Abstract
Ancestry informative single nucleotide polymorphisms (SNPs) can identify biogeographic ancestry (BGA); however, population substructure and relatively recent admixture can make differentiation difficult in heterogeneous Hispanic populations. Utilizing unrelated individuals from the Genomic Origins and Admixture in Latinos dataset (GOAL, n = 160), we designed an 80 SNP panel (Setser80) that accurately depicts BGA through STRUCTURE and PCA. We compared our Setser80 to the Seldin and Kidd panels via resampling simulations, which models data based on allele frequencies. We incorporated Admixed American 1000 Genomes populations (1000 G, n = 347), into a combined populations dataset to determine robustness. Using multinomial logistic regression (MLR), we compared the 3 panels on the combined dataset and found overall MLR classification accuracies: 93.2% Setser80, 87.9% Seldin panel, 71.4% Kidd panel. Naïve Bayesian classification had similar results on the combined dataset: 91.5% Setser80, 84.7% Seldin panel, 71.1% Kidd panel. Although Peru and Mexico were absent from panel design, we achieved high classification accuracy on the combined populations for Peru (MLR = 100%, naïve Bayes = 98%), and Mexico (MLR = 90%, naïve Bayes = 83.4%) as evidence of the portability of the Setser80. Our results indicate the Setser80 SNP panel can reliably classify BGA for individuals of presumed Hispanic origin.
Similar content being viewed by others

Introduction
It is important to study the genetics of Hispanic populations to avoid oversimplifying this heterogeneous ethnicity into a single conglomerate. The identification of specific biogeographic ancestries (BGA) has implications both in clinical1 and forensic2 genetics. Clinically, a more complete description of the various Hispanic BGAs may result in identification of rare variants that may not have been previously described when grouping all Hispanic populations together3, or for controlling for population substructure in clinical trials4,5. Hispanic individuals are known to have differential predispositions for various diseases and ignoring this diversity restricts the generalizability of the results6. In forensics, BGA data could be used to investigate the origin of unidentified human remains (UHR)7, or locate the rightful parents/guardians of a child who is unable to identify where she/he is from8. It is the heterogeneous nature of Hispanic populations that has previously deterred full characterization of their substructure. However, in the past decade, there has been a movement to explore global human diversity and a variety of genetic panels have been designed for this purpose.
Early ancestry informative marker (AIMs) panels are “continental” in nature, focused on admixture mapping to determine from which of the six inhabited continents an individual has ancestry; these include: Seldin1289, Galanter et al.’s 44610, Kidd5511, EUROFORGEN12, Genetic Atlas13, Genographic Project14, Cuba by Marcheco-Teruel et al.15, and Cuba by Fortes-Lima et al.16. Although these studies assessed continental ancestry proportions (e.g. Seldin128)9, highly differentiated populations may be detected within continental panels, even identifying admixed populations such as Gujarati Indians in Houston, TX and Mexican ancestry from Los Angeles, CA17. The ability to separate small admixed populations among larger more homogenous populations supports the notion that continental SNPs with high genetic differentiation may still be informative on a more specific country level. The simultaneous description of highly divergent populations alongside less specific populations using the same SNP panel is central to the goals of our study. However, dual level analysis of admixed populations within continental panels is rare, as it tends to decrease the panel’s performance2,17.
Other panels target more specific, country BGA beginning in European populations before extending to other regions of the world (e.g. Denmark within Northern Europe). Although the Genographic Project14 assessed populations worldwide (though sparsely in the Americas), their in-house geographic population structure (GPS) algorithm is capable of identifying country of origin. EASTASAIMS was one of the first non-European AIMs panels focusing on 22 East Asian populations using 1,500 AIMs and was able to separate the five largest populations in the region18. Zeng et al.19 created a panel of 23 AIMs using FST focusing on the four major US populations from HapMap 320: African ancestry from Southwest United States (ASW), Utah residents with Northern and Western European ancestry (CEU), Chinese from Metropolitan Denver, Colorado (CHD), and Mexican ancestry from Los Angeles, CA (MEX). And more recently, Huerta-Chagoya et al.21 reported 32 AIMs within Mexican mestizo populations, to estimate admixture proportions in various regions of Mexico.
Highly accurate BGA predictions are possible with up to 83% accuracy, but at the expense of panel size, requiring 40,000–130,000 SNPs as used in the Genographic Project14. Additionally, of the 12,476 reference samples used to select 40,000+ SNPs in their panel, only 9% were from American/Amerindian populations22, which limits the utility of their panel for resolving Hispanic ancestry. The size of this panel14, the proprietary nature of the SNPs on their Genochip22, and poor representation of the Western hemisphere, has prompted us to create a small, efficient, and publicly available SNP panel concentrated on BGA of Central America, South America, and the Caribbean.
Within one country, both Great Britain23 and Cuba15,16 have attempted to describe the diversity of their populations. The British Isles were ideal candidates for national differentiation due to their relative homogeneity and the presence of a geographic barrier which has historically restricted continuous gene flow with continental Europe and other island populations. In contrast, studies by Marcheco-Teruel et al.15, and Fortes-Lima et al.16 superficially appear to differentiate between the fifteen Cuban provinces on a national level, but their real focus was measuring admixture proportions using a subset of Galanter et al.’s 446 SNPs10, making their studies better described as continental and highlighting the need for a within country panel. Overall, at least 21 AIMs panels have been reported; however, of the 1,397 SNPs identified by Soundararajan et al.24, only 46 Consensus SNPs were in common to three or more SNP panels.
