Abstract
Mechanosensitive ion channels are pore-forming transmembrane proteins that allow ions to move down their electrochemical gradient in response to mechanical stimuli. They participate in many plant developmental processes including the maintenance of plastid shape, pollen tube growth, etc. Herein, a total of 11, 10, 6, 30, 9, and 8 MSL genes were identified in Aegilops tauschii, Hordeum vulgare, Sorghum bicolor, Triticum aestivum, Triticum urartu, and Zea mays, respectively. These genes were located on various chromosomes of their respective cereal, while MSLs of T. urartu were found on scaffolds. The phylogenetic analysis, subcellular localization, and sequence homology suggested clustering of MSLs into two classes. These genes consisted of cis-regulatory elements related to growth and development, responsive to light, hormone, and stress. Differential expression of various MSL genes in tissue developmental stages and stress conditions revealed their precise role in development and stress responses. Altered expression during CaCl2 stress suggested their role in Ca2+ homeostasis and signaling. The co-expression analysis suggested their interactions with other genes involved in growth, defense responses etc. A comparative expression profiling of paralogous genes revealed either retention of function or pseudo-functionalization. The present study unfolded various characteristics of MSLs in cereals, which will facilitate their in-depth functional characterization in future studies.
Similar content being viewed by others
Introduction
All organisms including bacteria, plants and animals, receive mechanical signals from both internal (osmotic pressure and plasma membrane distortion) and external environment (gravity, touch, sound)1. They respond to these signals by activating various channel proteins. A special type of channels that open and close in response to mechanical forces are known as mechanosensitive (MS) ion channels or stretch-gated ion channels1. They are pore-forming transmembrane (TM) proteins, which allow the movement of ions across the plasma membrane down to their electrochemical gradient. However, in plants, MS channels also act in response to signals related to developmental processes, such as cell wall damage, pollen tube growth, plant–pathogen interactions, and lateral root emergence2,3,4. Initially, MS channels were recognized during the electrophysiological characterization of the Escherichia coli cell envelope5,6,7. Based on the ionic conductance, the MS channels were divided into two main groups: mechanosensitive channels of large conductance (MscL) and mechanosensitive channels of small conductance (MscS)8,9. The ionic conductance of MscL and MscS corresponds to about three and one nanosiemens, respectively1. Previous studies suggested that MscL and MscS have no ion specificity while showing permeability to very small charged molecules such as potassium glutamate, proline10, etc. During hypo-osmotic shock, the bacterial MS channels act as a safety valve by allowing the diffusion of ions from the cell and prevent its bursting5,11,12.
Both MscL and MscS homologs are distributed in archaea, bacteria, and fungi, while MscS channels homologs are also found in algae and plants. A maximum of 10 MscS-Like (MSL) channels are found in model plant Arabidopsis thaliana, which are classified into two main classes: Class I MSL and Class II MSL13,14. Class I MSLs are found to be localized either in mitochondrial membrane or plastid envelop and show sequence similarity with MscS channels of bacteria. However, Class II MSLs are predicted to be present in the plasma membrane. Further, the characterization studies of MSL channels of A. thaliana suggested that AtMSL9 and AtMSL10 perform mechanosensitive activities in the plasma membrane of root cells15. Upon exposure to various stimuli, the AtMSL9 and AtMSL10 may increase cytosolic Ca2+ concentration in a small amount or they may depolarize the membrane, leading to activation of other Ca2+ channels16. In addition to it, AtMSL2 and AtMSL3 play a significant role in the regulation of plastid size and Z- rings formation17,18. Besides, AtMSL8 promotes pollen grain hydration and germination19. The MSL genes have been reported in other crops such as Oryza sativa, Phaseolus vulgaris, Cicer arietinum, etc.20,21,22. Due to their crucial role in plants during stress conditions and developmental stages, it is important to explore this gene family in other crop plants. In the current study, we identified and characterized MSL genes in six cereals such as Aegilops tauschii, Hordeum vulgare, Sorghum bicolor, Triticum aestivum, Triticum urartu and Zea mays. The identified genes and proteins were analyzed for structural configurations, chromosomal and subcellular distribution, cis-regulatory elements, duplication events, and phylogenetic relatedness. The expression profiling of MSL genes was carried out in abiotic and biotic stress conditions and, various tissue developmental stages using high throughput RNA sequence data. Further, we performed qRT-PCR for the expression study of a representative TaMSL gene from each phylogenetic group under CaCl2 stress conditions. Co-expression studies of MSL genes provided information about their interacting partners and their contribution to various biological processes of plants. This study provided valuable information about MSL genes, which will be useful in the future for their functional characterization.
Results and discussion
Identification and chromosomal distribution
A total of 74 putative MSL genes were identified after an extensive search, out of which diploid crops including Ae. tauschii, H. vulgare , S. bicolor, T. urartu, and Z. mays consisted of 11, 10, 6, 9, and 8 genes respectively (Supplementary File S1). S. bicolor have less number of genes as compared to other studied crops, which might be due to its small genome size23. We identified a maximum of 30 MSL genes in T. aestivum (Supplementary File S1), which could be due to its allohexaploid nature24. The previous findings have suggested 10 MSL genes in A. thaliana13,14, six in each O. sativa20 and C. arietinum22, and nine in P. vulgaris21. Therefore, the number of genes identified in the present study in six cereal plant species showed consistency with previous studies.
The chromosomal map of MSL genes indicates their variable distribution on six, five, four, and seven chromosomes of Ae. tauschii, H. vulgare, S. bicolor, and Z. mays, respectively (Fig. 1A–C,E). The MSL genes of T. aestivum were distributed on various chromosomes and subgenomes (A, B, and D) except chromosome 1B, 3A, 3B, 3D, and 4A. The highest number of TaMSL genes were found on chromosome 5A (Fig. 1D). However, in the case of T. urartu, scaffold locations of MSL genes were predicted. The distribution of MSL genes on various chromosomes has also been reported in other crops such as O. sativa, P. vulgaris, C. arietinum20,21,22.
Chromosomal localization and duplication analysis of MSL genes in five cereals. The AeMSL genes are distributed on 6 chromosomes of Ae. tauschii (A), HvMSL genes on 5 chromosomes of H. vulgare (B), SbMSL genes on 4 chromosomes of S. bicolor (C), TaMSL genes on 16 chromosomes of A, B and D subgenomes of T. aestivum (D), and ZmMSL genes on 7 chromosomes of Z. mays (E). Blue and Red dotted lines point toward the existence of tandemly and segmentally duplicated genes, respectively.
Prediction of homeologs and duplication events
According to previous studies, each progenitor subgenome (i.e. A, B, and D) has contributed to the formation of a large number of gene families in T. aestivum25,26,27 . We have identified 10 clusters of homeologous TaMSL genes and found that most of these clusters consisted of at least one gene from each subgenome. However, the homeologous group TaMSL1 was found on only A and D subgenome, while TaMSL9 was located on only A subgenome. The number of homeologous groups of TaMSL genes was found comparable to the number of MSL genes identified in the diploid plant species such as A. thaliana, P. vulgaris, and C. arietinum13,14,21,22. .
