Abstract
Retinitis pigmentosa is a retinal degenerative disease that leads to blindness through photoreceptor loss. Rhodopsin is the most frequently mutated protein in this disease. While many rhodopsin mutations have well-understood consequences that lead to cell death, the disease association of several rhodopsin mutations identified in retinitis pigmentosa patients, including F220C and F45L, has been disputed. In this study, we generated two knockin mouse lines bearing each of these mutations. We did not observe any photoreceptor degeneration in either heterozygous or homozygous animals of either line. F220C mice exhibited minor disruptions of photoreceptor outer segment dimensions without any mislocalization of outer segment proteins, whereas photoreceptors of F45L mice were normal. Suction electrode recordings from individual photoreceptors of both mutant lines showed normal flash sensitivity and photoresponse kinetics. Taken together, these data suggest that neither the F220C nor F45L mutation has pathological consequences in mice and, therefore, may not be causative of retinitis pigmentosa in humans.
Similar content being viewed by others
Introduction
Retinitis pigmentosa (RP) is a retinal degenerative disease characterized by a progressive loss of photoreceptors1. With a worldwide prevalence of about 1 in 4,000 people, there are an estimated 2 million people affected2. Approximately one third of autosomal dominant RP (adRP) cases are caused by mutations in the G protein-coupled receptor (GPCR) rhodopsin, with over 150 different causative mutations currently identified3. The majority of these mutations lead to defects in rhodopsin folding, trafficking, chromophore binding or transducin activation; yet, there are mutations that are currently unclassified with no known biochemical or cellular defects4.
One unclassified rhodopsin mutation, F220C, was described in 19935 in a study analyzing the sequences of rhodopsin genes from 88 patients/families with a positive family history of adRP. Another retinitis pigmentosa patient bearing the F220C mutant allele was subsequently identified6. However, a different study7 found that a similar F220L rhodopsin mutation did not co-segregate with adRP. Further, both F220C and F220L mutant rhodopsins have been predicted to have normal protein stability and folding in a computational study8, although more recent work found that the F220C mutant has a minor trafficking defect in mammalian cell culture9.
Another unclassified rhodopsin mutation, F45L, was first described in 199110 in a study analyzing 161 unrelated patients with adRP, where it was present in a single patient and not in any of 118 normal subjects. Seven family members of this patient were subsequently analyzed and the F45L mutation completely co-segregated with disease in all of them (including four affected and three unaffected family members). The presence of the F45L mutation was subsequently identified in a 34 year-old adRP patient (although the defects in visual function reported in this patient were relatively mild)11, a 73 year-old patient with severely affected vision12 and two additional patients whose age and pathology were not reported6. However, there are two observations that conflict with reports that the F45L allele is pathogenic. First, an adRP patient bearing the F45L mutation inherited this mutation from an asymptomatic father13. Second, eight carriers of the F45L mutation were all free of any signs of adRP14. When analyzed in cell culture, F45L mutant rhodopsin appeared to traffic normally to the plasma membrane13,15. One property that F45L rhodopsin shares with F220C rhodopsin is that they both behaved as monomers in in vitro assays in which WT rhodopsin behaved as a dimer16.
Given the conflicting evidence surrounding the molecular defects and pathogenicity of F220C and F45L rhodopsin mutations, we generated knockin mice bearing each mutation. We conducted a comprehensive analysis of both heterozygous and homozygous mice from each line, which included various microscopic techniques and single cell suction electrode recordings. Our experiments revealed no evidence of photoreceptor degeneration associated with either mutation, and rod photoreceptors of both lines had normal light sensitivities and photoresponse kinetics. These data challenge the causative role of each mutation in human adRP patients.
Results
Generation of F220C and F45L knockin mice
To study the pathogenicity of these rhodopsin mutations, we generated knockin mice using CRISPR/Cas9 methodology for both F220C (c.659 T > G mutation in exon 3 of Rho) and F45L (c.133 T > C mutation in exon 1 of Rho) rhodopsin mutations (Fig. 1). We used pronuclear injections of short guide RNAs along with both Cas9 mRNA and/or Cas9 protein and repair oligonucleotides containing either the c.659 T > G or c.133 T > C mutation. Potential founders were sequenced to verify the desired nucleotide substitution and the integrity of the remaining gene. Mice were then outcrossed with C57BL/6-J mice for at least three generations before analysis.
Generation of knockin mouse lines bearing the F220C and F45L rhodopsin mutations. The coding sequence of mouse Rho consists of five exons. (Left) The F45L mutation is caused by a c.133 T > C mutation in exon 1. A 200-nt long repair oligonucleotide (oligo) was used (purple) in conjunction with the depicted (in green) single small guide RNA (gRNA) and Cas9 mRNA and protein to generate this allele. (Right) The F220C mutation is caused by a c.659 T > G mutation in exon 3. A 102-nt long repair oligonucleotide (oligo) was used (purple) in conjunction with the depicted (in green) single small guide RNA (gRNA) and Cas9 protein to generate this allele. Sequencing of DNA from WT, heterozygous, and homozygous mice reveals the correct knockin mutation generated for both the F45L and F220C alleles.
Characterization of the F220C mouse
To address whether F220C mutant rhodopsin can cause photoreceptor degeneration, we analyzed thin retinal cross-sections from heterozygous F220C knockin mice (F220C/+ ; as in adRP patients), homozygous mice (F220C/F220C) and their WT littermates (Fig. 2). We observed no major difference in either gross photoreceptor morphology or the number of photoreceptor nuclei across these genotypes at either 1 month or 15 months of age. The only subtle difference was the occasional appearance of empty spaces between outer segments of mutant mice. These data show that the F220C rhodopsin mutation does not cause photoreceptor degeneration in mice.
