Abstract
Altered branched-chain amino acids (BCAAs) metabolism is a distinctive feature of various cancers and plays an important role in sustaining tumor proliferation and aggressiveness. Despite the therapeutic and diagnostic potentials, the role of BCAA metabolism in cancer and the activities of associated enzymes remain unclear. Due to its pivotal role in BCAA metabolism and rapid cellular transport, hyperpolarized 13C-labeled α-ketoisocaproate (KIC), the α-keto acid corresponding to leucine, can assess both BCAA aminotransferase (BCAT) and branched-chain α-keto acid dehydrogenase complex (BCKDC) activities via production of [1-13C]leucine or 13CO2 (and thus H13CO3−), respectively. Here, we investigated BCAA metabolism of F98 rat glioma model in vivo using hyperpolarized 13C-KIC. In tumor regions, we observed a decrease in 13C-leucine production from injected hyperpolarized 13C-KIC via BCAT compared to the contralateral normal-appearing brain, and an increase in H13CO3−, a catabolic product of KIC through the mitochondrial BCKDC. A parallel ex vivo 13C NMR isotopomer analysis following steady-state infusion of [U-13C]leucine to glioma-bearing rats verified the increased oxidation of leucine in glioma tissue. Both the in vivo hyperpolarized KIC imaging and the leucine infusion study indicate that KIC catabolism is upregulated through BCAT/BCKDC and further oxidized via the citric acid cycle in F98 glioma.
Similar content being viewed by others
Introduction
In tumors, energy dependence on substrates is significantly altered to compensate elevated biosynthesis and bioenergetics needed for tumor proliferation1. For instance, most cancer cells rely on energy-inefficient aerobic glycolysis, known as ‘the Warburg effect’2, to facilitate the uptake and incorporation of nutrients into the biomass for rapid proliferation. The biosynthetic demands in cancer also stimulate utilization of other nutrients such as glutamine3, acetate4,5, and branched-chain amino acids (BCAAs: leucine, valine, and isoleucine). In particular, BCAAs are reportedly essential for tumor proliferation6,7,8 and altered BCAA utilization has been demonstrated in multiple cancers9. Overexpressed branched-chain amino acid aminotransferase 1 (BCAT1), a cytosolic enzyme that transfers α-amino groups to α-ketoglutarate to yield the respective branched-chain α-keto acid (BCKA), has been suggested as a prognostic cancer indicator6,10,11,12. Moreover, a recent study showed increased expression of mitochondrial BCAT (BCAT2) in pancreatic adenocarcinoma13. Despite the emerging roles of BCAAs, however, the molecular mechanism of associated enzyme activities in cancer still remains elusive8,9,14.
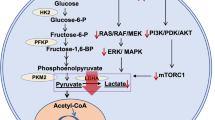
In the brain, BCAAs serve as nitrogen donors in the glutamine/glutamate cycle as well as sources for energetic and biosynthetic demands15,16 (Fig. 1). As the first step of BCAA catabolism, BCAAs are transaminated to BCKAs by BCAT isoenzymes. Although the BCATs catalyze reversible transamination reaction between BCAA and BCKA, the catabolic pathway of BCAAs is the predominant direction in most cell types15. α-ketoglutarate is converted to glutamate via BCAT1 in neurons and via BCAT2 in astrocytes to yield the major excitatory neurotransmitter of central nervous system17. In the tripartite synapse, astrocytes convert excess glutamate to α-ketoglutarate by reanimating BCKAs to maintain the concentration of glutamate below 5–10 mM as a glutamate-buffering mechanism for brain nitrogen homeostasis. As a result of this regulatory mechanism, for instance, physiological concentration of α-ketoisocaproic acid (KIC) is maintained at 0.7 ± 0.1 μM in rat brain, which is significantly lower than KIC in rat blood plasma (18.9 ± 1.4 μM)18. Subsequently, the BCKAs are converted to branched-chain acyl-CoA’s via branched-chain α-keto acid dehydrogenase complex (BCKDC) at an irreversible oxidative decarboxylation step. Thus, CO2 produced in this step can serve as a direct indicator of BCAA oxidation. The ensuing enzymatic reactions are unique for each BCAA with end products from each degradative pathway eventually being oxidized through the citric acid cycle (CAC).
In brain malignancies (e.g., glioma), BCAA uptake via L–amino acid transporter-1 (LAT1) is enhanced compared to normal astrocytes19 and BCAT1 activity is augmented in primary gliomas with wild-type isocitrate dehydrogenase (IDHwt). In particular, the BCATs are suggested as cancer-specific biomarkers, but the directionality of the BCAT-mediated reaction and how the BCAT enzyme levels are associated with metabolic phenotypes in tumor have been addressed in different manners9. In glioblastoma, BCAA catabolism is upregulated to support cell proliferation6. In contrast, increased reverse direction was observed in non-small cell lung cancer for tissue protein synthesis14 and in leukemia for cancer progression8. As the tumor metabolism is progressively remodeled by adapting to the surrounding tumor microenvironment, it is important to understand the environmental factors and tumor heterogeneities, and therefore necessary to assess comprehensive tumor characteristics in vivo20.