The duplication events lead to the origin of paralogous genes during evolution and play a major role in the expansion of gene families. They are essential for genetic variability and in gaining additional roles of genes, which further promote adaptation and speciation28,29,30. We analyzed duplication events (DEs) to study their contribution in the evolution of the MSL gene family. A total of 16 DEs were identified, out of which two were tandem duplication events (TDEs) and 13 were segmental duplication events (SDEs) (Supplementary File S2). One duplication event (TuMSL4-TuMSL7) found in T. urartu could not be categorized as TDE or SDE due to scaffold location of these genes (Supplementary File S2). Two TDEs (AeMSL1-AeMSL2; AeMSL5-AeMSL6) were found in Ae. tauschii, while one (HvMSL2-HvMSL7), nine (TaMSL3-A-TaMSL6-A; TaMSL3-B-TaMSL6-B; TaMSL3-D-TaMSL6-D, etc.) and three (ZmMSL3-ZmMSL8; ZmMSL4-ZmMSL5; ZmMSL6-ZmMSL7) SDEs were found in H. vulgare, T. aestivum and Z. mays, respectively (Fig. 1A,B,D,E). However, no duplication event was found to occur in the genome of S. bicolor. The DEs in MSL genes were earlier reported in some plants such as O. sativa, P. vulgaris and C. arietinum20,21,22. The variation in number of duplication events might be due to different genome size and ploidy level. For instance, the complex genome of T. aestivum is responsible for the occurrence of a large number of duplication events. The numerous incidences of SDEs have been attributed to the enlargement of gene families in T. aestivum that also supported the results obtained in the present study31,32,33.
Ka/Ks analysis
We have calculated the values of Ka, Ks and Ka/Ks ratio to know about the evolutionary divergence between the paralogous MSL genes (Table 1). The Ka/Ks ratio of all the duplicated gene pairs was less than one, which suggested negative or purifying selection pressure throughout the evolution. Due to non-availability of complete sequence of AeMSL2, it was not possible to perform Ka/Ks analysis of duplicated gene pair AeMSL1-AeMSL2.
Further, the divergence time of DEs was also calculated and it was predicted to be in the range of 16 to 110 Million years ago (MYA). The maximum and minimum divergent time of DEs were found in T. aestivum and Z. mays, respectively (Table 1). In previous study, it was estimated that main DEs were occurred ~ 67 MYA and ~ 11 MYA in H. vulgare and Zea mays, respectively34. The predicted duplications in these plants were in similar range with slight deviation, as it was ~ 36MYA in HvMSLs, and ~ 16–24 MYA in ZmMSLs (Table 1). In case of TaMSL paralogs, the DEs were predicted in the diverse range from ~ 30 to 110 MYA. The results suggested the occurrence of these DEs before the hybridization of A, B and D subgenomes that occurred in the range of ~ 0.5–6 MYA24. Moreover, higher rate of sequence divergence among the duplicated gene pairs during evolution could also be the region for the deviation in the years of predicted duplication events, as it was completely based upon sequence analysis.
Phylogenetic analysis
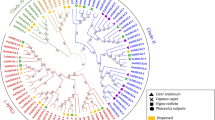
The phylogenetic tree was prepared to study the evolutionary relatedness among the MSL proteins of Ae. tauschii, A. thaliana, H. vulgare, O. sativa, S. bicolor, T. aestivum, T. urartu, and Z. mays using the full length protein sequences. In the phylogenetic tree, MSL proteins were clustered into two main classes i.e. class I and II (Fig. 2A). Similar classification of proteins has also been reported in A. thaliana17 O. sativa20, P. vulgaris21 and C. arietinum22. The class I and class II consisted of 31 and 59 MSL proteins, respectively. Two MSL proteins of Z. mays, three of each Ae. tauschii, A. thaliana, O. sativa, S. bicolor and T. urartu, four of H. vulgare, and 10 of T. aestivum were clustered in class I. While, three MSL proteins of each O. sativa and S. bicolor, six of each H. vulgare, T.urartu and Z. mays, seven of A. thaliana, eight of Ae. tauschii, and 20 of T. aestivum showed close relationships and found in class II of phylogenetic tree (Fig. 2A).
Phylogenetic tree, cis-regulatory element analysis, and gene structure (exon–intron organization and intron phases) analysis. (A) Shows a phylogenetic tree constructed by the neighborhood joining method with 1000 bootstraps using MEGA7.0 software. The tree shows two major classes i.e. class I and class II, which further divided into sub-classes highlighted with different colors. (B–G) Show the percentage of Cis-regulatory elements including light-responsive elements, stress-responsive elements, hormone-responsive elements, and growth and development responsive elements of MSL genes of Ae. tauschii (B), H. vulgare (C), S. bicolor (D), T. aestivum (E), T. urartu (F), and Z. mays (G). (H–M) Show the structure of MSL genes of A. tauschii (H), H. vulgare (I), T. aestivum (J), S. bicolor (K), T. urartu (L), and Z. mays (M) prepared by using GSDS 2.0. server.
The majority of paralogous genes were found close to each other, for instance, ZmMSL3-ZmMSL8, ZmMSL4-ZmMSL5, AeMSL1-AeMSL2, TaMSL3-A-TaMSL6-A, etc. The structural relatedness among the paralogous genes might be responsible for close clustering. The homeologous TaMSL of T. aestivum were tightly clustered due to their sequence resemblance at high bootstrap value (Fig. 2A). This was also in confirmation with the used nomenclature of TaMSL genes.
Promoter analysis
Promoter analysis suggested the occurrence of a large number of cis-regulatory elements in the promoter regions of MSL genes, which were responsive to light, hormone, stress, and growth and development (Fig. 2B–G). Some light-responsive elements including Sp1, G-box, G-Box, ACE, and GT1-motif were identified in most of MSL genes. STRE, as-1, MYB, MYC, WRE3, GC-motif, DRE core, TC-rich repeats, MYB recognition site, etc. were commonly occurring stress-responsive elements in MSL genes. Hormone-responsive elements such as ABRE, CGTCA-motif, TGACG-motif, ABRE3a, and ABRE4 were most frequently found in MSL genes. Furthermore, the growth and development related cis-regulatory elements including; CAT-box, Myb-binding site, CCGTCC-box, CCGTCC motif, AAGAA-motif was detected in most of the MSL genes. In Ae. tauschii, 37% cis-elements were found to be responsive to stress, 26% to light, 22% to growth and development, and 15% to hormones (Fig. 2B). In H. vulgare, 34% cis-elements were responsive to stress, 28% to light and 23% to growth and development, and 15% to hormone (Fig. 2C). In S. bicolor, 34% of cis-elements were found as light-responsive, 31% as stress-responsive, 21% as hormone-responsive, and 14% as growth and development responsive (Fig. 2D). In T. aestivum, 29% cis-elements were predicted to be involved in growth and development, and 28% responsive to light, 25% responsive to stress, 18% to hormone (Fig. 2E). In T. urartu, 31% elements were responsive to stress, 29% to light, 26% to growth and development, and 14% hormone-responsive (Fig. 2F). Similarly, in the case of Z. mays, we found that 32% cis-elements were responsive to stress, 29% to light, 22% to growth and development, and 17% to hormone (Fig. 2G). Besides, we found that the arrangement of cis-regulatory elements varies in each MSL gene. Enrichment of cis-regulatory elements in the MSL gene promoters revealed their numerous functions in each cereal.