F220C mice do not exhibit photoreceptor degeneration. (a) Light microscopy images of 0.5 μm thin retinal plastic sections stained with methylene blue. Depicted are WT, heterozygous (F220C/+), and homozygous (F220C/F220C) retinas at 1 month and 15 months of age. Scale bar is 10 μm. (b) The number of photoreceptor nuclei are quantified over a 100 μm length of retina at 500 μm intervals away from the optic nerve (ON) at 1 month and 15 months of age (n = 3 for each genotype at each age). There are no statistically significant differences among genotypes with two-way ANOVA at either age.
We next sought to identify whether there may be any defects in localization of rhodopsin or other outer segment proteins within mutant rods. Figure 3 shows that rhodopsin localization was normal in both hetero- and homozygous mice, as was localization of several other representative outer segment proteins – CNGβ1, ABCA4, R9AP and PRCD – the latter previously shown to rely on rhodopsin for transport to the outer segment17.
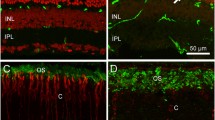
F220C mice do not exhibit mislocalization of photoreceptor outer segment proteins. Immunofluorescent staining of photoreceptor outer segment-specific proteins in WT, heterozygous (F220C/+), and homozygous (F220C/F220C) retinas at 1 month of age. Both disc-specific (rhodopsin, ABCA4, PRCD, R9AP) and outer segment plasma membrane-specific (CNGβ1) proteins are analyzed (red). Nuclei are stained with Hoechst (blue). Scale bar is 10 μm.
To analyze photoreceptor ultrastructure, we performed transmission electron microscopy (TEM) (Fig. 4), focusing primarily on photoreceptor outer segments because rhodopsin is the main protein component of these organelles18 and outer segment disc organization is often affected in other mouse adRP models19,20. We used tannic acid as a membrane-contrasting reagent, as it preferentially stains membranes of newly formed “open” discs21, thereby allowing us to reveal any defect in disc morphogenesis that could be affected by abnormal rhodopsin transport or incorporation into the outer segment membranes. However, TEM analysis showed that outer segments, including new disc membranes, appeared essentially normal in both hetero- and homozygous mice. The only trend we observed was a slightly smaller outer segment diameter of homozygous F220C rods.
Rod outer segments of F220C mice do not exhibit any gross ultrastructural defects. TEM images of the base of photoreceptor outer segments from WT, heterozygous (F220C/+), and homozygous (F220C/F220C) mice are shown at 1 month and 15 months of age. Sections are stained with tannic acid to intensely label nascent, open discs at the base of the outer segment. Scale bar is 0.5 μm.
To follow-up on the latter observation, we performed TEM on tangential sections of outer segments (Fig. 5a). This allowed us to measure the outer segment diameters of a very large number of rods (750 outer segments across 5 WT and homozygous F220C animals) to reliably identify a possible reduction in outer segment size. This quantification revealed that, while the diameter of WT outer segments was 1.40 ± 0.01 µm, the diameter of F220C/F220C outer segments was 1.34 ± 0.01 µm (p < 0.0001). Therefore, the F220C mutation causes a small reduction in outer segment diameter.
Rod outer segments of F220C mice are slightly thinner and longer. (a) TEM of a tangential section through the photoreceptor outer segment layer of WT and homozygous F220C mice. Scale bar is 1 μm. Diameters of 150 outer segments were measured in 5 different mice for a total of 750 outer segments per genotype. Diameters were grouped within 0.02 μm bins to plot the relative frequency of outer segment diameters ranging from 1.0 μm to 1.8 μm. The relative frequency plot is overlaid with Gaussian distribution curves for WT (solid line) and F220C/F220C (dashed line) datasets. Unpaired t-test reveals a statistically significant difference (p < 0.0001) between the outer segment diameters of WT (1.40 ± 0.01 µm) and F220C/F220C (1.34 ± 0.01 µm) mice. (b) Total rhodopsin content of dissected eyecups was determined by difference spectroscopy22. Unpaired t-test depicts no statistically significant difference (p = 0.7109) in rhodopsin content between WT and homozygous F220C mice at 10 months of age (n = 3 for each genotype). (c) Photoreceptor outer segment lengths were measured from five different regions of each of three different mice per genotype at 1 month of age. Unpaired t-test reveals a statistically significant difference (p = 0.011) between the outer segment lengths of WT (34.7 ± 0.6 µm) and F220C/F220C (31.1 ± 0.5 µm) mice.
To determine how this small reduction in outer segment diameter affected the total rhodopsin content of the retina, we dissected dark-adapted eyecups under dim red light and used difference spectroscopy22 to quantify rhodopsin in F220C/F220C and WT eyecups (Fig. 5b). Interestingly, the total rhodopsin content of the F220C/F220C eyecups was no different than that of wild-type (p = 0.7109). We then sought to measure outer segment lengths to determine whether there may be a compensatory effect that offsets the smaller diameter of F220C homozygous outer segments and thus keeps rhodopsin content the same (Fig. 5c). Indeed, when measured on plastic retinal cross-sections, outer segments of F220C/F220C mice did appear slightly longer than in WT mice (34.7 ± 0.6 µm vs. 31.1 ± 0.5 µm; p = 0.011).
We next performed single cell suction electrode recordings on rods of 8–13 week old age-matched wildtype, heterozygote and homozygote littermates. Rod responses to calibrated light flashes were very similar in mice of all three genotypes (Table 1; Fig. 6). Rod sensitivity to light was unaffected by the F220C mutation: the amplitude of the response to a single activated rhodopsin (SPR) was indistinguishable between lines and the flash strength needed to generate a half-maximal response (Io) was virtually identical (Table 1). The time constants of recovery for dim flash responses (𝜏rec) and bright flash responses (𝜏D) were likewise indistinguishable (Table 1; Fig. 6). Importantly, there was no significant difference in the effective collecting areas of wild-type and homozygous F220C rods (Table 1), which is likely due to the compensatory effect created by an increase in length and decrease in diameter, as reported above. Taken together, we observed no significant differences between WT, heterozygous and homozygous F220C rods.