Non-invasive assessment of in vivo metabolism is crucial to understand the altered BCAA metabolism and to identify potential imaging biomarkers in cancer. Dynamic nuclear polarization (DNP or ‘hyperpolarization’) in combination with a rapid dissolution and 13C MR spectroscopy allows in vivo investigation of multiple metabolic pathways by detecting metabolic flux of 13C labeled substrates21. It has been a useful imaging tool for assessing cancer-specific enzyme-catalyzed metabolic phenotypes22,23. In particular, hyperpolarized (HP) [1-13C]KIC was used to assess BCAT-mediated leucine metabolism in the brain in vivo24. [1-13C]KIC has a relatively long longitudinal relaxation time (T1), a key parameter that determines the length of the observable time-window for in vivo metabolism. KIC is rapidly transported into the cell via monocarboxylic transporter (MCT)24 and can assess both BCAT and BCKDC activities via production of [1-13C]leucine or 13CO2 (and thus H13CO3−), respectively. Indeed, previous studies showed that the HP [1-13C]KIC conversion to [1-13C]leucine was correlated with BCAT activity in prostate cancer cells, lymphoma, and normal rat brain. In the prostate cancer cells, elevated BCAT levels resulted in increased production of HP [1-13C]leucine from HP [1-13C]KIC25. Another HP [1-13C]KIC study observed increased [1-13C]leucine conversion in EL4 lymphoma26. Moreover, Butt, et al. demonstrated that 13C metabolic imaging using HP 13C KIC is a practical method to access the BCAT activity in vivo in rat brains24.
We hypothesized that BCAA oxidation would be increased in glioma-bearing rats as a unique feature of tumor, which can be visualized by 13C MRI in combination with DNP. In this study, we investigated the increase of in vivo leucine oxidation in F98 glioma and the feasibility of HP [1-13C]KIC as an imaging agent to measure altered BCAA metabolism in glioma by assessing BCAT- and BCKDC-catalyzed in vivo enzyme reactions. Moreover, 13C NMR isotopomer analysis was performed after a steady-state infusion of [U-13C]leucine into glioma-bearing rats to elucidate the downstream metabolism of BCAA. These results were further evaluated by subsequent ex vivo tissue analysis of BCAT/BCKDC enzyme activities and its expression levels.
Results and Discussion
GBM animal model
We used F98 cell line for glioma model. It is histologically classified as an anaplastic glioma, which is pathologically undifferentiated malignant glioma27,28, and the characteristics of tumor growth are similar with human glioblastoma multiforme (GBM) in overexpression of the epidermal growth factor receptor (EGFR)28. The growth of tumors was confirmed 15–18 days after the cell implantation using both T2-weighted and contrast-enhanced (CE) T1-weighted proton MRI. The tumor regions identified by postmortem hematoxylin and eosin (H&E) stain was better matched to the hyperintense regions of the CE T1-weighted-MRI rather than those of T2-weighted images (Fig. S1).
In vivo 13C imaging of branched-chain amino acid metabolism in glioma using hyperpolarized [1-13C] α-ketoisocaproate to assess BCAT/BCKDC activity of glioma
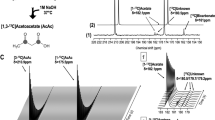
We first tested HP [1-13C]KIC in F98 glioma cells in vitro. A time-resolved pulse-and-acquire 13C sequence with a non-selective radiofrequency (RF) pulse was used to monitor the injected [1-13C]KIC (δ13C = 172.6 ppm) and products every 3 sec. BCAT and BCKDC activities were detected by the appearance of [1-13C]leucine (δ13C = 176.8 ppm) and H13CO3− (δ13C = 163.1 ppm), respectively. The signal of HP [1-13C]leucine and H13CO3− reached a maximum about 10–15 sec after the injection of HP [1-13C]KIC (Fig. 2a–c). All the KIC samples were polarized by a SPINlab™ DNP polarizer. The polarization level and the T1 at 3 T were measured as 24% (35 sec after the dissolution) and 77 sec from independent calibration experiments.
(a) Metabolic pathway of hyperpolarized (HP) [1-13C]KIC. 13C-labeled KIC and its metabolic products are highlighted in red. (b) An in vitro time-averaged spectrum from F98 cells showed the injected HP [1-13C]KIC and produced [1-13C]leucine and H13CO3− peaks. (c) The corresponding time courses of HP KIC and the products. (d–e) In vivo chemical shift imaging of a F98 glioma-bearing rat using HP [1-13C]KIC and (f) the contrast-enhanced (CE) T1-weighted 1H image. Metabolite distributions of [1-13C]leucine and H13CO3− in a tumor-bearing rat brain slice after an injection of HP [1-13C]KIC. (g–h) The reconstructed spectra in the glioma (solid red) and the contralateral normal-appearing brain (NAB; dotted blue) and (i) in vivo metabolite ratio of [1-13C]leucine to H13CO3−.
