Abstract
Photobioreactors heat up significantly during the day due to irradiation by sunlight. High temperatures affect cell physiology negatively, causing reduced growth and productivity. To elucidate the microalgal response to stressful supra-optimal temperature, we studied the physiology of Picochlorum sp. (BPE23) after increasing the growth temperature from 30 °C to 42 °C, whereas 38 °C is its optimal growth temperature. Cell growth, cell composition and mRNA expression patterns were regularly analyzed for 120 h after increasing the temperature. The supra-optimal temperature caused cell cycle arrest for 8 h, with concomitant changes in metabolic activity. Accumulation of fatty acids was observed during this period to store unspent energy which was otherwise used for growth. In addition, the microalgae changed their pigment and fatty acid composition. For example, palmitic acid (C16:0) content in the polar fatty acid fraction increased by 30%, hypothetically to reduce membrane fluidity to counteract the effect of increased temperature. After the relief of cell cycle arrest, the metabolic activity of Picochlorum sp. (BPE23) reduced significantly over time. A strong response in gene expression was observed directly after the increase in temperature, which was dampened in the remainder of the experiment. mRNA expression levels associated with pathways associated with genes acting in photosynthesis, carbon fixation, ribosome, citrate cycle, and biosynthesis of metabolites and amino acids were downregulated, whereas the proteasome, autophagy and endocytosis were upregulated.
Similar content being viewed by others
Introduction
Photobioreactors heat up significantly as a result of irradiation by sunlight, leading to shifts in temperature throughout the day up to levels that can be stressful for microalgae. Heat stress poses a significant challenge in the cultivation of microalgae for the production of food, feed and chemicals. Non-optimal temperatures reduce microalgal growth, thereby reducing productivity and photosynthetic efficiency1. The effect of excessive heat on the cellular physiology has been studied in photosynthetic model organisms such as Arabidopsis thaliana and Chlamydomonas reinhardtii. While many aspects of the response to heat have been studied, much still has to be learned on how exactly microalgae cope with heat2,3,4. In addition, detailed research on the effect of heat stress on other photosynthetic (micro)organisms, is scarce.
Many cellular processes are impacted by supra-optimal temperature which ultimately unbalances cellular homeostasis5. Several effects are; failure of correct protein folding and protein complex assembly, destabilization of cell membranes through increased membrane fluidity, a reduction in photosynthesis, changes in DNA replication and repair, changes in enzymatic activity, and reduced metabolic activity2,6,7. As a response to temperature-induced physiological changes, heat shock factors (HSF’s) and heat shock proteins (HSP’s) are expressed to mediate and protect against adverse effects through chaperoning after exposure to heat stress4,5,8. Excessive heat kills microalgae quickly due to denaturation of proteins and destabilization of membranes. However, in this paper we will focus on a supra-optimal temperature. Supra optimal temperatures fall between the optimal and maximal temperature for growth. Here, growth rate is reduced, but cells still survive.
Photosynthesis, carbon fixation and the central energy metabolism form a delicate system with consecutive enzymatic reactions3,9. Photosystem II is considered the most thermosensitive complex within the photosynthetic pathway while several components of the carbon fixation pathway are thermosensitive2. However, Photosystem II is not considered a bottleneck between the optimal and lethal growth temperature9. When one or more reactions in the energy metabolism are inhibited by temperature, the entire energy flux through the pathway will slow down. This ultimately results in excessive energy in the photosystems, which will overflow and lead to the formation of harmful reactive oxygen species (ROS)3. To prevent the accumulation of ROS, cells suppress photosynthesis by downregulation of key genes. Next to suppression of photosynthesis to prevent ROS formation, mechanisms for ROS scavenging are activated. ROS scavenging in microalgae is done through several mechanisms: enzymatically (by catalases, superoxide dismutases, and peroxidases) and by production of antioxidants for ROS scavenging such as carotenoids10,11. A third protection mechanism, to prevent formation of ROS, is de novo production and accumulation of energy storage compounds such as lipids and carbohydrates as an energy sink2,4. As a result of the before mentioned impact of temperature and the response mechanisms, growth rate decreases12,13.
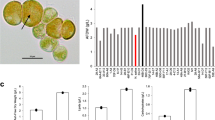
In this study, we investigated the effect of supra-optimal temperature on the physiology of Picochlorum sp. (BPE23) to elucidate how this microalga copes and acclimates to temperature stress. Picochlorum sp. (BPE23) exhibits a high maximal growth rate of 5 d−1 under non-limiting growth conditions which together with its robustness and its relatively small genome size of ~ 13 Mbp makes it an interesting model platform for both fundamental studies, and industrial application14,15,16,17. This chlorophyte has a cell size of 3–4 µm. Currently, five genome assemblies for various species of Picochlorum are available and tools for genetic modification have been developed17. A temperature increase from 30 to 42 °C was studied, which is 4 °C above the optimal growth temperature of this strain14. To elucidate the effect of the temperature shift to 42 °C, the growth rate, cell volume, quantum yield, pigment composition, fatty acid composition, and mRNA expression were periodically measured for 120 h. The results show how Picochlorum sp. (BPE23) copes with the immediate adverse effects of supra-optimal temperatures to prevent cell damage and how the onset to acclimatization to this temperature takes place.
Results
Temperature increase to supra-optimal level leads to cell-cycle arrest and decreased growth
The effect of the increase in temperature from 30 to 42 °C for a time span of 120 h on Picochlorum sp. (BPE23) can be seen in Fig. 1. The specific growth rate, quantum yield, and cell volume were measured over time, before and after temperature increase. At the onset of temperature increase (t = 0), the cell culture was at steady state, in a photobioreactor operated in turbidostat mode. Since only temperature was changed, it is safely assumed that the observed changes in the culture are solely due to the temperature shift. Two distinct responses were observed; an immediate short-term response, followed by a long-term response in which the microalgae gradually acclimated.