Gene structure analysis
The exon/intron organization and intron phases were examined to get information about the structural features of MSL genes. The number of exons ranged from 3 to 12 in Ae. tauschii, 4 to 13 in H. vulgare and T .aestivum, 5 to 13 in S. bicolor and Z. mays, and 5 to 10 in T. urartu. (Supplementary File S3, Fig. 2H–M). The majority of MSL genes carried five exons for each cereal except T. urartu, where six exons were present in the majority of genes. Further, we found that the class I MSL genes consisted of 6–13 exons, while class II MSL genes consisted of 3 to 10 exons in all the studied crops. The maximum number of introns were present in 0 phase followed by 2 phase and then 1 phase, which indicated the conserved architecture of MSL genes (Fig. 2H–M). The previous studies regarding gene structure of MSLs suggested the number of exons from 5 to 19 in O. sativa20, 4 to 12 in P. vulgaris21 and 5 to 8 in C. arietinum22 which showed consistency with the structural organization of MSLs in our studied crops.
Protein characterization
The physicochemical properties of MSL proteins were analyzed using computational approaches. The predicted average length, molecular weight (MW) and isoelectric point (pI) of MSL proteins from Ae. tauschii, H. vulgare, S. bicolor, T. aestivum, T. urartu and Z. mays ranged from 680 to 782 AA, 73.4 to 90.6 kDa and, 9.1 to 9.7, respectively (Supplementary File S3). These findings were similar to the previously reported MSL proteins in other plants20,21,22.
In each cereal crop, class I MSL proteins consisted of five or six TM helices, while class II MSL proteins carried five to eight TM helices (Supplementary File S3). However, AeMSL2 and TuMSL5 protein exhibited the occurrence of only one and two TM helices, respectively, which could be due to the partial sequence of these genes. Previous studies have suggested the presence of five TMs in class I and six TMs in class II MSLs of A. thaliana14 and O. sativa22, which were comparable to our findings.
The subcellular localization prediction of MSL proteins of each cereal suggested that class I MSL proteins were either located on chloroplastic envelop or mitochondrial membrane, while class II proteins were located on the plasma membrane (Supplementary File S3). Similar results have been reported in other crops including A. thaliana, O. sativa , P. vulgaris and C. arietinum14,20,21,22.
The functionality of a protein depends upon the characteristic features of its domain. Pfam analysis suggested that the MS-channel domain (PF00924) was present in all MSL proteins of Ae. tauschii, H. vulgare, S. bicolor, T. aestivum, T. urartu and Z. mays (Supplementary File S4). Further, the occurrence of 15 conserved motifs was examined among MSL proteins of each cereal. Motif 3 present in MS channel domain was found to be highly conserved, due to its occurrence in all the MSL proteins. Motif 10 was also distributed in all proteins except AeMSL2. The majority of identified motifs were present in class II MSL proteins, while motifs 3, 7, 8, 10, and 13 were present in the members of class I MSL (Fig. 3A). Furthermore, protein kinase C phosphorylation site, casein kinase 2 phosphorylation site, tyrosine kinase site could be detected in the motifs 1, 2, 4, 5,7, 8, and 9. These motifs were predominately found in most of class II proteins and few of class I proteins. These finding suggested that these proteins might also be responsible for phosphorylation. Similar results have been reported in other crops such as O. sativa20, P. vulgaris21 and C. arietinum22.
Motif analysis and multiple sequence alignment of MSL proteins. (A) Shows the distribution of 15 conserved motifs in MSL proteins. The colored boxes represent these conserved motifs constructed by the MEME suite. (B) Shows multiple sequence alignments with the conserved transmembrane region (TM) and a conserved motif in each class of MSL proteins.
The identified MSL proteins in our present study were aligned along with MSL protein sequences of A. thaliana and O. sativa, to study the conserved residues among them (Fig. 3B). A conserved TM region was found in the C-terminal half of the MSL proteins of both the classes. Further, motifs PF(X12–16)GXV(X20–21)PN(X9)N and F(X3)P(X3)GD(X10–14)V(X20–21)PN(X7)IXNXXR were also found conserved at C-terminus in class I and class II MSL proteins, respectively (Fig. 3B). The C-terminal conserved TM domain of class I MSLs exhibited similarity with TM3 of E. coli MscS and similar findings has also been described in other crops13,20,21,22. Moreover, the TM domains of class I MSL proteins consisted of high proportion of glycine and alanine residues, while the class II MSL proteins had large hydrophobic side chains (Fig. 3B). Similar conserved domain and motifs were reported in MSL proteins of A. thaliana, O. sativa, P. vulgaris, and C. arietinum14,20,21,22.
Expression analysis in tissue development stages
Expression analysis of MSL genes was carried out to study their involvement in the developmental processes of plants. We performed expression analysis in H. vulgare, S. bicolor, T. aestivum and Z. mays (Supplementary File S5, Fig. 4A–D). We selected those MSL genes for analysis, which showed expression ≥ 2 FPKM in a minimum of one developmental stage (Supplementary File S5). In H. vulgare, most of the HvMSLs exhibited expression in various development stages. HvMSL1, HvMSL8, HvMSL9 and HvMSL10 showed prominent expression in reproductive tissues including inflorescence, caryopsis, germinating embryo that suggested its involvement in the development of reproductive tissues (Supplementary File S5, Fig. 4A). However, higher expression of HvMSL3 in shoot, root and internode indicated their role in vegetative tissue development.
Expression profiling in various tissue developmental stages and co-expression analysis of MSL genes of H. vulgare, S. bicolor, T. aestivum, and Z. mays. Heat maps (A–D) show the expression profiling of MSL genes of H.vulgare (A), S. bicolor (B), T. aestivum (C) and, Z. mays (D). (E–H) Show the gene interaction networks of MSL genes of H. vulgare (E), S. bicolor (F), T. aestivum, and (G) Z. mays (H), based on co-expression analyses and generated using Gephi 0.9.1.
In S. bicolor, SbMSL1 exhibited significant expression in shoot, leaf, flower, anther, and emerging inflorescence, which revealed its involvement in the development of both vegetative and reproductive tissues. Moreover, SbMSL3, SbMSL5, and SbMSL6 showed expression in most of the tissues or organs. SbMSL3 and SbML6 exhibited elevated expression in the vegetative and floral meristem, while SbMSL3 also exhibited enhanced expression in plant embryo and seed 5 day after pollination (DPA). SbMSL2 showed less expression in almost all the tissues. SbMSL4 was found to be expressed only in anther and flower suggested their participation in the development of reproductive organs (Supplementary File S5, Fig. 4B).