For in vivo imaging, a two-dimensional spiral chemical shift imaging pulse sequence with a variable RF scheme was used to acquire single time-point metabolite maps of glioma-bearing rat brain following an intravenous injection of HP [1-13C]KIC (injection-to-scan = 30 sec, scan time = 0.5 sec)29. Upon intravenous bolus injection of [1-13C]KIC into F98 glioma rats, [1-13C]leucine and H13CO3− were detected in the rat brain tissue (Fig. 2d–f). Each metabolite was quantified by integrating the corresponding peak in the absorption mode after spectral tomosynthesis30 and phase correction. Figure 2g–h are the reconstructed spectra from the tumor and the normal-appearing brain (NAB) for leucine and bicarbonate (HCO3−), respectively. KIC has been found to cross the blood-brain barrier (BBB) predominantly by a carrier-mediated transport system and subsidiary by simple diffusion under physiological condition31. Exogenously injected HP KIC was taken up by MCTs and metabolized to leucine and HCO3− by BCAT and BCKDC, respectively32. Reversible BCAT reaction enables a carbon isotope exchange between KIC and leucine in addition to the net flux of HP 13C from KIC to leucine33. Therefore, HP 13C KIC to leucine conversion is determined by endogenous 12C leucine pool size as well as membrane transport of KIC, delivery to the tumor, and BCAT enzyme activities. On the other hand, conversion of HP KIC to HCO3−, a decarboxylated product of [1-13C]KIC, is irreversibly mediated by BCKDC and, therefore, indicates a pure metabolic catabolic flux. 13C-leucine, the transamination product of KIC was smaller in the tumor as compared to the contralateral NAB ([1-13C]leucine/total 13C = 0.0053 in tumor vs. 0.0081 in NAB, p < 0.05). The low 13C conversion from KIC to leucine in the tumor implies predominant net conversion of leucine to KIC or suppressed isotope exchange between KIC and leucine (e.g., depletion of intrinsic leucine pool). Conversely, more H13CO3− was detected in the tumor (H13CO3−/total 13C = 0.005, p < 0.05) compared to the NAB (0.0027), suggesting elevated KIC catabolism by BCKDC towards isovaleryl-CoA.
Increased of [U-13C]leucine oxidation in F98 glioma
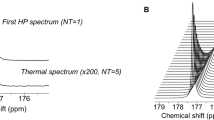
BCAA can be partially oxidized for protein synthesis or can be fully oxidized as an energy source. Although major BCAA oxidative capacity is high in skeletal muscle or liver, brain has 20% proportion of the whole-body capacity to oxidize BCAA34 and this rate is greater than incorporation rate of BCAA into protein15. Previous studies have shown increased uptake of BCAA by C6 glioma cell19 and breast cancer cell lines (MDA-MB-231, MCF-7) via LAT135. To elucidate the metabolic fate of BCAA in glioma, leucine metabolism was further explored in F98 glioma using an ex vivo 13C NMR isotopomer analysis. The isotopomer analysis can track down multiple metabolic pathways by accessing the distribution of 13C isotopomers in tissue after infusion of 13C-labeled substrates. We evaluated downstream metabolism of leucine in brain tumor after a [U-13C]leucine infusion into F98 glioma-bearing rats. Oxidation of [U-13C]leucine in the glioma produced distinct labeling in several CAC-intermediates as downstream products. In particular, 13C-glutamate labeling patterns enable direct comparison of leucine utilization between glioma and NAB. The glutamate doublet in 13C NMR spectra were also analyzed to determine the quantity of 13C labeled acetyl-CoA entering the CAC. The protocol for 13C-leucine infusion study is described in Fig. 3a. Both glioma and contralateral NAB tissues were harvested and freeze-clamped after a steady-state infusion of [U-13C]L-leucine. 13C NMR spectra from glioma and NAB tissue extracts were compared by examining the distribution of 13C isotopomers. The data showed that [U-13C]leucine was converted to [U-13C]KIC and further catabolized to [1,2,3,4,5-13C5]isovaleryl-CoA and H13CO3− via mitochondrial BCKDC, followed by conversion to [2,3,4-13C3]acetoacetate and [1,2-13C2]acetyl-CoA. Both [1,2-13C2] and [2-13C]acetyl-CoA are oxidized in the CAC and form various 13C isotopomers in 13C-glutamate. 13C NMR spectra reflected higher quantities of 13C-labeled leucine in glioma tissue versus NAB tissue (Fig. 3b). Glutamate derived from leucine was estimated from C4 doublet of [4,5-13C2]glutamate (23.1 ± 6.2% of total glutamate pool in glioma, 15.5 ± 5.9% in NAB, p < 0.05, Fig. 3d,e). Moreover, the multiplets appearing in the lactate C2 resonance were more abundant in the spectrum of glioma tissue, reflecting oxidation of leucine in the CAC followed by cycling of three carbon units out of the CAC and into pyruvate (pyruvate recycling) (Fig. 3c). Based on the 1H MR spectroscopy, leucine concentration in brain tissue was significantly higher in glioma (0.278 ± 0.035 μmol/g of wet tissue) than in contralateral NAB (0.075 ± 0.001 μmol/g). Lactate pool was also larger in the glioma (9.989 ± 1.413 μmol/g) compared to the NAB (7.639 ± 1.506 μmol/g) (Fig. S2). Together, these results indicate that leucine uptake and oxidation are increased in glioma. The total leucine plasma level was within the physiological condition during the infusion (91.7 ± 16 μmol/L in tumor-bearing rat)36. Increased catabolism of [U-13C]leucine in tumor tissue is consistent with the in vivo 13C imaging results using HP [1-13C]KIC in F98 glioma that showed higher decarboxylation product of HCO3− from [1-13C]KIC in the tumor compared to NAB.
