Abstract
The differences in artificial and natural selection have been some of the factors contributing to phenotypic diversity between Chinese and western pigs. Here, 830 individuals from western and Chinese pig breeds were genotyped using the reduced-representation genotyping method. First, we identified the selection signatures for different pig breeds. By comparing Chinese pigs and western pigs along the first principal component, the growth gene IGF1R; the immune genes IL1R1, IL1RL1, DUSP10, RAC3 and SWAP70; the meat quality-related gene SNORA50 and the olfactory gene OR1F1 were identified as candidate differentiated targets. Further, along a principal component separating Pudong White pigs from others, a potential causal gene for coat colour (EDNRB) was discovered. In addition, the divergent signatures evaluated by Fst within Chinese pig breeds found genes associated with the phenotypic features of coat colour, meat quality and feed efficiency among these indigenous pigs. Second, admixture and genomic introgression analysis were performed. Shan pigs have introgressed genes from Berkshire, Yorkshire and Hongdenglong pigs. The results of introgression mapping showed that this introgression conferred adaption to the local environment and coat colour of Chinese pigs and the superior productivity of western pigs.
Similar content being viewed by others
Introduction
Pigs were independently domesticated in Europe and China approximately 9000 years ago1,2,3,4. Since then, various pig breeds have been subjected to different forces of natural and artificial selection, which have been some of the contributory factors to the distinct phenotypes of different pig breeds5. Chinese pig breeds are famous for high prolificacy6, good adaptability to local environment7, high resistance to disease8,9,10 and desirable meat quality11,12. However, there is still variation in these characteristics among different Chinese pig breeds. Compared with Chinese local breeds, European pig breeds are renowned for their fast growth rate13, high feed efficiency14 and superior meat yield15.
Therefore, it is possible to use selection signature detection methods to elucidate the genetic background of the distinct phenotypes of pig breeds, which were influenced by different selection pressures. Moreover, the availability of genomic data facilitates the identification of the genomic regions affecting the specific characteristics among different pig breeds. For instance, by comparing the genomes of Tibetan pigs with low-land pigs, belted and non-belted pigs, Ai et al.16 discovered ADAMTS12, SIM1 and NOS1 as candidate genes contributing to high-altitude adaption and EDNRB as a gene affecting coat colour16. Based on a comparison between Chinese and European pigs, Yang et al.17 found that the JAK2 gene was associated with immune response in Chinese pigs and that the IGF1R gene was associated with growth in European pigs17.
In addition, there has been gene flow from Chinese pigs to European pig breeds since the nineteenth century18 aiming to improve the productivity of local breeds. A study has shown that Asian pig haplotypes have been introgressed into European pig breeds to improve traits of commercial interest19. For example, reproduction20 and carcass and meat quality traits21 in European pigs have been improved by the introduction of Asian haplotypes. Conversely, European haplotypes might also have been introgressed into Chinese pig breeds. To obtain the superior characteristics of western pigs, human-mediated hybridization and introgression were performed to improve the productivity and environmental adaptability of Chinese pig breeds. This is similar to the formation of the hybrid nature of Chinese Sutai pigs cultivated from Chinese Erhualian and Duroc pigs16,22. However, except the hybrid breed, Sutai, introgression from western pigs into other Asian pigs has not been reported so far.
In this study, we collected samples from western pig breeds and different Chinese pig breeds with distinct phenotypic characteristics in the Yangtze River Delta (YRD) area in China. By comparing the genomes between Chinese and western pig breeds and among Chinese pig breeds, we were able to identify genes associated with the distinct phenotypes in these pig breeds. Because of the history and location of the YRD, it plays an important role in international communication, as well as in gene flow. Some Chinese pig breeds in this region might have gene flow from western breeds.
Therefore, the objective of the study is to (1) identify genes associated with the distinct phenotypic characteristics between western and Chinese pig breeds and within different Chinese pig breeds and (2) characterize the introgression from western into Chinese pig breeds.
Methods
Ethics statement
All experimental procedures were approved by the Institutional Animal Care and Use Committee of Shanghai Jiao Tong University, and all methods involving pigs were in accordance with the agreement of the Institutional Animal Care and Use Committee of Shanghai Jiao Tong University (contract no. 2011-0033).
Populations and Data
A total of 830 pigs were used in this study, including 156 western pigs, i.e., Duroc (D), Landrace (L), Yorkshire (Y), Berkshire (B) and Pietrain (P) pig breeds, and 674 Chinese indigenous pigs within the YRD region (covering Jiangsu and Zhejiang provinces and Shanghai municipality). Figure 1 shows the location where the samples of Chinese pigs were collected. Table 1 lists detailed information of the sampled pig breeds in this study, including breed name, abbreviations and sample size. Most of the reduced-representation genotyping data has been described in previous studies23,24,25,26, except for that from the Pietrain and Berkshire populations.
Sample location of Chinese indigenous pig populations within the Yangtze River Delta used in this study. For abbreviations, see Table 1.
The individuals of the Pietrain and Berkshire populations were genotyped according to the protocol of GGRS (genotyping by genome reducing and sequencing)27. In the process of GGRS, genomic DNA was extracted from ear tissue using a commercial kit (Lifefeng Biotech Co., Ltd, Shanghai, China). After the DNA samples were digested with AvaII enzyme and ligated with a unique adapter barcode, the samples were pooled and enriched to construct a sequencing library. DNA libraries (fragment lengths ranging from 300 to 400 bp) were sequenced using an Illumina HiSeq4000 platform according to the manufacturer’s protocol. Quality control of sequences was performed using NGS QC Toolkit28 v2.3, and the parameters were set according to the report from Chen et al.27. Sequencing reads were aligned to the Sscrofa10.2 pig reference genome using BWA29.
