of the Global Fund's Malaria Project “PAMAFRO” (2005–2011)6. This project aimed to increase access and quality to microscopy diagnosis, treatment and capacity building. Due to PAMAFRO activities, the number of malaria cases reported per year in Peru declined from 87,805 cases in 2005 to 23,060 cases in 2011. The prevalence reduction was higher for P. falciparum (85%) than P. vivax (63%)7. However, after PAMAFRO ended, interventions were not sustained, and malaria cases substantially increased from 23,060 cases in 2011 to 56,530 cases in 20167.
In 2017, the Peruvian government launched a malaria elimination initiative called Plan Malaria Cero (PMC) aiming to eliminate malaria within 25 years from the Loreto region. PMC has three phases: Phase I, malaria control; II, towards malaria elimination; and III, residual malaria elimination8. During Phase I, the malaria cases in 2020 were reduced by more than 75% compared to 2017 (15,519 vs 55,227, respectively). PMC's phase II has ahead multiple challenges, e.g., increasing focal cases distribution, transmission shift to rural and remote settings, an increment in the prevalence of asymptomatic and submicroscopic cases among others7. Thanks to the successful control outcome achieved by PMC, last January 2022, the government extended it to a national level initiative for malaria elimination for 2030.
As PMC's elimination efforts continue, they will impact circulating parasite populations as previously shown9,10,11,12. In this regard, malaria population genetic studies conducted in the Peruvian Amazon showed a higher population heterogeneity in P. vivax transmission than in P. falciparum13. Furthermore, communities with variable connectedness showed high diversity and polyclonal infections ranging between 44 and 70%14 compared to isolated communities characterized by limited genetic diversity and low frequency of polyclonal infections (14–19.7%)15,16,17. Regarding P. falciparum, parasite populations probably expanded from bottlenecked populations after malaria eradication efforts from 1966 to 198910. These Peruvian P. falciparum populations consisted of at least five clonal lineages and presented a low proportion of polyclonal infections10.
Plasmodium genomic plasticity is an important threat for malaria elimination as it can lead to the relatively rapid development of drug resistance against antimalarials18. In addition, migration, climate change and human-driven activities can enhance drug resistance spread or introduce new strains across regions11,12,19. Therefore, it is key to implement stronger surveillance platforms in endemic settings that can allow for early detection of drug resistance or emergence of virulent strains. This is particularly important in Peru due to the lack of data after the implementation of new malaria elimination policies in 2001.
This study, explored the population genetics and molecular surveillance of mutations associated with drug-resistance in P. falciparum and P. vivax parasites circulating in the city of Iquitos and surrounding communities of the Loreto region, Peru, between 2015 and 2019. The information provided by this study will contribute with relevant information to guide malaria elimination efforts in Peru.
Materials and methods
Study sites, sample selection
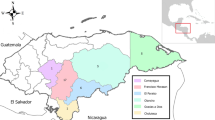
The Loreto region is located in the Northeast Peruvian Amazon and is characterized by an equatorial climate with annual temperatures between 24 and 33 °C. Blood spot samples for this study were collected from sixteen study sites in Iquitos (Loreto's capital city) and surrounding communities as part of two surveillance projects conducted between 2015 and 2019 (Fig. 1). The sample collection was conducted by U.S. Naval Medical Research Unit 6 (NAMRU-6) and Universidad Peruana Cayetano Heredia (UPCH).
Map of the study sites located in the region of Loreto. The inset shows the location of the field sites in and around Iquitos city depicted in blue marks. The map was created using ArcGIS online (ESRI Inc. Redlands, CA, USA. https://www.esri.com/) using open data obtained from GADM database of Global Administrative Areas, version 3.6. www.gadm.org.
The NAMRU-6 project was a passive surveillance study that enrolled people > 1-year-old with of fever or history of fever during the previous 72 h in different health centers across the city. The UPCH project was an active surveillance study carried out in 2018 to screen high-risk malaria populations in various communities. Inhabitants from Quistococha, Santo Tomas and Rumococha communities were enrolled in April from Gamitanacocha, Libertad, 1 de Enero, Salvador, Lago Yuracyacu, Puerto Alegre, and Urcomiraño communities in July 2018.
Ethics
The samples tested for this study were selected from two sources: i) the NAMRU-6 project which was approved by the Institutional Review Board of the U.S Naval Medical Research Unit 6 (NAMRU-6) in compliance with all applicable federal regulations governing the protection of human subjects (protocol NMRCD.2007.0004) and ii) The UPCH projects which was approved by the Institutional Ethical Review Board of the Universidad Peruana Cayetano Heredia (protocols SIDISI 101645/2017 and SIDISI 101518/2018). Informed consent was obtained from all participants and/or their legal guardians. All methods were performed in accordance with the relevant guidelines and regulations.
Sample processing and malaria diagnostics
Two thin and thick smears were prepared for each participant and stained with Giemsa. Slides were read by two microscopists whereas a third microscopist reviewed slides with discordant results.