13C NMR spectra acquired from glioma and contralateral normal-appearing brain (NAB) ex vivo after a steady-state [U-13C]leucine infusion. (a) Protocol for the [U-13C]leucine infusion. (b) Elevated leucine uptake and significantly higher (c) lactate and (d) glutamine labeling were observed in the tumor as compared to NAB, indicating an increased oxidation of leucine in the tumor. (e) 13C-labeled [4-13C]glutamate derived from [U-13C]leucine relative to the total glutamate pool size (%) from the leucine infusion study (*p < 0.05). D indicates doublet.
Ex vivo assay of BCAT and BCKDC activities in brain tissues
To verify the results of the in vivo imaging and the steady-state infusion study, enzyme activities and expression levels of BCAT and BCKDC were measured in brain tissues. BCAT activity assays were performed as previously described using in vitro continuous spectrophotometric method. L-alanine aminotransferase and L-lactate dehydrogenase were used as coupled enzyme in this assay37. BCKDC subunit activity was assayed in the brain tissue according to the methods of radiochemical assay by [1-14C]KIC substrate and measure the [14C]-decarboxylation by BCKDC38. The previous study reported that BCAT1 is a major isoenzyme in rat brain responsible for 60 - 70% of total BCAT activity39. This is congruent with our western blot analysis; The expression of BCAT1, rather than BCAT2, is relatively predominant in both NAB and glioma. While the total BCAT activities were comparable between glioma (3.34 ± 0.36 U/gram of protein) and NAB (3.16 ± 0.05), the expression levels of isoenzyme BCAT1 and BCAT2 were significantly tumor-specific: lower BCAT1 and higher BCAT2 in glioma than in NAB (Fig. 4). The distinct levels of BCAT1 and 2 in glioma might be due to the characteristics of its cellular origin, glial cell28,40, as tumors conserve the metabolic phenotype of their tissue origin41. Indeed, BCAT2 is predominant in glial cells whereas BCAT1 is primarily in cortical neurons17,42,43. In contrast, both the activity and the expression level of the BCKDC-E1α subunit were at similar levels in glioma (3.80 ± 1.87 nmol CO2/min/g tissue) and NAB (4.41 ± 0.86) (Fig. 4). Increased expression level of BCAT2 and unaltered BCKDC level in glioma together suggests that BCAT2 is a key enzymatic step that regulates BCAA catabolism in glioma. Islam, M.M., et al.39 recently reported that BCAT2 forms a protein-protein complex with BCKDC such that the substrates like KIC are channeled to the adjacent E1 subunit of BCKDC for further catabolism. Such channeling of substrates could explain the enhanced H13CO3− production observed in glioma versus normal brain tissue.
Outlook
GBM is one of the most aggressive primary brain tumors. Nevertheless, diagnosis and prognosis is limited by poor understanding of molecular biomarker44 and biopsy is restricted if the tumor is inoperable. Although the role of BCAA in cancer needs further investigation, multiple enzymatic steps of BCAA metabolism can be important prognosis and diagnosis biomarkers in tumor. In particular, BCAT1 expression is dependent on the concentration of α-ketoglutarate substrate in glioma cell lines and could be suppressed by ectopic overexpression of mutant IDH isoform 1 (IDH1) in immortalized human astrocytes, providing a link between IDH1 function and BCAT1 expression. This link is supported by IDHwt astrocytic gliomas being characterized by high BCAT1 expression6. The overexpression of mutant IDH1 associated with reduced BCAT1 expression and reduced invasiveness and tumor growth provides a plausible rationale for the aggressiveness of primary GBM and the relatively less aggressive low grade or secondary gliomas. The HP KIC imaging method can assess activities of two key enzymes in BCAA catabolism, BCAT and BCKDC, and therefore potentially play a crucial role in tumor characterization, drug development and therapeutic decision.
Conclusion
HP [1-13C]KIC is a promising substrate to investigate in vivo brain tumor metabolism. KIC can be delivered to intracellular space via MCT and instantly converted to [1-13C]leucine by BCAT24 or to CO2 (and thus HCO3−) by BCKDC. The detection of BCKDC activity is crucial for assessing BCAA oxidation since it irreversibly catabolizes BCKAs and regulates the BCAA oxidation. Here we demonstrated the feasibility of HP [1-13C]KIC to assess BCAT/BCKDC activity in F98 glioma in vivo and investigated that leucine oxidation was increased in the tumor as a unique feature of glioma. In addition, we showed the increase of in vivo leucine oxidation in F98 glioma as a unique metabolic feature of brain tumor using steady-state infusion [U-13C]leucine. Both the in vivo HP KIC imaging and the leucine infusion study indicate that KIC catabolism is upregulated through BCAT/BCKDC and further oxidized via the CAC in F98 glioma. To our knowledge, this is the first report of HP [1-13C]KIC in vivo imaging in rat glioma model. Besides glioma and other cancer applications, we expect that this technique can be utilized to investigate BCAT/BCKDC-associated pathological features in multiple other diseases with altered BCAA metabolism such as traumatic brain injury45.