The BAM files from alignments were used to call SNPs. To improve the precision of SNP detection, SNP calling was performed by both SAMtools30 v0.1.9 (set 1) and GATK UnifiedGenotyper31 with “hard filters” (QD > 20.0 && FS < 60.0 && MQRankSum > −12.5 && ReadPosRankSum > −8.0) by the VariantFiltration tool (set 2) simultaneously. The SNPs found in both set 1 and set 2 were retained for further steps. Beagle32 v4.1 was utilized to impute the missing genotypes in the present study with default parameters. After imputation, SNPs were filtered out if their minor allele frequencies (MAFs) were less than 0.05. Non-autosomal SNPs were also discarded because the demographic patterns of sex chromosomes are different, which may cause distortion in the subsequent analysis33. PLINK34 v1.07 was used to filter the SNPs with extreme deviations (p-value ≤ 1 × 10−6) from Hardy-Weinberg equilibrium proportions for each population, and the union set of SNPs that failed to pass the test within at least one population were also discarded. In total, 129,882 high-confidence SNPs were retained for further analysis. Generally, these SNPs were roughly distributed uniformly across the genome, which can represent the information of the whole genome (Supplementary Fig. S1).
Population structure and genetic diversity
To illustrate the population structure and evaluate the genetic diversity within and between these populations, the following steps were performed: (1) principal component analysis (PCA) was conducted using SMARTPCA integrated in EIGENSOFT35 v6.1.4, which transformed the genetic variation into continuous axes (principle components) by singular value decomposition. (2) A total of 41,118 SNPs, which discarded ones that were in LD (linkage disequilibrium) larger than 0.5 across these populations (command: PLINK–indep 50 5 2), were kept for population structure analysis using ADMIXTURE36 v1.3.0. The number of ancestral clusters (K) was set from 2 to 40, and five-fold cross-validation was run to determine the K value with the lowest cross-validation error. The result was shown by DISTRUCT37 v1.1. (3) The allelic richness was calculated by ADZE38 v1.0, which can correct for unequal sample size using a rarefaction procedure39,40,41.
Effective population size
The historical effective population size (Ne) was estimated based on the SNP data used in admixture analysis by the software SNeP42 v1.1, which can estimate Ne at different t generations based on LD between SNPs with the distance of c, where \(t={(2c)}^{-1}\), and c is the distance measured in Morgan43 (assuming 100 Mb = 1Morgan). Some options were also used for SNeP software: (1) sample size correction for unphased genotypes; (2) correction to account for mutation; (3) Sved & Feldman’s recombination rate modifier44.
EigenGWAS analysis
Inspired by the PCA result that the first principal component (PC1) clearly separated the Chinese and western pigs, a method called EigenGWAS45, which considered PC1 as the phenotype, was used to identify loci associated with the pattern of PC1. Additionally, as the result of the third principal component (PC3) showed, the Pudong White pig population (PD) was obviously separated from the other populations on this axis. Given that PD is a unique Chinese indigenous pig breed covered by a wholly white coat, PC3 was also considered as a phenotype for EigenGWAS analysis. This method corrected for genetic drift by using a genomic inflation factor46. After this correction, the p-values of SNPs were further corrected by Bonferroni correction, and the cut-off was 0.05/129882. Moreover, in order to validate whether the EigenGWAS analysis with PC1 as the phenotype could identify differentiation between Chinese and western pigs, Weir and Cockerham’s Fst47 between them was calculated using VCFtools48. Finally, the Spearman’s rank correlation coefficient between Weir and Cockerham’s Fst and the negative logarithm of p-values of the EigenGWAS with PC1 as the phenotype was calculated in R49.
Identification of selection signatures among Chinese indigenous pigs by F st
To identify highly differentiated genomic regions among Chinese indigenous pig breeds, an Fst outlier approach implemented in the R package OutFLANK50 was used to find significant differentiation loci among these pig breeds. Firstly, the near-independent SNPs were identified using R package bigsnpr51 according to the tutorial of OutFLANK. Based on this near-independent SNP set, OutFLANK fit a chi-square distribution to the core distribution of Fst (that is, trimming the top and bottom 5%) to estimate the mean and degree of freedom, so that this core distribution would not be affected by strong balancing and diversifying selection. Then the p-values of all the SNPs with heterozygosity greater than 0.1 were calculated based on the core distribution of Fst. OutFLANK adjusted multiple p-values to q-values52, and the threshold of 0.01 was used53.
Three-population test
To investigate the statistical significance of admixture among these pig populations, TreeMix54 software was used to perform the three-population (f3) test55. In the f3 test with the form of f3 (A; B, C), an extreme negative f3 statistic indicates that significant gene flow to population A from populations B and C exists. All 24 populations were included in the f3 test, and this would generate \((\begin{array}{c}24\\ 3\end{array})=6072\) different combinations. The SNP set that had been LD filtered for the ADMIXTURE analysis was used in this step. A block jackknife56 implemented in TreeMix with a window of 200 SNPs that excluded the dependence between different windows was used to calculate the standard deviation of the test. Then, the Z scores were calculated, and combinations that had Z scores less than −2 were regarded as significant.
Mapping of admixture along the genome using PCAdmix
The results of the f3 test only showed that the Shan pig (SZ) was significantly admixed by Hongdenglong (HD), B and Y. PCAdmix57 v1.0 was used to identify probable significant admixed fragments due to genomic introgression from the three ancestral populations into SZ. Based on PCA, PCAdmix can infer the ancestry of admixed genomes from ancestral individuals using a sliding window along the genome. Then, the posterior probability of ancestry affiliation for each window can be determined by a hidden Markov model. PCAdmix requires phased genotypes, therefore the genotype data used in the ADMIXTURE analysis was phased by fastPHASE58 v1.2 with default parameters. According to Barbato et al.59, the window size was set to be a fixed value of 5 SNPs due to no available linkage map and a low density of markers59. PCAdmix inferred the posterior probability (PP) of ancestry from the HD, B and Y populations for each individual haploid genome of SZ for each window. Then, the PP for each reference population was added up across all the haploid genomes of SZ to calculate the scores of affiliation of different ancestry for each window. The windows with the top 1% of scores for each ancestry affiliation were selected to be candidate genomic introgression regions, and genes within these regions were extracted using the biomaRt60 package.