F220C rhodopsin mutant photoreceptors do not exhibit any electrophysiological defects. (a) Suction electrode recordings of families of responses to flashes that ranged from 13–51,000 photons/μm2 by factors of 2–4. Flashes were delivered at t = 0 s. (b) Population average single photon responses calculated from WT (n = 22), F220C/+ (n = 14) and F220C/F220C (n = 11) rods. Light shading represents SEM. (c) Relationship between the time that a bright flash response remained in saturation and the natural log of the number of photoexcited rhodopsins (R*) produced by the flash. The initial slope reflects the dominant time constant of recovery for bright flashes, which was the same for mutant and WT rods. Error bars represent SEM.
Characterization of the F45L mouse
To address whether F45L mutant rhodopsin can cause photoreceptor degeneration, we analyzed thin retinal cross-sections from heterozygous F45L knockin mice (F45L/+ ; as in adRP patients), homozygous mice (F45L/F45L) and their WT littermates (Fig. 7). We observed no major difference in the number of photoreceptor nuclei across these genotypes at either 1 month or 6 months of age. Further, we did not observe any gross morphological differences between genotypes of any age. These data show that the F45L rhodopsin mutation does not cause photoreceptor degeneration in mice. We next investigated localization of rhodopsin and two other representative outer segment proteins (ABCA4 and PRCD) within mutant rods (Fig. 8) and found that their localization was normal in both hetero- and homozygous F45L mice.
F45L mice do not exhibit photoreceptor degeneration. (a) Light microscopy images of 0.5 μm thin retinal plastic sections stained with methylene blue. Depicted are WT, heterozygous (F45L/+), and homozygous (F45L/F45L) retinas at 1 month and 6 months of age. Scale bars are 10 μm. (b) The number of photoreceptor nuclei are quantified over a 100 μm length of retina at 500 μm intervals away from the optic nerve (ON) at 1 month and 6 months of age (n = 3 for each genotype at each age). There are no statistically significant differences among genotypes with two-way ANOVA at either age.
F45L mice do not exhibit mislocalization of photoreceptor outer segment proteins. Immunofluorescent staining of photoreceptor outer segment disc-specific proteins (red) in WT, heterozygous (F45L/+), and homozygous (F45L/F45L) retinas at 1 month of age. Nuclei are stained with Hoechst (blue). Scale bar is 10 μm.
To analyze photoreceptor ultrastructure, we performed transmission electron microscopy (TEM) (Fig. 9), focusing primarily on photoreceptor outer segments. This analysis showed that outer segments, including new disc membranes, appeared normal in both F45L/+ and F45L/F45L mice.
Rod outer segments of F45L mice do not exhibit any gross ultrastructural defects. TEM images of the base of photoreceptor outer segments from WT, heterozygous (F45L/+), and homozygous (F45L/F45L) mice are shown at 1 month of age. Sections are stained with tannic acid to intensely label nascent, open discs at the base of the outer segment. Scale bar is 0.5 μm.
Lastly, we performed single cell suction electrode recordings on rods of 9–12 week old age-matched wildtype, heterozygote and homozygote littermates, as described above. Rod responses to calibrated light flashes were identical in mice of all three genotypes (Table 2; Fig. 10).
F45L rhodopsin mutant photoreceptors do not exhibit any electrophysiological defects. (a) Suction electrode recordings of families of responses to flashes that ranged from 6–50,000 photons/μm2 by factors of 2–4. Flashes were delivered at t = 0 s. (b) Population average single photon responses calculated from WT (n = 22), F45L/+ (n = 14) and F45L/F45L (n = 19) rods. Light shading represents SEM. (c) Relationship between the time that a bright flash response remained in saturation and the natural log of the number of photoexcited rhodopsins (R*) produced by the flash. The initial slope reflects the dominant time constant of recovery for bright flashes, which was the same for mutant and WT rods. Error bars represent SEM.
Discussion
The data obtained in this study demonstrate that neither the F220C nor the F45L mutation in rhodopsin has pathological consequences in knockin mouse models. The F220C mutation displayed a minor abnormality in the outer segment size, while F45L photoreceptors were completely normal. Since all well-characterized disease-causing rhodopsin mutations lead to photoreceptor degeneration in mice23,24, it is unlikely that the F220C and F45L mutations are directly causal in producing adRP. The apparent discrepancy between our mouse data and the notion that these two mutations are causative of adRP can be resolved in a number of ways. As described in the introduction, neither mutation has complete co-segregation with adRP6,13,14. The fact that both mutations have been found in healthy humans has been consistently used as an argument against the notion that these mutations are causative of disease, which to a great degree motivated our study.
There are two potential explanations as to why the F45L or F220C mutant alleles were identified in adRP patients. A trivial explanation is that neither allele is pathogenic and they were coincidently present in the individuals analyzed in the original genetic studies13,25. This explanation is consistent with a relatively high frequency of these alleles in the general population25 and a modest number of subjects analyzed in the original studies. Another possibility is that F45L and F220C are modifying alleles, which interact with other non-rhodopsin mutations to cause retinal degeneration in adRP patients. Examples of gene interactions affecting photoreceptor degeneration and other ciliopathies have been reported in several studies26,27,28,29,30. Indeed, most studies of patients bearing the F45L and F220C alleles only analyzed the sequence of the rhodopsin gene5,10,11,13,14. Therefore, in conjunction with our data, it is reasonable to suggest that the F220C and F45L substitutions in rhodopsin are most likely polymorphisms that are either benign or modifying alleles.