In T. aestivum, 16 TaMSL genes showed expression in various tissue developmental stages (Supplementry File S5, Fig. 4C). The majority of TaMSL genes were highly expressed in spike followed by the stem, leaf, grain, and root tissues. Moreover, these TaMSL genes exhibited higher expression in the later stages of tissue development. On the other hand, it was found that TaMSL5-B and TaMSL5-D showed higher expression in all the tissue development stages as compared to the rest of TaMSL genes. TaMSL2 and TaMSL8 group genes exhibited higher expression in all the developmental stages of root tissue, which suggested that they might be preferable involved in absorptive transportation of related ions in root.
In case of Z. mays, ZmMSL1, ZmMSL2, ZmMSL5 and ZmMSL8 showed significant expression in all the tissues development stages, which suggested their role throughout the plant development. Moreover, ZmMSL5 exhibited quite higher expression in some tissues like tip of ear primordium, embryo 20 DPA, etc. and ZmMSL7 was highly expressed in various reproductive tissues (Supplementary File S5, Fig. 4D).
The significant expression of the majority of MSL genes in various tissue developmental stages suggested their roles in the growth and development. In the case of both A. thaliana and O. sativa, the majority of MSL genes were expressing in various tissue/organs throughout the plant life14,35. OsMSL3 specifically shows a characteristically high expression in almost all the tissues and organs of O. sativa20. In the present study, the orthologous TaMSL5 group genes were also high expressing in one or more stages of the tissue development in leaf, stem, spike, and grain. Furthermore, in the case of H. vulgare, S. bicolor, Z. mays, the orthologous genes HvMSL3, SbMSL1, and ZmMSL1 were also high expressing in all the developmental stages (Fig. 4A–D). Moreover, the majority of the OsMSL genes of rice are significantly expressing in reproductive stages20 . In the present study, several MSLs including HvMSL8, SbMSL1, SbMSL4, SbMSL6, ZmMSL7 and TaMSL5 and TaMSL7 group genes were highly expressed in reproductive stages. The putative role of MSL genes in plant development was further reinforced through the occurrence of various growth and development responsive cis-regulatory elements like CCGTCC-box, CCGTCC motif, CTAG-motif, Circadian, CAT-box, etc.
Co-expression analysis
We analyzed the co-expression of MSL genes with other genes of their respective crop in tissue development stages of H. vulgare, S. bicolor, T. aestivum and Z. mays (Supplementary File S6, Fig. 4E–H). In H. vulgare, three HvMSL genes including HvMSL3, HvMSL10, and HvMSL8 were found to be co-expressed with 65 transcripts. HvMSL8 showed co-expression with the maximum number of transcripts (56) which encoded Histone H4, a serine-threonine kinase, DNA topoisomerase 2, E3 ubiquitin- ligase, etc. (Supplementary File S6, Fig. 4E).
In the case of S. bicolor, four SbMSL genes revealed their co-expression with a total of 96 transcripts encoding ethylene receptor 2, Zinc-finger homeodomain, subtilisin-like protease, callose synthase 7, etc. (Supplementary File S6, Fig. 4F). In T. aestivum, six TaMSL genes showed co-expression with 21 transcripts encoding early nodulin 2, elongation factor 2, dehydration-induced 19, Beta-glucosidase 31, etc. (Supplementary File S6, Fig. 4G). Similarly, in Z. mays, four ZmMSL genes were co-expressing with 53 transcripts, among them ZmMSL7 exhibited co-expression with 36 transcripts. These transcripts encoded LRR receptor-like serine-threonine kinase, L-ascorbate oxidase, E3 ubiquitin-ligase, Peroxidase 1, etc. (Supplementary File S6, Fig. 4H). The gene ontology (GO) mapping of co-expressed transcripts of H. vulgare, S. bicolor, T. aestivum and Z. mays, suggested their involvement in oxidative stress (GO: 0006979), oxidation–reduction process (GO: 0055114), iron ion binding (GO: 0005506), Zinc ion binding (GO: 0008270), electron carrier activity (GO: 0009055), peroxidase activity (GO: 0004601), defense response to fungus (GO: 0050832), voltage-gated anion channel activity (GO: 0008308), catalytic activity (GO: 0003824), response to salicylic acid (GO:0009751), etc. (Supplementary File S6).
We have concluded that the co-expression of MSLs with other genes involved in numerous biological processes suggested their diverse functions. The GO mapping of co-expressed transcripts suggested the participation of MSL genes in growth and development. For instance, one of MSL gene i.e. HvMSL8 found to be co-expressed with ABC transporter, which was reported to be involved in pollen growth development36. Thus, HvMSL8 can have probable role in pollen growth and development. We analyzed that MSL genes were co-expressed with cellulose synthase, suggested their involvement in cell wall development. Further, the co-expression of MSL genes with transcripts encoding kinases such as serine-threonine kinase, cysteine rich receptor kinase etc. suggested their roles in signaling pathways, and stress responses.
Comparative expression profiling of paralogous genes
The duplication events play a significant contribution to the origin of paralogous genes during evolution. Due to their participation in the formation of similar protein complexes, it is important to coordinate their expression within the cell37. Therefore, we performed comparative tissue expression profiling of paralogous MSL gene pairs using available high throughput RNA seq data38,39,40. The paralogous gene pairs could be divided into three classes; retention of expression, neo-functionalization, and pseudo-functionalization, based on their expression patterns. However, MSL paralogues genes exhibited only pseudo-functionalization and retention of functions, which suggested functional conservation after duplication. The segmentally duplicated gene pair of H. vulgare (HvMSL2 and HvMSL7) exhibited pseudo-functionalization, due to the insignificant expression of one gene (Fig. 5A). However, in the case of Z. mays, three segmentally duplicated genes pairs such as ZmMSL3: ZmMSL8; ZmMSL4: ZmMSL5 and ZmMSL6: ZmMSL7 revealed their existence in class retention of expression due to their comparable expression pattern (Fig. 5K–M). Similarly, in T. aestivum, six segmentally duplicated gene pairs (TaMSL3-A: TaMSL6-A; TaMSL3-B: TaMSL6-B; TaMSL3-D: TaMSL6-D; TaMSL5-A: TaMSL10-A; TaMSL5-B: TaMSL10-B; TaMSL5-D: TaMSL10-D) also exhibited retention of function (Fig. 5B–D,H–J). However, three segmentally duplicated gene pairs (TaMSL4-A1: TaMSL7-A; TaMSL4-B2: TaMSL7-B and TaMSL4-D1: TaMSL7-D) showed pseudofunctionalization (Fig. 5E–G). Due to the non-availability of expression data, we could not perform the comparative expression analysis for paralogous genes of Ae. tauschii and T. urartu. Moreover, the results from above analysis revealed that despite of duplication during evolution, the function of paralogous MSL genes are still conserved.
Comparative expression profiling of paralogous genes. (A,B–J,K–M) shows the comparative expression profiling of duplicated genes of H. vulgaris, T. aestivum, and Z. mays, respectively. Based on expression pattern, duplicated gene pairs have been classified into the retention of expression (B–D,H–J,K–M) and pseudo-functionalization (A,E–G).