Functional annotation of candidate genes
ANNOVAR61 was used to identify candidate SusScr3 Ensembl genes near the significant SNPs based on EigenGWAS and Fst (within 150 kb). Functional gene set enrichment analysis was then performed for these gene sets and candidate genes within the top genomic introgression regions from the PCAdmix analysis. The R package org.Ss.eg.db was used to annotate the pig genes. Enriched Gene Ontology62 (GO) terms were then identified using the R package GOstats63. Then, the p-value of each GO term was calculated by GOstats using a hypergeometric test. The results with a p-value ≤ 0.05 were reported to identify potential biological processes influenced by these genes.
Results and Discussion
Population structure and genetic diversity
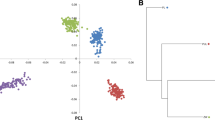
An overview of the relationships among these populations is presented in Fig. 2. PC1, which accounted for 13.0% of the total variance, separated the Chinese and western pigs. Except for Duroc pigs, all individuals from the western pig breeds were clustered together. This is consistent with the breed’s history, since Duroc pigs were developed in the United States, while other western pig breeds were originated from the European continent. Western pig populations were clustered more compactly than those from Chinese pig populations. PC2, which explained 4.0% of the total variance, separated most of Chinese indigenous pig breeds (Fig. 2). The first two components together still could not separate some Chinese pig populations. For example, the points representing the PD pig breed, overlapped with some points of the pig breeds from Jiangsu Province (points in the shape of cross). However, along the third component (PC3), PD was clearly separated from the other pig breeds (Supplementary Fig. S2). The allelic richness results (Table 1) reflected that more genetic diversity existed within Chinese pig populations. Chinese pig populations had more allelic richness compared with western pig populations, except for those in Dongchuan (DC), Fengjing (FJ) and Lanxi (LX), with sample sizes of 16 or less (Table 1). This is due to the tendency for populations with lower sample sizes to have fewer distinct alleles, although ADZE can correct for sample size.
Scatter plot for the first and second principal components. The x-axis represents the first principal component, and the y-axis represents the second principal component. Points of individuals from the same region (Table 1) are the same shape but different colours.
The estimated K for the ADMIXTURE analysis with the lowest cross-validation error was 25, nearly the same as the number of actual populations in this study. When two ancestors were assumed, Chinese indigenous pigs and western pigs were clearly distinguished (Fig. 3), but some Chinese pig populations contained some genetic ancestry that was similar to western populations, especially for SZ, which was always the most admixed population over different K values. Compared with a previous study22 which exhibited approximately 20% Chinese admixture in most European breeds (with K = 2) using 60 K porcine SNP array, no such much admixture could be observed in this study. This might be due to ascertainment bias resulting from the development of the SNP array mainly based on the polymorphisms distributed in western pig breeds, which would overestimate the shared ancestry between Chinese and western pig breeds. When K was 25, all of the populations roughly had their own ancestry, except for Small Meishan (SMS), Chunan (CA) and SZ. SMS and CA contained new ancestry, which might not be part of the populations included in this study. More pig populations are needed to identify the distinct genetic components not shared with other breeds in the study. Compared with Chinese pigs, the extents of admixture within western pig populations across different K values were low and stable.
Effective population size
The estimation of Ne of each pig breed across the generations is shown in Supplementary Fig. S3. The past Ne was reflected by LD over shorter recombination distances, and the longer distances provided recent Ne43. From 900 generations to approximately 50 generations ago, all the breeds exhibited a decrease in Ne estimates over time. However, between 900 and 1000 generations ago, there were some obvious inflection points in several lines. The nearest anti-climax point indicated the nearest starting point of artificial selection, which caused the bottleneck in the population64. In general, the Ne of western pig breeds was smaller than that of Chinese pig populations, which was due to the higher LD of western pig breeds65. Admixture is a potential confounding factor for the estimation of Ne that can cause bias66. Therefore, among Chinese indigenous pigs, the populations with a low extent of admixture tend to have a smaller Ne67. The Ne estimates did not get stable even in recent generations. This sort of trend could also be found in another study for Landrace and Yorkshire pigs68. For western pigs, the reason of this phenomenon might be the ongoing strong selection on production traits5, whereas for Chinese local pig breeds, it might be due to the inbreeding caused by small population size69.
EigenGWAS for PC1 and PC3
To find loci that were related to the pattern of PC1, which separated the western and Chinese pigs, EigenGWAS with PC1 as the phenotype was performed. Further, this association study was also performed to explain why PD was separated from the other populations on PC3. The Manhattan plots for PC1 and PC3 are shown in Fig. 4A,B, respectively. There were as many as 353 and 414 significant SNPs associated with PC1 and PC3, respectively. EigenGWAS aims to find ancestry informative markers (AIMs), which can be found in huge quantities if the genetic backgrounds of populations are very different. The Spearman’s correlation coefficient between Weir and Cockerham’s Fst and the negative logarithm of p-values of the EigenGWAS with PC1 as the phenotype was 0.969. Some studies have suggested that individual-level eigenvectors are measures of population differentiation reflecting Fst among subpopulations35,70,71. Therefore, the high value of this correlation coefficient validated that the EigenGWAS with PC1 as the phenotype could reflect the differentiation between Chinese and western pigs. In this study, it is difficult to determine whether these differentiated signals were formed in the pre-domestication or during the divergent post-domestication selection. However, both of these different kinds of significant signals could help explain the genetic background of distinct phenotypes between Chinese and western pigs. In addition, the genomic inflation factors were 100.81 and 17.75 for the EigenGWAS with PC1 and PC3 as the phenotype, respectively. According to the original paper on EigenGWAS45, the genomic inflation factors are highly correlated to eigenvalues, which were 107.58 and 24.26 for PC1 and PC3, respectively. A large eigenvalue indicates underlying population structure. Therefore, correction for genomic inflation factors will filter out signals due to population stratification, allowing loci under selection to be identified45.