Despite a lack of photoreceptor degeneration in either F220C or F45L mice, we did observe a small morphological defect associated with the F220C mutation. Outer segments of these mice have a slightly reduced diameter (1.34 µm vs. 1.40 µm in WT). One potential explanation for this phenomenon is a reduced ability of F220C rhodopsin to correctly fold in the biosynthetic membranes, such that the amount of rhodopsin delivered to the outer segment is slightly reduced. Based on the measured reduction of the outer segment diameter and the corresponding reduction in the disc surface area in F220C mice, we estimate that the amount of rhodopsin transported to the outer segment is approximately 92% of WT. This explanation is consistent with a recent finding that the F220C rhodopsin mutant has a minor trafficking defect in cultured mammalian cells, with only 86% of mutant protein trafficked to the plasma membrane9.
The phenotype of F220C mice can be compared to that of rhodopsin hemizygous mice whose outer segment diameter is only 1.1–1.2 µm, corresponding to about 60% of the WT outer segment rhodopsin content31,32. Rhodopsin hemizygous mice exhibit slow photoreceptor degeneration with 1–2 rows of photoreceptor nuclei reported to be lost by 3 months of age33. Together, these results suggest that while photoreceptors can tolerate a small decrease in the rhodopsin content of discs (e.g. the 8% decrease observed in F220C mutant mice), there is some level of reduced rhodopsin content that ultimately causes pathology.
The reduced diameter of F220C homozygous outer segments is offset by an increase in length by approximately 11%, which results in no detectable change in the total rhodopsin content of the retina as measured by difference spectroscopy or collecting area measurements obtained from suction electrode recordings. These data support the idea that the photoreceptor outer segment has some innate capacity to regulate its propensity to capture photons, perhaps related to other reported mechanisms of outer segment length control34 or through regulation of disc shedding35. Interestingly, such compensation is not observed in the rhodopsin hemizygous mouse whose outer segments are both thinner and shorter than in WT mice32,33.
Lastly, a recent study found that both F220C and F45L mutant rhodopsin behave as monomers in in vitro reconstitution assays, whereas WT rhodopsin behaves as a dimer. Given that GPCR dimerization has previously been shown to regulate receptor folding36,37,38,39,40,41 and trafficking42,43,44,45, it is intriguing to believe that the phenotype of the F220C mutant could be due to issues with rhodopsin dimerization. Indeed, the only study to address the effect of disrupted rhodopsin dimerization in intact photoreceptors reported that inhibitory peptides46, shown to mildly disrupt rhodopsin dimerization in vitro47, caused rhodopsin mislocalization46. Yet, another recent study in mammalian cells suggested that F220C mutant rhodopsin does not affect its dimerization9. While rhodopsin dimerization within disc membranes has been shown through atomic-force microscopy48 and cryo-electron microscopy49, there is currently no assay for analyzing rhodopsin dimerization during folding and trafficking, precluding us for identifying whether the F220C mutation in rhodopsin does indeed affect its dimerization in vivo, and, subsequently, what the role of rhodopsin dimerization in intact photoreceptors is.
Materials and Methods
Animals
All methods with regard to mice handling were carried out in accordance with relevant guidelines and regulations. F220C mutant mice were generated through the Duke Transgenic and Knockout Mouse Facility through pronuclear injections of C57BL/6 J mouse embryos with 2 ng/µl of a short guide RNA (5′ TCGTCATCTTCTTCTGCTAT GGG 3′), 8 ng/µl of a 102-nucleotide repair oligonucleotide containing the c.659 T > G mutation to knock-in the F220C allele (5′ T GTC ATC TAC ATG TTC GTG GTC CAC TTC ACC ATT CCT ATG ATC GTC ATC TGC TTC TGC TAT GGC CAG CTG GTC TTC ACA GTC AAG GAG GTA TGA GCA GGG GG 3′), 2 ng/µl Cas9 mRNA, and 2 ng/µl Cas9 protein. Potential founders were genotyped for successful knock-in through sequencing of a 491-base pair PCR product of the rhodopsin locus using primers (5′ GCAGAGCTGCGTGGTCAAGTGG 3′; 5′ CCTTCTGAGTGGTGGCTGACTCC 3′). F45L mutant mice were generated through the Duke Transgenic and Knockout Mouse Facility through pronuclear injections of C57BL/6 J mouse embryos with 200 ng/µl of a short guide RNA (5′ GAACATGTACGCTGCCAGCA TGG 3′), 0.649 ng/µl of a 200-nucleotide repair oligonucleotide containing the c.133 T > C mutation to knock-in the F220C allele (5′ CCC TTC TCC AAC GTC ACA GGC GTG GTG CGG AGC CCC TTC GAG CAG CCG CAG TAC TAC CTG GCG GAA CCA TGG CAG TTC TCT ATG CTG GCA GCG TAC ATG CTC CTG CTC ATC GTG CTG GGC TTC CCC ATC AAC TTC CTC ACG CTC TAC GTC ACC GTA CAG CAC AAG AAG CTG CGC ACA CCC CTC AAC TAC ATC CTG CTC AA 3′), and 8 µM Cas9 protein. Potential founders were genotyped for successful knock-in through sequencing of a 596-base pair PCR product of the rhodopsin locus using primers (5′ GCCTCCACCCGATGTCACC 3′; 5′ CCCTCGAGATTACAGCCTG 3′). Mice bearing successful knock-ins were outcrossed to C57BL/6 J mice for several generations. All mice were genotyped to ensure that they did not contain either the rd850 or rd151 mutations commonly found in inbred mouse strains. All mice were housed under a 12/12 hour diurnal light cycle. Littermate WT, heterozygous, and homozygous mutant mice were used for all experiments.