Effect of temperature increase to supra-optimal value, on the specific growth rate (d−1), quantum yield (Fv/Fm), and average cell volume (μm3 cell−1) in Picochlorum sp. (BPE23). Temperature was increased from steady state growth at 30–42 °C at Time = 0 h. Data represents the mean ± standard deviation of biological triplicate experiments.
One hour after the temperature shift all monitored growth parameters had changed (Fig. 1). The specific growth rate decreased during the first hour, followed by a temporary increase to higher levels than those observed at 30 °C. It remained high until 8 h after the temperature shift point at which it started to decrease until the end of the experiment (120 h). During the first 8 h the average cell volume increased by 30%, from 10.6 to 13.76 µm3. The combination of a decreased growth rate and an increase in cell volume due to halted cell division indicates a cell cycle arrest. The cell cycle arrest was released after 8 h and cell volume returned to the initial value after 24 h. The cell volume gradually kept increasing from 24 h after the temperature shift until the end of the experiment.
Quantum yield (Fv/Fm) is a measure for the maximum photosynthetic capacity of photosystem II and can be used to monitor photo inhibitory damage in response to stress, in this study as a result of high temperature18. The observed quantum yield shortly increased from 0.72 to 0.75 at 42 °C, after which a gradual decrease was observed to a level of 0.45 at 72 h and remained stable until the end of the experiment (120 h).
Pigment content increased after exposure to supra-optimal temperature
Pigment content was measured over time and is displayed in Fig. 2. The pigment concentrations were also measured through optical density measurements which show a comparable trend (Supplementary material 5). The concentration of chlorophyll-a and chlorophyll-b decreased slightly in the first hour after the temperature shift. However, from 4 h onwards an increase in concentration was observed with a peak at 72 h. The concentration for both chlorophyll-a and chlorophyll-b increased with a factor of 1.75, indicating an upregulation of photosystems. The ratio between chlorophyll-a and chlorophyll-b increased slightly between 8 and 24 h, but returned to the values as found at the start of the experiment at 72 h.
Concentrations of chlorophyll-a, chlorophyll-b, β-carotene, canthaxanthin, violaxanthin, zeaxanthin, and lutein in algal biomass as measured over time, starting at the moment of the temperature increase from 30 to 42 °C. Data represents the mean ± standard deviation of biological triplicate experiments.
Carotenoids, in addition to a light-harvesting function within the photosystems, also have a photoprotective role in non-photochemical quenching, ROS scavenging, and/or filtering of light19. The xanthophyll pigments Violaxanthin and Zeaxanthin show a peak concentration at 72 h after temperature increase. Antheraxanthin was measured but not detected. The concentration of zeaxanthin increased directly after exposure to supra-optimal temperature, from 0.18 to 1.84 mg g−1 after 72 h, whereas the concentration of violaxanthin initially decreases after 1 h, followed by a gradual increase during the first 24 h. Xanthophyll pigments accommodate energy dissipation through non-photochemical quenching19. Also, β-carotene, Canthaxanthin and Lutein, known as strong antioxidants, were found at increased concentrations in stressed cells after exposure to supra-optimal temperatures19.
Picochlorum sp. (BPE23) accumulated fatty acids as a response to the increased temperature
Fatty acids can be distributed into two general groups; neutral and polar, which are present as lipid bodies and in the cell membranes, respectively (Fig. 3). The concentration of neutral fatty acids increased from 3.0 mg g−1 at 0 h to 11.5 mg g−1 at 24 h, whereas the total content of polar fatty acids increased from 79.0 to 88 mg g−1. Within the neutral fatty acids, all fatty acid species showed an increased concentration at approximately the same ratio. In the polar fatty acids the increase in content was mainly caused by C16:0 content, which C16:0 increased from 21.0 mg g−1 after 0 h to 30.0 mg g−1 after 72 h. At the same time, a decrease in C16:3 was observed over time.
A large transcriptional change was observed after the increase in temperature
Comparative transcriptome analysis provided further insight into the stress response of Picochlorum sp. (BPE23) to supra-optimal temperatures. In total 6990 genes were identified in the transcriptome, of which 5930 were annotated through BLASTP and Orthofinder search and inferring domains. In addition, 55 novel genes were identified. 4427 of these genes matched to A. thaliana proteins (62.8%). Iteratively, the remaining unidentified genes were matched to other species of which 678 genes matched to C. Variabilis orthologs (9.6%), 116 genes matched to A. protothecoides orthologs (1.6%), 3 genes matched to Helicosporidium sp. orthologs (0.04%) and 60 genes matched to C. reinhardtii orthologs (0.9%). 643 out of the remaining 1761 genes had a domain annotation (9.1%).
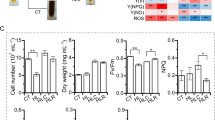
Differential gene expression levels and enrichment analyses are displayed in two ways in Figs. 4C,D, 5, and 6. The left-hand side of Fig. 4 shows the number of genes that are differentially expressed in the individual timepoints compared to the control condition at timepoint zero, which represents steady state growth at 30 °C. The right-hand side of the figure shows the number of genes that are differentially expressed between consecutive timepoints.
(A,B) PCA bi-plots of the RNA-Seq samples collected at 0 h, 1 h, 4 h, 8 h, 24 h and 120 h and for samples collected at 4 h, 8 h, 24 h and 120 h after the increase in temperature from 30 to 42 °C. The first two PCs are displayed. (C,D) Percentage of DEGs over time, observed in Picochlorum sp. (BPE23) after an increase in temperature from 30 to 42 °C. The total pool of genes was 6990. The time after temperature shift is displayed on the x-axis, as is the sampling time that was used as control. Up-regulated genes are indicated in green whereas down-regulated DEGs are indicated in red. DEGs were selected based on an FDR-adjusted p value ≤ 0.05 and an absolute log2 fold change value > 1.
Module hub genes associated with processes affected by temperature stress in Picochlorum sp. (BPE23). Modules are defined from network inference by WGCNA of genes in the temperature-shock experiment. A heatmap represents differential expression levels of mRNA. Module color is displayed on the left. LFC represents the log2 fold change.