Expression profiling of TaMSL under biotic and abiotic stresses
Various environmental stresses such as abiotic and biotic stresses affect plant growth and development, which leads to the loss of yield and quality41,42. The expression patterns of TaMSL genes were analyzed to understand their role in biotic and abiotic stress responses.
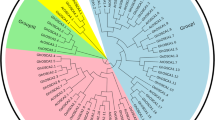
Two fungal pathogens i.e. Puccinia striiformis f. sp. tritici (Pst) and Blumeria graminis f. sp. tritici (Bgt) were used for expression analysis of MSL genes under biotic stresses using high throughput RNA seq data43. The MSL genes, which showed a fold change of ≥ 2, were selected for the study (Supplementary File S7). TaMSL genes were clustered in two groups, based on their relatively similar expression. All TaMSL genes of group I was down regulated after Bgt and Pst infestation except TaMSL1-A which was up-regulated after 24 h of Pst attack. However, in group II, the majority of TaMSL genes were up-regulated after Bgt and down-regulated after Pst infestation. Moreover, TaMSL2-B and TaMSL2-D were down-regulated after 24 h of Bgt infestation and TaMSL2-A, TaMSL5-D and TaMSL2-D were up-regulated after 24, 48 and 72 h of Pst inoculation, respectively (Supplementary File S7, Fig. 6A).
Expression profiling of TaMSL genes under biotic and abiotic stress conditions in T. aestivum. Heat maps (A–C) show the expression profile of up and downregulated genes under (A) biotic, (B) DS, HS, and HD and, (C) salt stress. Bar graphs (D–G) show the qRT-PCR results of four TaMSL genes under CaCl2 stress at 6, 12, 24, and 48 h of treatments. The bar graphs indicate the fold change of each TaMSL gene expression and a vertical line indicate standard deviation. A significant mean difference (≤ 0.05) level is shown by different letters on the top of each vertical bar.
We also analyzed the expression profiling of TaMSL genes under abiotic stress conditions such as drought stress (DS), heat stress (HS), heat drought (HD), and salt stress by using high throughput RNA seq data44,45. A total of 14 TaMSL genes were differentially expressed (≥ 2) under these stresses (Fig. 6B,C, Supplementary File S7). The majority of TaMSL genes were upregulated at later stages (6 h) of DS, HS, and HD treatment. However, a few genes such as TaMSL7-A, and TaMSL7-B were upregulated at 1 h of DS. TaMSL7-D and TaMSL8-D were upregulated at each hour of DS, HS, and HD treatment. In contrary, TaMSL1-D and TaMSL4-D2 were downregulated in each treatment. Similarly, OsMSL6 and OsMSL3 of O. sativa show significant expression in response to drought stress20. Their orthologs, TaMSL8-D, TaMSL5-A and TaMSL5-D were also upregulated in drought stress (Fig. 6B).
Based on the similar expression patterns, TaMSL genes were clustered into two groups under salt stress (Fig. 6C). All TaMSL genes of group I showed upregulation during every hour of salt stress except TaMSL4-A1, which was downregulated at 6 and 12 h. In group II, the majority of TaMSL genes were downregulated, while certain genes such as TaMSL2-A and TaMSL2-B were upregulated at 6 h and 12 h of salt stress (Fig. 6C). TaMSL5-B was upregulated at every hour of salt stress treatment similar to its orthologs OsMSL3 of O. sativa, which was also significantly upregulated during salt stress20.
The differential expression of TaMSL genes during various biotic and abiotic stresses along with the occurrence of various stress-responsive cis-regulatory elements including as-1, MYB, MYC, WRE3, GC-motif, DRE core etc., suggested their involvement in stress tolerance. All these stresses result in an increase in cytoplasmic Ca2+. Since the MSL proteins are also involved in Ca2+ transport, the differential expression of TaMSL genes along with their co-expression with various signaling related genes, suggested their roles in Ca2+ homeostasis and signaling during these stress conditions.
Expression analysis during CaCl2 stress
Ca2+ is an important secondary messenger involved in signaling46. During stress conditions, the cytoplasmic Ca2+ concentration elevated by usually tenfolds, resulted in activation of signaling process and activation of various calcium transport elements to restore the cytoplasmic Ca2+ concentration by the process called calcium homeostasis47. Since the MSL proteins are also associated with the influx of Ca2+ and its homeostasis16, expression analysis of TaMSL genes was performed under CaCl2 stress using qRT-PCR. A total of four TaMSL genes (TaMSL5-B from class I, and TaMSL4-D2, TaMSL7-D, and TaMSL8-D from class II) were selected for qRT-PCR that represented each major phylogenetic group (Fig. 6D–G). The expression pattern of TaMSL4-D2, TaMSL5-B, TaMSL7-D, and TaMSL8-D was similar at every hour of CaCl2 stress. TaMSL5-B, TaMSL7-D, and TaMSL8-D were significantly upregulated (> tenfolds) in the initial hours (6 and 12 h) of treatment. However, all these genes get normalized at later stages of treatment (Fig. 6D–G).
The elevated expression of class I TaMSL gene; TaMSL5-B, which was predicted to be localized in chloroplastic membrane, might be associated with the transport of Ca2+ from this organelle to the cytoplasm and stress signaling, as well. On the other hand, enhanced expression of class II TaMSL genes suggested that they might initially be activated and subsequently stimulated other Ca2+ channels through depolarization of the plasma membrane. These Ca2+ channels further help in Ca2+influx into the cytosol, which in turn activates various signaling pathways for the adaptation of plants under stress conditions. Further, the occurrence of various Ca2+ and phytohormone responsive cis-regulatory elements such ABRE, DRE, W-box etc. in the promoter region of MSL genes also suggested their involvement in Ca2+ homeostasis and signaling.
Conclusions
The present study involved identification, classification and detailed characterization of MSL channels from six cereal crops. The occurrence of cis-regulatory elements in MSLs suggested their roles in the diverse physiological phenomenon. Expression profiling in various tissue developmental stages highlighted the role of MSLs in the growth and development. Co-expression analyses and gene ontology mapping revealed the plausible role of MSLs in signaling, stress responses, etc. Comparative expression profiling of paralogous genes showed either retention of expression or pseudo-functionalization. Abiotic and biotic stress responsive differential expression of TaMSL genes revealed their role in stress response. Enhanced expression of MSLs under CaCl2 stress suggested their involvement in Ca2+ homeostasis and signaling. However, the specific role of the individual MSL gene should be functionally validated in future studies.
Methods
Identification and nomenclature of MSL genes
To identify the MSL proteins in Ae. taushii, H. vulgare, S. bicolor, T. aestivum, T. urartu and Z. mays, a BLASTp search was performed at Ensembl Plants (plants.ensembl.org/index.html). The MSL protein sequences of A. thaliana and O.sativa were used as a query against the protein model sequences of these six cereals. Further, the MS-channel (PF00924) domain was confirmed in all the extracted MSL proteins by performing Pfam BLAST search (10–10) with Hidden Markov Model (HMM)48. This domain was further ensured within each MSL protein through the SMART49 and the NCBI Conserved Domain Database (CDD) BLAST servers50.