Immunofluorescence
Anesthetized mice were transcardially perfused with a fixative solution containing 4% paraformaldehyde in 80 mM PIPES (pH 6.8), 5 mM EGTA, and 2 mM MgCl2. Eyes were enucleated and post-fixed in the same solution for two hours at room temperature. After fixation, dissected eyecups were embedded in 2.5% low-melt agarose (Precisionary) and cut by a Vibratome (VT1200S; Leica) into 100 µm thick slices as described previously52. Agarose sections were blocked in PBS containing 5% donkey serum and 0.5% Triton X-100 for 1 h at room temperature before staining with primary antibody in blocking buffer overnight at 4 °C. After primary antibody staining, sections were washed three times in PBS and incubated with secondary antibody in blocking buffer overnight at 4 °C. Finally, sections were washed three times in PBS and nuclei were stained with 10 µg/ml Hoechst (H3569; Thermo Fisher Scientific) for 30 min at room temperature. Finally, sections were washed three times in PBS, and mounted onto slides with Immu-Mount (Thermo) and coverslipped. Images were taken with a confocal microscope (Eclipse 90i and A1 confocal scanner; Nikon) with a 60× objective (1.4 NA Plan Apochromat VC; Nikon) using Nikon NIS-Elements software. Image analysis and processing was performed with ImageJ.
Antibodies
For immunofluorescence imaging, commercial primary antibodies included: polyclonal goat anti-human ABCA4, 1:2000 (Everest Biotech) and monoclonal mouse anti-bovine RHO (1D4), 1:2000 (Abcam)53. The polyclonal rabbit anti-mouse CNGβ1, 1:500, was a generous gift from Dr. Steven Pittler. The polyclonal rabbit anti-mouse R9AP, 1:2500, was a generous gift from Dr. Stefan Heller54. The polyclonal rabbit anti-mouse PRCD, 1:2000, was previously generated by the Arshavsky lab55. The commercial secondary antibodies included donkey anti-goat, donkey anti-mouse, and donkey anti-rabbit conjugated to Alexa Fluor 488 or 568, 1:1000 (Invitrogen).
Histological techniques and transmission electron microscopy
Fixation and processing of mouse eyes for light microscopy of plastic sections was performed as described previously21. Anesthetized mice were transcardially perfused with 2% paraformaldehyde, 2% glutaraldehyde, and 0.05% calcium chloride in 50 mM MOPS (pH 7.4) resulting in exsanguination. Enucleated eyes were fixed for an additional 2 h in the same fixation solution at room temperature. To obtain semi-thin plastic retinal sections, eyecups were cut in half through the optic nerve, dehydrated with ethanol, and embedded in EMbed 812 (Electron Microscopy Sciences). Embedded retinal cross sections were cut through the optic nerve in 500 nm slices and stained with methylene blue for light microscopy as previously described56. Images were taken with a confocal microscope (Eclipse 90i and A1 confocal scanner; Nikon) with a 60× objective (1.4 NA Plan Apochromat VC; Nikon) using Nikon NIS-Elements software. Image analysis and processing was performed with ImageJ.
For TEM involving tannic acid staining, eyes were fixed as described above. Eyecups were dissected from fixed eyes, embedded in 2.5% low-melt agarose (Precisionary), and cut into 200 µm thick slices on a Vibratome (VT1200S; Leica)21. Agarose sections were stained with 1% tannic acid (Electron Microscopy Sciences) and 1% uranyl acetate (Electron Microscopy Sciences), gradually dehydrated with ethanol, and infiltrated and embedded in Spurr’s resin (Electron Microscopy Sciences). For TEM involving tangential sections of outer segments, eyes were fixed as described above. Dissected eyecups were flatmounted, dehydrated with ethanol, and embedded in EMbed 812 (Electron Microscopy Sciences). For all TEM, 70 nm sections were cut, placed on copper grids, and counterstained with 2% uranyl acetate and 3.5% lead citrate (19314; Ted Pella). The samples were imaged on a JEM-1400 electron microscope (JEOL) at 60 kV with a digital camera (Orius; Gatan).
Suction electrode recordings
Animals used for electrophysiological analysis were cared for and handled in accordance with a protocol approved by the Institutional Animal Care and Use Committees of UC Davis. Suction electrode recordings from the outer segments of intact mouse rods were performed as previously described57. Briefly, mice 2–3 months of age were dark-adapted overnight, euthanized, and their retinas were dissected and stored on ice in L-15 medium supplemented with 10 mM glucose. Recordings were performed in oxygenated, bicarbonate buffered Locke’s solution supplemented with 10 mM glucose at 35–37 °C. A suction pipette containing HEPES-buffered Locke’s solution (pH 7.4) recorded the electrical responses to brief (10 ms, 500 nm) flashes of calibrated strength, which were amplified (Axopatch 200B; Molecular Devices), filtered at 30 Hz with an eight-pole Bessel (Frequency Devices), and digitized at 200 Hz using custom-written acquisition procedures in IgorPro (Wavemetrics). Responses to a large number (>30) of dim flashes were averaged and used to determine the mean time to peak and integration time (time integral of the response divided by the peak amplitude). Variance-to-mean analysis of the dim flash responses was used to calculate the average single-photon response (SPR) and effective collecting areas for each cell, and saturating flash responses were used to calculate the dominant time constant of recovery (τD), as previously described57,58,59.
Rhodopsin difference spectra
Mice were dark-adapted overnight before removal of cornea and lens during dim red light. Eyecups were lysed in 600 µl of 2% octyl glucoside with protease inhibitor (cOmplete; Sigma Aldrich) and sonicated 3 times each of 5 seconds. Lysates were centrifuged at 10,000 × g for 10 minutes and the rhodopsin concentration of the supernatant was measured by difference spectroscopy22. Hydroxylamine (25 mM; pH 7.5) was added to 100 μl of lysate and the initial spectra recorded from 300 to 700 nm using a DU800 spectrophotometer (Beckman). The cuvette sample was then bleached with a 100-W halogen lamp for 60 s before taking the second spectrum reading. The difference in the absorbance at 500 nm was used to calculate rhodopsin concentration, using the extinction coefficient of 40,500 M−1 ·cm−1.