39% of all genes showed an log2 fold change value larger than 1 after 1 h, which was largely reversed between 1 and 4 h (Fig. 4). The number of differentially expressed genes (DEGs) decreased from 4 to 8 h, and subsequently to 24 h after the temperature increase. This indicates that the initial heat stress subsided and that Picochlorum sp. (BPE23) was reverting to a stable state once again. Between 24 and 120 h only 2% of genes were differentially expressed. The changes in log2 fold change after 8 h, 24 h and 120 h look very similar when compared to 0 h. However, when compared to the previous sampling time then differences in expression levels can be observed.
A PCA was conducted to identify similarities among biological replicates (Fig. 4). mRNA expression at 0 h and 1 h showed a significant distance from samples taken at 4 h, 8 h, 24 h, and 120 h. A second PCA was then done using only samples at 4 h, 8 h, 24 h, and 120 h. The samples taken at 4 h, 8 h, 24 h, and 120 h cluster together by biological triplicate and display distance to sample moments as expected. The clustering of biological replicates confirms that samples were processed and sequenced correctly.
Temperature stress affects protein processing, HSP expression, and DNA replication
KEGG pathway enrichment was performed to identify which metabolic pathways were affected by the temperature increase (Fig. 5). The RNA degradation pathway was upregulated directly after temperature increase. Other upregulated pathways involved in the immediate response were the proteasome, autophagy, and endocytosis pathways. Downregulated pathways were DNA replication, ribosome, and base/nucleotide excision repair. In addition, pathways in the central metabolism such as biosynthesis of amino acids, biosynthesis of secondary metabolites, and citrate cycle were downregulated 1 h after the temperature increase. A reversal of this initial downregulation was observed after 4 h. Despite the observed upregulation between sampling moments, the processes were never enriched in the upregulated pathways when compared to growth at 0 h (30 °C). The ribosome pathway remained enriched in the downregulated pathways compared to timepoint zero throughout the experiment. Photosynthesis pathway was downregulated after 8 h. Despite enrichment in the upregulated pathways at 24–8 h, the photosynthesis was still downregulated compared to 0 h. From 24 to 120 h after the temperature shift, downregulation increased further.
GO-term enrichment analysis was conducted in addition to the KEGG enrichment to further identify enriched cellular processes (Supplementary material 2). Various GO-terms related to protein processing such as Protein folding, Proteolysis and Proteasome were upregulated throughout the experiment. These GO-terms showed enrichment in the upregulated GO-terms compared to the control at 0 h. In addition, cellular response to heat and heat shock protein binding show enrichment in the upregulated GO-terms. On the contrary, DNA replication, photosynthesis, peptide biosynthesis, translation, and three ribosome related GO-terms are enriched in the downregulated genes. The observations from the GO-term enrichment indicate similar changes in the mRNA expression as the observations from the KEGG pathway enrichment indicated.
Network analysis verifies the results of the KEGG enrichment analysis
Weighted gene co-expression network analysis (WGCNA) was performed to describe correlation patterns among genes. Genes were clustered into 18 modules based on co-expression patterns (Fig. 6, Supplementary material 3). Each module was functionally annotated with GO-terms and KEGG pathways to assess differences between modules (Supplementary material 4). Each module displayed a unique functional annotation pattern which consisted of multiple GO-terms. The hubs were grouped roughly by similarities in functionality. The different module colours and their phylogenetic relationships are displayed in Fig. 6. The brown, cyan, green, grey and yellow modules were made up of genes associated with proteolysis, protein ubiquitination, autophagy, endocytosis and protein processing in ER. These processes are involved in the protein-related protection response to temperature stress4. Furthermore, the light-green, black, green–yellow and turquoise modules were associated with processes to energy fixation and metabolism, such as ATP binding, oxidative phosphorylation, photosynthesis, thylakoid membrane, carbon fixation, glycolysis and gluconeogenesis. Other interesting modules which were affected by temperature stress are the blue, light-cyan, magenta, pink and red module. These modules are associated with RNA-, and DNA-related processes such as replication, binding and repair. Lastly, all hubs except for grey and yellow were associated to the chloroplast envelope, chloroplast stroma, and plastid.
The average module expression profile was summarised through the module’s eigengene. The eigengene is a hypothetical gene that is highly correlated with expression profiles of genes in the module and can therefore function to represent the average expression of the module20. Genes with the highest eigengene-based connectivity were selected as hub genes, and presented together with their expression levels (Fig. 6). The largest changes in differential expression were observed during the first hours of the experiment. The transcriptional response in the first hour after the temperature increase displayed the largest log2 fold change value, a large reversal was observed for all modules after 4 h. Despite this reversal in differential expression, about half of the hub genes stay up or downregulated to a lower extent throughout the experiment. Hub genes are representative for their associated hub and therefore these hubs and their associated functions, as annotated through GO-term and KEGG pathway enrichment, hypothetically display a comparable expression pattern (Supplementary material 4). In general, the results from the WGCNA correspond to enrichment analysis results. Several interesting hub genes were identified which play a role in the response to an increase in temperature.
Discussion
Cell physiology changes severely after exposure to supra-optimal temperature
In an industrial photobioreactor, temperature can raise to levels that significantly affect the growth and productivity of microalgae. Microalgae are continuously exposed to changing environmental conditions throughout the day and between days. We aimed to characterize the effect of supra-optimal temperature on the physiology of the green microalga Picochlorum sp. (BPE23). We subjected Picochlorum sp. (BPE23) to a supra-optimal temperature of 42 °C for 120 h, whereas 38 °C is the optimal temperature for growth14. Cell growth, cell volume, pigment composition, fatty acid composition and mRNA expression were measured throughout the experiment. While such long exposure to high temperature does not commonly occur in nature or in photobioreactors, this study generated knowledge on how microalgae immediately cope and eventually acclimate to supra-optimal temperatures.