At present, there is no AIMs panel that focuses on the determination of BGA between countries in the Americas. Despite the overlap of our region of interest with the Galanter et al.’s 446 Latin American AIMs10, our purpose was to classify BGA, not to estimate the ancestral proportions contributed from 3–4 continental populations. The majority of AIMs panels and genetic ancestry studies have a heavy concentration of populations in Europe and Asia and far fewer in Central America, South America, and the Caribbean13,14,18. Our country panel addresses this gap in knowledge and focuses on these same populations.
Results
Setser80 SNP panel evaluation
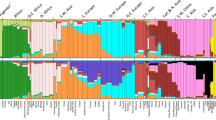
We evaluated the ability of a newly developed Hispanic AIMs panel (the Setser80) versus the Seldin1289 and Kidd5511 to separate heterogeneous Hispanic populations in the GOAL dataset (from Moreno-Estrada et al.25) using STRUCTURE26 and principal components analysis (PCA). With the STRUCTURE26 results, we applied the Evanno method27 which optimized the computer-determined (K) populations; the highest likelihood for the Setser80 was at K = 4 while Seldin1289 and Kidd5511 were optimized at K = 3 (Fig. 1a,1c,1e). The genetic proportions from STRUCTURE26 indicated that the Setser80 clearly separates HUR (Cluster 1 = 0.8290), DOM (Cluster 2 = 0.6976), and COL (Cluster 3 = 0.6562) (Table 1); but CUB (Cluster 2 = 0.2892, Cluster 4 = 0.6125) and PUR (Cluster 2 = 0.2048, Cluster 4 = 0.4145) remain indistinguishable (Fig. 1c). Using the genetic proportions from STRUCTURE26 for the Seldin1289 and Kidd5511 panels, HUR and COL separated predominately into Cluster 1 (HUR: Seldin128 = 0.7274, Kidd55 = 0.7258)(COL: Seldin128 = 0.5370, Kidd55 = 0.5311) (Table 1), but the remaining populations did not separate into distinct clusters.
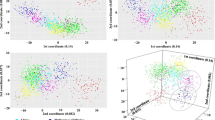
Comparison to other panels. Each plot represents 160 unrelated GOAL individuals and their respective populations. Figures a, c, and e are STRUCTURE plots where each vertical line represents one person. Figures b, d, and f are PCA plots created through EIGENSOFT where the first three principal components are plotted. Figures a and b use the Kidd55 SNP panel (K = 3), c and d use the Setser80 (K = 4), and e and f use the Seldin128 (K = 3). Abbreviations used: HUR = Honduras, DOM = Dominican Republic, COL = Colombia, CUB = Cuba, PUR = Puerto Rico, PCA = principal components analysis.
We performed a principal components analysis (PCA) for the AIMs panels in the GOAL population (Fig. 1b,1d,1f). In the PCA of the Setser80, HUR clearly separated across PC1 and PC2, DOM separated from HUR across PC2, and COL separated from HUR across PC1 and from DOM across PC1 and PC2, which occupies three separate quadrants of the PCA (Fig. 1d). Seldin1289 PCA showed HUR and COL separated together but apart from the other populations across PC1, and CUB and DOM separated together along PC2 (Fig. 1f). The Kidd5511 performed poorly in PCA (Fig. 1b), not forming recognizable clusters, consistent with the genetic proportions generated in STRUCTURE26 (Fig. 1a) (Table 1). The Setser80 was able to differentiate HUR, DOM, and COL by the two different algorithms underlying STRUCTURE26 and PCA.
Classification of unknowns
Based on the GOAL25 and 1000 Genomes Project28 (1000 G) allele frequencies, we modeled populations to determine classification accuracy using the Snipper 2.5 app suite29. Snipper uses naïve Bayesian likelihood ratios and multinomial logistic regression (MLR) for prediction of unknowns via −log(likelihood)29. Despite the different algorithms, both analyses had similar results.
As expected, the Setser80 had the highest overall accuracy across the three panels in the simulated GOAL dataset (98.4%) by naïve Bayesian classification implemented via leave-one-out cross-validation. Additionally, the Setser80 achieved 90% accuracy in the 1000 G dataset and 91.5% in the 7 Populations Combined dataset, both of which include populations not involved in our SNP ascertainment (Table 2). In the latter, the Setser80 panel (98%) and the Seldin panel (98.8%) achieved approximately equal accuracy in PEL, a population on which the Setser80 was not trained. In the 1000 G simulations, the Seldin panel was more accurate overall (92.4%) in comparison to the Setser80 (90%).