DNA from whole blood was extracted using the Qiagen DNA extraction kit according to the manufacturer’s protocol. Malaria-positive cases were detected by Malachite green LAMP or real-time PCR as previously described20,21,22. For MG-LAMP, the reaction was performed in a 20 µL reaction volume that contained 5 µL of template DNA in 2X in-house reaction buffer (40 mM Tris–HCl pH 8.8, 20 mM KCl, 16 mM MgSO4, 20 mM (NH4)2SO4, 0.2% Tween -20, 1.6 M Betaine, 2 mM of dNTP's each), 0.25µL of 1:400 SYTO 9 dye, 8 units of Bst Polymerase (New England Biolabs, Ipswich, MA) and 0.004% Malachite Green dye. DNA amplification was carried out at 63 °C for 60 min and two independent readers visually inspected the results after 15 min post amplification. For the real-time PCR, the reaction was performed in a 25 µL reaction volume that contained 5 µL of template DNA in a 1X master mix of PerfeCTa SYBR® Green Fastmix, 0.3 μM of each primer PL1473F18 5′-TAACGAACGAGATCTTAA-3′ and PL1679R18 5′-GTTCCTCTAAGAAGCTTT-3′. The real-time PCR conditions consisted of an initial denaturation step at 95 °C for 2 min, followed by 45 cycles of 20 s at 95 °C, 20 s at 50 °C, and 20 s at 68 °C.
Sequencing and SNP genotyping
Blood spots from malaria-positive samples were sent to the Welcome Sanger Institute for sequencing, evaluation of drug resistance markers and barcodes generation23. A selective whole genome amplification was used to enrich target regions of DNA by multiplex PCR. Additionally, an extra round of PCR was done to incorporate adapters on both P. falciparum and P. vivax and subsequently, these were pooled and sequenced on Illumina MiSeq sequencer. Finally, reads from each sample were aligned onto the P. falciparum 3D7 or P. vivax P01 reference sequences.
Sample barcodes were generated for P. falciparum (101 SNPs) and P. vivax (38 SNPs). The barcodes consist of biallelic SNPs selected from the malaria genome variation database based on their usefulness to evaluate parasite interrelationship. Barcodes are distributed across the P. vivax and P. falciparum genomes and have not been associated to drug resistance23,24.
Drug resistance markers
In P. vivax, three drug-resistance markers were evaluated: Pvdhfr (positions 57, 58, 61 and 117) and Pvdhps (380, 382, 383, 385 and 553) associated with sulfadoxine-pyrimethamine resistance and Pvmdr1 (976) that is putatively associated with CQ resistance25,26,27. In the case of P. falciparum, mutations in nine markers were evaluated: arps10 (positions 127 and 128), ferredoxin (193), Pfcrt (326 and 356), k13 (BTB/POZ and propeller) and Pfmdr2 (484) associated with artemisinin resistance28,29, Pfdhfr (51, 59, 108 and 164) and Pfdhps (436, 437, 540, 581 and 613) associated with SP resistance30,31,32,33, the exonuclease gene, Pfexo (415) associated with piperaquine resistance, Pfcrt (72, 73, 74, 75 and 76) associated with chloroquine resistance and Pfmdr1 (86, 184 and 1246) associated with CQ, amodiaquine, lumefantrine and MQ resistance34,35,36,37,38,39. Markers were combined for each sample in order to assess the prevalence of common drug resistance haplotypes.
Data analysis, multiplicity of infection and population diversity
Complexity of infection (COI) was analyzed by a Markov Chain Monte Carlo (MCMC) method using COIL and Real McCOIL which estimate the proportion of heterozygous calls40,41.
Barcodes were subsequently filtered using the R package poppr42 to exclude samples with > 20% missing calls and positions with missing calls in > 20% of the samples. The resulting barcoding data was used to estimate the population barcode diversity (π) using 10,000 iterations of nonparametric bootstrapping24. Moreover, an Analysis of Molecular Variance (AMOVA) was performed to estimate the strength of genetic differentiation among Plasmodium populations using Arlequin 3.5 by a locus-by-locus analysis with 1000 permutations43.
Phylogenetic and Principal Component Analysis
Phylogenetic reconstruction was carried out using only monoclonal samples using a maximum likelihood approach in PhyML v3.044 with 1000 bootstrap and under the best-fit model defined by Bayesian information criterion obtained from jModelTest 2.1.545. The P. falciparum 3D7 and P. vivax P01 strains were used to root the phylogenetic trees visualized in iTOL46.
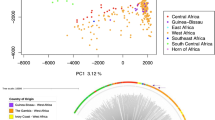
In addition, principal components analysis (PCA) was carried out on monoclonal samples using the R package adegenet47 to estimate the parasite population structure. To evaluate the relationship among P. falciparum haplotypes, a phylogenetic network was built adding control samples from Africa (n = 11), South East Asia (n = 10) and South America (n = 15) on PopART48 using the median-joining algorithm. This method was also employed for P. vivax adding control samples from Colombia (n = 26) and Honduras (n = 18).