The international guidelines for gene symbolization were followed for the nomenclature of MSL of T. aestivum (https://wheat.pw.usda.gov/ggpages/wgc/98/Intro.htm). However, in case of Ae. taushii, H. vulgare, S. bicolor, T. urartu and Z. mays names were given to MSL genes according to their location on respective chromosome.
Chromosomal localization, homeologs and paralogs analysis
Ensembl Plants was used to retrieve the information regarding the chromosomal and genome-wide distribution of MSL genes (https://plants.ensembl.org/Triticum_aestivum/). Chromosomal maps were prepared using Map Inspect (http://mapinspect.software.informer.com/), to depict the location of each gene on the respective chromosome. TaMSL genes of T. aestivum were grouped as homeologs based on their locations on the homeologous chromosomes and by performing bidirectional BLAST search with ≥ 90% sequence similarity. The paralogous genes formed at the time of duplication events were identified by using bidirectional blast search (10−10) with ≥ 80% sequence similarity. Based on the distance between the chromosomal localization of paralogous genes, they were considered as either segmentally or tandemly duplicated51.
Ka/Ks calculations
The nucleotide and protein sequences of the duplicated MSL gene pairs were aligned using ClustalOmega server (https://www.ebi.ac.uk/Tools/msa/clustalo). Further, PAL2NAL server was used for the calculation of non-synonymous substitution per non-synonymous site (Ka), synonymous substitution per synonymous site (Ks), and Ka/Ks ratio52. The divergence time of each duplicated MSL gene pair was calculated by applying the formula T = Ks/2r, whereas T indicates the divergence time and r indicates the divergence rate. The value of divergence rate was implicated as 6.5 × 10–9 in cereals53.
Multiple sequence alignment and phylogenetic relationship
Full-length amino acid (AA) sequences of each MSL protein were aligned via Multalin and MUSCLE programs54 to predict the conserved residues. The phylogenetic tree was generated by MEGA 7 software using the neighbor-joining method with 1000 bootstrap replicates55.
Promoter analysis
An upstream promoter sequence (~ 1.5 kb) of each MSL was extracted from genomic sequences of all the studied cereal plants. By using the PlantCARE database, cis-regulatory elements were investigated among the extracted promoter sequences56.
Gene structure analysis
The MSL gene structures were constructed by Gene structure display server (GSDS2.0) using genomic and coding sequences (CDS) of each cereal57. The structural features were visualized in form of intron/exon organization and presence of various intron phases.
Protein characterization
Various physicochemical properties of MSL proteins such as MW and pI were calculated using standard parameters by the Expasy MW/pI tool58. Phyre2 tool was used to predict the number of TM regions in MSL proteins59. To determine the subcellular distribution of MSL proteins, the Prot comp 9.0 tool (https://linux1.softberry.com/berry.phtml) was used. Moreover, the Chlorop v1.1 program was used for the re-confirmation of chloroplastic localization of MSL proteins60. Multiple Expectation Maximization for Motif Elicitation (MEME- suite version 5.0.1), was used for the investigation of conserved motifs within MSL proteins61.
Expression analysis of MSL genes
To study the expression patterns of MSL genes in various tissue development stages of H. vulgare, S. bicolor and Z. mays, the RNA seq data was retrieved from the expression ATLAS database38. However, in the case of T. aestivum, the publically available RNA seq data were obtained from NCBI and URGI databases39,40 (wheaturgi.versailles.inra.fr/files/RNASeqWheat/). Then, trinity package was used for the calculation of expression values in the form of FPKM (fragment per kilobase per million reads) in duplicates (n = 2)62.
The expression patterns were analyzed under biotic and abiotic stress conditions in T. aestivum. The expression of TaMSL genes was studied under biotic stresses by using publically available RNA seq data43 from leaves after 24, 48 and 72 h of fungal infestation of Blumeria graminis f. sp. tritici (Bgt) and Puccinia striiformis f. sp. tritici (Pst) in triplicates (n = 3). Expression study under abiotic stresses such as heat (HS), drought (DS), and their combination (HD) was done by using the available RNA seq data generated in duplicates (n = 2) after 1 and 6 h of these stresses from leaves44. Moreover, RNA seq data generated in triplicates (n = 3) by Zhang et al. in 2016, after 6, 12, 24, and 48 h of salt treatment from root tissues were used for expression profiling of TaMSL genes under salt stress45. Heat maps were prepared using Hierarchical Clustering Explorer 3.563.
Co-expression analysis and gene ontology mapping
The CoExpress v.1.5. tool was used for the analysis of co-expression of MSL genes of H. vulgare, S. bicolor, T. aestivum and Z. mays. The co-expression values were calculated by using the Pearson correlation coefficient64 with a correlation power 1 and threshold filter ≥ 0.9. Gene ontology (GO) mapping and functional annotation of co-expressed genes were done using the Blast2GO tool65. Interaction networks of co-expressed genes were made by the Gephi 0.9.1 tool66.
qRT-PCR
Firstly, the surface sterilization of seeds of T. aestivum (cv. chinese spring) was done using sodium hypochlorite (1.2%) in 10% ethanol. Then, sterilized seeds were kept overnight at 4 °C for stratification after washing with double autoclaved water. The seeds were allowed to germinate at room temperature. After germination, these seedlings were placed in fresh phytaboxes and were grown in the plant growth chamber using autoclaved water. CaCl2 (20 mM) treatment was given to 7-day-old seedlings with growth media i.e. Murashige and Skoog (MS) and then, samples were collected in liquid nitrogen after 6, 12, 24, and 48 h. RNA was extracted using the Spectrum TM Plant Total RNA kit (Sigma, USA) from root tissues. Traces of DNA contamination were removed using the TURBO DNA-free™ Kit (Invitrogen, USA). RNA samples were examined both qualitatively and quantitatively through agarose gel electrophoresis and Nanodrop spectrophotometer, respectively. Superscript III First-Strand Synthesis Super-mix (Invitrogen, USA) was used for cDNA synthesis from one microgram of RNA. By using SYBR Green and gene-specific primers (Supplementary Table S1) of chosen TaMSL genes, qRT-PCR was done at CFX96 Real-Time PCR (BioRad, USA) by tracking already established protocol67. An ADP-ribosylation factor (TaARF1) was taken for the internal control and the delta-delta CT method (2-ΔΔCT) was used for calculating the expression values68. All the experiments were performed in triplicates (n = 3) and represented in term of mean ± standard deviation (SD). After Analysis of Variance, we performed the post hoc Tukey’s test at p-value < 0.05, to analyze the significant mean difference between control and treatments using SPSS 16.0 software.
References
Peyronnet, R., Tran, D., Girault, T. & Frachisse, J. M. Mechanosensitive channels: Feeling tension in a world under pressure. Front. Plant Sci. 5, 558 (2014).