Two different sequential response phases were observed after the temperature shift from 30 to 42 °C. In the first phase (0–8 h), the cell cycle was arrested and an increase in cell volume was observed. The cell cycle resumed after 8 h as cells started to divide and returned to their original cell size. During the second phase (8–120 h), Picochlorum sp. (BPE23) acclimated towards a new homeostatic phase. The average cell volume gradually increased between 24 and 120 h. This increase in cell size was unexpected as increased temperature commonly causes a decreased cell volume in other microalgal species to counteract the imbalance between catabolic and anabolic processes1. The growth rate and cell composition stabilized after 24–72 h. The changes in mRNA transcription were very severe between 0 and 4 h, while between 4 and 120 h differential expression levels rapidly became smaller.
Interestingly, the hub gene in the red module, minichromosome maintenance (MCM2/3/5) family protein, was downregulated 5 times log2 fold during the first hours after temperature increase. The MCM complex is critical for cell division and essential for DNA replication as it is a target of various checkpoint pathways required for S-phase entry21. This confirms the hypothesis that the cell cycle was arrested to protect the DNA from temperature-induced damage during the replication cycle. In literature, the cell cycle of Chlamydomonas reinhardtii was found to be inhibited after a heat shock treatment4,12. Full recovery of the cell cycle and recovery of the cell physiology in C. reinhardtii was only observed after the temperature was decreased.
Heat shock proteins were upregulated directly after the temperature increase
A heat shock response was observed in the first hours, and to a lesser extent throughout the experiment. HSP20 and HSP70 were found among the strongest upregulated genes in the dataset. HSP70 was the main cause for enrichment in the endocytosis, protein processing and the endoplasmic reticulum pathways. Moreover, genes encoding for HSP40, HSP60, and HSP90, and HSFs showed upregulated gene expression. HSFs are transcriptional activators that regulate expression of heat shock proteins. The heat shock proteins protect cells from adverse effects of thermal stress through chaperoning and refolding of polypeptides and by stabilizing protein complexes, cellular membranes and key cellular processes5,6,22. Their regulatory role provides thermotolerance and the ability to gradually acclimate to a new environmental temperature by mediating the part of the cellular stress response that deals with protein homeostasis23. Next to the high upregulation of heat shock proteins, most of the other genes in the endocytosis pathway were also upregulated (~ 0.5–1 times log2 fold). In addition to regulation by heat shock proteins, the 5.3 times log2 fold upregulation of the ubiquitin-like superfamily protein-encoding genes is the major cause for the enrichment of the autophagy pathway and is involved in numerous stress-related functions such as the cell cycle, DNA repair, transcription, autophagy, and post-translational protein modification24. Proteins can still become damaged despite chaperoning and stabilization. Degradation of damaged proteins and proteins that are no longer required for cell functioning is essential to maintain cellular homeostasis2. The 20S proteasome alpha subunit F2 (PAF2) encoding gene was upregulated 7.5 times log2 fold and is involved in the degradation of proteins with partially unfolded regions25.
DNA repair in microalgae is still poorly understood2,6. However, heat shock proteins are known to initiate DNA repair after stressful events. HSP70 and HSP27 in particular are associated to this activity26. DNA replication, base excision repair, Nucleotide excision repair, and mismatch repair were enriched in the downregulated genes at 1 h, whereas DNA replication was enriched in the downregulated genes at 24 h and 120 h. The DNA associated network interference module (red) shows a similar pattern of downregulation, followed by almost complete reversal towards a log2 fold change value of ~ 0. Hypothesizing from the mRNA expression data, DNA was not damaged after exposure to supra-optimal temperatures, nor by formation of ROS.
Supra-optimal temperature downregulates photosynthesis but increases pigment concentrations.
The quantum yield (Fv/Fm) increased during the first hours after the temperature increase, indicating an increased efficiency of photosystem II at 42 °C. After 8 h, the quantum yield decreased until the end of the experiment. Moreover, between 8 and 120 h, gene expression of nuclear genes encoding subunits of both photosystem I and photosystem II was enriched in the downregulated genes. mRNA expression for genes associated to both photosystems was downregulated despite the fact that photosystems are not considered a bottleneck at survivable but supra-optimal culture temperatures. In addition, several genes in the Porphyrin and chlorophyll metabolism show up to fourfold downregulation, of which the most severely downregulates genes are: Coproporphyrinogen III, Uroporphyrinogen decarboxylase, and NAD(P)-binding Rossman-fold superfamily protein. Genes enriched in the turquoise hub, associated with chloroplasts, plastids, and the thylakoid membrane, showed downregulation throughout the experiment. Research on Dunaliella bardawil reported that photosynthesis was severely downregulated and that chlorophyll content decreased dramatically after exposure to a heat treatment of 42 °C for 2 h, after which temperature was returned to the optimal temperature13. In Picochlorum sp. (BPE23), the mRNA transcript levels indicate overall decreased photosynthetic activity. However, an unexpected increase in chlorophyll-a and chlorophyll-b was observed nonetheless.
Downregulation of photosynthesis is done to reduce the formation of ROS13. When metabolism becomes unbalanced due to temperature stress the excessive energy flux from photosynthesis results in the formation of ROS. Unfortunately, too few genes in the carotenoid biosynthesis KEGG pathway gained a functional annotation. Therefore, little insight on the transcriptional response of the carotenoid biosynthesis pathway was gained. However, data on pigment concentration in the biomass show that the concentration of the carotenoids: zeaxanthin, violaxanthin, lutein, β-carotene and canthaxanthin increased as a result of exposure to supra-optimal temperature. An increase in carotenoid content in response to temperature increase was also found in the literature for D. bardawil and Haematococcus pluvialis13,27. Increased levels of carotenoids are often induced when microalgae are exposed to abiotic stress conditions such as salinity, light, and temperature for quenching of ROS10,28.