There were 286 Ensembl genes located near the significant SNPs of the EigenGWAS for PC1 (Supplementary Table S1). The first and second significant SNPs on chromosome 1 were located near and within the IGF1R gene. This gene plays an important role in pig production traits, such as post-natal growth72 and carcass and meat content73. In addition, a previous study verified that different alleles of the gene IGF1R are highly correlated with pig performance based on litter size74, indicating that pleiotropy of the IGF1R gene can be a potential explanation of the genetic relationship between traits of production and reproduction75. Therefore, the differentiated variants in the IGF1R gene may be considered a potential quantitative trait locus (QTL), which can account for the phenotypic differences of growth and reproduction between Chinese and western pig breeds. Besides, Chinese pigs are well adapted to their local environments7 and are known for their desirable meat quality11,12. These phenotypic characteristics could be explained by genes near significant signals on different chromosomes (Fig. 4A). Among them was the olfactory gene, OR1F1. A sharp sense of smell is very important for pigs to improve their appetite for roughage feed, and may also help increase their preference for specific food to produce human-desired meat under captivity76. In addition, the gene SNORA50, located near the top significant SNP on chromosome 8, was identified as a candidate gene for meat quality in a previous GWAS study12. Two immunity-related genes, IL1RL1 and IL1R1, were identified on chromosome 3. IL1R1 is a mediator gene involved in many cytokine-induced immune and inflammatory responses. IL1RL1 plays an important role in some human diseases such as rheumatoid arthritis and asthma77. In human, asthma is a counterpart disease of swine mycoplasmal pneumonia. Chinese indigenous pigs, especially pigs in the YRD, are very sensitive to Mycoplasma hyopneumoniae78,79,80. Intriguingly, the most significant GO term (Supplementary Table S2) was “regulation of respiratory burst (GO:0060263)”, which might be related to Mycoplasmal pneumonia. In addition to respiration-related terms, there were also some mast cell-related GO terms at the top of the list, such as “regulation of mast cell chemotaxis (GO:0060753)”, “mast cell chemotaxis (GO:0002551)” and “mast cell migration (GO:0097531)”. Mast cells are found to participate in the early recognition of pathogens, which plays an important role in immunity81. These mast cell- or respiratory-related terms were all enriched by the genes DUSP10, RAC3 and SWAP70, together with IL1R1 and IL1RL1, which might explain the high resistance to disease in Chinese pigs8,9,10.
The appearance of the PD pig breed is very distinct from that of other Chinese indigenous pigs due to its wholly white coat. Chinese indigenous pigs are often black and sometimes belted or spotted, but never wholly white like PD pigs. Along PC3, PD pigs were separated from the other populations. Performing EigenGWAS with PC3 as the phenotype might therefore help to explain some particular characteristics of PD pigs, such as its uniqueness of coat colour. The coat colour-related gene EDNRB82 was identified near significant signals on chromosome 11 (Supplementary Table S3). In a previous study about Chinese raccoon dogs, a SNP in EDNRB gene was identified as the causal variant for the determinant of white colour in this animal83. Therefore, the identification of this gene might help account for the distinct coat colour of PD pigs. Except for some general biological processes, the enriched GO terms were mainly related to pigmentation (“pigmentation (GO:0043473)” and “pigment cell differentiation (GO:0050931)”), some behaviour- and cognition-related processes (“adult locomotory behavior (GO:0008344)”, “learning (GO:0007612)” and “cognition (GO:0050890)”) and nervous system-related functions (“nervous system process (GO:0050877)”, “neuron apoptotic process (GO:0051402)” and “neuron death (GO:0043524)”) (Supplementary Table S4). PD has long been suspected to be formed by admixture between Chinese and western pig breeds because of its white coat colour. The suspicion of its admixture origin could be overturned by the facts that it belonged to the Chinese indigenous pig cluster along the PC1 axis and that there was no evident admixture from western pigs when K = 25 in the ADMIXTURE analysis.
F st among Chinese indigenous pigs
The core distribution of Fst based on 42,746 near-independent SNPs were shown in Supplementary Fig. S4. The estimated mean of the fitted chi-square distribution was 0.23, and the degree of freedom was 14.24. The mean value was not high, indicating that there was not extreme differentiation among Chinese indigenous pigs. The Manhattan plot of q-values of 109, 451 SNPs with heterozygosity greater than 0.1 is shown in Fig. 5. After correcting for multiple-testing, there were 129 SNPs identified as significant (Fig. 5). Table 2 lists all the Fst candidate genes that have been verified to be related to pig traits by other studies.