Epidemiological analysis
Clinical and epidemiological data were analyzed in Stata 16. The potential association of clinical data, disease severity, sociodemographic data and laboratory results with specific P. vivax parasite populations was analyzed using Fisher's exact test or Kruskal–Wallis test.
Results
Sample collection and sociodemographic data
A total of 67 participants positive for P. falciparum and 170 positives for P. vivax were enrolled in both studies (Table 1). Up to 61.5% (217 out of 237) of all participants were enrolled in 2018. The median participant age was 27 years old (IQR: 18–46 y.o) and mainly males (66.2%) for P. vivax and 37 years old (IQR: 22–58 y.o) and 62.7% males in P. falciparum.
Regarding households, 45.3% of the P. vivax and 38.6% of participants carrying P. falciparum reported having a family member with malaria in the last year. More than 50% of participants carrying P. vivax presented at least three malaria episodes in the previous 10 years whereas 50% of participants carrying P. falciparum presented at least two episodes during the same period.
The most frequent clinical symptoms in the study population were headache, malaise, chills, sweating and fever. There were not significant differences in clinical symptoms between people infected with P. vivax or with P. falciparum.
Haplotypes and drug resistance polymorphisms
SNPs genotyping of P. falciparum showed that 100% of the samples (n = 52) carried the 184F/1246Y double Pfmdr1 mutation. The SVMNT haplotype on Pfcrt was present in 98.4% of the samples (63/64). None of the samples presented parasite genetic background mutations on Pfarp10. One sample had a 617S mutation in Pfk13 whereas the rest were wild-type.
Most P. falciparum samples (98%; 53/54) had quadruple mutations at Pfdhfr and Pfdhps (51I/108N + 437G/540E), which is a combination that has been associated with SP treatment failure32. In addition, our study did not find mutations on PfEXO associated with piperaquine resistance nor FERREDOXIN and Pfmdr2 associated with artemisinin resistance (Tables S1 and S2)29,33. The most frequent haplotype for this species based on Pfdhfr, Pfdhps, Pfcrt, PfEXO, Pfmdr1, Pfarps-10, FERREDOXIN and Pfmdr2 (ICNISGEGASVMNTDLENFYVDDT) accounted for more than 57% of the samples (Table S2).
The 976F mutation on Pvmdr1 was prevalent on 4.1% P. vivax infections (7 out of 170). For Pvdhfr, 0.6% cases (1/161) presented the 57L mutation, 97.5% (154/158) were mutants on position 58 (R, K or L), and 97% (165/170) presented the 117 N mutation with 48% (76 out of 158) harboring the double mutation S58R and S117N. For Pvdhps, 72.3% (123/170) of the infections presented the 383G mutation. All infections carried wildtype parasites for Pvdhps for positions 385, 553 (Tables S3 and S4). The most frequent P. vivax haplotype based on Pvdhfr, Pvdhps and Pvmdr1 (FKTNECGYAY) accounted for 31% of the samples and the next prevalent haplotypes accounted for 15,9% (FRTNECGYAY) and 12.4% (FRTNESAYAY) of the samples (Table S3).
Population diversity and complexity of infection
All P. falciparum samples (n = 67) were monoclonal by COIL and real McCOIL methods. The barcode intrapopulation diversity for this species was π = 0.02 (95%CI: 0.007–0.039). In P. vivax, 10 out of 170 samples were polyclonal infections (10.6%) and the overall intrapopulation diversity was higher than P. falciparum with π = 0.34 (95%CI: 0.309–0.381).
Population structure
Data cleaning resulted in 46 P. falciparum and 145 P. vivax samples passing the quality filters for population analysis and two loci in P. falciparum were removed. One sample (MDP4546) was excluded from DAPC due to being very different from the rest of P. falciparum.
DAPC on the rest of the samples did not show population sub-structuring related to geographic origin for both P. vivax (Fig. 2A) and P. falciparum (Fig. 2B). AMOVA's showed that 98% and 100% of the genetic variation relied within P. vivax and P. falciparum’s populations (p-value = 0.13 and 0.69, respectively) (Tables S5 and S6).
Discriminant analysis of principal components (DAPC) P. falciparum and P. vivax. The figure shows the population structure of 46 P. falciparum (a) and 145 P. vivax samples (b). The color scheme corresponds to each of the sites where samples were collected. DAPC shows the absence of clustering due to geographical origin.
In P. falciparum, median-joining network showed an apparent clustering of parasites according to the country of isolation (Fig. 3A). Phylogenetic analyses showed that sample MDP4546 was distant from the rest of Peruvian strains (Fig. 3B).
Phylogenetics results. (a) Median-joining network of Peruvian P. falciparum strains and control samples from South America (Honduras and Colombia). The network shows the distribution of lineages according to their barcode and the geographical clustering of P. falciparum lines. Each circle represents an independent haplotype, the lines connect nearby haplotypes and the cross line represents one non-synonymous mutation. Figure created in PopART48. (b) Maximum likelihood phylogenetic analysis of P. falciparum samples. Circles in tree nodes represent bootstrap support values, 3D7 denotes P. falciparum 3D7 strain. Figure generated in Figtree (http://tree.bio.ed.ac.uk/software/figtree/).