The metabolism and protein synthesis were downregulated to conserve energy
As expected, connected to downregulation of genes associated to photosynthesis, the carbon fixation pathway and central energy metabolism were downregulated as determined through KEGG pathway enrichment. All analyses point towards downregulation of the metabolism of Picochlorum sp. in response to temperature stress. Lower photosynthetic activity ultimately results in depletion of the cell’s energy reserves. Suppression of ribosome biogenesis reduces the energy demand of the cell through reduction of protein synthesis to conserve energy25. Ribosomal activity was significantly downregulated throughout the experiment. In a study with a thermosensitive Arabidopsis Thaliana strain it was hypothesized that the ribosomal activity was downregulated in response to stressful temperature to prevent errors in translation and protein folding29. Production of misfolded proteins would not only lead to wasted energy, but could also lead to formation of toxic compounds in the cell that cause further damage8. Therefore, reduced ribosomal activity indirectly relieves the workload of chaperones. Other protein processing related processes were affected throughout the experiment. The rate of protein synthesis was adapted to the reduced growth rate that was caused by the increased temperature. At the same time, the conserved energy was available for damage repair caused by oxidative stress and acclimation to the new temperature6. Downregulation of protein production and photosynthesis after temperature stress was also observed in Picochlorum costavermella29. We observed that downregulation of these vital cellular processes led to a severe decrease in growth rate.
Picochlorum sp. (BPE23) accumulated fatty acids under heat stress
The concentration of neutral fatty acids is initially low (3.0 mg g−1) as cells are grown in nitrogen replete growth medium30. However, 24 h after temperature increase, a fatty acid concentration of 11.5 mg g−1 was measured. This significant 383% increase shows the funnelling of excess energy into de novo synthesis of lipid bodies as energy storage. In a study on Chlamydomonas reinhardtii and Chlorella vulgaris a screening was done for neutral lipid accumulation at elevated temperature, after which about 25% of the mutant strains showed similar elevated lipid accumulation31. Apparently, induction of neutral lipids through increased temperature is strain specific. Next to neutral lipids, also the polar membrane fatty acid concentration increased, from 79.0 to 88.5 mg g−1. Especially C16:0 increased significantly. A study on Chlamydomonas reinhardtii showed that translocation of unsaturated fatty acids from cell membranes to neutral lipids happened after a temperature increase, to maintain optimal membrane fluidity4. Especially plastid membranes are heat sensitive, and quick modification of these membranes is necessary to keep photosynthesis running32. Almost all hubs found through the WGCNA analysis show gene ontology annotation to the plastid membranes, indicating that these membranes are also severely affected in Picochlorum sp. (BPE23).
Genes in the fatty acid biosynthesis pathway exhibited downregulation as an overall trend from 0 to 1 h, although the pathway itself was not significantly enriched due to the presence of few severely upregulated genes, up to 2.6 times log2 fold (Supplementary material 6). Pathview pathway analysis indicated increased mRNA expression of genes encoding for Malonyl-CoA synthase, which catalyses the reaction from malonate into malonyl-CoA. In addition expression of mRNA of acyl carrier protein and long chain acyl-CoA synthase were upregulated. However, drawing accurate conclusions is challenging as the fatty acid biosynthesis pathway is only partly annotated in the used reference genome of Picochlorum sp. SENEW3 (assembly ASM87641v1)33. Another possible reason for the fatty acid accumulation could be high stability of enzymes involved in fatty acid biosynthesis which resulted in the continuation of fatty acid biosynthesis, whereas other metabolic activity and cell growth decreased. Although the total increase in fatty acid content was just 18 mg g−1 at 24 h and 24 mg g−1 at 72 h, this increase corresponds to a percentual gain in fatty acid content of 22% and 38%, respectively. The ability to induce an increased fatty acid concentration by increasing temperature for only 24 h opens up the possibility to boost the lipid content of microalgal biomass before harvest.
Conclusions
In this study, the mechanisms underlying temperature stress and acclimation were determined through assessment of cell physiology and transcriptome analysis. A cell cycle arrest and a transcriptomic heat shock response were observed directly after an increase in temperature to 42 °C, 4° above the optimal growth temperature. During the first 8 h, various regulatory and chaperoning mechanisms were differentially expressed to protect the cells from heat-induced cell damage. Between 8 and 120 h after temperature increase, an acclimation process towards a new homeostatic phase started. Photosynthesis, carbon fixation, ribosomal activity, and central metabolism were downregulated, hypothetically to reduce oxidative cell damage. At the same time, pigment concentration increased significantly and the concentration of fatty acids increased by 22% over between 0 and 24 h. The combination of these effects led to a decrease in growth rate throughout the experiment. This study contributes knowledge on the effect of supra-optimal temperature on the physiology of the industrially relevant green microalgae Picochlorum sp. (BPE23).
Materials and methods:
Cell cultivation
Growth media and inoculum preparation
Picochlorum sp. (BPE23), isolated from a saltwater body of Bonaire was pre-cultivated in shake flasks in an orbital shaker incubator (Multitron, Infors HT) with a 12/12 h day/night cycle and a light intensity of 100 μmolph m−2 s−114. The temperature was 30 °C during night and 40 °C during day. Furthermore, the relative humidity of the air in the incubator was set to 60% and enriched with 2% CO2. Cells were cultured in artificial seawater enriched with nutrients and trace elements. Elements were provided at the following concentrations (in g L−1): NaCl, 24.5; MgCl2.6H2O, 9.80; Na2SO4, 3.20; NaNO3 2.12; K2SO4, 0.85; CaCL2·2H2O, 0.80; KH2PO4, 0.23; Na2EDTA·2H2O, 0.105; Na2EDTA, 0.06; FeSO4·7H2O, 0.0396; MnCl2·2H2O, 1.71 × 10−3; ZnSO4·7H2O, 6.60 × 10−4; Na2Mo4·2H2O, 2.42 × 10−4; Co(NO3)2·6H2O, 7.00 × 10−5; NiSO4·6H2O, 2.63 × 10−5; CuSO4·5H2O, 2.40 × 10−5; K2CrO4, 1.94 × 10−5; Na3VO4, 1.84 × 10−5; H2SeO3, 1.29 × 10−5. HEPES (4.77 g L−1) was added for shake flask cultures as a pH buffer. The medium pH was adjusted to 7.4 after and filter sterilized before use. During photobioreactor cultivation, Antifoam B (J.T.Baker, Avantor, USA) was added at a concentration of 0.5 mL L−1 out of a 1% w/w% stock. In addition, 0.168 g L−1 Sodium bicarbonate (NaHCO3) was added at the time of inoculation to provide sufficient CO2 at the start of the cultivation. The photobioreactor was inoculated at a starting OD density of 0.2.