Unlike the EigenGWAS of PC1, which could identify common features of Chinese pigs through the comparison with western pig breeds, Fst signal detection within Chinese pig breeds enabled us to find the differentiated features among these breeds. A total of 75 genes were identified near significant Fst signals (Supplementary Table S5). The first four significant SNPs were all located near the JPH3 gene. JPH3 was identified to be associated with boar taint by affecting skatole levels84. A previous study verified that some Chinese pigs, such as JH pigs, had a significantly lower level of skatole than Landrace pigs85. Boar taint can affect the flavour of pork, which is important in Chinese cuisine. In addition, in another study, this gene was also identified as a candidate target of meat quality traits86. Chinese indigenous pigs, such as FJ, JH and DC pigs in this study, are well known for their desirable meat quality and flavor; thus, the JPH3 gene might have undergone selection to improve meat quality by reducing skatole levels. The most significant signal on chromosome 4 was located in the ZFPM2 gene. This gene is important in the development of diaphragmatic hernia and a previous study has found it to be significantly associated with pig scrotal hernias87. The second significant SNP on chromosome 15 was near CNTNAP5 gene, which was identified as a candidate gene of pig vertebra number in a previous study88. Vertebra number is a trait associated with carcass and meat production. Chinese pigs perform well in the vertebra number trait, and western pigs have also benefited from Chinese pigs in this trait by introgression21. The first and second significant signals on chromosome 1 were both located in the HMCN2 gene, which is related to stimulus response in Chinese pig breeds76. As expected, given many pig breeds covered with different kinds of coloured coats, some pigmentation-related genes, such as KIT and EDNRB, were also identified89. In terms of GO enrichment analysis (Supplementary Table S6), the most significant terms were related to pigmentation, such as “pigmentation (GO:0043473)”, “melanocyte differentiation (GO:0030318)” and “pigment cell differentiation (GO:0050931)”. Some GO terms were related to behaviour, such as “locomotion (GO:004001)”. Chinese pigs often have low locomotion and low behavioural reactivity90. A previous study has also shown that Chinese pigs have become timid and tame due to selection pressure on behavioural traits during the long time of domestication76.
Three population test and PCAdmix analysis for SZ population
There were only two extreme Z scores for the f3 test: −2.23 and −4.62 for the combination of (SZ; Y, HD) and (SZ; B, HD), respectively. This result is consistent with the admixture of SZ (Fig. 3). The admixture might be deliberately human mediated to improve productivity and adaptability to the environment. Mapping the specific regions of admixture can help understand breeders’ agronomic interests as well as the direction of natural selection. Therefore, the PCAdmix analysis was performed to localize the potential regions of gene introgression. The top potential introgression regions along the SZ genome from each of three breeds are listed in Supplementary Table S7. Based on the GO enrichment results enriched by genes introgressed from Y (Supplementary Table S8), the top terms were mainly related to growth (“negative regulation of cell growth (GO:0030308)”, “negative regulation of growth GO:0045926”) and bone development (“BMP signaling pathway (GO:0030509)” and “regulation of ossification (GO:0030278)”). Several of the top GO processes enriched by genes introgressed from B (Supplementary Table S9) were related to growth and development, such as “heart development (GO:000750)”and “biomineral tissue development (GO:0031214)”, and some were related to response to external stimulus (“cellular response to corticosteroid stimulus (GO:0071384)”, “cellular response to glucocorticoid stimulus (GO:0071385)” and “sensory perception of pain (GO:0019233)”). The second significant GO term enriched by genes introgressed from HD (Supplementary Table S10) was “toll-like receptor 4 signaling pathway (GO:0034142)”, which is related to immunity. Like the coat colour-related findings mentioned above, another two melanin metabolism processes (“regulation of melanin biosynthetic process (GO:0048021)” and “positive regulation of melanin biosynthetic process (GO:0048023)”) were identified, and ASIP was found to be the causal gene. This gene was identified to be related to human pigmentation diversity91,92. Given these results, it can be summarized that the introgression regions from western pigs were mainly related to growth and development. On the other hand, the major introgression regions from Chinese pig breeds were related to immunity and pigmentation. Breeders in China preferred the high adaptability and black coat colour of Chinese pigs, while western pigs were chosen for their high productivity. Given the hypothesis that these introgressions were deliberately human mediated, the breeding goal of making good use of characteristics of Chinese and western pigs helped explain these results. Moreover, these results contributed to explaining the genetic basis of the phenotypic distinctions between Chinese and western pigs and within different Chinese pigs.
In this study, the genetic basis of phenotypic differences between Chinese and western pig breeds was studied from the viewpoint of selection signal detection. Numerous genes related to growth, immunity, reproduction and meat quality were identified as candidate differentiated genes, which might contribute to the distinct phenotypes of western and Chinese pigs. In addition, the coat colour-related gene EDNRB was identified as a candidate gene for the white colour of the PD pig breed. The significant divergent genetic signals among these Chinese pig populations were related to various economically important traits. Based on admixture and genomic introgression analysis, we observed that there was introgression from western pigs and other Chinese pigs into SZ pigs. The mapping of the introgression also helped to elucidate the genetic basis of phenotypic features, namely, that western pigs are good at production traits, while Chinese pigs do well in adaption to their environments.
Data Availability
All BAM data were deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA). 264 samples are available under the Bioproject number PRJNA436152. 128 samples are available under Bioproject number PRJNA281578. 438 samples are available under the Bioproject number PRJNA471328.
References
Giuffra, E. et al. The origin of the domestic pig: independent domestication and subsequent introgression. Genetics 154, 1785–1791 (2000).
Kijas, J. & Andersson, L. A phylogenetic study of the origin of the domestic pig estimated from the near-complete mtDNA genome. Journal of Molecular Evolution 52, 302–308 (2001).
Larson, G. et al. Worldwide phylogeography of wild boar reveals multiple centers of pig domestication. Science 307, 1618–1621 (2005).
Xiang, H. et al. Origin and dispersal of early domestic pigs in northern China. Scientific Reports 7, 5602 (2017).
Amaral, A. J. et al. Genome-wide footprints of pig domestication and selection revealed through massive parallel sequencing of pooled DNA. Plos One 6, e14782 (2011).
White, B. R., Barnes, J. & Wheeler, M. B. Advances in Swine in Biomedical Research: Volume 2 (ed. Mike E. Tumbleson & Lawrence B. Schook) 503–521 (Springer US, 1996).
Li, M. et al. Genomic analyses identify distinct patterns of selection in domesticated pigs and Tibetan wild boars. Nature Genetics 45, 1431 (2013).
Clapperton, M., Bishop, S. & Glass, E. Innate immune traits differ between Meishan and Large White pigs. Veterinary Immunology and Immunopathology 104, 131–144 (2005).