Appel, H. M. & Cocroft, R. B. Plants respond to leaf vibrations caused by insect herbivore chewing. Oecologia 175, 1257–1266 (2014).
Jayaraman, D., Gilroy, S. & Ane, J. M. Staying in touch: Mechanical signals in plant–microbe interactions. Curr. Opin. Plant Biol. 20, 104–109 (2014).
Lucas, M. et al. Lateral root morphogenesis is dependent on the mechanical properties of the overlaying tissues. Proc. Natl. Acad. Sci. 110, 5229–5234 (2013).
Martinac, B., Buechner, M., Delcour, A. H., Adler, J. & Kung, C. Pressure-sensitive ion channel in Escherichia coli. Proc. Natl. Acad. Sci. 84, 2297–2301 (1987).
Sukharev, S. I., Martinac, B., Arshavsky, V. Y. & Kung, C. H. Two types of mechanosensitive channels in the Escherichia coli cell envelope: Solubilization and functional reconstitution. Biophys. J . 65, 177–183 (1993).
Berrier, C., Besnard, M., Ajouz, B., Coulombe, A. & Ghazi, A. Multiple mechanosensitive ion channels from Escherichia coli, activated at different thresholds of applied pressure. J. Membr. Biol. 151, 175–187 (1996).
Edwards, M. D. et al. Characterization of three novel mechanosensitive channel activities in Escherichia coli. Channels 6, 272–281 (2012).
Cox, C. D., Nakayama, Y., Nomura, T. & Martinac, B. The evolutionary ‘tinkering’ofMscS-like channels: Generation of structural and functional diversity. PflügersArchiv-Eur. J. Physiol. 467, 3–13 (2015).
Hamill, O. P. & Martinac, B. Molecular basis of mechanotransduction in living cells. Physiol. Rev. 81, 685–740 (2001).
Levina, N. et al. Protection of Escherichia coli cells against extreme turgor by activation of MscS and MscLmechanosensitive channels: Identification of genes required for MscS activity. EMBO J. 18, 1730–1737 (1999).
Booth I.R., Edwards, M.D., Murray E. & Miller, S. The role of bacterial channels in cell physiology. In Bacterial Ion Channels and Their Eukaryotic Homologs 291–312 (2005).
Pivetti, C. D. et al. Two families of mechanosensitive channel proteins. Microbiol. Mol. Biol. Rev. 67, 66–85 (2003).
Haswell, E. S. MscS-like proteins in plants. Curr. Top. Membr. 58, 329–359 (2007).
Haswell, E. S., Peyronnet, R., Barbier-Brygoo, H., Meyerowitz, E. M. & Frachisse, J. M. Two MscS homologs provide mechanosensitive channel activities in the Arabidopsis root. Curr. Biol. 18, 730–734 (2008).
Peyronnet, R., Haswell, E. S., Barbier-Brygoo, H. & Frachisse, J. M. AtMSL9 and AtMSL10: Sensors of plasma membrane tension in Arabidopsis roots. Plant Signal. Behav. 3, 726–729 (2008).
Haswell, E. S. & Meyerowitz, E. M. MscS-like proteins control plastid size and shape in Arabidopsis thaliana. Curr. Biol. 16, 1–11 (2006).
Wilson, M. E., Jensen, G. S. & Haswell, E. S. Two mechanosensitive channel homologs influence division ring placement in Arabidopsis chloroplasts. Plant Cell 23, 2939–2949 (2011).
Hamilton, E. S. et al. Mechanosensitive channel MSL8 regulates osmotic forces during pollen hydration and germination. Science 350, 438–441 (2015).
Saddhe, A. A. & Kumar, K. In silico identification and expression analysis of MscS like gene family in rice. Plant Gene 1, 8–17 (2015).
de Souza, S. G. H., da Silva, A. P., Ito, T. M., da Silva, E. G. & Cantú, T. Genome-wide analysis of mechanosensitive channel of small conductance (MscS)-like gene family in common bean. Afr. J. Biotechnol. 15, 580–592 (2016).
Basumatary, N., Kumari, A. & Sarmah, J. Genome-wide identification of MscS like gene family in Cicer arietinum using bioinformatics approach. Ann. Plant Sci. 7(2), 2002–2012 (2018).
Paterson, A. H. et al. The Sorghum bicolor genome and the diversification of grasses. Nature 457, 551–556 (2009).
Marcussen, T. et al. Ancient hybridizations among the ancestral genomes of bread wheat. Science 345, 1250092–1250092. https://doi.org/10.1126/science.1250092 (2014).
Shumayla. et al. Genomic dissection and expression profiling revealed functional divergence in Triticum aestivum leucine rich repeat receptor like kinases (TaLRRKs). Front. Plant Sci. 7, 1374. https://doi.org/10.3389/fpls.2016.01374 (2016).
Shumayla. et al. Molecular characterization and global expression analysis of lectin receptor kinases in bread wheat (Triticum aestivum). PLoS ONE 11, e0153925. https://doi.org/10.1371/journal.pone.0153925 (2016).
Zeng, L., Deng, R., Guo, Z., Yang, S. & Deng, X. Genomewide identification and characterization of glyceraldehyde-3-phosphate dehydrogenase genes family in wheat (Triticum aestivum). BMC Genomics 17, 240. https://doi.org/10.1186/s12864-016-2527-3 (2016).
Kondrashov, F. A. Gene duplication as a mechanism of genomic adaptation to a changing environment. Proc. R. Soc. B Biol. Sci. 279, 5048–5057. https://doi.org/10.1098/rspb.2012.1108 (2012).
Magadum, S., Banerjee, U., Murugan, P., Gangapur, D. & Ravikesavan, R. Gene duplication as a major force in evolution. J. Genet. 92, 155–161 (2013).
Panchy, N., Lehti-Shiu, M. & Shiu, S.-H. Evolution of gene duplication in plants. Plant Physiol. 171, 2294–2316. https://doi.org/10.1104/pp.16.00523 (2016).
Daron, J. et al. Organization and evolution of transposable elements along the bread wheat chromosome 3B. Genome Biol. 15, 546. https://doi.org/10.1186/s13059-014-0546-4 (2014).
Cao, J., Lv, Y., Hou, Z., Li, X. & Ding, L. Expansion and evolution of thaumatin-like protein (TLP) gene family in six plants. Plant Growth Regul. 79, 299–307. https://doi.org/10.1007/s10725-0150134-y (2015).
Appels, R. et al. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361, eaar7191. https://doi.org/10.1126/science.aar7191 (2018).
Paterson, A. H., Bowers, J. E. & Chapman, B. A. Ancient polyploidization predating divergence of the cereals, and its consequences for comparative genomics. Proc. Natl. Acad. Sci. 101, 9903–9908 (2004).
Kimbrough, J. M., Salinas-Mondragon, R., Boss, W. F., Brown, C. S. & Sederoff, H. W. The fast and transient transcriptional network of gravity and mechanical stimulation in the Arabidopsis root apex. Plant Physiol. 136, 2790–2805 (2004).