Naïve Bayes analysis of the actual 1000 G genotypes revealed the Setser80 had the highest specificity in CLM (98.4%), the highest sensitivity in MXL (84.4%), and similar specificity in PUR (85.2%) and PEL (97.7%) in comparison to the Seldin (86.8%, 95.4%) and Kidd (85.2%, 94.7%) panels (Table 3). In all three SNP panels, the micro-simulations underestimated the positive predictive value of CLM. The positive predictive value of Setser80 for PUR (simulated = 69.8%, real = 70.2%) and PEL (simulated = 91.8%, real = 89.8%) was concordant between the simulated and real data where it was either under or overestimated by the Seldin and Kidd panels. Both the Setser80 (simulated = 59.3%, real = 36.7%) and the Seldin (simulated = 80.1%, real 54.1%) panels overestimated positive predictive value in MXL while the Kidd panel values were concordant between the simulations (47.3%) and real genotypes (45.7%).
Utilizing the MLR algorithm, Setser80 had the highest accuracy in GOAL and 7 Populations Combined (99% and 93.2%, respectively); the Setser80 and Seldin panel had equal accuracy in 1000 G (93.8%); and the Kidd panel had 80.5% in GOAL, 71.4% in 7 Populations Combined, and 82.2% overall in 1000 G (Table 4). As expected, HUR achieved >95% accuracy in the Setser80 and the Seldin panel across all datasets. Surprisingly, PEL also achieved >95% and MXL > 90% accuracies using the Setser80, although the Setser80 had not been trained on these populations.
Despite performing best overall, the Setser80 did misclassify COL 22.5% of the time in the 7 Populations Combined dataset (Supplemental Table S3). When it misclassified COL, the individual was classified as MXL 77.8% and PUR 22.2% of the time. Conversely, even though MXL classified correctly 90% of the time, when individuals were misclassified they were misclassified as COL 100% of the time. In comparison, the Seldin panel misclassified COL 17.5% of the time spread across four countries, primarily into PUR (10%). The Kidd panel exhibited a similar trend where COL misclassified into five countries: PUR (15%), MXL (10%), HUR (7.5%), CUB (7.5%), and DOM (2.5%) in addition to one individual which could not be classified. When MXL was misclassified using the Kidd panel, it misclassified into PEL (7.5%), HUR (5%), and COL (5%). Additionally, the Kidd panel had high misclassification of HUR into MXL (20%), COL (15%), and PUR (7.5%).
Discussion
We report a panel of 80 AIMs for Hispanic BGA classification using Weir & Cockerham’s estimator30 of Wright’s FST31. Honduras (HUR) and DOM emerged first in STRUCTURE26 and PCA, followed by COL at K = 4, which separated from CUB & PUR, indicating three distinct populations (Table 1). Based on the allele frequencies, we created a series of micro-simulations to compare the BGA classification of the Setser, Seldin, and Kidd panels. Overall, the Setser80 outperformed the Seldin and Kidd panels in naïve Bayesian classification and MLR classification accuracies in the GOAL dataset (naïve Bayes = 98.4%, MLR = 99%) and the 7 Populations Combined (naïve Bayes = 91.5%, MLR = 93.2%). Notably, PEL and MXL were classified with >95% and >80% accuracy, respectively, indicating the Setser80 panel is portable into other Hispanic datasets and populations.
Many panels have sought country-level ancestry determination, using a variety of SNP ascertainment methods19,21,28,29. Continentally, the EUROFORGEN Global AIMs12 and the Kidd5511 panel used allele frequency differentials (δ). Within a country, the United States HapMap 3 populations20 used PCA with receiver operating characteristics curve (ROC)19, and the Mexican mestizos panel used nested subsets with high SNP weights followed by the lowest number of SNPs with the highest PC121. Similar to Kidd et al.11, we prioritized SNPs that distinguished populations with lower mean FST per country. However, we focused on differentiating Hispanic instead of continental populations. Kosoy et al.9 (Seldin128) also concentrates on continental differentiation, but they also evaluated their AIMs on African American, Puerto Rican, and Mexican/Mexican American populations.
We used the Snipper 2.5 app suite29 that provided two classification methods: a naive Bayesian classifier and MLR32. This web-based classifier was designed for classification of externally visible characteristics33,34,35,36,37 and ancestry12,38,39,40,41, particularly in forensics. Snipper has successfully analyzed admixed South American populations34,42,43, similar to those used here.
The classification accuracy of the Seldin and Kidd panels is due to both the composition of their SNP ascertainment datasets and the size of the panels. The Seldin panel (96.2%, 96.3%) was more accurate in MXL than the Setser80 (83.4%, 89.8%) in the 7 Populations Combined and 1000 G datasets, respectively. Its success is likely because 199 of their 825 samples were from admixed Latin American and Amerindian individuals (Mexico and Puerto Rico especially)9. The Kidd panel emphasized capturing diversity by using 63 global populations11 including seven isolated Amerindian populations; they continue to add more populations via ALFRED44. The size of the Kidd panel and the ratio of SNPs to the number of samples (Kidd55 = 55 SNPs / 3071 samples = 0.0179; Seldin128 = 128 SNPs / 825 samples = 0.1552) suggests the number of SNPs, rather than SNP ascertainment population size, is the higher contributing factor to population differentiation. However, the number of individuals per population may also be a factor.