Duchet-Suchaux, M., Bertin, A. & Menanteau, P. Susceptibility of Chinese Meishan and European large white pigs to enterotoxigenic Escherichia coli strains bearing colonization factor K88, 987P, K99, or F41. American Journal of Veterinary Research 52, 40–44 (1991).
Ma, X., Zhang, X., Wang, L. & Liu, Z. Studies on difference of immune and production indexes between Songliao black pig and large white pig. China Animal Husbandry & Veterinary Medicine 38, 52–55 (2011).
Dai, F. et al. Developmental differences in carcass, meat quality and muscle fibre characteristics between the Landrace and a Chinese native pig. South African Journal of Animal Science 39, 267–273 (2009).
Ma, J. et al. Genome-wide association study of meat quality traits in a White Duroc × Erhualian F2 intercross and Chinese Sutai pigs. Plos One 8, e64047 (2013).
Wilkinson, S. et al. Signatures of diversifying selection in European pig breeds. PLOS Genetics 9, e1003453 (2013).
Wang, K. et al. Detection of Selection Signatures in Chinese Landrace and Yorkshire Pigs Based on Genotyping-by-SequencingData. Frontiers in Genetics 9, 119 (2018).
Rubin, C. J. et al. Strong signatures of selection in the domestic pig genome. Proceedings of the National Academy of Sciences 109, 19529–19536 (2012).
Ai, H., Huang, L. & Ren, J. Genetic diversity, linkage disequilibrium and selection signatures in Chinese and Western pigs revealed by genome-wide SNP markers. Plos One 8, e56001 (2013).
Yang, S., Li, X., Li, K., Fan, B. & Tang, Z. A genome-wide scan for signatures of selection in Chinese indigenous and commercial pig breeds. BMC Genetics 15, 7 (2014).
White, S. From globalized pig breeds to capitalist pigs: a study in animal cultures and evolutionary history. Environmental History 16, 94–120 (2011).
Bosse, M. et al. Artificial selection on introduced Asian haplotypes shaped the genetic architecture in European commercial pigs. Proc. R. Soc. B 282, 20152019 (2015).
Bosse, M. et al. Genomic analysis reveals selection for Asian genes in European pigs following human-mediated introgression. Nature Communications 5, 4392 (2014).
Yang, J. et al. Possible introgression of the VRTN mutation increasing vertebral number, carcass length and teat number from Chinese pigs into European pigs. Scientific Reports 6, 19240 (2016).
Chen, M. et al. Population admixture in Chinese and European Sus scrofa. Scientific Reports 7, 13178 (2017).
Li, Z. et al. Detection of selection signatures of population‐specific genomic regions selected during domestication process in Jinhua pigs. Animal Genetics 47, 672–681 (2016).
Wang, Z. et al. Genetic diversity and population structure of six Chinese indigenous pig breeds in the Taihu Lake region revealed by sequencing data. Animal Genetics 46, 697–701 (2015).
Xiao, Q., Zhang, Z., Sun, H., Wang, Q. & Pan, Y. Pudong White pig: a unique genetic resource disclosed by sequencing data. Animal 11, 1117–1124 (2017).
Xiao, Q. et al. Genetic variation and genetic structure of five Chinese indigenous pig populations in Jiangsu Province revealed by sequencing data. Animal Genetics 48, 596–599 (2017).
Chen, Q. et al. Genotyping by genome reducing and sequencing for outbred animals. Plos One 8, e67500 (2013).
Trivedi, U. H. et al. Quality control of next-generation sequencing data without a reference. Frontiers in Genetics 5, 111 (2014).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Research 20, 1297–1303 (2010).
Browning, B. L. & Browning, S. R. Genotype imputation with millions of reference samples. The American Journal of Human Genetics 98, 116–126 (2016).
Bosse, M. et al. Regions of homozygosity in the porcine genome: consequence of demography and the recombination landscape. PLOS Genetics 8, e1003100 (2012).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. The American Journal of Human Genetics 81, 559–575 (2007).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLOS Genetics 2, e190 (2006).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Research 19, 1655–1664 (2009).
Rosenberg, N. A. DISTRUCT: a program for the graphical display of population structure. Molecular Ecology Resources 4, 137–138 (2004).
Szpiech, Z. A., Jakobsson, M. & Rosenberg, N. A. ADZE: a rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24, 2498–2504 (2008).
Hurlbert, S. H. The Nonconcept of Species Diversity: A Critique and Alternative Parameters. Ecology 52, 577–586 (1971).
Petit, R. J., El Mousadik, A. & Pons, O. Identifying populations for conservation on the basis of genetic markers. Conservation Biology 12, 844–855 (1998).
Kalinowski, S. T. Counting alleles with rarefaction: private alleles and hierarchical sampling designs. Conservation Genetics 5, 539–543 (2004).
Barbato, M., Orozco-terWengel, P., Tapio, M. & Bruford, M. W. SNeP: a tool to estimate trends in recent effective population size trajectories using genome-wide SNP data. Frontiers in Genetics 6, 109 (2015).
Hayes, B. J., Visscher, P. M., McPartlan, H. C. & Goddard, M. E. Novel multilocus measure of linkage disequilibrium to estimate past effective population size. Genome Research 13, 635–643 (2003).
Sved, J. & Feldman, M. Correlation and probability methods for one and two loci. Theoretical Population Biology 4, 129–132 (1973).
Chen, G.-B., Lee, S. H., Zhu, Z.-X., Benyamin, B. & Robinson, M. R. EigenGWAS: finding loci under selection through genome-wide association studies of eigenvectors in structured populations. Heredity 117, 51–61 (2016).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Weir, B. S. & Cockerham, C. C. Estimating F‐statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Core Team, R. R. A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria, ISBN 3-900051-07-0. (2016).