After removing sample MDP4546, K-means clustering showed two subpopulations with no evidence of admixed individuals (Fig. 4). One of these subpopulations accounted for 95.5% of all P. falciparum samples (43 out of 45 samples).
In the case of P. vivax, K-means clustering and phylogenetic analysis revealed the presence of three subpopulations with 21 admixed individuals (Fig. 5). The median-joining network for P. vivax showed that some samples from Colombia were closer to Peru than to the rest of samples from Colombia (Supplementary Fig. 1).
Phylogenetic relationship and population structure of P. vivax. Inset A shows the Phylogenetic three for P. vivax. Black colored circles denote bootstrap support > 700. Colored circles on tree tips match the three clusters defined by DPCA. P01 denotes the Plasmodium vivax P01 reference. Figure generated in Figtree (http://tree.bio.ed.ac.uk/software/figtree/). Inset B Shows a DAPC analysis with K-means clustering for P. vivax. The y-axis denotes the membership probability of each sample to one of the three clusters defined by this analysis. Figure created musing the adegenet package47.
Malaria population structure matched with the drug resistance haplotypes for both P. vivax and P. falciparum. In this sense, most samples of the P. falciparum predominant cluster carried point mutations on genes associated with SP, MQ and CQ resistance.
In P. vivax, the three subpopulations matched with three haplotypes associated with SP resistance as previously shown. In this regard, significant differences were found between the three P. vivax genetic clusters for putative drug-resistance associated SNPs for positions Pvdhfr:58, Pvmdr1:976, Pvdhps:382 and Pvdhps:383 (Table 2) with populations 2 and 3 presenting higher prevalences for these SNPs than population 1.
Discussion
In response to the malaria threat, Peru’s launched in 2017 a malaria elimination plan called Plan Malaria Cero 8, and in 2022 the program has expanded to a national level49. PMC advances caused significant malaria transmission disruptions and a potential parasite population bottleneck. In this regard, our results show low genetic diversity in both species with a single predominant cluster of P. falciparum and sub-structured P. vivax populations in the peri-urban settings of Iquitos (Loreto, Peru). PMC control activities in Peru had a higher impact against the P. falciparum population than P. vivax as previously shown in other regions50. This difference could be due to the lower prevalence and absence of hypnozoites in P. falciparum51,52. In addition, the lower treatment adherence for P. vivax in Peru53,54 may have also played a role in the differential scenarios for both species51,52.
Clinical and epidemiological data derived from our study showed no significant differences between participants carrying P. vivax and P. falciparum parasites. In this regard, most subjects were male with the most common symptoms being a headache, malaise, chills, fever and sweating which were present in more than 85% of all subjects. This lack of severity in P. falciparum and P. vivax is characteristic in the region compared to Africa or Southeast Asia where P. falciparum is associated with more severe symptoms55.
Previous studies have shown that antifolate drugs have exerted strong selective pressures on P. vivax56,57. In this regard, our results show a high proportion of putative SP resistance markers 58R and 117N on Pvdhfr and 383G Pvdhps25,26,27. This high proportion could result from indirect selection pressure from P. falciparum treatment. Noteworty, SP was removed as first line treatment against P. falciparum in the study region in 200158. Therefore, Pvdhfr and Pvdhps mutations remain fixed for more than 20 years. In the case of Pvmdr1, there is a low prevalence of the putative CQ resistant marker 976F in the P. vivax population (4.7%). However, it is important to notice that the role of the Y976F on CQ resistance remains under debate27,59,60.
Currently, there is concern regarding the spread of artemisinin resistance which could halter malaria elimination efforts around the globe61. Our study did not find evidence of artemisinin resistance nor related mutations on P. falciparum in Iquitos. This result and the lack of reports of clinical resistance nor delayed parasite clearance indicate that artemisinin remains effective in the region. Although continuing surveillance and increased sample sizes are needed to detect early resistance signals.
Regarding other drug resistance markers and haplotypes, our results show that most P. falciparum parasites presented a profile of drug resistance mutations compatible with the foreign PfBv1 lineage. This profile includes the CQ resistance 184F/1246Y double mutation on Pfmdr162 and the SVMNT haplotype on Pfcrt34 and the SP resistant quadruple mutation 51I/108N + 437G/540E at Pfdhfr and Pfdhps32. This lineage was found on a P. falciparum outbreak in the North Coast of Peru between 2010 and 201212 and in another outbreak in the region of Cusco in 201311. In this regard, our population structure results for P. falciparum indicate that up to 95% of our samples belong to PfBv1 which appears to have become the predominant lineage in the study region.