Our study’s limitations include: genechip design, sample size and its effect on allele frequencies. The GOAL25 study genechip45 was built on 270 African (YRI), Caucasian (CEU), and East Asian population (CHB and JPT) samples from HapMap 146, without any Amerindian component. Although, our SNP ascertainment dataset was small it was not inconsistent with other studies11,18,20 where the larger overall size was coupled with small sub-populations. Therefore, we combined the GOAL25 dataset with the 1000 Genomes Admixed American dataset (n = 347)28, merging COL with CLM (n = 147) and PUR with PUR (n = 122) due to negligible allele frequency differences, to create the 7 Populations Combined.
The design of the Setser80 is based on the balance of the countries via country attributable mean FST and selection of SNPs with LD < 0.7. Using a dilution series of 234 to 44 SNPs, we evaluated the effect of panel size on classification accuracy in relation to Seldin and Kidd sized panels and found 80 SNPs to be sufficient. Therefore we chose 80 SNPs from 247 candidates by selecting SNPs such that ~20% could be attributed to each country. It is possible that other panels informative of Hispanic ancestry could be selected from the same candidates, but testing multiple different panels was beyond the scope of this study. Residual LD is possible despite our threshold where four pairs of SNPs had r2 > 0.5; however, removing one of each pair and classifying two separate 76 SNP subsets had negligible effect on classification accuracy via naïve Bayes (Supplemental Table S4) or MLR (Supplemental Table S5). By treating these loci as independent, we may underestimate accuracy as Kidd et al. 2013 has shown that diplotypes are effective predictors of ancestry47.
We used micro-simulations in this study in order to normalize the size of each population and expand the analysis to seven Hispanic populations instead of the four publicly available through the 1000 Genomes Project28. Although real genotypes would have been preferable, widely variable population sizes could disproportionately affect the classification accuracy for smaller populations, as may have been the case with the real MXL genotypes. Our analysis of additional populations is a more realistic representation of the challenges of a more granular classification of heterogeneous populations. Forensic labs may not have access to a sizeable Hispanic database of individuals from multiple different countries; therefore, we simulated datasets based on readily available allele frequencies from multiple sources. By doing so, we have allowed MXL to misclassify into HUR which otherwise do not exist within the same dataset.
Additionally, our use of a static model for BGA determination may have overestimated classification success; despite reasonable success by other research groups48. Finally, our imputation of the Seldin1289 and Kidd5511 into the GOAL25 dataset required removal of ~30 loci to comply with the Setser80 QC filters. Missingness was not detrimental here because STRUCTURE disregards it49,50, and at 10% MLR is robust49. Alternatively, some missingness in micro-simulations may approximate the degraded forensic samples51.
Our findings indicate that the Setser80 can predict BGA of individuals of presumed Hispanic origin with high confidence. By selecting additional SNPs attributed to countries with lower average country attributable FST (COL, CUB, and PUR) to create the panel, the Setser80 had similar accuracy overall in GOAL25 and 7 Populations Combined. The Setser80 is robust as it clusters well with Bayesian model-based clustering and PCA, and classifies equally well in naïve Bayes classification and MLR. The Setser80 is portable and, to our knowledge, is the first BGA AIMs panel specifically for the Caribbean and surrounding mainland countries. In comparison to Seldin1289, Kidd5511, and 46 Consensus SNPs24, our 80 AIMs for Hispanic BGA is unique, both exact and by linkage disequilibrium. Therefore, it is our intention that the Setser80 be integrated into a future Western Hemisphere panel.
Materials and Methods
Genomic Origins and Admixture in Latinos (GOAL) dataset
Here we downloaded the GOAL dataset and used 160 unrelated individuals including Honduran (HUR, n = 13), Dominican Republican (DOM, n = 21), Colombian (COL, n = 53), Cuban (CUB, n = 55), and Puerto Rican (PUR, n = 18) populations with three of four grandparents from the same country25. These samples were collected in South Florida and genotyped using the Affymetrix 6.0 gene chip of 906,600 predetermined SNPs45.
The Genomic Origins and Admixture in Latinos (GOAL) dataset analyzed during the current study is available in the dbGaP repository, accession number phs000750.v1.p1, found at: https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000750.v1.p1&phv=202273&phd=4443&pha = &pht=3936&phvf = &phdf = &phaf = &phtf = &dssp=1&consent = &temp=1. Funding support for the GOAL Study was provided by the National Institute of General Medical Sciences (1R01GM090087). Additional support for sample collection was provided by a grant from the Stanley J. Glaser Foundation and the Dr. John T. Macdonald Foundation Department of Human Genetics.
Genomes (1000 G) dataset
For further comparison, we used fully sequenced individuals from the 1000 Genomes Project Phase 3 Admixed American populations (n = 347)28, accessed through the UCSC Genome Browser52. These include Colombia in Medellin (CLM, n = 94), Peru in Lima (PEL, n = 85), Puerto Rico (PUR, n = 104), and Mexican Living in Los Angeles (MXL, n = 64)28. The 1000 Genomes Project dataset is available via the UCSC Genome Browser, found at: http://genome.ucsc.edu/.