Whitlock, M. C. & Lotterhos, K. E. Reliable detection of loci responsible for local adaptation: inference of a null model through trimming the distribution of F ST. The American Naturalist 186, S24–S36 (2015).
Privé, F., Aschard, H., Ziyatdinov, A. & Blum, M. G. Efficient analysis of large-scale genome-wide data with two R packages: bigstatsr and bigsnpr. Bioinformatics 1, 7 (2018).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proceedings of the National Academy of Sciences 100, 9440–9445 (2003).
Lotterhos, K. E. & Whitlock, M. C. Evaluation of demographic history and neutral parameterization on the performance of FST outlier tests. Molecular Ecology 23, 2178–2192 (2014).
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLOS Genetics 8, e1002967 (2012).
Reich, D., Thangaraj, K., Patterson, N., Price, A. L. & Singh, L. Reconstructing Indian population history. Nature 461, 489–494 (2009).
Kunsch, H. R. The Jackknife and the bootstrap for general stationary observations. The Annals of Statistics, 1217–1241 (1989).
Brisbin, A. et al. PCAdmix: principal components-based assignment of ancestry along each chromosome in individuals with admixed ancestry from two or more populations. Human Biology 84, 343–364 (2012).
Scheet, P. & Stephens, M. A fast and flexible statistical model for large-scale population genotype data: applications to inferring missing genotypes and haplotypic phase. The American Journal of Human Genetics 78, 629–644 (2006).
Barbato, M. et al. Genomic signatures of adaptive introgression from European mouflon into domestic sheep. Scientific Reports 7, 7623 (2017).
Durinck, S. et al. BioMart and Bioconductor: a powerful link between biological databases and microarray data analysis. Bioinformatics 21, 3439–3440 (2005).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Research 38, e164–e164 (2010).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nature Genetics 25, 25 (2000).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2006).
Orozco-terWengel, P. et al. Revisiting demographic processes in cattle with genome-wide population genetic analysis. Frontiers in Genetics 6, 195 (2015).
Amaral, A. J., Megens, H.-J., Crooijmans, R. P., Heuven, H. C. & Groenen, M. A. Linkage disequilibrium decay and haplotype block structure in the pig. Genetics 179, 569–579 (2008).
Orozco‐terWengel, P. A. & Bruford, M. W. Mixed signals from hybrid genomes. Molecular Ecology 23, 3941–3943 (2014).
Makina, S. O. et al. Extent of linkage disequilibrium and effective population size in four South African Sanga cattle breeds. Frontiers in Genetics 6, 337 (2015).
Uimari, P. & Tapio, M. Extent of linkage disequilibrium and effective population size in Finnish Landrace and Finnish Yorkshire pig breeds. Journal of Animal Science 89, 609–614 (2011).
Wang, X. et al. Genetic diversity, population structure and phylogenetic relationships of three indigenous pig breeds from Jiangxi Province, China, in a worldwide panel of pigs. Animal Genetics 49, 275–283 (2018).
Bryc, K., Bryc, W. & Silverstein, J. W. Separation of the largest eigenvalues in eigenanalysis of genotype data from discrete subpopulations. Theoretical Population Biology 89, 34–43 (2013).
McVean, G. A genealogical interpretation of principal components analysis. PLOS Genetics 5, e1000686 (2009).
Pierzchała, M. et al. Study of the differential transcription in liver of growth hormone receptor (GHR), insulin-like growth factors (IGF1, IGF2) and insulin-like growth factor receptor (IGF1R) genes at different postnatal developmental ages in pig breeds. Molecular Biology Reports 39, 3055–3066 (2012).
Yang, Y. et al. Genome-wide analysis of DNA methylation in obese, lean, and miniature pig breeds. Scientific Reports 6, 30160 (2016).
Terman, A. The IGF1R gene: A new marker for reproductive performance traits in sows? Acta Agric Scand A 61, 67–71 (2011).
Bolormaa, S. et al. A multi-trait, meta-analysis for detecting pleiotropic polymorphisms for stature, fatness and reproduction in beef cattle. PLOS Genetics 10, e1004198 (2014).
Zhu, Y. et al. Signatures of selection and interspecies introgression in the genome of Chinese domestic pigs. Genome Biology and Evolution 9, 2592–2603 (2017).
Akhabir, L. & Sandford, A. Genetics of interleukin 1 receptor-like 1 in immune and inflammatory diseases. Current Genomics 11, 591–606 (2010).
Fang, X. et al. Difference in susceptibility to Mycoplasma pneumonia among various pig breeds and its molecular genetic basis. Scientia Agricultura Sinica 48, 2839–2847 (2015).
Liu, W. et al. Complete genome sequence of Mycoplasma hyopneumoniae strain 168. Journal of Bacteriology 193, 1016–1017 (2011).
Liu, W. et al. Comparative genomic analyses of Mycoplasma hyopneumoniae pathogenic 168 strain and its high-passaged attenuated strain. BMC Genomics 14, 80 (2013).
Urb, M. & Sheppard, D. C. The role of mast cells in the defence against pathogens. PLOS Pathogens 8, e1002619 (2012).
Gariepy, C. E., Cass, D. T. & Yanagisawa, M. Null mutation of endothelin receptor type B gene in spotting lethal rats causes aganglionic megacolon and white coat color. Proceedings of the National Academy of Sciences 93, 867–872 (1996).
Yan, S. et al. Cloning and association analysis of KIT and EDNRB polymorphisms with dominant white coat color in the Chinese raccoon dog (Nyctereutes procyonoides procyonoides). Genetics and Molecular Research 14, 6549–6554 (2015).
Ramos, A. M. et al. The distal end of porcine chromosome 6p is involved in the regulation of skatole levels in boars. BMC Genetics 12, 35 (2011).