Our results show that most samples were monoclonal and there was a low intrapopulation diversity (π) for both species. However, P. falciparum had a much lower value than P. vivax which indicates that circulating P. falciparum are less admixed and predominantly clonal. This result is consistent with the population structure results for both species which did not show clustering according to geographical locations and the presence of a single P. falciparum cluster that accounted for 95% of all infections. The differential collection types of the samples plus the restricted number of samples could mask sub-structuring due to geographical origin. In the case of P. vivax, our results reflect the higher diversity with three different circulating genetic clusters, which share the same habitats and 21 admixed individuals. This could reflect a scenario with more gene flow between the current Amazonian communities compared to results from previous years probably due to higher connectivity between communities15,16. Furthermore, the haplotype network for P. vivax showed a closer relationship with some Colombian samples. This finding points towards genetic exchange across these countries derived from human migration and high connectivity across the Peruvian/Colombian borders.
Malaria elimination programs face the challenge of drug resistance7 and introduction of new parasite lineages63. In this regard, the P. falciparum predominant cluster (Bv1) could have been introduced from Brazil or Bolivia where the genotypes Pfdhr 50R, 51I and 108N and Pfcrt 72S, 74M, 75N, 76T, 326D and 356L were initially described9,34,63. There is evidence that parasites harboring this haplotype have displaced local populations and caused outbreak in different endemic regions in South America10,11,12,63.
In P. vivax, the three most frequent haplotypes represented more than 59% of all our samples and the prevalence of these populations was associated with Pvdhfr and Pvdhps mutations. In this regard, the presence of these mutations could have provided a selective advantage over other circulating populations during the period of SP use for P. falciparum and mixed infections in the Peruvian Amazon (1996–2001) and in the Peruvian North Coast (1996–2015)58,64.
Unfortunately, analyses of P. vivax in the context of our study are limited due to the short barcode for this species (38 SNPs). Also, our study was restricted to a brief period of time and therefore we cannot assess the dynamics of circulating parasite populations over time. Finally, our study was executed prior the COVID 19 pandemic and it is highly likely that disruptions derived from the pandemic could have impacted the dynamics of malaria transmission in the region.
In summary, our study shows a low genetic diversity for both species, sub-structured P. vivax populations, clonal propagation in P. falciparum and absence of artemisinin resistance mutations in the peri-urban settings of Iquitos. The lack of P. vivax sub-structuring due to geographic origin reflects a higher rate of gene flow among the geographic units compared to previous years. Therefore, the new malaria elimination plan must consider developing strategies that address human mobility as a high risk for malaria transmission and persistence. Moreover, our findings support the continuity of ACTs in the region and highlight the need to adapt the elimination strategies to decreased P. vivax incidence.
Data availability
The dataset generated during and/or analyzed during the current study is available from the corresponding author on reasonable request. Raw sequence data has been deposited at the European Nucleotide Archive (https://www.ebi.ac.uk/ena/browser/home) under primary accession numbers from ERS2741604 to ERS3516726.
References
WHO. World malaria report 2021. Geneva: WHO (2021).
Recht, J. et al. Malaria in Brazil, Colombia, Peru and Venezuela: Current challenges in malaria control and elimination. Malar. J. 16, 273. https://doi.org/10.1186/s12936-017-1925-6 (2017).
CDC-Peru. Numero de Casos de Malaria, Peru 2015–2020, https://www.dge.gob.pe/portal/ (2020).
Branch, O. et al. Clustered local transmission and asymptomatic Plasmodium falciparum and Plasmodium vivax malaria infections in a recently emerged, hypoendemic Peruvian Amazon community. Malar. J. 4, 27. https://doi.org/10.1186/1475-2875-4-27 (2005).
Mejia Torres, R. E. et al. Efficacy of chloroquine for the treatment of uncomplicated Plasmodium falciparum malaria in Honduras. Am. J. Trop. Med. Hygiene 88, 850–854. https://doi.org/10.4269/ajtmh.12-0671 (2013).
Feo Istúriz, O. et al. Compartiendo lecciones aprendidas. Proyecto control de malaria en zonas fronterizas de la región andina: un enfoque comunitario-PAMAFRO. (2009).
Rosas-Aguirre, A. et al. Epidemiology of Plasmodium vivax Malaria in Peru. Am. J. Trop. Med. Hyg. 95, 133–144. https://doi.org/10.4269/ajtmh.16-0268 (2016).
Pardo, K. Plan de Eliminación de la Malaria en Loreto (Plan Malaria Cero 2017–2021). (Dirección de Prevención y Control de Enfermedades Metaxenicas y Zoonosis, 2021).
Bacon, D. J. et al. Dynamics of malaria drug resistance patterns in the Amazon basin region following changes in Peruvian national treatment policy for uncomplicated malaria. Antimicrob. Agents Chemother. 53, 2042–2051. https://doi.org/10.1128/AAC.01677-08 (2009).
Griffing, S. M. et al. South American Plasmodium falciparum after the malaria eradication era: Clonal population expansion and survival of the fittest hybrids. PLoS ONE 6, e23486. https://doi.org/10.1371/journal.pone.0023486 (2011).
Okoth, S. A. et al. Molecular investigation into a Malaria outbreak in Cusco, Peru: Plasmodium falciparum BV1 lineage is linked to a second outbreak in recent times. Am. J. Trop. Med. Hyg. 94, 128–131. https://doi.org/10.4269/ajtmh.15-0442 (2016).