Methods
GBM animal model and tumor implantation
F98 glioma cell line was used for both in vitro and in vivo studies. It is histologically classified as an anaplastic glioma, which is pathologically undifferentiated malignant glioma, and the characteristics of tumor growth are similar with human GBM in overexpression of the EGFR28. The F98 glioma cells (CRL-2397TM) were obtained from ATCC (Manassas, VA, USA). The cells were cultured at 37 °C in a monolayer using Dulbecco’s Modified Eagle Medium (DMEM) with 10% fetal bovine serum (FBS) and streptomycin with 5% CO229. The cells were harvested at 85~90% confluency. Approximately 1 × 104 cells were prepared for each animal. After a Trypan blue exclusion test for cell viability, the cells were implanted into the right striatum of male Fisher rats (n = 12, body weight = 240 ± 10 g). The tumor implantation was performed at the Neuro-Models Core Facility of the University of Texas Southwestern Medical Center. The growth of tumors was confirmed after 15–18 days of implantation by T2-weighted and CE T1-weighted proton MRI. The tumor regions identified by postmortem H&E stain was better matched to the hyperintense regions of the CE T1-weighted-MRI rather than those of T2-weighted images (Fig. S1).
When the tumors were established the animals were either injected with HP [1–13C]KIC for MRSI or infused with [U-13C]leucine for isotopomer analysis. Tumor volumes were estimated as 93.5 ± 10.3 mm3 by measuring hyperintense regions in a 2D T2-weighted MRI using OsiriX dicom viewer. Tumor sizes were further confirmed by a 2D CE T1-weighted MRI (50.0 ± 26.5 mm3). Immediately after the imaging or infusion studies, the glioma and NAB tissues were collected for enzyme assays, histology or isotopomer analysis. For comparison, healthy male Fisher rats (n = 3, body weight = 252 ± 10 g) were also examined. All the animal protocols were approved by the local Institutional Animal Care and Use Committee and all experiments were performed in accordance with relevant guidelines and regulations.
Hyperpolarization study of [1-13C] α-ketoisocaproic acid
[1-13C] α-ketoisocaproic acid (KIC) was prepared from [1-13C] α-ketoisocaproic acid sodium salt (Cambridge Isotope Laboratories, Andover, MA, USA) by dissolving in 1-M HCl (pH < 1) and extracting the aqueous layer with diethyl ether. The combined organic layer was dried over anhydrous sodium sulfate, filtered and evaporated to afford [1-13C]KIC (7 M, colorless oil, purity >95%, 94% yield)25. For hyperpolarization, a 70-μL sample of the [1-13C]KIC mixed with 11-mM trityl radical (OX063, Oxford Instruments Molecular Biotools Ltd, Oxfordshire, UK) was placed in a sample vial, which was then assembled with a research fluid path (GE Healthcare, Waukesha, WI, USA) containing 16 mL of dissolution media (0.1 g/L Na2EDTA) in the dissolution syringe. After placed in a SPINlab® polarizer (GE Healthcare), the sample was polarized at ~0.8 K of temperature in a 5 T magnet by irradiating microwaves at a frequency of 139.93 GHz. The polarized sample (3 - 4 hrs) was dissolved and mixed with neutralization media (0.72 M NaOH, 0.4 M Trizma and 0.1 g/L Na2EDTA), resulting in 6.5–7.0 mL of 80-mM [1-13C]KIC solution (pH 7.4–8.2).
Hyperpolarized 13C MRS study of F98 cell
HP 13C MR cell studies were performed on a clinical GE 3 T Discovery 750 W MR scanner (∅bore = 70 cm) and the GE SPINlab™. F98 cells were cultured as described above and collected (1.1 ± 0.1 × 108, n = 3) and washed with PBS twice. The cells suspended in 2 mL of DMEM media were placed in a FalconTM 50 mL centrifuge tube at 37 °C, and 2 mL of HP [1-13C]KIC was injected into the tube within 15–18 sec following the dissolution. Dynamic and time-average 13C spectra were acquired using a custom-made 13C transmit/receive surface coil (∅inner = 28 mm).