SNP ascertainment
We created our AIMs panel by applying a series of quality control algorithms. Beginning with 897,336 autosomal SNPs on the genechip45, we filtered the GOAL dataset by linkage disequilibrium (LD) ≤ 0.7, missingness ≤ 0.1, and minor allele frequency (maf) ≥ 0.01 using PLINK v.1.953,54 and retained 494,886 SNPs. After calculating FST31 by Weir & Cockerham’s algorithm30 in PLINK v.1.9 (https://www.cog-genomics.org/plink/1.9/basic_stats#fst)55, 1509 SNPs with FST ≥ 0.15 for at least one pairwise comparison were retained.
We calculated the mean FST for each of the five countries and assigned each SNP to a country based on the highest mean FST. The next highest mean FST was designated the 2nd country mean FST. For example, rs3777908 is attributed to HUR because the average of HUR vs. DOM, HUR vs. COL, HUR vs. CUB, and HUR vs. PUR is [(0.27318 + 0.19754 + 0.19560 + 0.28808)/4] = 0.23860, which was the highest country mean FST value for rs3777908. The 2nd highest country mean FST = 0.07442, corresponded to PUR (see Supplemental Table S1 for example calculations).
We binned the 1509 SNPs by the 1st and 2nd highest country attributable mean FST and removed SNPs where the 1st country mean FST < 0.11 and 2nd country mean FST < 0.09, resulting in 437 SNPs. Since 63.3% of the 1509 candidate SNPs were attributable to HUR or DOM, we removed SNPs where HUR and DOM had the 1st and 2nd highest country mean FST, where HUR had the 2nd highest country mean FST, and the 100 lowest ranked SNPs where HUR or DOM had the highest country mean FST. From the remaining 247 SNPs, we chose a subset of 80 in order to maintain ~20% contribution of SNPs for each country across 1st and 2nd country attribution. Therefore, we proceeded with the Setser80 (Supplemental Table S2), which has the following country attributable mean FST values: HUR (mean FST = 0.21228), DOM (mean FST = 0.16901), COL (mean FST = 0.14212), CUB (mean FST = 0.10803), and PUR (mean FST = 0.10272).
To assess the value of our panel, we compared it to two commonly sited AIMs panels9,11. Here, we refer to the panel developed by Kosoy et al., 2009 as the Seldin1289, and the 55 ancestry informative SNPs developed by Kidd et al., 2014 as the Kidd5511. We performed each analysis on the Setser80 in parallel with the Kidd and Seldin panels to evaluate the utility of our Hispanic AIMs panel.
Imputation
The SNPs on the Affymetrix 6.0 gene chip45 were pre-determined and not all SNPs were included in the ABI Taqman assay used to genotype the Seldin1289 and Kidd5511; therefore, we imputed these two panels into the GOAL dataset25 using IMPUTE256 on the full 250 individuals using a 5 Mb window centered on each SNP and an effective population size of 20,000 as seen in Instructions for IMPUTE version 257. We used 2,504 individuals from 1000G28 for the genetic map and legend and the strand alignment from dbSNP batch query. Given the use of genome builds NCBI35/hg17 to GRCh38/hg38, we converted all components to GRCh37/hg19 for analysis.
However, the gene chip used45 was based on an early genome build (NCBI35/hg17) which did not have all the tag SNPs necessary (in comparison to the 1000 G Project) to reliably impute ~30 of the SNPs from Seldin1289 and 11 from Kidd5511 for each individual. We assessed the accuracy of the imputation using the concordance tables provided by IMPUTE2; of the ~160 imputed SNPs from 20 chromosomes the mean concordance = 92.6% and range = 85.3% to 96.4%. Of the ~30 SNPs with missingness >10%, there was no obvious pattern between missingness proportion and concordance. Despite multiple attempts with different intervals, rs10954737 from the Seldin1289 was unable to be imputed due to the lack of Panel 2 SNPs. Because STRUCTURE and PCA ignore missing data49,50, the full Seldin1289 and Kidd5511 were used in these analyses. However, since the resampling approach to simulations is dependent upon the reliability of allele frequencies in our real data58, we applied the same <10% missingness filter used in the development of the Setser80; this resulted in 96 SNPs in the Seldin panel and 44 SNPs in the Kidd panel after imputation.
STRUCTURE
We evaluated ancestry by the Bayesian model-based clustering method used in STRUCTURE v.2.3.426 to compare the self-reported to computer-determined (K) populations. We performed STRUCTURE analysis at K = 2 to K = 7 for each dataset/panel at 10 iterations each using the admixture model, no LOCPRIOR, 10,000 burn-in, and 100,000 Markov Chain Monte Carlo (MCMC) repetitions. The final STRUCTURE diagrams for each SNP panel were optimized and averaged through STRUCTURE Harvester59, CLUMPP60, and Distruct61 to create the diagrams in Fig. 1.
Principal components analysis (PCA)
We analyzed the Setser80, Seldin1289, and Kidd5511 on the GOAL dataset by PCA using EIGENSOFT v.6.1.462 and plotted the first three eigenvectors. Genesis63 was used for improved visualization of clustering as seen in Fig. 1.
Linkage disequilibrium (LD) analysis
Using the web-based tool LDmatrix64, we compared the Setser80 to the Seldin1289 and Kidd5511, and the 46 Consensus SNPs described in a review article by Soundararajan et al.24. We used r2 > 0.7 as the threshold to evaluate whether any SNP in the Setser80 was in strong LD with SNP(s) from Seldin1289 and Kidd5511 (tested together) or the 46 Consensus SNPs appearing in more than 3 of 21 panels of AIMs24.