Baldeviano, G. C. et al. Molecular epidemiology of Plasmodium falciparum Malaria Outbreak, Tumbes, Peru, 2010–2012. Emerg. Infect. Dis. 21, 797–803. https://doi.org/10.3201/eid2105.141427 (2015).
Sutton, P. L., Neyra, V., Hernandez, J. N. & Branch, O. H. Plasmodium falciparum and Plasmodium vivax infections in the Peruvian Amazon: Propagation of complex, multiple allele-type infections without super-infection. Am. J. Trop. Med. Hyg. 81, 950–960. https://doi.org/10.4269/ajtmh.2009.09-0132 (2009).
Van den Eede, P. et al. Multilocus genotyping reveals high heterogeneity and strong local population structure of the Plasmodium vivax population in the Peruvian Amazon. Malar. J. 9, 151. https://doi.org/10.1186/1475-2875-9-151 (2010).
Delgado-Ratto, C. et al. Population genetics of Plasmodium vivax in the Peruvian Amazon. PLoS Negl. Trop. Dis. 10, e0004376. https://doi.org/10.1371/journal.pntd.0004376 (2016).
Manrique, P. et al. Microsatellite analysis reveals connectivity among geographically distant transmission zones of Plasmodium vivax in the Peruvian Amazon: A critical barrier to regional malaria elimination. PLoS Negl. Trop. Dis. 13, e0007876. https://doi.org/10.1371/journal.pntd.0007876 (2019).
Delgado-Ratto, C. et al. Population structure and spatio-temporal transmission dynamics of Plasmodium vivax after radical cure treatment in a rural village of the Peruvian Amazon. Malar. J. 13, 8. https://doi.org/10.1186/1475-2875-13-8 (2014).
Rathod, P. K., McErlean, T. & Lee, P. C. Variations in frequencies of drug resistance in Plasmodium falciparum. Proc. Natl. Acad. Sci. USA 94, 9389–9393 (1997).
Larranaga, N. et al. Genetic structure of Plasmodium falciparum populations across the Honduras-Nicaragua border. Malar. J. 12, 354. https://doi.org/10.1186/1475-2875-12-354 (2013).
Lucchi, N. W., Ljolje, D., Silva-Flannery, L. & Udhayakumar, V. Use of malachite green-loop mediated isothermal amplification for detection of Plasmodium spp. parasites. PLoS ONE 11, e0151437. https://doi.org/10.1371/journal.pone.0151437 (2016).
Barazorda, K. A., Salas, C. J., Bishop, D. K., Lucchi, N. & Valdivia, H. O. Comparison of real time and malachite-green based loop-mediated isothermal amplification assays for the detection of Plasmodium vivax and P. falciparum. PLoS ONE 15, e0234263. https://doi.org/10.1371/journal.pone.0234263 (2020).
Mangold, K. A. et al. Real-time PCR for detection and identification of Plasmodium spp. J. Clin. Microbiol. 43, 2435–2440. https://doi.org/10.1128/JCM.43.5.2435-2440.2005 (2005).
Jacob, C. G. et al. Genetic surveillance in the Greater Mekong subregion and South Asia to support malaria control and elimination. Elife https://doi.org/10.7554/eLife.62997 (2021).
Baniecki, M. L. et al. Development of a single nucleotide polymorphism barcode to genotype Plasmodium vivax infections. PLoS Negl. Trop. Dis. 9, e0003539. https://doi.org/10.1371/journal.pntd.0003539 (2015).
Marfurt, J. et al. Molecular markers of in vivo Plasmodium vivax resistance to amodiaquine plus sulfadoxine-pyrimethamine: Mutations in pvdhfr and pvmdr1. J. Infect. Dis. 198, 409–417. https://doi.org/10.1086/589882 (2008).
Korsinczky, M. et al. Sulfadoxine resistance in Plasmodium vivax is associated with a specific amino acid in dihydropteroate synthase at the putative sulfadoxine-binding site. Antimicrob. Agents Chemother. 48, 2214–2222. https://doi.org/10.1128/AAC.48.6.2214-2222.2004 (2004).
Suwanarusk, R. et al. Chloroquine resistant Plasmodium vivax: In vitro characterisation and association with molecular polymorphisms. PLoS ONE 2, e1089. https://doi.org/10.1371/journal.pone.0001089 (2007).
Ariey, F. et al. A molecular marker of artemisinin-resistant Plasmodium falciparum malaria. Nature 505, 50–55. https://doi.org/10.1038/nature12876 (2014).
Miotto, O. et al. Genetic architecture of artemisinin-resistant Plasmodium falciparum. Nat. Genet. 47, 226–234. https://doi.org/10.1038/ng.3189 (2015).
Peterson, D. S., Walliker, D. & Wellems, T. E. Evidence that a point mutation in dihydrofolate reductase-thymidylate synthase confers resistance to pyrimethamine in falciparum malaria. Proc. Natl. Acad. Sci. USA 85, 9114–9118. https://doi.org/10.1073/pnas.85.23.9114 (1988).