Photobioreactor operation
Microalgae were cultivated in heat-sterilized flat panel photobioreactors (Labfors 5 Lux, Infors HT, Switzerland) with a working volume of 1.8 L, an optical depth of 20.7 mm and a surface area for irradiation of 0.08 m2. Continuous irradiation (24/24 h) was applied from one side by 260 warm white LED lamps at 813 μmolph m−2 s−1 (PAR). To remove variation in gene expression due to the circadian cycle we grew the microalgae under continuous irradiation while maintaining all other growth conditions stable at the same time. The biomass density was maintained at approximately 2.3 g L−1 by continuous light controlled dilution of the cell culture (turbidostat mode). The turbidostat control was set to maintain an outgoing light level of 10 μmolph m−2 s−1 (PAR). The dilution rate of the photobioreactor was logged continuously by weighing the ingoing medium vessel. Compressed air was supplied at a rate of 980 mL min−1. CO2 was provided on-demand by pH-controlled addition. The pH level in the photobioreactor was set at 7. The photobioreactor temperature was set at 30 °C from the start of cell cultivation. When steady state was reached, the temperature was increased to 42 °C in one step, which took approximately 15 min. Samples were taken daily at 0 h, 1 h, 4 h, 8 h, 24 h after the temperature increase, and once a day for every day onwards to monitor changes in cell physiology.
Biomass analysis
Dry weight
Biomass concentration (g L−1) was measured in duplicate by dry weight determination. Empty Whatman glass microfiber filters (θ 55 mm, pore size 0.7 μm) were dried overnight at 95 °C and placed in a desiccator for 2 h. Filters were then weighed and placed in the mild vacuum filtration setup. Cell culture containing 1–10 mg of microalgae biomass was diluted in 25 mL 0.5 M ammonium formate and filtered. The filter was washed twice with 25 mL 0.5 M ammonium formate to remove residual salts. The wet filter was dried overnight at 95 °C, placed in a desiccator for 2 h, and weighed. Biomass concentration was calculated from the difference in filter weight before and after filtration and drying.
Cell volume and number
Cell size and cell number were measured in duplicate with the Multisizer III (Beckman Coulter Inc., USA, 50 μm aperture). Samples were diluted in two steps before analysis, initially by dilution of 5× in fresh medium, followed by dilution of 100× in Coulter Isoton II. Cell volume was then derived from the cell size by assuming that cells were shaped spherical.
Quantum yield
The cell culture’s quantum yield (Fv/Fm), representing the maximum photosynthetic capacity of photosystem II was determined. Cells were measured after dark adaption at room temperature for 15 min (AquaPen-C 100, PSI; excitation light 455 nm (blue), saturating light pulse: 3000 µmol m−2 s−1).
Biomass harvest and lyophilizing
Biomass samples for compositional analysis were taken at the same moment as when offline measurements were performed. Microalgae cells were pelleted by centrifugation at 4000g for 5 min and washed with 0.5 M ammonium formate. The centrifugation/washing cycle was repeated twice more after which the cell pellet was frozen at − 20 °C. Samples were then lyophilised for 24 h and stored at − 20 °C until further processing.
Pigment analysis
Pigment content was determined through extraction and HPLC analysis 34. 10 mg of lyophilized biomass was disrupted by bead beating (Precellys 24, Bertin Technologies, France) at 5000 rpm for three cycles of 90 s with 60-s breaks on ice between each cycle. The extraction was done through five washing steps with methanol containing 0.1% butylhydroxytoluene. Separation, identification and quantification of pigments were performed using a Shimadzu (U)HPLC system (Nexera X2, Shimadzu, Japan), equipped with a pump, degasser, oven (25 °C), autosampler, and photodiode array (PDA) detector. Separation of pigments was achieved using a YMC Carotenoid C30 column (250 × 4.6 mm 5 μm ID) coupled to a YMC C30 guard column (20 × 4 mm, 5 μm ID)(YMC, Japan) at 25 °C with a flow rate of 1 mL min−1. A sample injection volume of 20µL was used. The mobile phases consisted of Methanol (A), water/methanol (20/80 (v/v%)) containing 0.2% ammonium acetate (B), tert-methyl butyl ether (C) (all solvents were purchased at Sigma Aldrich). The elution protocol started with 0–12 min isocratic A:95% B:5% C:0%, with at 12 min a step to A:80%, B:5%, C:15%, followed by a linear gradient 12–30 min to A:30%, B:5%, C:65%, finally followed by a conditioning phase 30–40 min at the initial concentration. Analytical HPLC standards for chlorophyll a, chlorophyll b, β-carotene, canthaxanthin, violaxanthin, antheraxanthin, zeaxanthin, and lutein had a purity of > 99% (Carotenature, Switzerland).
Fatty acid analysis
Fatty acids within the triacylglycerol (TAG) and polar lipids (PL) fraction were quantified through GC-FID analysis according to35. 10 mg of lyophilised biomass was disrupted by bead beating. The fatty acids were extracted from the disrupted biomass in a mixture of chloroform/methanol (1:1.25, v:v) containing Glyceryl tripentadecanoate (C15:0 TAG) (T4257, Sigma-Aldrich) and 1,2-didecanoyl-sn-glycero-3-phospho-(1′-rac-glycerol) (sodium salt) C10:0 PG (840434, Avanti Polar Lipids Inc) as internal standards for the TAG and the PL fraction, respectively. Separation of TAG and PL was done by use of Sep-Pak Vac silica cartridge (6 cc, 1000 mg; Waters). TAG was eluted from the column with a solution of hexane:diethylether (7:1, v:v) and the PL were eluted with a solution of methanol:acetone:hexane (2:2:1, v:v:v). The extracts were methylated for 3 h at 70 °C in methanol containing 5% H2SO4.