Ariizumi, T. & Toriyama, K. Genetic regulation of sporopollenin synthesis and pollen exine development. Annu. Rev. Plant Biol. 62, 437–460 (2011).
Makova, K. D. & Li, W. H. Divergence in the spatial pattern of gene expression between human duplicate genes. Genome Res. 13, 1638–1645. https://doi.org/10.1101/gr.1133803 (2003).
Papatheodorou, I. et al. Expression Atlas: Gene and protein expression across multiple studies and organisms. Nucleic Acids Res. 46, D246–D251. https://doi.org/10.1093/nar/gkx1158 (2018).
Pingault, L. et al. Deep transcriptome sequencing provides new insights into the structural and functional organization of the wheat genome. Genome Biol. 16, 29. https://doi.org/10.1186/s13059-015-0601-9 (2015).
Choulet, F. et al. Structural and functional partitioning of bread wheat chromosome 3B. Science 345, 1249721–1249721. https://doi.org/10.1126/science (2014).
Prasad, P. V., Pisipati, S. R., Momčilović, I. & Ristic, Z. Independent and combined effects of high temperature and drought stress during grain filling on plant yield and chloroplast EF-Tu expression in spring wheat. J. Agron. Crop Sci. 197, 430–441. https://doi.org/10.1111/j.1439-037X.2011.00477.x (2011).
Izadi, M. H., Rabbani, J., Emam, Y., Pessarakli, M. & Tahmasebi, A. Effects of salinity stress on physiological performance of various wheat and barley cultivars. J. Plant Nutr. 37, 520–523 (2014).
Zhang, H. et al. Large scale transcriptome comparison reveals distinct gene activations in wheat responding to stripe rust and powdery mildew. BMC Genomics 15, 898. https://doi.org/10.1186/1471-216415-898 (2014).
Liu, Z. et al. Temporal transcriptome profiling reveals expression partitioning of homeologous genes contributing to heat and drought acclimation in wheat (Triticum aestivum L.). BMC Plant Biol. 15, 152. https://doi.org/10.1186/s12870-015-0511-8 (2015).
Zhang, Y. et al. Expression partitioning of homeologs and tandem duplications contribute to salt tolerance in wheat (Triticum aestivum L.). Sci. Rep. 6, 21476. https://doi.org/10.1038/srep21476 (2016).
Sanders, D., Pelloux, J., Brownlee, C. & Harper, J. F. Calcium at the crossroads of signaling. Plant Cell 14, S401–S417. https://doi.org/10.1105/tpc.002899 (2002).
Axelsen, K. B. & Palmgren, M. G. Inventory of the superfamily of P-type ion pumps in Arabidopsis. Plant Physiol. 126, 696–706. https://doi.org/10.1104/pp.126.2.696 (2001).
El-Gebali, S. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, D427–D432 (2019).
Letunic, I., Doerks, T. & Bork, P. SMART: Recent updates, new developments and status in 2015. Nucleic Acids Res. 43, D257–D260 (2015).
Marchler-Bauer, A. et al. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 45, D200–D203 (2017).
Sharma, A., Tyagi, S., Alok, A., Singh, K. & Upadhyay, S. K. Thaumatin-like protein kinases: Molecular characterization and transcriptional profiling in five cereal crops. Plant Sci. 290, 110317 (2019).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: Robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 34, W609–W612 (2006).
Gaut, B. S., Morton, B. R., McCaig, B. C. & Clegg, M. T. Substitution rate comparisons between grasses and palms: Synonymous rate differences at the nuclear gene Adh parallel rate differences at the plastid gene rbcL. Proc. Natl. Acad. Sci. 93, 10274–10279 (1996).
Edgar, R. C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797. https://doi.org/10.1093/nar/gkh340 (2004).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874. https://doi.org/10.1093/molbev/msw054 (2016).
Lescot, M. et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 30, 325–327 (2002).
Hu, B. et al. GSDS 2.0: An upgraded gene featurevisualization server. Bioinformatics 31, 1296–1297 (2015).
Gasteiger, E. et al. Protein identification and analysis tools on the expasy server. In The Proteomics Protocols Handbook, Vol. 112, 571–607, https://doi.org/10.1385/1-59259-890-0:571 (2005).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocol 10, 845 (2015).
Emanuelsson, O., Nielsen, H. & von Heijne, G. ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci. 8, 978–984 (1999).
Bailey, T. L. et al. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208. https://doi.org/10.1093/nar/gkp335 (2009).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protocol 8, 1494–1512. https://doi.org/10.1038/nprot.2013.084 (2013).
Seo, J., Gordish-Dressman, H. & Hoffman, E. P. An interactive power analysis tool for microarray hypothesis testing and generation. Bioinformatics 22, 808–814. https://doi.org/10.1093/bioinformatics/btk052 (2006).
Nazarov, P., Muller, A., Khutko, V. & Vallar, L. Co-Expression Analysis of Large Microarray Data Sets Using CoExpress Software Tool (2010).
Conesa, A. & Götz, S. Blast2GO: A comprehensive suite for functional analysis in plant genomics. Int. J. Plant Genomics https://doi.org/10.1155/2008/619832 (2008).
Bastian, M. Heymann, S. & Jacomy, M. Gephi: An open source software for exploring and manipulating networks visualization and exploration of large graphs. In Third international AAAI conference on weblogs and social media. https://doi.org/10.13140/2.1.1341.1520 (2009)
Shumayla, Tyagi, S., Sharma, A., Singh, K. & Upadhyay, S. K. Genomic dissection and transcriptional profiling of Cysteine-rich receptor-like kinases in five cereals and functional characterization of TaCRK68-A. Int. J. Biol. Macromol. 134, 316–329 (2019).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408. https://doi.org/10.1006/meth.2001.1262 (2001).
Acknowledgements
Authors are grateful to the Panjab University, Chandigarh, India for research facilities, Ensembl Plants, URGI and NCBI for data availability. SKU are grateful to the Science and Engineering Board (SERB), Government of India for Early Career Research Award (ECR/2016/001270). AK and AS are thankful to CSIR, and MT is thankful to UGC for their research fellowships.
Author information
Authors and Affiliations
Contributions
S.K.U. conceived the idea and designed the experiments. A.K., M.T., S.T. and A.S. performed the experiments. S.K.U., K.S. and A.K. analyzed the data. A.K. and S.K.U. wrote the manuscript. A.K. and S.K.U. finalized the manuscript. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kaur, A., Taneja, M., Tyagi, S. et al. Genome-wide characterization and expression analysis suggested diverse functions of the mechanosensitive channel of small conductance-like (MSL) genes in cereal crops. Sci Rep 10, 16583 (2020). https://doi.org/10.1038/s41598-020-73627-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73627-7
- Springer Nature Limited
This article is cited by
-
Origin, evolution and diversification of plant mechanosensitive channel of small conductance-like (MSL) proteins
BMC Plant Biology (2023)
-
Plants detect and respond to sounds
Planta (2023)
-
Molecular response and evolution of plant anion transport systems to abiotic stress
Plant Molecular Biology (2022)