MR Imaging protocol
In vivo imaging studies were also carried out at the clinical GE 3 T MR scanner. For 1H imaging, a quadrature birdcage volume RF coil (∅inner = 80 mm) was used for both RF excitation and data acquisition. After locating the rat brain at the center of the birdcage coil using a 3-plane fast gradient-recalled echo (FGRE) sequence, high-resolution axial brain images were acquired using a dual-echo T2-weighted fast spin echo (FSE; echo times [TEs] = 11.3 msec/64.0 msec, TR = 5,000 msec, 7–13 slices, field of view [FOV] = 96 × 96 mm2, matrix size = 256 × 192, slice thickness = 2 mm, echo train length = 8). B0 field inhomogeneity over the brain region was minimized using a single voxel point-resolved spectroscopy (PRESS) sequence by adjusting linear shim currents. For 13C MRSI, a 2D spiral chemical shift imaging pulse sequence described in previous study29 using the 13C surface coil with a variable flip angle scheme46, \({\theta }_{n}={\tan }^{-1}(\frac{1}{\sqrt{4-n}}),\,n=1,\,\ldots ,\,4\), was used to acquire single time-point metabolite maps of rat brain (four spatial interleaves of spiral k-space readout, spectral bandwidth = 210.8 Hz, #echoes = 64, FOV = 50 × 50 mm2, matrix size = 16 × 16, slice thickness = 7.7 mm, acquisition time = 1.5 sec) following an intravenous injection of 80-mM HP [1-13C]KIC (injection-to-scan time = 25 sec, 1 mmol/kg body weight up to 4.0 mL). Finally, a CE T1-weighted spin echo (SE) images were acquired at the end of each imaging session (TE = 12 msec, TR = 700 msec, 96 × 96 mm2, 7–13 slices, matrix size = 256 × 192, slice thickness = 2 mm).
Image reconstruction
The HP 13C MRSI were reconstructed for individual peak similarly as the described in previous study30. After a 5-Hz Gaussian line broadening and 4-fold zero-fillings along the time domain, an inverse FFT was applied to the raw data. Prior to a spatial apodization (hanning function) and a zero-filling (4x) along each spatial domain, chemical shift artifact was removed by correcting the off-resonance and aliasing pattern of each peak (spectral tomosynthesis), followed by a gridding onto the Cartesian coordinate and a spatial 2D inverse FFT.
Metabolic maps of HP [1-13C]KIC, [1-13C]leucine, and H13CO3− were produced by integrating the signal around each peak in absorption mode. The metabolic images were normalized by the maximum signal. Regions of interest (ROIs) of tumor and NAB were drawn manually to calculate the signal intensities for the metabolites. All the 13C data were reconstructed and analyzed using MATLAB (Mathworks, Natick MA, USA).
Infusion of [U-13C]leucine
13C isotope tracer infusion studies were performed as previously descried4. After 15–18 days from the tumor implantation, rats were fasted (n = 3) for 15 hrs prior to leucine infusion (U-13C; 99%, Sigma-Aldrich Isotec). After a tail vein catheterization under anesthesia (<10 min), all the rats stayed awake during the infusion. Each infusion started with a bolus injection of [U-13C]leucine (0.25 mg/g per body weight in 4 mL saline solution over 1 min), continued by a slow infusion (0.0069 mg/g of body weight/min in saline solution at 4.8 mL/hr for 150 min). Following the completion of the infusion, each animal was fully anesthetized and 4–6 mL of blood sample was collected by cardiac puncture. After decapitation, the whole brain was harvested and sliced to 2-mm coronal sections using brain matrices (Ted Pella, Inc., Redding CA, USA). Tumor (112.2 ± 24 mg) and contralateral NAB tissues (92.0 ± 10.8 mg) were rapidly collected from the brain slices, immediately frozen by liquid nitrogen and stored at −80 °C. The quantification of 13C fractional enrichment on glutamate methods are in supporting material.
Statistical analysis
Statistical significance between tumor and contralateral NAB was evaluated by a paired t-test (α = 0.05, one-tailed) using GraphPad Prism version 7.0 (GraphPad Software, Inc., La Jolla CA, USA). Unpaired t-tests were used for comparison between tumor-bearing rats and healthy controls. All the data were presented as mean ± standard deviation.
References
Kurhanewicz, J. et al. Analysis of Cancer Metabolism by Imaging Hyperpolarized Nuclei: Prospects for Translation to Clinical Research. Neoplasia 13, 81–97 (2011).
Warburg, O. über den Stoffwechsel der Carcinomzelle. Klinische Wochenschrift 4, 534–536 (1925).
Hosios, A. M. et al. Amino Acids Rather than Glucose Account for the Majority of Cell Mass in Proliferating Mammalian Cells. Developmental cell 36, 540–549 (2016).
Mashimo, T. et al. Acetate Is a Bioenergetic Substrate for Human Glioblastoma and Brain Metastases. Cell 159, 1603–1614 (2014).
Comerford, S. A. et al. Acetate dependence of tumors. Cell 159, 1591–1602 (2014).
Tonjes, M. et al. BCAT1 promotes cell proliferation through amino acid catabolism in gliomas carrying wild-type IDH1. Nat Med 19, 901–908 (2013).
Zhou, W. et al. Over-expression of BCAT1, a c-Myc target gene, induces cell proliferation, migration and invasion in nasopharyngeal carcinoma. Mol Cancer 12, 53 (2013).
Hattori, A. et al. Cancer progression by reprogrammed BCAA metabolism in myeloid leukaemia. Nature 545, 500–504 (2017).
Ananieva, E. A. & Wilkinson, A. C. Branched-chain amino acid metabolism in cancer. Current opinion in clinical nutrition and metabolic care 21, 64–70 (2018).
Panosyan, E. H., Lin, H. J., Koster, J. & Lasky, J. L. In search of druggable targets for GBM amino acid metabolism. BMC Cancer 17, 162 (2017).