Modeling for the prediction of unknowns
To model the data for BGA prediction of unknown individuals, we used a resampling approach based on calculated allele frequencies of the three SNP panels on each dataset58. We simulated a randomly mating population of 100–125 individuals within each country. Next, we assigned a genotype to individuals by generating a random number between 1 and 0 and comparing this number to the maf for the country at the specified locus. Any random number above the maf was assigned the major allele. All genotypes were created from 2 separate allele generations for each locus. The simulation of each population was performed at least 5 times for the GOAL and 1000 G countries. The 7 Populations Combined dataset was created by merging the countries from the 1000 G and GOAL simulations without regard to simulation number. We verified our model using a chi-square test for each panel and found the allele frequencies from the simulation sets were not significantly different from the true allele frequencies at α = 0.05 after Bonferroni correction.
Classification of unknowns
Snipper 2.5 app suite29 is a web-based Naïve Bayes classifier, found here (http://mathgene.usc.es/snipper/), which calculates −log(likelihood) with leave-one-out cross-validation and multinomial logistic regression (MLR) options. Cross-validation divides a set of data into a training set and a testing set, and rotates the samples until all samples have been in the testing set. Using the “Thorough analysis of population data with a custom Excel file” option, Snipper calculated likelihood ratios (LR) of population vs. not the population and selected the country that corresponded to the highest LR. MLR is similar to STRUCTURE26,32, which calculated genetic proportions of individuals (as percent admixture) instead of whole populations, and categorized individuals based on those probabilities. We used 100–125 micro-simulations (individuals) each from population for references and selected 10% of profiles from a separate set of micro-simulations to predict unknowns. We evaluated potential overlap of MLR classification using the confusion matrix and assessed the validity of our classification by sensitivity, specificity, and positive predictive value from the naïve Bayes classification of the actual 1000 G genotypes (n = 347; CLM = 94, PUR = 104, PEL = 85, and MXL = 64).
Ethical approval and informed consent
This research study using the Genomic Origins and Admixture in Latinos (GOAL) from Moreno-Estrada, A. et al. (2013)25, and the 1000 Genomes Project28 datasets was approved under University of North Texas Health Science IRB 2013-201. As this manuscript only used pre-existing genetic data from Moreno-Estrada, A. et al. (2013)25, where their “Informed consent was obtained from all participants under approval by the University of Miami Institutional Review Board (study no. 20081175)”. The 1000 Genomes Project data was only included in the International Genome Sample Resource if the submission was in accordance with the Consent, Ethics Review and Sampling Process of the 1000 Genomes Project28.
References
Gao, C. A comprehensive analysis of common and rare variants to identify adiposity loci in Hispanic Americans: The IRAS family study (IRASFS). PLoS ONE. 10(11) e0134649; https://doi.org/10.1371/journal.pone.0134649 (2015 November 1).
Phillips, C. Forensic genetic analysis of bio-geographical ancestry. Forensic Sci. Int-Gen. 18, 49–65 (2015).
Burkart, K. M. et al. A genome-wide association study in Hispanics/Latinos identifies novel signals for lung function – The Hispanic Community Health Study/Study of Latinos. Am J Resp Crit Care Med. 198 (2), 208–219 (2018 July 15).
Manichaikul, A. et al. Population structure of Hispanics in the United States: The multi-ethnic study of atherosclerosis. PLoS Genet. 8 (4), e1002640; https://doi.org/10.1371/journal.pgen.1002640 (2012 April).
MacArthur, J. et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS catalog). Nucleic Acids Res. 45(D1), D896-D901 (2017 January 1).
Norris, E. T. et al. Genetic ancestry, admixture and health determinants in Latin America. BMC Genomics. 19(Suppl 8), 861 (2018 December).
Ambers, A. D. et al. Comprehensive forensic genetic marker analysis for accurate human remains identification using massively parallel DNA sequencing. BMC Genomics. 17(Suppl 9), 750 (2016 October 17).
Lorente, J. A. Trafficking in human beings: modern slavery. EndSlavery. Workshop 2-3, November 2013. URL, http://www.endslavery.va/content/endslavery/en/publications/scripta_varia_122/lorente.html (2019).
Kosoy, R. et al. Ancestry informative marker sets for determining continental origin and admixture proportions in common populations in America. Hum. Mutat. 30(1), 69–78 (2009 January).
Galanter, J. M. et al. Development of a panel of genome-wide ancestry informative markers to study admixture throughout the Americas. PLoS Genet. 8 (3), e1002554; https://doi.org/10.1371/journal.pgen.1002554 (2012 March).
Kidd, K. K. et al. Progress toward an efficient panel of SNPs for ancestry inference. Forensic Sci. Int-Gen. 10(1), 23–32 (2014 May).
Phillips, C. et al. Building a forensic ancestry panel from the ground up: the EUROFORGEN Global AIM-SNP set. Forensic Sci. Int-Gen. 11(1), 13–25 (2014 July).
Hellenthal, G. et al. A genetic atlas of human admixture history. Science. 343(6172), 14 (2014 February 14).