Transcriptome analysis
mRNA extraction and sequencing
mRNA-Seq analysis was done for samples taken at 0 h, 1 h, 4 h, 8 h, 24 h and 120 h after the temperature increase from 30 to 42 °C. Biomass was directly put on ice and centrifuged for 5 min at 4255 g at 2 °C. The cell pellet was then immediately frozen in liquid nitrogen and stored at − 80 °C until further processing. mRNA was extracted from ~ 200 ul frozen cell pellet by automated mRNA extraction (Maxwell® 16 LEV simplyRNA, Promega, USA). Extracted mRNA was tested for integrity (Qsep100, GCbiotech, Netherlands) and quantity (Qubit fluorometer, ThermoFisher, USA). Sequencing libraries were prepared using the NEB Next® Ultra™ mRNA Library Prep Kit. Fragments of the mRNA library in the size range of 250–300 bp were sequenced using the Illumina Novoseq PE150 platform, yielding paired-end reads of 150 nt (Novogene, China). The quality of mRNA-Seq reads was assessed using FastQC v0.11.5 36.
Transcript assembly and annotation
Paired-end reads were mapped to the genome of Picochlorum sp. SENEW3 (assembly ASM87641v1) using HISAT2 v2.2.1 with the -very-sensitive pre-set 33,37. Transcript were assembled and predicted using StringTie v2.1.4, which was guided by the structural annotation of Picochlorum sp. SENEW3. StringTie’s prepDE3 python script was used to extract gene counts and predict genes de novo. Functional annotation was initiated by a BLASTP search of the translated Picochlorum sp. (BPE23) coding sequences against the protein sequences of Arabidopsis thaliana with an E-value threshold of 1E−10. Unannotated genes were filtered and orthology inference was conducted using OrthoFinder v2.5.2 against the protein sequences of the microalgae Auxenochlorella protothecoides, Chlorella variabilis, Chlamydomonas reinhardtii, and Helicosporidium sp. Ortholog gene identifiers were then matched to their gene description. Functional annotation was concluded by matching unannotated genes to their inferred domains, as derived from Pico-PLAZA.
Differential expression analysis and GO, and KEGG enrichment analysis
Pairwise differential expression (DE) analysis was performed using the DESeq2 v1.30.0. R package. Sample-level quality control consisted of pairwise correlation clustering, hierarchical clustering, and Principal Component Analysis (PCA). Fold change values were generated on a log2 scale (LFC). Two designs for data display were implemented; first, where a control condition after 0 h was used for each sample to compare DE between stressed and non-stressed growth. Second, where the previous sampling moment was used as a control condition to compare DE over time. Genes with a false discovery rate (FDR) adjusted p value ≤ 0.05 and an LFC > 1 were considered as significantly differentially expressed.
Arabidopsis thaliana gene identifiers that matched to Picochlorum sp. (BPE23) genes were linked to their corresponding DE-analysis results and used for GO and KEGG pathway enrichment analysis38. Enrichment analyses were conducted by use of clusterProfiler v3.18.1. and org.At.tain.db v3.12.0. packages. GO-terms and KEGG pathways with an FDR-adjusted p value ≤ 0.05 and a positive or negative enrichment score were considered as significantly enriched and visualized with the ggplots2 package.
Network interference analysis
Weighted gene co-expression network analysis (WGCNA) was conducted by applying the WGCNA R package v1.6920. The correlation network was inferred from a correlation matrix of normalized counts. The optimal value of power was determined through scale-free topology analysis. The network was restricted to genes with informative connectivity, referring to a connectivity higher than the median connectivity of the entire network. Modules were then constructed with average linkage hierarchical clustering using distances in the topological overlap construction. Networks were constructed in a hybrid adaptive tree with a deepSplit of 1, a power of 12, a minimum cluster size of 30, a cut height of 0.8, and no PAM-like stage filtering. Subsequently, the gene with the highest eigengene-based connectivity was identified for each module and considered the module’s hub gene. Furthermore, modules were annotated with GO-terms and KEGG pathways to infer functional properties.
References
Ras, M., Steyer, J.-P. & Bernard, O. Temperature effect on microalgae: A crucial factor for outdoor production. Rev. Environ. Sci. Biotechnol. 12, 153–164 (2013).
Schroda, M., Hemme, D. & Mühlhaus, T. The Chlamydomonas heat stress response. Plant J. 82, 466–480 (2015).
Allakhverdiev, S. I. et al. Heat stress: An overview of molecular responses in photosynthesis. Photosynth. Res. 98, 541 (2008).
Hemme, D. et al. Systems-wide analysis of acclimation responses to long-term heat stress and recovery in the photosynthetic model organism Chlamydomonas reinhardtii. Plant Cell 26, 4270–4297 (2014).
Kobayashi, Y. et al. Algae sense exact temperatures: Small heat shock proteins are expressed at the survival threshold temperature in Cyanidioschyzon merolae and Chlamydomonas reinhardtii. Genome Biol. Evol. 6, 2731–2740 (2014).
Richter, K., Haslbeck, M. & Buchner, J. The heat shock response: Life on the verge of death. Mol. Cell 40, 253–266 (2010).
Velichko, A. K., Petrova, N. V., Kantidze, O. L. & Razin, S. V. Dual effect of heat shock on DNA replication and genome integrity. Mol. Biol. Cell 23, 3450–3460 (2012).
Balchin, D., Hayer-Hartl, M. & Hartl, F. U. In vivo aspects of protein folding and quality control. Science 353, aac4354 (2016).
Mathur, S., Agrawal, D. & Jajoo, A. Photosynthesis: Response to high temperature stress. J. Photochem. Photobiol. B Biol. 137, 116–126 (2014).
Mittler, R. Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci. 7, 405–410 (2002).
Bowler, C., Montagu, M. V. & Inze, D. Superoxide dismutase and stress tolerance. Annu. Rev. Plant Biol. 43, 83–116 (1992).