Zhang, L. & Han, J. Branched-chain amino acid transaminase 1 (BCAT1) promotes the growth of breast cancer cells through improving mTOR-mediated mitochondrial biogenesis and function. Biochemical and Biophysical Research Communications 486, 224–231 (2017).
Zheng, Y. H. et al. BCAT1, a key prognostic predictor of hepatocellular carcinoma, promotes cell proliferation and induces chemoresistance to cisplatin. Liver International 36, 1836–1847 (2016).
Dey, P. et al. Genomic deletion of malic enzyme 2 confers collateral lethality in pancreatic cancer. Nature 542, 119–123 (2017).
Mayers, J. R. et al. Tissue-of-origin Dictates Branched-Chain Amino Acid Metabolism in Mutant Kras-driven Cancers. Science 353, 1161–1165 (2016).
Hutson, S. M., Sweatt, A. J. & LaNoue, K. F. Branched-Chain Amino Acid Metabolism: Implications for Establishing Safe Intakes. The Journal of nutrition 135, 1557S–1564S (2005).
Yudkoff, M. et al. Brain Amino Acid Requirements and Toxicity: The Example of Leucine. The Journal of nutrition 135, 1531S–1538S (2005).
Bixel, M., Shimomura, Y., Hutson, S. & Hamprecht, B. Distribution of key enzymes of branched-chain amino acid metabolism in glial and neuronal cells in culture. The journal of histochemistry and cytochemistry: official journal of the Histochemistry Society 49, 407–418 (2001).
Keen, R. E., Nissenson, C. H. & Barrio, J. R. Analysis of Femtomole Concentrations of α-Ketoisocaproic Acid in Brain Tissue by Precolumn Fluorescence Derivatization with 4,5-Dimethoxy-1,2-diaminobenzene. Analytical Biochemistry 213, 23–28 (1993).
Kim, D. K. et al. System L-amino acid transporters are differently expressed in rat astrocyte and C6 glioma cells. Neuroscience research 50, 437–446 (2004).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Ardenkjær-Larsen, J. H. et al. Increase in signal-to-noise ratio of >10,000 times in liquid-state NMR. Proceedings of the National Academy of Sciences 100, 10158–10163 (2003).
Golman, K., Zandt, R. I. T., Lerche, M., Pehrson, R. & Ardenkjaer-Larsen, J. H. Metabolic Imaging by Hyperpolarized 13C Magnetic Resonance Imaging for In vivo Tumor Diagnosis. Cancer Research 66, 10855–10860 (2006).
Brindle, K. M., Bohndiek, S. E., Gallagher, F. A. & Kettunen, M. I. Tumor imaging using hyperpolarized 13C magnetic resonance spectroscopy. Magnetic Resonance in Medicine 66, 505–519 (2011).
Butt, S. A. et al. Imaging cerebral 2-ketoisocaproate metabolism with hyperpolarized (13)C magnetic resonance spectroscopic imaging. Journal of cerebral blood flow and metabolism: official journal of the International Society of Cerebral Blood Flow and Metabolism 32, 1508–1514 (2012).
Billingsley, K. L. et al. The Feasibility of Assessing Branched-Chain Amino Acid Metabolism in Cellular Models of Prostate Cancer with Hyperpolarized [1-(13)C]-Ketoisocaproate. Magnetic resonance imaging 32, 791–795 (2014).
Karlsson, M. et al. Imaging of branched chain amino acid metabolism in tumors with hyperpolarized 13C ketoisocaproate. International journal of cancer 127, 729–736 (2010).
Barth, R. F. & Kaur, B. Rat brain tumor models in experimental neuro-oncology: the C6, 9L, T9, RG2, F98, BT4C, RT-2 and CNS-1 gliomas. Journal of Neuro-Oncology 94, 299–312 (2009).
Mathieu, D., Lecomte, R., Tsanaclis, A. M., Larouche, A. & Fortin, D. Standardization and detailed characterization of the syngeneic Fischer/F98 glioma model. The Canadian journal of neurological sciences. Le journal canadien des sciences neurologiques 34, 296–306 (2007).
Park, J. M. et al. Metabolic response of glioma to dichloroacetate measured in vivo by hyperpolarized (13)C magnetic resonance spectroscopic imaging. Neuro Oncol 15, 433–441 (2013).
Mayer, D., Levin, Y. S., Hurd, R. E., Glover, G. H. & Spielman, D. M. Fast metabolic imaging of systems with sparse spectra: Application for hyperpolarized 13C imaging. Magnetic Resonance in Medicine 56, 932–937 (2006).
Steele, R. D. Blood-brain barrier transport of the alpha-keto acid analogs of amino acids. Federation proceedings 45, 2060–2064 (1986).
Vannucci, S. J. & Simpson, I. A. Developmental switch in brain nutrient transporter expression in the rat. American journal of physiology. Endocrinology and metabolism 285, E1127–1134 (2003).
Day, S. E. et al. Detecting tumor response to treatment using hyperpolarized 13C magnetic resonance imaging and spectroscopy. Nat Med 13, 1382–1387 (2007).