Elhaik, E. et al. Geographic population structure analysis of worldwide human populations infers their biogeographical origins. Nat Commun. 5, 3513; https://doi.org/10.1038/ncomms4513 (2014 April 29).
Marcheco-Teruel, B. et al. Cuba: Exploring the history of admixture and the genetic basis of pigmentation using autosomal and uniparental markers. PLoS Genet. 10(7), e1004488, https://doi.org/10.1371/journal.pgen.1004488 (2014 July 14).
Fortes-Lima, C. et al. Exploring Cuba’s population structure and demographic history using genome-wide data. Sci Rep. 8(1), 11422, https://doi.org/10.1038/s41598-018-29851-3 (2018 December 1).
Jia, J. et al. Developing a novel panel of genome-wide ancestry informative markers for bio-geographical ancestry estimates. Forensic Sci. Int-Gen. 8, 187–194 (2014).
Tian, C. et al. Analysis of East Asia genetic substructure using genome-wide SNP arrays. PLoS ONE. 3 (12), e3862, https://doi.org/10.1371/journal.pone.0003862 (2008 December 5).
Zeng, X. et al. Selection of highly informative SNP markers for population affiliation of major US populations. Int. J. Leg. Med. 130(2), 341–352 (2016 March).
Altshuler, D. M. et al. Integrating common and rare genetic variation in diverse human populations. Nature. 467(7311), 52–58 (2010 September 2).
Huerta-Chagoya, A. et al. A panel of 32 AIMs suitable for population stratification correction and global ancestry estimation in Mexican mestizos. BMC Genetics. 20(1) 5; https://doi.org/10.1186/s12863-018-0707-7 (2019 January 8).
Elhaik, E. et al. The GenoChip: A new tool for genetic anthropology. Genome Biol Evol. 5 (5), 1021–1031 (2013 May).
Leslie, S. et al. The fine-scale genetic structure of the British population. Nature. 519(7543), 309–314 (2015 March 19).
Soundararajan, U., Yun, L., Shi, M. & Kidd, K. K. Minimal SNP overlap among multiple panels of ancestry informative markers argues for more international collaboration. Forensic Sci Int-Gen. 23, 25–32 (2016 July 1).
Moreno-Estrada, A. et al. Reconstructing the population genetic history of the Caribbean. PLoS Genet. 9 (11), e1003925; https://doi.org/10.1371/journal.pgen.1003925 (2013 November 14).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics. 155(2), 945–59 (2000 June).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 14(8), 2611–2620 (2005 July).
Auton, A. et al. A global reference for human genetic variation. Nature. 526 (7571), 68–74 (2015 October 1).
Phillips, C. et al. Inferring ancestral origin using a single multiplex assay of ancestry-informative marker SNPs. Forensic Sci. Int-Gen. 1(3-4), 273–280 (2007 December).
Weir, B. S. & Cockerham, C. C. Estimation of gene flow from F-statistics. Evolution. 47(3), 855–863 (1993).
Wright, S. The genetical structure of populations. Ann. Eugenic. 15(4), 323–354 (1951 March).
McNevin, D. et al. An assessment of Bayesian and multinomial logistic regression classification systems to analyse admixed individuals. Forensic Sci. Int-Gen. Suppl. Ser. 4, e63–e64, https://doi.org/10.1016/j.fsigss.2013.10.032 (2013).
Ruiz, Y. et al. Further development of forensic eye color predictive tests. Forensic Sci. Int-Gen. 7(1), 28–40 (2013 January).
Freire-Aradas, A. et al. Exploring iris colour prediction and ancestry inference in admixed populations of South America. Forensic Sci. Int-Gen. 13, 3–9 (2014 Novemeber).
Maroñas, O. et al. Development of a forensic skin colour predictive test. Forensic Sci. Int-Gen. 13, 34–44 (2014 November).
Söchtig, J. et al. Exploration of SNP variants affecting hair colour prediction in Europeans. Int. J. Leg. Med. 129(5), 963–975 (2015 September).
Pośpiech, E. et al. The common occurrence of epistasis in the determination of human pigmentation and its impact on DNA-based pigmentation phenotype prediction. Forensic Sci. Int-Gen. 11, 64–72 (2014 July).
Fondevila, M. et al. Revision of the SNPforID 34-plex forensic ancestry test: Assay enhancements, standard reference sample genotypes and extended population studies. Forensic Sci. Int-Gen. 7(1), 63–74 (2013 January).
Pereira, R. et al. Straightforward inference of ancestry and admixture proportions through ancestry-informative insertion deletion multiplexing. PLoS ONE. 7(1), e29684; https://doi.org/10.1371/journal.pone.0029684 (2012 January 17).
De la Puente, M. et al. The Global AIMs Nano set: A 31-plex SNaPshot assay of ancestry-informative SNPs. Forensic Sci. Int-Gen. 22, 81–88 (2016 May).
Eduardoff, M. et al. Inter-laboratory evaluation of the EUROFORGEN Global ancestry-informative SNP panel by massively parallel sequencing using the Ion PGM™. Forensic Sci. Int-Gen. 23, 178–189 (2016 July 1).