Quantification and statistical analysis
To assess photoreceptor degeneration, we counted photoreceptor nuclei in 100 μm boxes at 500 μm intervals from the optic nerve spanning 2000 μm in each direction for three mice of each genotype at each age, as previously described60. Two-way ANOVA (with respect to genotype and location) was performed to assess any statistical differences between genotypes. To measure photoreceptor diameter, the diameter of 150 outer segments was measured in each of 5 mice for each genotype for a total of 750 outer segments measured per genotype. A normal Gaussian distribution was fit to each genotype. Unpaired, two-tailed t-test (assuming equal variances) t-test was performed to assess any statistical differences between genotypes. To determine the total rhodopsin content of eyecups, difference spectra were taken from each of three dissected eyecups per genotype. Unpaired, two-tailed t-test (assuming equal variances) was performed to assess any statistical differences between genotypes. To quantify outer segment length, the lengths of outer segments in five regions across the retina were measured of each of three different retinal sections per genotype. Unpaired, two-tailed t-test (assuming equal variances) was performed to assess any statistical differences between genotypes. For all experiments, significance is labeled as *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001. Non-significant values are unlabeled. Data is represented as mean ± standard error of the mean.
Data availability
All data generated or analyzed during this study are included in this published article and available from authors on reasonable request.
References
Farrar, G. J., Kenna, P. F. & Humphries, P. On the genetics of retinitis pigmentosa and on mutation-independent approaches to therapeutic intervention. Embo j 21, 857–864, https://doi.org/10.1093/emboj/21.5.857 (2002).
Hartong, D. T., Berson, E. L. & Dryja, T. P. Retinitis pigmentosa. Lancet 368, 1795–1809, https://doi.org/10.1016/s0140-6736(06)69740-7 (2006).
Stenson, P. D. et al. The Human Gene Mutation Database: building a comprehensive mutation repository for clinical and molecular genetics, diagnostic testing and personalized genomic medicine. Hum Genet 133, 1–9, https://doi.org/10.1007/s00439-013-1358-4 (2014).
Mendes, H. F., van der Spuy, J., Chapple, J. P. & Cheetham, M. E. Mechanisms of cell death in rhodopsin retinitis pigmentosa: implications for therapy. Trends Mol Med 11, 177–185, https://doi.org/10.1016/j.molmed.2005.02.007 (2005).
Bunge, S. et al. Molecular analysis and genetic mapping of the rhodopsin gene in families with autosomal dominant retinitis pigmentosa. Genomics 17, 230–233 (1993).
Stone, E. M. Finding and interpreting genetic variations that are important to ophthalmologists. Trans Am Ophthalmol Soc 101, 437–484 (2003).
Dryja, T. P., McEvoy, J. A., McGee, T. L. & Berson, E. L. Novel rhodopsin mutations Gly114Val and Gln184Pro in dominant retinitis pigmentosa. Invest Ophthalmol Vis Sci 41, 3124–3127 (2000).
Rakoczy, E. P., Kiel, C., McKeone, R., Stricher, F. & Serrano, L. Analysis of disease-linked rhodopsin mutations based on structure, function, and protein stability calculations. J Mol Biol 405, 584–606, https://doi.org/10.1016/j.jmb.2010.11.003 (2011).
Mallory, D. P. et al. The Retinitis Pigmentosa-Linked Mutations in Transmembrane Helix 5 of Rhodopsin Disrupt Cellular Trafficking Regardless of Oligomerization State. Biochemistry 57, 5188–5201, https://doi.org/10.1021/acs.biochem.8b00403 (2018).
Sung, C. H. et al. Rhodopsin mutations in autosomal dominant retinitis pigmentosa. Proc Natl Acad Sci USA 88, 6481–6485, https://doi.org/10.1073/pnas.88.15.6481 (1991).
Berson, E. L., Rosner, B., Weigel-DiFranco, C., Dryja, T. P. & Sandberg, M. A. Disease progression in patients with dominant retinitis pigmentosa and rhodopsin mutations. Invest Ophthalmol Vis Sci 43, 3027–3036 (2002).
Matias-Florentino, M., Ayala-Ramirez, R., Graue-Wiechers, F. & Zenteno, J. C. Molecular screening of rhodopsin and peripherin/RDS genes in Mexican families with autosomal dominant retinitis pigmentosa. Curr Eye Res 34, 1050–1056, https://doi.org/10.3109/02713680903283169 (2009).
Davies, W. I. et al. Next-generation sequencing in health-care delivery: lessons from the functional analysis of rhodopsin. Genet Med 14, 891–899, https://doi.org/10.1038/gim.2012.73 (2012).
Vincent, A. L. et al. Rhodopsin F45L Allele Does Not Cause Autosomal Dominant Retinitis Pigmentosa in a Large Caucasian Family. Transl Vis Sci Technol 2, 4, https://doi.org/10.1167/tvst.2.2.4 (2013).
Behnen, P. et al. A Small Chaperone Improves Folding and Routing of Rhodopsin Mutants Linked to Inherited Blindness. iScience 4, 1–19, https://doi.org/10.1016/j.isci.2018.05.001 (2018).
Ploier, B. et al. Dimerization deficiency of enigmatic retinitis pigmentosa-linked rhodopsin mutants. Nat Commun 7, 12832, https://doi.org/10.1038/ncomms12832 (2016).
Spencer, W. J. et al. Progressive Rod-Cone Degeneration (PRCD) Protein Requires N-Terminal S-Acylation and Rhodopsin Binding for Photoreceptor Outer Segment Localization and Maintaining Intracellular Stability. Biochemistry 55, 5028–5037, https://doi.org/10.1021/acs.biochem.6b00489 (2016).