Zachleder, V., Ivanov, I., Vítová, M. & Bišová, K. Cell cycle arrest by Supraoptimal temperature in the alga Chlamydomonas reinhardtii. Cells 8, 1237 (2019).
Liang, M.-H., Jiang, J.-G., Wang, L. & Zhu, J. Transcriptomic insights into the heat stress response of Dunaliella bardawil. Enzym. Microb. Technol. 132, 109436 (2020).
Barten, R. J., Wijffels, R. H. & Barbosa, M. J. Bioprospecting and characterization of temperature tolerant microalgae from Bonaire. Algal Res. 50, 102008 (2020).
Krishnan, A. et al. Picochlorum celeri as a model system for robust outdoor algal growth in seawater. Sci. Rep. 11, 1–13 (2021).
Weissman, J. C. et al. High-light selection produces a fast-growing Picochlorum celeri. Algal Res. 36, 17–28 (2018).
Dahlin, L. R. et al. Development of a high-productivity, halophilic, thermotolerant microalga Picochlorum renovo. Commun. Biol. 2, 1–9 (2019).
Maxwell, K. & Johnson, G. N. Chlorophyll fluorescence—A practical guide. J. Exp. Bot. 51, 659–668 (2000).
Jahns, P. & Holzwarth, A. R. The role of the xanthophyll cycle and of lutein in photoprotection of photosystem II. Biochim. Biophys. Acta BBA Bioenerg. 1817, 182–193 (2012).
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinform. 9, 1–13 (2008).
Bochman, M. L. & Schwacha, A. The Mcm complex: Unwinding the mechanism of a replicative helicase. Microbiol. Mol. Biol. Rev. 73, 652–683 (2009).
Vega, V. L., Charles, W. & De Maio, A. A new feature of the stress response: Increase in endocytosis mediated by Hsp70. Cell Stress Chaperones 15, 517–527 (2010).
de Vries, J. et al. Heat stress response in the closest algal relatives of land plants reveals conserved stress signaling circuits. Plant J. 103, 1025–1048 (2020).
Streich, F. C. Jr. & Lima, C. D. Structural and functional insights to ubiquitin-like protein conjugation. Annu. Rev. Biophys. 43, 357–379 (2014).
Kumar Deshmukh, F., Yaffe, D., Olshina, M. A., Ben-Nissan, G. & Sharon, M. The contribution of the 20S proteasome to proteostasis. Biomolecules 9, 190 (2019).
Sottile, M. L. & Nadin, S. B. Heat shock proteins and DNA repair mechanisms: An updated overview. Cell Stress Chaperones 23, 303–315 (2018).
Tjahjono, A. E. et al. Hyper-accumulation of astaxanthin in a green alga Haematococcus pluvialis at elevated temperatures. Biotechnol. Lett. 16, 133–138 (1994).
Ambati, R. R. et al. Industrial potential of carotenoid pigments from microalgae: Current trends and future prospects. Crit. Rev. Food Sci. Nutr. 59, 1880–1902 (2019).
Krasovec, M. et al. Genome analyses of the microalga Picochlorum provide insights into the evolution of thermotolerance in the green lineage. Genome Biol. Evol. 10, 2347–2365 (2018).
Janssen, J. H., Lamers, P. P., de Vos, R. C., Wijffels, R. H. & Barbosa, M. J. Translocation and de novo synthesis of eicosapentaenoic acid (EPA) during nitrogen starvation in Nannochloropsis gaditana. Algal Res. 37, 138–144 (2019).
Yao, S., Brandt, A., Egsgaard, H. & Gjermansen, C. Neutral lipid accumulation at elevated temperature in conditional mutants of two microalgae species. Plant Physiol. Biochem. 61, 71–79 (2012).
Hu, S., Ding, Y. & Zhu, C. Sensitivity and responses of chloroplasts to heat stress in plants. Front. Plant Sci. 11, 375 (2020).
Foflonker, F. et al. Genome of the halotolerant green alga Picochlorum sp. reveals strategies for thriving under fluctuating environmental conditions. Environ. Microbiol. 17, 412–426 (2015).
Barten, R., Djohan, Y., Evers, W., Wijffels, R. & Barbosa, M. Towards industrial production of microalgae without temperature control: The effect of diel temperature fluctuations on microalgal physiology. J. Biotechnol. 336, 56–63 (2021).
Breuer, G. et al. Analysis of fatty acid content and composition in microalgae. JoVE J. Vis. Exp. https://doi.org/10.3791/50628 (2013).
Andrews, S. et al. (2010).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucl. Acids Res. 28, 27–30 (2000).
Acknowledgements
We would like to acknowledge the Water and Energiebedrijf Bonaire (WEB) as project partner.
Funding
This work was supported by the Dutch Research Council (NWO) within NWO Caribbean Program with project number ALWCA.2016.029.
Author information
Authors and Affiliations
Contributions
R.J.P.B.: Conceptualization, Methodology, Data acquisition, Data analysis, writing—original manuscript. M.K.: methodology, data analysis, writing—review and editing G. D’Ermo: Methodology, data acquisition. H.N.: methodology, writing—review and editing. R.H. Wijffels: Writing—review and editing, funding acquisition. M.J.B.: conceptualization, writing—review and editing, methodology, funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Barten, R., Kleisman, M., D’Ermo, G. et al. Short-term physiologic response of the green microalga Picochlorum sp. (BPE23) to supra-optimal temperature. Sci Rep 12, 3290 (2022). https://doi.org/10.1038/s41598-022-06954-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-06954-6
- Springer Nature Limited
This article is cited by
-
UV mutagenesis improves growth potential of green algae in a green algae–yeast co-culture system
Archives of Microbiology (2024)
-
Global omics study of Tetraselmis chuii reveals time-related metabolic adaptations upon oxidative stress
Applied Microbiology and Biotechnology (2024)
-
Genetic mechanisms underlying increased microalgal thermotolerance, maximal growth rate, and yield on light following adaptive laboratory evolution
BMC Biology (2022)