Heinz, T. et al. Ancestry analysis reveals a predominant Native American component with moderate European admixture in Bolivians. Forensic Sci. Int-Gen 7, 537–542 (2013).
Taboada-Echalar, P. et al. The genetic legacy of the pre-colonial period in contemporary Bolivians. PLoS ONE. 8 (3), e58980; https://doi.org/10.1371/journal.pone.0058980 (2013 March).
Rajeevan, H., Soundararajan, U., Kidd, J. R., Pakstis, A. & Kidd, K. K. ALFRED: An allele frequency resource for research and teaching. Nucleic Acids Res. 40(D1), D1010–D1015 (2012 January).
McCarroll, S. A. et al. Integrated detection and population genetic analysis of SNPs and copy number variation. Nat Genet. 40 (10), 1166–74 (2008 October).
The International HapMap Consortium. The International HapMap Project. Nature. 426(6968), 789–796 (2003 December 18).
Kidd, K. K. et al. Microhaplotype loci are a powerful new type of forensic marker. Forensic Sci. Int-Gen. Supp Ser. 4, e123–e124, https://doi.org/10.1016/j.fsigss.2013.10.063 (2013).
Kusev, P., van Schaik, P., Tsaneva-Atanasova, K., Juliusson, A. & Chater, N. Adaptive anchoring model: How static and dynamic presentation of time series influence judgment predictions. Cogn. Sci. 42(1), 77–102 (2018 January).
Cheung, E. Y. Y., Gahan, M. E. & McNevin, D. Prediction of biogeographical ancestry from genotype: a comparison of classifiers. Int. J. Leg. Med. 131(4), 901–912 (2017 July 1).
Pritchard, J. K., Wen, X. & Falush, D. Documentation for STRUCTURE software: Version 2.3. (2010 February 2).
Butler, J. M. (2nd ed.) Forensic DNA typing: Biology, technology, and genetics of STR markers. (Elsevier, 2005).
Kent, W. J. et al. UCSC Genome Browser: The human genome browser at UCSC. Genome Res. 12(6), 996–1006 (2002 June). URL, http://genome.ucsc.edu
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81(3), 559–575 (2007 September).
Chang, C. C. et al. Second-generation PLINK: Rising to the challenge of larger and richer datasets. GigaScience. 4(1), 7; (2015 February 25).
Purcell, S. & Chang, C. PLINK 1.9. URL, http://www.cog-genomics.org/plink/1.9/.
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5(6), e1000529, https://doi.org/10.1371/journal.pgen.1000529 (2009).
Howie, B. & Marchini, J. Instructions for IMPUTE version 2. (2009 June 18). URL, https://mathgen.stats.ox.ac.uk/impute/impute_v2_instructions.pdf
Yuan, X., Miller, D. J., Zhang, J., Herrington, D. & Wang, Y. An overview of population genetic data simulation. J. Comput. Biol. 19(1), 42–54 (2012).
Earl, D. A. & vonHoldt, B. M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4(2), 359–361 (2012).
Jakobsson, M. & Rosenberg, N. A. CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics. 23 (14), 1801–1806 (2007 July 15).
Rosenberg, N. A. Distruct: A program for the graphical display of population structure. Mol. Ecol. Notes. 4(1), 137–138 (2004 March).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2(12), e190, https://doi.org/10.1371/journal.pgen.0020190 (2006).
Buchmann, R.W. Genesis: Copyright (C) 2014, University of the Witwatersrand.
National Cancer Institute, Division of Cancer Epidemiology & Genetics. LD Matrix. URL, https://ldlink.nci.nih.gov/?tab=ldmatrix (2019).
Acknowledgements
Funding support for the Genomic Origins and Admixture in Latinos (GOAL) Study was provided by the National Institute of General Medical Sciences (1R01GM090087). Additional support for sample collection was provided by a grant from the Stanley J. Glaser Foundation and the Dr. John T. Macdonald Foundation Department of Human Genetics. The dataset used for the analyses described in this manuscript was obtained from dbGaP through accession number phs000750.v1.p1. The authors would like to thank the late Dr. Arthur Eisenberg for the inspiration behind this project based on the needs of the Center for Human Identification and DNA ProKids. We would also like to thank the late Dr. Ranajit Chakraborty who was instrumental in the design of this research. We thank Dr. Carlos Bustamante for making the GOAL dataset available via dbGaP. Dr. Xiangpei Zeng helped with STRUCTURE and Dr. Frank Wendt helped with access and use of data from 1000 Genomes. Dr. Gita Pathak helped in many small but significant ways in discussing concepts and troubleshooting software.
Author information
Authors and Affiliations
Contributions
C.H.S. designed the project alongside R.C., performed the analyses, interpreted results alongside J.V.P. and R.B., troubleshot with input from N.P., and prepared the manuscript. D.S.C. has provided substantial guidance in experimental design, interpretation, troubleshooting, and especially crafting research into a publishable manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Setser, C.H., Planz, J.V., Barber, R.C. et al. Differentiation of Hispanic biogeographic ancestry with 80 ancestry informative markers. Sci Rep 10, 7745 (2020). https://doi.org/10.1038/s41598-020-64245-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-64245-4
- Springer Nature Limited