Li, C. Y., Wu, C., Liu, J. X., Wang, Y. Z. & Wang, J. K. Spatial variation of intestinal skatole production and microbial community in Jinhua and Landrace pigs. Journal of the Science of Food and Agriculture 89, 639–644 (2009).
Jeong, H. et al. Exploring evidence of positive selection reveals genetic basis of meat quality traits in Berkshire pigs through whole genome sequencing. BMC Genetics 16, 104 (2015).
Zhao, X. et al. Association of HOXA10, ZFPM2, and MMP2 genes with scrotal hernias evaluated via biological candidate gene analyses in pigs. American Journal of Veterinary Research 70, 1006–1012 (2009).
Rohrer, G. A., Nonneman, D. J., Wiedmann, R. T. & Schneider, J. F. A study of vertebra number in pigs confirms the association of vertnin and reveals additional QTL. BMC Genetics 16, 129 (2015).
Moller, M. J. et al. Pigs with the dominant white coat color phenotype carry a duplication of the KIT gene encoding the mast/stem cell growth factor receptor. Mammalian Genome 7, 822–830 (1996).
Chu, Q., Liang, T., Fu, L., Li, H. & Zhou, B. Behavioural genetic differences between Chinese and European pigs. Journal of Genetics 96, 707–715 (2017).
Bonilla, C. et al. The 8818G allele of the agouti signaling protein (ASIP) gene is ancestral and is associated with darker skin color in African Americans. Human Genetics 116, 402–406 (2005).
Sturm, R. A. Molecular genetics of human pigmentation diversity. Human Molecular Genetics 18, R9–R17 (2009).
Schachtschneider, K. M. et al. Impact of neonatal iron deficiency on hippocampal DNA methylation and gene transcription in a porcine biomedical model of cognitive development. BMC Genomics 17, 856 (2016).
Tan, C. et al. Genome-wide association study and accuracy of genomic prediction for teat number in Duroc pigs using genotyping-by-sequencing. Genetics Selection Evolution 49, 35 (2017).
Yu, L. et al. Comparative analyses of long non-coding RNA in lean and obese pigs. Oncotarget 8, 41440 (2017).
Ayuso, M. et al. Comparative analysis of muscle transcriptome between pig genotypes identifies genes and regulatory mechanisms associated to growth, fatness and metabolism. PLos One 10, e0145162 (2015).
Borowska, A., Reyer, H., Wimmers, K., Varley, P. F. & Szwaczkowski, T. Detection of pig genome regions determining production traits using an information theory approach. Livestock Science 205, 31–35 (2017).
Zambonelli, P., Gaffo, E., Zappaterra, M., Bortoluzzi, S. & Davoli, R. Transcriptional profiling of subcutaneous adipose tissue in Italian Large White pigs divergent for backfat thickness. Animal Genetics 47, 306–323 (2016).
Reyer, H. et al. Exploring the genetics of feed efficiency and feeding behaviour traits in a pig line highly selected for performance characteristics. Molecular Genetics and Genomics 292, 1001–1011 (2017).
Wang, X., Liu, X., Deng, D., Yu, M. & Li, X. Genetic determinants of pig birth weight variability. BMC Genetics 17, S15 (2016).
Chung, H. et al. A genome-wide analysis of the ultimate pH in swine. Genetics and Molecular Research 14, 15668–15682 (2015).
Le, T. H., Christensen, O. F., Nielsen, B. & Sahana, G. Genome-wide association study for conformation traits in three Danish pig breeds. Genetics Selection Evolution 49, 12 (2017).
Schneider, J. et al. Genomewide association analysis for average birth interval and stillbirth in swine. Journal of Animal Science 93, 529–540 (2015).
Do, D. N. et al. Genome-wide association study reveals genetic architecture of eating behavior in pigs and its implications for humans obesity by comparative mapping. Plos One 8, e71509 (2013).
Bai, C. et al. Genome‐wide association analysis of residual feed intake in Junmu No. 1 White pigs. Animal Genetics 48, 686–690 (2017).
Acknowledgements
This study was supported by the 2011–2016 Animal Germplasm Resources Conservation Project from the Ministry of Agriculture of China, the National Natural Science Foundation of China (Grant No. 31472069, U1402266, 31370043, 31272414,), the Agriculture Development Through Science and Technology Key Project of Shanghai (Grant No. TuiZi (2016) 1-1-4 and ChanZi (2014-2016) 6), and the National 948 Project of China (2014-Z29, 2012-Z26, 2011-G2A).
Author information
Authors and Affiliations
Contributions
Y.C.P. and S.Q.W. conceived and designed the whole study. Z.Z., X.Q., H.S., C.J.C. and C.Z.L. collected samples. Q.X., H.S., C.J.C. performed the experiments. Z.Z., X.Q., Q.Q.Z. and P.P.M. analyzed the data. M.X., J.H.Y. and Y.N.X. contributed to reagents, materials and analysis tools. Z.Z. and Q.Q.Z. wrote the manuscript. C.Y.P. revised the article. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Z., Xiao, Q., Zhang, Qq. et al. Genomic analysis reveals genes affecting distinct phenotypes among different Chinese and western pig breeds. Sci Rep 8, 13352 (2018). https://doi.org/10.1038/s41598-018-31802-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31802-x
- Springer Nature Limited
This article is cited by
-
Genetic introgression from commercial European pigs to the indigenous Chinese Lijiang breed and associated changes in phenotypes
Genetics Selection Evolution (2024)
-
A genome-wide association study for loin depth and muscle pH in pigs from intensely selected purebred lines
Genetics Selection Evolution (2023)
-
Whole-genome sequence analysis reveals selection signatures for important economic traits in Xiang pigs
Scientific Reports (2022)
-
Whole-genome sequencing of European autochthonous and commercial pig breeds allows the detection of signatures of selection for adaptation of genetic resources to different breeding and production systems
Genetics Selection Evolution (2020)