Rakshit, T. & Park, P. S. Impact of reduced rhodopsin expression on the structure of rod outer segment disc membranes. Biochemistry 54, 2885–2894, https://doi.org/10.1021/acs.biochem.5b00003 (2015).
McNally, N. et al. Murine model of autosomal dominant retinitis pigmentosa generated by targeted deletion at codon 307 of the rds-peripherin gene. Hum Mol Genet 11, 1005–1016, https://doi.org/10.1093/hmg/11.9.1005 (2002).
Sakami, S., Kolesnikov, A. V., Kefalov, V. J. & Palczewski, K. P23H opsin knock-in mice reveal a novel step in retinal rod disc morphogenesis. Hum Mol Genet 23, 1723–1741, https://doi.org/10.1093/hmg/ddt561 (2014).
Ding, J. D., Salinas, R. Y. & Arshavsky, V. Y. Discs of mammalian rod photoreceptors form through the membrane evagination mechanism. J Cell Biol 211, 495–502, https://doi.org/10.1083/jcb.201508093 (2015).
Bownds, D., Gordon-Walker, A., Gaide-Huguenin, A. C. & Robinson, W. Characterization and analysis of frog photoreceptor membranes. J Gen Physiol 58, 225–237, https://doi.org/10.1085/jgp.58.3.225 (1971).
Nour, M. & Naash, M. I. Mouse models of human retinal disease caused by expression of mutant rhodopsin. A valuable tool for the assessment of novel gene therapies. Adv Exp Med Biol 533, 173–179, https://doi.org/10.1007/978-1-4615-0067-4_22 (2003).
Rossmiller, B., Mao, H. & Lewin, A. S. Gene therapy in animal models of autosomal dominant retinitis pigmentosa. Mol Vis 18, 2479–2496 (2012).
Athanasiou, D. et al. The molecular and cellular basis of rhodopsin retinitis pigmentosa reveals potential strategies for therapy. Prog Retin Eye Res 62, 1–23, https://doi.org/10.1016/j.preteyeres.2017.10.002 (2018).
Khanna, H. et al. A common allele in RPGRIP1L is a modifier of retinal degeneration in ciliopathies. Nat Genet 41, 739–745, https://doi.org/10.1038/ng.366 (2009).
Badano, J. L. et al. Dissection of epistasis in oligogenic Bardet-Biedl syndrome. Nature 439, 326–330, https://doi.org/10.1038/nature04370 (2006).
Tory, K. et al. High NPHP1 and NPHP6 mutation rate in patients with Joubert syndrome and nephronophthisis: potential epistatic effect of NPHP6 and AHI1 mutations in patients with NPHP1 mutations. J Am Soc Nephrol 18, 1566–1575, https://doi.org/10.1681/asn.2006101164 (2007).
Leitch, C. C. et al. Hypomorphic mutations in syndromic encephalocele genes are associated with Bardet-Biedl syndrome. Nat Genet 40, 443–448, https://doi.org/10.1038/ng.97 (2008).
Venturini, G., Rose, A. M., Shah, A. Z., Bhattacharya, S. S. & Rivolta, C. CNOT3 is a modifier of PRPF31 mutations in retinitis pigmentosa with incomplete penetrance. PLoS Genet 8, e1003040, https://doi.org/10.1371/journal.pgen.1003040 (2012).
Makino, C. L. et al. Rhodopsin expression level affects rod outer segment morphology and photoresponse kinetics. PLoS One 7, e37832, https://doi.org/10.1371/journal.pone.0037832 (2012).
Liang, Y. et al. Rhodopsin signaling and organization in heterozygote rhodopsin knockout mice. J Biol Chem 279, 48189–48196, https://doi.org/10.1074/jbc.M408362200 (2004).
Lem, J. et al. Morphological, physiological, and biochemical changes in rhodopsin knockout mice. Proc Natl Acad Sci USA 96, 736–741, https://doi.org/10.1073/pnas.96.2.736 (1999).
Penn, J. S. & Williams, T. P. Photostasis: regulation of daily photon-catch by rat retinas in response to various cyclic illuminances. Exp Eye Res 43, 915–928, https://doi.org/10.1016/0014-4835(86)90070-9 (1986).
Lewis, T. R., Kundinger, S. R., Link, B. A., Insinna, C. & Besharse, J. C. Kif17 phosphorylation regulates photoreceptor outer segment turnover. BMC Cell Biol 19, 25, https://doi.org/10.1186/s12860-018-0177-9 (2018).
White, J. H. et al. Heterodimerization is required for the formation of a functional GABA(B) receptor. Nature 396, 679–682, https://doi.org/10.1038/25354 (1998).
Terrillon, S. et al. Oxytocin and vasopressin V1a and V2 receptors form constitutive homo- and heterodimers during biosynthesis. Mol Endocrinol 17, 677–691, https://doi.org/10.1210/me.2002-0222 (2003).
Lee, S. P. et al. Inhibition of cell surface expression by mutant receptors demonstrates that D2 dopamine receptors exist as oligomers in the cell. Mol Pharmacol 58, 120–128, https://doi.org/10.1124/mol.58.1.120 (2000).
Zhu, X. & Wess, J. Truncated V2 vasopressin receptors as negative regulators of wild-type V2 receptor function. Biochemistry 37, 15773–15784, https://doi.org/10.1021/bi981162z (1998).
Salahpour, A. et al. Homodimerization of the beta2-adrenergic receptor as a prerequisite for cell surface targeting. J Biol Chem 279, 33390–33397, https://doi.org/10.1074/jbc.M403363200 (2004).