Foote, S. J., Galatis, D. & Cowman, A. F. Amino acids in the dihydrofolate reductase-thymidylate synthase gene of Plasmodium falciparum involved in cycloguanil resistance differ from those involved in pyrimethamine resistance. Proc. Natl. Acad. Sci. USA 87, 3014–3017. https://doi.org/10.1073/pnas.87.8.3014 (1990).
Picot, S. et al. A systematic review and meta-analysis of evidence for correlation between molecular markers of parasite resistance and treatment outcome in falciparum malaria. Malar. J. 8, 89. https://doi.org/10.1186/1475-2875-8-89 (2009).
Amato, R. et al. Genetic markers associated with dihydroartemisinin-piperaquine failure in Plasmodium falciparum malaria in Cambodia: A genotype-phenotype association study. Lancet Infect Dis. 17, 164–173. https://doi.org/10.1016/S1473-3099(16)30409-1 (2017).
Fidock, D. A. et al. Mutations in the P. falciparum digestive vacuole transmembrane protein PfCRT and evidence for their role in chloroquine resistance. Mol. Cell. 6, 861–871. https://doi.org/10.1016/s1097-2765(05)00077-8 (2000).
Foote, S. J. et al. Several alleles of the multidrug-resistance gene are closely linked to chloroquine resistance in Plasmodium falciparum. Nature 345, 255–258. https://doi.org/10.1038/345255a0 (1990).
Venkatesan, M. et al. Polymorphisms in Plasmodium falciparum chloroquine resistance transporter and multidrug resistance 1 genes: parasite risk factors that affect treatment outcomes for P. falciparum malaria after artemether-lumefantrine and artesunate-amodiaquine. Am. J. Trop. Med. Hygiene 91, 833–843. https://doi.org/10.4269/ajtmh.14-0031 (2014).
Veiga, M. I. et al. Globally prevalent PfMDR1 mutations modulate Plasmodium falciparum susceptibility to artemisinin-based combination therapies. Nat. Commun. 7, 11553. https://doi.org/10.1038/ncomms11553 (2016).
Malmberg, M. et al. Plasmodium falciparum drug resistance phenotype as assessed by patient antimalarial drug levels and its association with pfmdr1 polymorphisms. J. Infect. Dis. 207, 842–847. https://doi.org/10.1093/infdis/jis747 (2013).
Reed, M. B., Saliba, K. J., Caruana, S. R., Kirk, K. & Cowman, A. F. Pgh1 modulates sensitivity and resistance to multiple antimalarials in Plasmodium falciparum. Nature 403, 906–909. https://doi.org/10.1038/35002615 (2000).
Chang, H. H. et al. THE REAL McCOIL: A method for the concurrent estimation of the complexity of infection and SNP allele frequency for malaria parasites. PLoS Comput. Biol. 13, e1005348. https://doi.org/10.1371/journal.pcbi.1005348 (2017).
Galinsky, K. et al. COIL: A methodology for evaluating malarial complexity of infection using likelihood from single nucleotide polymorphism data. Malar. J. 14, 4. https://doi.org/10.1186/1475-2875-14-4 (2015).
Kamvar, Z. N., Tabima, J. F. & Grunwald, N. J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2, e281. https://doi.org/10.7717/peerj.281 (2014).
Excoffier, L. & Lischer, H. E. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567. https://doi.org/10.1111/j.1755-0998.2010.02847.x (2010).
Criscuolo, A. morePhyML: Improving the phylogenetic tree space exploration with PhyML 3. Mol. Phylogenet. Evol. 61, 944–948. https://doi.org/10.1016/j.ympev.2011.08.029 (2011).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: More models, new heuristics and parallel computing. Nat. Methods 9, 772. https://doi.org/10.1038/nmeth.2109 (2012).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296. https://doi.org/10.1093/nar/gkab301 (2021).
Jombart, T. adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405. https://doi.org/10.1093/bioinformatics/btn129 (2008).
Leigh, J. W. & Bryant, D. POPART: Full-feature software for haplotype network construction. Methods Ecol. Evol. 6, 1110–1116 (2015).
MINSA. Resolucion Ministerial 034–2022-MINSA (2024).
Ome-Kaius, M. et al. Differential impact of malaria control interventions on P. falciparum and P. vivax infections in young Papua New Guinean children. BMC Med. 17, 220. https://doi.org/10.1186/s12916-019-1456-9 (2019).
Betuela, I. et al. Relapses contribute significantly to the risk of Plasmodium vivax infection and disease in Papua New Guinean children 1–5 years of age. J. Infect Dis. 206, 1771–1780. https://doi.org/10.1093/infdis/jis580 (2012).
Rovira-Vallbona, E. et al. Predominance of asymptomatic and sub-microscopic infections characterizes the Plasmodium gametocyte reservoir in the Peruvian Amazon. PLoS Negl. Trop. Dis. 11, e0005674. https://doi.org/10.1371/journal.pntd.0005674 (2017).