Suryawan, A. et al. A molecular model of human branched-chain amino acid metabolism. The American Journal of Clinical Nutrition 68, 72–81 (1998).
Shennan, D. B., Thomson, J., Gow, I. F., Travers, M. T. & Barber, M. C. L-leucine transport in human breast cancer cells (MCF-7 and MDA-MB-231): kinetics, regulation by estrogen and molecular identity of the transporter. Biochimica et biophysica acta 1664, 206–216 (2004).
Holecek, M., Siman, P., Vodenicarovova, M. & Kandar, R. Alterations in protein and amino acid metabolism in rats fed a branched-chain amino acid- or leucine-enriched diet during postprandial and postabsorptive states. Nutrition & metabolism 13, 12 (2016).
Schadewaldt, P. & Adelmeyer, F. Coupled Enzymatic Assay for Estimation of Branched-Chainl-Amino Acid Aminotransferase Activity with 2-Oxo Acid Substrates. Analytical Biochemistry 238, 65–71 (1996).
Chuang, J. L. & Chuang, D. T. Diagnosis and mutational analysis of maple syrup urine disease using cell cultures. In Methods in Enzymology, Vol. 324, 413–423 (Academic Press, 2000).
Islam, M. M. et al. A novel branched-chain amino acid metabolon. Protein-protein interactions in a supramolecular complex. The Journal of biological chemistry 282, 11893–11903 (2007).
Zong, H., Verhaak, R. G. W. & Canoll, P. The cellular origin for malignant glioma and prospects for clinical advancements. Expert Review of Molecular Diagnostics 12, 383–394 (2012).
Hu, J. et al. Heterogeneity of tumor-induced gene expression changes in the human metabolic network. Nature Biotechnology 31, 522 (2013).
Castellano, S., Casarosa, S., Sweatt, A. J., Hutson, S. M. & Bozzi, Y. Expression of cytosolic branched chain aminotransferase (BCATc) mRNA in the developing mouse brain. Gene Expression Patterns 7, 485–490 (2007).
Garcia-Espinosa, M. A., Wallin, R., Hutson, S. M. & Sweatt, A. J. Widespread neuronal expression of branched-chain aminotransferase in the CNS: implications for leucine/glutamate metabolism and for signaling by amino acids. Journal of neurochemistry 100, 1458–1468 (2007).
Huse, J. T., Phillips, H. S. & Brennan, C. W. Molecular subclassification of diffuse gliomas: seeing order in the chaos. Glia 59, 1190–1199 (2011).
Sharma, B., Lawrence, D. W. & Hutchison, M. G. Branched Chain Amino Acids (BCAAs) and Traumatic Brain Injury: A Systematic Review. The Journal of head trauma rehabilitation 33, 33–45 (2018).
Zhao, L. et al. Gradient-Echo Imaging Considerations for Hyperpolarized 129Xe MR. Journal of magnetic resonance. Series B 113, 179–183 (1996).
Acknowledgements
We thank to Ian Corbin for sharing the F98 cells. We acknowledge the funding support: National Institute of Health (P41 EB015908, S10 OD018468, R37 HL034557, R00 CA168746, R01 DK62306); Cancer Prevention & Research Institute of Texas (RP140021-P2, RR140036); UT Southwestern Mobility Foundation Center; The Texas Institute of Brain Injury and Repair; The Welch Foundation (I-1903, I-1286, AT-584); Susan G Komen® (CCR16376227).
Author information
Authors and Affiliations
Contributions
E.H.S. and J.M.P. designed the research; E.H.S., E.P.H. and J.M.P. performed the hyperpolarized 13C imaging experiments; E.H.S. performed F98 cell culture; E.P.H. was responsible of tumor implantations and animal care; E.H.S. and E.P.H. performed steady-state infusion and tissue extractions; E.H.S., A.D.S. and J.M.P. analyzed 13C NMR spectra; E.H.S. analyzed tissue histology; E.H.S., R.M.W., and D.T.C. performed enzyme assays; B.Z. and W.L. performed Western blot analysis; E.H.S., R.M.W., D.T.C., W.L., A.D.S. and J.M.P. contributed on writing the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Suh, E.H., Hackett, E.P., Wynn, R.M. et al. In vivo assessment of increased oxidation of branched-chain amino acids in glioblastoma. Sci Rep 9, 340 (2019). https://doi.org/10.1038/s41598-018-37390-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-37390-0
- Springer Nature Limited
This article is cited by
-
CHIP-mediated ubiquitin degradation of BCAT1 regulates glioma cell proliferation and temozolomide sensitivity
Cell Death & Disease (2024)
-
Tissue and plasma free amino acid detection by LC-MS/MS method in high grade glioma patients
Journal of Neuro-Oncology (2023)
-
Pancreatic cancer: branched-chain amino acids as putative key metabolic regulators?
Cancer and Metastasis Reviews (2021)
-
Plasma amino acids indicate glioblastoma with ATRX loss
Amino Acids (2021)
-
In vivo detection of γ-glutamyl-transferase up-regulation in glioma using hyperpolarized γ-glutamyl-[1-13C]glycine
Scientific Reports (2020)