Wilson, S., Wilkinson, G. & Milligan, G. The CXCR1 and CXCR2 receptors form constitutive homo- and heterodimers selectively and with equal apparent affinities. J Biol Chem 280, 28663–28674, https://doi.org/10.1074/jbc.M413475200 (2005).
Prinster, S. C., Holmqvist, T. G. & Hall, R. A. Alpha2C-adrenergic receptors exhibit enhanced surface expression and signaling upon association with beta2-adrenergic receptors. J Pharmacol Exp Ther 318, 974–981, https://doi.org/10.1124/jpet.106.106526 (2006).
Uberti, M. A., Hague, C., Oller, H., Minneman, K. P. & Hall, R. A. Heterodimerization with beta2-adrenergic receptors promotes surface expression and functional activity of alpha1D-adrenergic receptors. J Pharmacol Exp Ther 313, 16–23, https://doi.org/10.1124/jpet.104.079541 (2005).
Chalothorn, D. et al. Differences in the cellular localization and agonist-mediated internalization properties of the alpha(1)-adrenoceptor subtypes. Mol Pharmacol 61, 1008–1016, https://doi.org/10.1124/mol.61.5.1008 (2002).
Uberti, M. A., Hall, R. A. & Minneman, K. P. Subtype-specific dimerization of alpha 1-adrenoceptors: effects on receptor expression and pharmacological properties. Mol Pharmacol 64, 1379–1390, https://doi.org/10.1124/mol.64.6.1379 (2003).
Zhang, T. et al. Dimerization of visual pigments in vivo. Proc Natl Acad Sci USA 113, 9093–9098, https://doi.org/10.1073/pnas.1609018113 (2016).
Jastrzebska, B. et al. Disruption of Rhodopsin Dimerization with Synthetic Peptides Targeting an Interaction Interface. J Biol Chem 290, 25728–25744, https://doi.org/10.1074/jbc.M115.662684 (2015).
Fotiadis, D. et al. Atomic-force microscopy: Rhodopsin dimers in native disc membranes. Nature 421, 127–128, https://doi.org/10.1038/421127a (2003).
Gunkel, M. et al. Higher-order architecture of rhodopsin in intact photoreceptors and its implication for phototransduction kinetics. Structure 23, 628–638, https://doi.org/10.1016/j.str.2015.01.015 (2015).
Mattapallil, M. J. et al. The Rd8 mutation of the Crb1 gene is present in vendor lines of C57BL/6N mice and embryonic stem cells, and confounds ocular induced mutant phenotypes. Invest Ophthalmol Vis Sci 53, 2921–2927, https://doi.org/10.1167/iovs.12-9662 (2012).
Pittler, S. J., Keeler, C. E., Sidman, R. L. & Baehr, W. PCR analysis of DNA from 70-year-old sections of rodless retina demonstrates identity with the mouse rd defect. Proc Natl Acad Sci USA 90, 9616–9619, https://doi.org/10.1073/pnas.90.20.9616 (1993).
Lobanova, E. S. et al. Mechanistic basis for the failure of cone transducin to translocate: why cones are never blinded by light. J Neurosci 30, 6815–6824, https://doi.org/10.1523/jneurosci.0613-10.2010 (2010).
Molday, R. S. & MacKenzie, D. Monoclonal antibodies to rhodopsin: characterization, cross-reactivity, and application as structural probes. Biochemistry 22, 653–660, https://doi.org/10.1021/bi00272a020 (1983).
Keresztes, G., Mutai, H., Hibino, H., Hudspeth, A. J. & Heller, S. Expression patterns of the RGS9-1 anchoring protein R9AP in the chicken and mouse suggest multiple roles in the nervous system. Mol Cell Neurosci 24, 687–695 (2003).
Skiba, N. P. et al. Proteomic identification of unique photoreceptor disc components reveals the presence of PRCD, a protein linked to retinal degeneration. J Proteome Res 12, 3010–3018, https://doi.org/10.1021/pr4003678 (2013).
Lobanova, E. S. et al. Transducin γ-subunit sets expression levels of α - and β-subunits and is crucial for rod viability. J. Neurosci. 28, 3510–3520 (2008).
Gross, O. P. & Burns, M. E. Control of rhodopsin’s active lifetime by arrestin-1 expression in mammalian rods. J Neurosci 30, 3450–3457, https://doi.org/10.1523/jneurosci.5391-09.2010 (2010).
Krispel, C. M. et al. RGS expression rate-limits recovery of rod photoresponses. Neuron 51, 409–416, https://doi.org/10.1016/j.neuron.2006.07.010 (2006).
Pepperberg, D. R. et al. Light-dependent delay in the falling phase of the retinal rod photoresponse. Vis Neurosci 8, 9–18 (1992).
Lobanova, E. S. et al. Increased proteasomal activity supports photoreceptor survival in inherited retinal degeneration. Nat Commun 9, 1738, https://doi.org/10.1038/s41467-018-04117-8 (2018).
Acknowledgements
The authors are grateful to Drs. Anant Menon and George Khelashvili (both at Cornell University) for ongoing discussions and inspiration to proceed with this project. This work was supported by the National Institutes of Health grants EY027969 (V.Y.A.; M.E.B.), EY005722 (V.Y.A.), EY029929 (T.R.L.), and an Unrestricted Award from Research to Prevent Blindness Inc. (Duke University).
Author information
Authors and Affiliations
Contributions
T.R.L., C.R.S., M.A.C., and Y.H. designed/performed experiments and analyzed data. V.Y.A. and M.E.B. oversaw the experimental design and data analysis. T.R.L. wrote the first draft of the manuscript. All authors edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lewis, T.R., Shores, C.R., Cady, M.A. et al. The F220C and F45L rhodopsin mutations identified in retinitis pigmentosa patients do not cause pathology in mice. Sci Rep 10, 7538 (2020). https://doi.org/10.1038/s41598-020-64437-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-64437-y
- Springer Nature Limited