Grietens, K. P. et al. Adherence to 7-day primaquine treatment for the radical cure of P. vivax in the Peruvian Amazon. Am. J. Trop. Med. Hygiene 82, 1017–1023. https://doi.org/10.4269/ajtmh.2010.09-0521 (2010).
Muela Ribera, J., Hausmann-Muela, S., Gryseels, C. & Peeters Grietens, K. Re-imagining adherence to treatment from the “other side”: Local interpretations of adverse anti-malarial drug reactions in the Peruvian Amazon. Malar. J. 15, 136. https://doi.org/10.1186/s12936-016-1193-x (2016).
Siqueira, A. M. et al. Characterization of Plasmodium vivax-associated admissions to reference hospitals in Brazil and India. BMC Med. 13, 57. https://doi.org/10.1186/s12916-015-0302-y (2015).
de Oliveira, T. C. et al. Genome-wide diversity and differentiation in New World populations of the human malaria parasite Plasmodium vivax. PLoS Negl. Trop. Dis. 11, e0005824. https://doi.org/10.1371/journal.pntd.0005824 (2017).
Hupalo, D. N. et al. Population genomics studies identify signatures of global dispersal and drug resistance in Plasmodium vivax. Nat. Genet. 48, 953–958. https://doi.org/10.1038/ng.3588 (2016).
Ruebush, T. K. 2nd., Neyra, D. & Cabezas, C. Modifying national malaria treatment policies in Peru. J. Public Health Policy 25, 328–345. https://doi.org/10.1057/palgrave.jphp.3190032 (2004).
Suwanarusk, R. et al. Amplification of pvmdr1 associated with multidrug-resistant Plasmodium vivax. J. Infect Dis 198, 1558–1564. https://doi.org/10.1086/592451 (2008).
Faway, E. et al. Plasmodium vivax multidrug resistance-1 gene polymorphism in French Guiana. Malar. J. 15, 540. https://doi.org/10.1186/s12936-016-1595-9 (2016).
Muller, O., Lu, G. Y. & von Seidlein, L. Geographic expansion of artemisinin resistance. J. Travel. Med. 26, 4. https://doi.org/10.1093/jtm/taz030 (2019).
Sidhu, A. B., Valderramos, S. G. & Fidock, D. A. pfmdr1 mutations contribute to quinine resistance and enhance mefloquine and artemisinin sensitivity in Plasmodium falciparum. Mol. Microbiol. 57, 913–926. https://doi.org/10.1111/j.1365-2958.2005.04729.x (2005).
Villena, F. E., Lizewski, S. E., Joya, C. A. & Valdivia, H. O. Population genomics and evidence of clonal replacement of Plasmodium falciparum in the Peruvian Amazon. Sci. Rep. 11, 21212. https://doi.org/10.1038/s41598-021-00806-5 (2021).
Durand, S., Lachira-Alban, A. & Sánchez, C. C. Impacto de diferentes esquemas terapéuticos sobre la malaria en la costa y amazonia peruana, en el marco de una política de medicamentos antimaláricos, 1994–2017. Revista Peruana de Medicina Experimental y Salud Publica 35, 497–504 (2018).
Acknowledgements
This publication uses data from the MalariaGEN SpotMalaria project as described in 'Jacob CG et al.; Genetic surveillance in the Greater Mekong Subregion and South Asia to support malaria control and elimination; eLife 2021;10:e62997 https://doi.org/10.7554/eLife.62997, https://doi.org/10.7554/eLife.62997. The project is coordinated by the MalariaGEN Resource Centre with funding from Wellcome (206194, 090770). The authors would like to thank the staff of Wellcome Sanger Institute Sample Management, Genotyping, Sequencing and Informatics teams for their contribution.
Disclaimer
The views expressed in this article are those of the author and do not necessarily reflect the official policy or position of the Department of the Navy, Department of Defense, nor the U.S. Government.
Funding
This work was supported by the Armed Forces Health Surveillance Division (AFHSD) and its Global Emerging Infections Surveillance and Response (GEIS) Branch (P0074_21_N6). Samples from UPCH were collected under the Projects U19AI089681 (NIH/NIAID) and FONDECYT-162-2016 (CONCYTEC). ON is supported by the FOGARTY training grant 2D43TW007120-11A1. The funders had no role in study design, data collection, analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
HOV, JFS, CDR, DG participated in study conception, design and data analysis; GB, LR, KB, CJS, FV and ON participated in data acquisition; FEV, ON, HOV, JFS participated in data analysis, CAJ, SEL, HOV, JFS, CDR and DG participated in critical manuscript revision. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Villena, F.E., Sanchez, J.F., Nolasco, O. et al. Drug resistance and population structure of Plasmodium falciparum and Plasmodium vivax in the Peruvian Amazon. Sci Rep 12, 16474 (2022). https://doi.org/10.1038/s41598-022-21028-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-21028-3
- Springer Nature Limited
This article is cited by
-
Genomic surveillance of malaria parasites in an indigenous community in the Peruvian Amazon
Scientific Reports (2024)
-
A new Plasmodium vivax reference genome for South American isolates
BMC Genomics (2023)